Abstract
Ten-eleven translocation (TET) proteins play key roles in the regulation of DNA-methylation status by oxidizing 5-methylcytosine (5mC) to generate 5-hydroxymethylcytosine (5hmC), which can both serve as a stable epigenetic mark and participate in active demethylation. Unlike the other members of the TET family, TET2 does not contain a DNA-binding domain, and it remains unclear how it is recruited to chromatin. Here we show that TET2 is recruited by the RNA-binding protein Paraspeckle component 1 (PSPC1) through transcriptionally active loci, including endogenous retroviruses (ERVs) whose long terminal repeats (LTRs) have been co-opted by mammalian genomes as stage- and tissue-specific transcriptional regulatory modules. We found that PSPC1 and TET2 contribute to ERVL and ERVL-associated gene regulation by both transcriptional repression via histone deacetylases and post-transcriptional destabilization of RNAs through 5hmC modification. Our findings provide evidence for a functional role of transcriptionally active ERVs as specific docking sites for RNA epigenetic modulation and gene regulation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Rasmussen, K. D. & Helin, K. Role of TET enzymes in DNA methylation, development, and cancer. Genes Dev. 30, 733–750 (2016).
Wu, X. & Zhang, Y. TET-mediated active DNA demethylation: mechanism, function and beyond. Nat. Rev. Genet. 18, 517–534 (2017).
Frye, M. & Blanco, S. Post-transcriptional modifications in development and stem cells. Development 143, 3871–3881 (2016).
Masiello, I. & Biggiogera, M. Ultrastructural localization of 5-methylcytosine on DNA and RNA. Cell. Mol. Life Sci. 74, 3057–3064 (2017).
Zhang, H. Y., Xiong, J., Qi, B. L., Feng, Y. Q. & Yuan, B. F. The existence of 5-hydroxymethylcytosine and 5-formylcytosine in both DNA and RNA in mammals. Chem. Commun. (Camb.) 52, 737–740 (2016).
Miao, Z. et al. 5-hydroxymethylcytosine is detected in RNA from mouse brain tissues. Brain Res. 1642, 546–552 (2016).
Delatte, B. et al. Transcriptome-wide distribution and function of RNA hydroxymethylcytosine. Science 351, 282–285 (2016).
Fu, L. et al. Tet-mediated formation of 5-hydroxymethylcytosine in RNA. J. Am. Chem. Soc. 136, 11582–11585 (2014).
Schlesinger, S. & Goff, S. P. Retroviral transcriptional regulation and embryonic stem cells: war and peace. Mol. Cell. Biol. 35, 770–777 (2015).
Macfarlan, T. S. et al. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487, 57–63 (2012).
Kim, J., Cantor, A. B., Orkin, S. H. & Wang, J. Use of in vivo biotinylation to study protein-protein and protein-DNA interactions in mouse embryonic stem cells. Nat. Protoc. 4, 506–517 (2009).
Wang, J. et al. A protein interaction network for pluripotency of embryonic stem cells. Nature 444, 364–368 (2006).
Huang, Y. et al. Distinct roles of the methylcytosine oxidases Tet1 and Tet2 in mouse embryonic stem cells. Proc. Natl. Acad. Sci. USA 111, 1361–1366 (2014).
Hon, G. C. et al. 5mC oxidation by Tet2 modulates enhancer activity and timing of transcriptome reprogramming during differentiation. Mol. Cell 56, 286–297 (2014).
Sigova, A. A. et al. Transcription factor trapping by RNA in gene regulatory elements. Science 350, 978–981 (2015).
Tsai, M. C. et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689–693 (2010).
Fox, A. H., Bond, C. S. & Lamond, A. I. P54nrb forms a heterodimer with PSP1 that localizes to paraspeckles in an RNA-dependent manner. Mol. Biol. Cell 16, 5304–5315 (2005).
Gilbert, S. L., Pehrson, J. R. & Sharp, P. A. XIST RNA associates with specific regions of the inactive X chromatin. J. Biol. Chem. 275, 36491–36494 (2000).
Chen, Q., Chen, Y., Bian, C., Fujiki, R. & Yu, X. TET2 promotes histone O-GlcNAcylation during gene transcription. Nature 493, 561–564 (2013).
Knott, G. J., Bond, C. S. & Fox, A. H. The DBHS proteins SFPQ, NONO and PSPC1: a multipurpose molecular scaffold. Nucleic Acids Res. 44, 3989–4004 (2016).
Ma, C. et al. Nono, a bivalent domain factor, regulates Erk signaling and mouse embryonic stem cell pluripotency. Cell Rep. 17, 997–1007 (2016).
Cao, S. et al. Specific gene-regulation networks during the pre-implantation development of the pig embryo as revealed by deep sequencing. BMC Genomics 15, 4 (2014).
Peaston, A. E. et al. Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev. Cell 7, 597–606 (2004).
Waterston, R. H. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Friedli, M. & Trono, D. The developmental control of transposable elements and the evolution of higher species. Annu. Rev. Cell Dev. Biol. 31, 429–451 (2015).
Chuong, E. B., Elde, N. C. & Feschotte, C. Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science 351, 1083–1087 (2016).
Leeb, M. et al. Polycomb complexes act redundantly to repress genomic repeats and genes. Genes Dev. 24, 265–276 (2010).
Maksakova, I. A. et al. Distinct roles of KAP1, HP1 and G9a/GLP in silencing of the two-cell-specific retrotransposon MERVL in mouse ES cells. Epigenetics Chromatin 6, 15 (2013).
Kigami, D., Minami, N., Takayama, H. & Imai, H. MuERV-L is one of the earliest transcribed genes in mouse one-cell embryos. Biol. Reprod. 68, 651–654 (2003).
Macfarlan, T. S. et al. Endogenous retroviruses and neighboring genes are coordinately repressed by LSD1/KDM1A. Genes Dev. 25, 594–607 (2011).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517, 583–588 (2015).
Rowe, H. M. & Trono, D. Dynamic control of endogenous retroviruses during development. Virology 411, 273–287 (2011).
Rowe, H. M. et al. KAP1 controls endogenous retroviruses in embryonic stem cells. Nature 463, 237–240 (2010).
Kowalska, E. et al. Distinct roles of DBHS family members in the circadian transcriptional feedback loop. Mol. Cell. Biol. 32, 4585–4594 (2012).
Li, Z. et al. Deletion of Tet2 in mice leads to dysregulated hematopoietic stem cells and subsequent development of myeloid malignancies. Blood 118, 4509–4518 (2011).
Williams, K. et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473, 343–348 (2011).
Huber, S. M. et al. Formation and abundance of 5-hydroxymethylcytosine in RNA. ChemBioChem 16, 752–755 (2015).
Amort, T. et al. Distinct 5-methylcytosine profiles in poly(A) RNA from mouse embryonic stem cells and brain. Genome Biol. 18, 1 (2017).
Zhang, X. et al. The tRNA methyltransferase NSun2 stabilizes p16INK4 mRNA by methylating the 3′-untranslated region of p16. Nat. Commun. 3, 712 (2012).
Warren, L. et al. Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7, 618–630 (2010).
Zhang, Q. et al. Tet2 is required to resolve inflammation by recruiting Hdac2 to specifically repress IL-6. Nature 525, 389–393 (2015).
Vella, P. et al. Tet proteins connect the O-linked N-acetylglucosamine transferase Ogt to chromatin in embryonic stem cells. Mol. Cell 49, 645–656 (2013).
Deplus, R. et al. TET2 and TET3 regulate GlcNAcylation and H3K4 methylation through OGT and SET1/COMPASS. EMBO J. 32, 645–655 (2013).
Chen, L. L. & Carmichael, G. G. Altered nuclear retention of mRNAs containing inverted repeats in human embryonic stem cells: functional role of a nuclear noncoding RNA. Mol. Cell 35, 467–478 (2009).
Fox, A. H. & Lamond, A. I. Paraspeckles. Cold Spring Harb. Perspect. Biol. 2, a000687 (2010).
He, C. et al. High-resolution mapping of RNA-binding regions in the nuclear proteome of embryonic stem cells. Mol. Cell 64, 416–430 (2016).
Lee, E. et al. Landscape of somatic retrotransposition in human cancers. Science 337, 967–971 (2012).
Li, W. et al. Human endogenous retrovirus-K contributes to motor neuron disease. Sci. Transl. Med. 7, 307ra153 (2015).
Choi, Y. J. et al. Deficiency of microRNA miR-34a expands cell fate potential in pluripotent stem cells. Science 355, eaag1927 (2017).
Gerdes, P., Richardson, S. R. & Faulkner, G. J. TET enzymes: double agents in the transposable element-host genome conflict. Genome Biol. 17, 259 (2016).
de la Rica, L. et al. TET-dependent regulation of retrotransposable elements in mouse embryonic stem cells. Genome Biol. 17, 234 (2016).
Ding, J., Xu, H., Faiola, F., Ma’ayan, A. & Wang, J. Oct4 links multiple epigenetic pathways to the pluripotency network. Cell Res. 22, 155–167 (2012).
Ivanova, N. et al. Dissecting self-renewal in stem cells with RNA interference. Nature 442, 533–538 (2006).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Lee, T. I. et al. Control of developmental regulators by Polycomb in human embryonic stem cells. Cell 125, 301–313 (2006).
Ule, J., Jensen, K., Mele, A. & Darnell, R. B. CLIP: a method for identifying protein-RNA interaction sites in living cells. Methods 37, 376–386 (2005).
Xue, Y. et al. Genome-wide analysis of PTB-RNA interactions reveals a strategy used by the general splicing repressor to modulate exon inclusion or skipping. Mol. Cell 36, 996–1006 (2009).
Elsässer, S. J., Noh, K. M., Diaz, N., Allis, C. D. & Banaszynski, L. A. Histone H3.3 is required for endogenous retroviral element silencing in embryonic stem cells. Nature 522, 240–244 (2015).
Ding, J. et al. Tex10 coordinates epigenetic control of super-enhancer activity in pluripotency and reprogramming. Cell Stem Cell 16, 653–668 (2015).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Acknowledgements
We thank Y. Kurihara for PSPC1 constructs, T. Macfarlan for the Zfp352-luciferase reporter construct, R. Jaenisch for the Tet-TKO ESC line, D. Trono for the Trim28-cKO line, and D.M. Gilbert and Y. Shinkai for the Ehmt2-cKO line. We also thank the medical illustrator J. Gregory from Icahn School of Medicine at Mount Sinai for the model drawing. This research was funded by the US National Institutes of Health (NIH) (grants 1R01-GM095942 and R21HD087722 to J.W.; grant R01HL112294 to M.X.) and the Empire State Stem Cell Fund through the New York State Department of Health (NYSTEM) (grants C028103 and C028121 to J.W.). J.W. is a recipient of an Irma T. Hirschl and Weill-Caulier Trusts Career Scientist Award, and M.F. is a recipient of a Ramón y Cajal contract (RYC-2014-16779) from the Ministerio de Economía y Competitividad of Spain. The research from the Shen laboratory is supported by the National Natural Science Foundation of China (31471219 and 31428010) and the Center for Life Sciences (CLS) at Tsinghua University. Additional support was provided by the Agencia Estatal de Investigación (BFU2016-80899-P to M.F.) (AEI/FEDER, UE) and the Consellería de Cultura, Educación e Ordenación Universitaria (ED431F 2016/016 to M.F.).
Author information
Authors and Affiliations
Contributions
D.G. conceived, designed and conducted the studies. D.G. and M.F. wrote the manuscript with contributions from all other authors. F.P. and M.X. generated the Tet2-knockin ESC line. J.D. and F.F. performed the TET2 interactome studies. X.H. conducted computational analysis. J.A.P., C.S., X. Shi, H.Z., P.H., F.Y., D.L., C.S.-P., A.S., M.G.B., L.C., H.W., J.S., and M.F. provided reagents and performed experiments. K.K. conducted embryo microinjections. V.J.V. provided technical advice and helpful discussion. X.B. and X. Shen conducted CLIP-seq experiments and provided helpful discussion. J.W. conceived, designed and supervised the project, and wrote and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Generation of a Tet2 3×Flag-tagged knock-in ESC line.
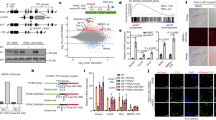
a, Targeting strategy for the generation of a Tet2:3xFlag knock-in allele in ESCs. The targeting vector contained a Neo cassette flanked by two FRT sites, followed by a 0.5 kb genomic fragment upstream of the Tet2 start codon and an ATG/3xFlag/V5 sequence. 2.2 kb 5′ and 4.8 kb 3′ arm genomic fragments were subcloned into the vector for gene targeting. Square bars indicate exons. E: EcoRI. S: ScaI. b, (Left) Southern blot of WT and Tet2:3xFlag ESC DNA digested with ScaI and hybridized with a genomic fragment external to the 5’ arm displaying a WT band of 6.3 kb and a recombinant band of 4.0 kb. (Right) ESC DNA digested with EcoRI and hybridized with a probe external to the 3’ arm displaying a WT band of 10.0 kb and a recombinant band of 8.9 kb. Representative images are shown. c, Quantitative expression of WT Tet2 and 3xFlag-Tet2 transcripts in Tet2:3xFlag ESCs. Data are reported as mean ± s.e.m. (n=3 technical replicates), relative to the WT Tet2 expression in WT ESCs using β-Actin as an internal calibrator. One representative experimental dataset is shown.
Supplementary Figure 2 PSPC1 and TET2 are coexpressed in various tissues, and their interaction in mESCs is not compromised in the absence of other TET2 partners.
a, Representative AP-MS peaks of TET2 and PSPC1 peptides identified in the SILAC IP-MS from Figure 1a, right panel. b, Western blot analysis of OGT, SIN3A, NONO, TET2 and PSPC1 in mESCs transduced with the indicated shRNAs, compared to control (shEV). GAPDH was used as a loading control. c, Validation of the interaction between endogenous PSPC1 and TET2 by IP-coIP in cells transduced with the indicated shRNAs. IgG was used as a negative control for the immunoprecipitation. Representative images of blots from 2 independent experiments are shown in b and c. d, Quantitative expression of Tet2 and Pspc1 in different mouse adult tissues, relative to the Gapdh housekeeping gene. Data are shown as mean±s.e.m. (n=2 independent experiments). e, Dot plots depicting relative RNA expression levels of Tet2 and its interacting partners determined in this study, analyzed from a previously published microarray [GSE14012] in MEFs versus pluripotent iPSCs and ESCs (log2 intensity). Data are presented as mean ± s.e.m. (replicate numbers “n” are specified for each gene). MEF, mouse embryonic fibroblast; iPSC, induced pluripotent stem cell; ESC, embryonic stem cell.
Supplementary Figure 3 PSPC1 depletion does not affect the self-renewal or pluripotency of ESCs.
a, Western blotting analysis of total lysates of ESCs with Pspc1 knock-down using three different small hairpin RNAs (shPspc1#1, shPspc1#2, shPspc1#3) and control empty vector knock-down (shEV). β-ACTIN (ACTB) blot is shown as a protein loading control. Representative images of blots from 3 independent experiments are shown. b, Cell cycle analysis of PSPC1-depleted ESCs using two different shRNAs (shPspc1#1 and shPspc1#3) compared to a control (shEV). Y-axis represents the percentages of cells in each cell cycle phase: G2/M, S and G1. c, Immunostaining of PSPC1 (red), POU5F1/OCT4 (green) and SOX2 (blue) in ESCs with knock-down of PSPC1 (shPspc1#1 and shPspc1#3) compared to the control knock-down (shEV). Brightfield is shown to visualize all cells. Representative images from 3 independent experiments are shown. d, Relative expression of pluripotency (Pou5f1, Sox2, Nanog, Zfp42) and lineage (Gata6, Sox17, Brachyury/T, Fgf5) markers in PSPC1 (shPspc1#1 and shPspc1#3) and TET2 (shTet2) knocked-down cells relative to the control knock-down (shEV) using β-Actin as an internal calibrator. Data are shown as mean ±s.e.m. from at least two independent experiments.
Supplementary Figure 4 Generation and characterization of a Pspc1-KO ESC line rescued with wild-type and RNA-binding-deficient PSPC1.
a, (Top) Scheme of the genomic position targeted by the sgRNA at the Pspc1 locus, showing the sgRNA recognition site (underlined) and the PAM sequence (in red). (Bottom) Sequence confirmation of the genomic deletion observed in Pspc1 CRISPR-Cas9 knock-out ESC line. b, Immunostaining of PSPC1 (red), OCT4 (green) and SOX2 (blue) of wild-type (WT) and Pspc1 knock-out (Pspc1 KO) ESCs. Representative images from 2 independent experiments are shown. c, (Left) Relative expression of Pspc1 in WT and Pspc1 KO cells rescued with EV, PSPC1WT and PSPC1Mut, determined by qPCR and normalized to β-Actin and WT cells. Data are shown as mean ± s.e.m. (n=3 independent experiments). Two-tailed student´s t-test was performed. ns, not significant. (Right) Western blotting analysis of whole cell extracts (WCE) showing PSPC1 expression levels in rescued Pspc1 KO cell lines. GAPDH is shown as a loading control. d, Agarose gel of RNA fraction bound by the wild-type (+3xFLPSPC1WT) and mutant (+3xFLPSPC1Mut) PSPC1 protein determined by iv-RIP. RNA is visualized by staining with ethidium bromide. e, Immunoprecipitation of MycPSPC1WT and MycPSPC1Mut and western blotting analysis of the coimmunoprecipitated 3xFLTET2 in HEK293T cells. f, Western blotting analysis of TET2 at the chromatin (Chromatin) (left top) or in whole cell extracts (WCE) (left bottom) in Pspc1 KO cells before (+3xFLEV) or after rescue with wild-type (+3xFLPSPC1WT) or RNA-binding mutant (+3xFLPSPC1Mut) PSPC1. Histone H3 was used as a loading control. (Right) Quantitation of the Western blots shown on left. Data are normalized to Histone H3 and calculated relative to EV. Representative images from 2 independent experiments are shown in d, e and f.
Supplementary Figure 5 TET2 interacts with RNA through PSPC1 both in vitro and in vivo.
a, Percentages of RNA recovered after incubation with PSPC1 (PSPC1 iv-RIP) or TET2 (TET2 iv-RIP) protein complexes in wild-type (WT) and Pspc1 KO cells. Half of the samples were treated with RNase A to validate that all the nucleic acids IP'ed are RNA and there is no DNA contamination. b, (Left) Diagram showing the protocol used for in vivo RNA interaction assay after formaldehyde crosslinking and IP with PSPC1, TET2 or IgG (control) antibodies. Immunoprecipitated (IPed) RNA was converted to cDNA to make it more stable for visualization. (Right) Silver staining of cDNA run in a SDS-PAGE gel before (Input) and after IP with PSPC1, TET2 or IgG (control) antibodies. Treatment with DNase I and RNase A provides a strong evidence that all IPed nucleic acids are indeed RNA but not DNA. Representative images from 2 independent experiments are shown. c, PSPC1 RIP-qPCR in Tet2 KO ESCs. IgG is a negative control for the IP. Data are shown as mean ± s.e.m. (n=3 independent experiments).
Supplementary Figure 6 PSPC1 regulates expression of developmental genes in ESCs.
a, PSPC1 CLIP-seq peaks at Adss and Ywhae transcripts. ChIP-qPCR analysis of TET2 at PSPC1 CLIP-seq target loci in Pspc1 KO cells rescued with a control empty vector (+EV), a wild-type 3xFLPSPC1 (PSPC1WT), or an RNA-binding mutant 3xFLPSPC1Mut (PSPC1Mut). Data are from one representative experiment (n=2); Data are presented as mean ± s.e.m (n=3 technical replicates). b, Scatter plot of gene expression (in log10 FPKM) in PSPC1 knock-down (shPspc1#3) versus control knock-down (shEV) ESCs determined by RNA-seq. Biological duplicates were analyzed. Genes upregulated more than 1.5-fold are shown in red and genes downregulated more than 1.5-fold are shown in blue. c, An overlap between PSPC1-bound (CLIP-seq) and PSPC1 de-regulated (RNA-seq) coding RNAs in ESCs. d, Gene ontology (GO) analysis of PSPC1 de-regulated genes. PSPC1-activated genes (downregulated after PSPC1 knock-down) are shown in blue and PSPC1-repressed genes (upregulated after PSPC1 knock-down) are shown in red. The P-value of each process is presented as log2. e, Box plot of expression levels (log10RPKM) of PSPC1 de-regulated genes determined by RNA-seq during preimplantation development22. Whiskers represent 10-90 percentile. OO, oocyte; 2C, 2-cell embryo; 4C, 4-cell embryo; 8C, 8-cell embryo; MO, morula; BL, blastocyst; TE, trophectoderm.
Supplementary Figure 7 PSPC1 binds and regulates repetitive elements in vivo and in vitro.
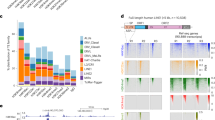
a, A doughnut chart showing the distribution (in percentage) of LTR-containing and non-LTR LINE and SINE elements among retrotransposable elements (RE) bound by PSPC1 according to CLIP-seq. b, Percentages of input recovery of U6, IAP and MERVL RNAs after PSPC1 CLIP determined by qPCR. Data are presented as mean ± s.e.m. (n=3 experimental replicates). c, Expression (in reads per million, RPM) of ERV1, ERVK, ERVL and MaLR elements in wild type ESCs. Box plot with whiskers extending to ±1.5 × IQR; line across the box indicates the median. Statistical analysis: Wilcoxon rank sum test with continuity correction. **P<0.01, ns: statistically not significant. d, Percentages of input recovery of RNA bound in vitro (iv-RIP) by PSPC1 in wild-type (Pspc1 WT) and Pspc1 knock-out (Pspc1 KO) ESCs. U6 is a negative control for the IP. Non-LTR and LTR-containing RNAs were assayed. e, Percentages of input recovery of RNA in vivo immunoprecipitated with anti-PSPC1 antibody after formaldehyde crosslinking of cells. U6 is a negative control for the IP. Non-LTR and LTR-containing RNAs were assayed. f, Relative expression of Non-LTR and LTR containing ERV elements in PSPC1 depleted (shPspc1) compared to control (shEV) knock-down cells. g, MERVL is bound and repressed by PSPC1. The read coverage along an internal MERVL transcript is shown by CLIP-seq reads (blue) and RNA-seq reads of shEV (dark pink) and shPspc1 (light pink) mapping to the genome. h, IAP and MusD expression in wild type (WT) and Pspc1 KO cells and the indicated rescued cell lines. Data in d, e, f and h are shown as mean ± s.e.m. (n=3 independent experiments). Statistical analysis: Two-tailed Student´s t-test was performed. ns, not significant.
Supplementary Figure 8 PSPC1 regulates 2C-specific gene expression through regulation of co-opted MERVL elements.
a, Frequency histogram of absolute distances from each gene upregulated by PSPC1 knock-down to the nearest LTR-containing element bound by PSPC1 (CLIP-seq). A similar amount of PSPC1 unbound LTR-containing RNAs was used as background control (n=14,220). 341 versus 138 LTRs bound and not-bound by PSPC1, respectively, were found within 50 kb. P < 2.2*10−16 calculated by binomial statistical two-tailed test. b, A ranked list of 2C embryo genes deregulated by PSPC1 knock-down based on the genomic distance of their TSSs to the closest ERVL element. c, (Top) Diagram of the luciferase reporter containing the MT2B MERVL element responsible for PSPC1-regulated Zfp352 locus and the MT2B-lacking reporter control. (Bottom) Luciferase assay of the Zfp352 reporter in PSPC1-expressing cells (Pspc1 WT) compared to Pspc1 knock-out cells (Pspc1 KO). Representative data from 3 independent experiments are shown. Data are presented as mean ± s.e.m. (n=3 technical replicates). Two-tailed Student's t-test was performed. d, Schematic representation of the CRISPR activation system used to activate MERVL (sgRNA MERVL) or a non-targeting control (sgRNA NT). e, Relative expression of MERVL and 2C-coregulated genes relative to the control (white dots) upon MERVL activation (blue dots). Data are shown as mean ± s.e.m. (n=3 technical replicates).
Supplementary Figure 9 TET2 binds retroelements through PSPC1 and regulates their expression.
a, Percentages of input recovery of RNA bound in vitro (iv-RIP) by TET2 in wild-type (Pspc1 WT) and Pspc1 knock-out (Pspc1 KO) ESCs. U6 is a negative control for the IP. Non-LTR and LTR-containing RNAs were assayed. b, Percentages of input recovery of RNA in vivo immunoprecipitated with anti-TET2 antibody after formaldehyde crosslinking of cells. U6 is a negative control for the IP. Samples were treated with RNase A to rule out potential DNA contamination. Non-LTR and LTR-containing RNAs were assayed. c, Relative expression of Non-LTR and LTR containing ERV elements in TET2 depleted (shTet2) cells compared to control (shEV) cells. Data in a, b and c are shown as mean ± s.e.m. (n=3 independent experiments). Statistical analysis: Two-tailed Student´s t-test was performed. ns, not significant.
Supplementary Figure 10 The contribution of PSPC1–TET2 to ERV regulation is similar to that of other ERV regulators such as TRIM28 and EHMT2.
a, Relative expression changes of different ERV families in Pspc1 knock-down, Trim28 knock-out and Ehmt2 knock-out ESCs relative to their control (shEV and WT, respectively). Center line, median; box and whisker plots: ± 10th–90th percentile range. Average expression is depicted with “+”. Statistical analysis: Two-tailed Student´s t-test on the means was calculated; ns, not significant. The number (n) of ERVs of each category analyzed is specified in the figure. b, Trim28 and MERVL expression in inducible Trim28 knock-out cells33 before (Vehicle or Veh.) and after 4-hydroxytamoxifen (4-OHT) treatment. c, Ehmt2 and MERVL expression in Ehmt2 inducible knock-out cell line before (Vehicle or Veh.) and after 4-hydroxytamoxifen (4-OHT) treatment. d, Relative expression of MERVL in Ehmt2 knock-out, Tet2 knock-out, Trim28 knock-out and Pspc1 knock-out ESCs compared to their corresponding controls. Representative data from 2 independent experiments are shown in b, c and d. Data are presented as mean ± s.e.m. (n=3 technical replicates).
Supplementary Figure 11 Depletion of PSPC1 or TET2 in early embryos results in deregulation of MERVL and MERVL-associated transcripts.
a, Representative pictures of embryos injected with non-targeting (siNT) control, Pspc1 (siPspc1) or Tet2 (siTet2) knock-down after 1.5, 2.5 and 4.5 days of in vitro culture. The experiment was repeated three times with similar results. b, Percentages of embryos reaching the corresponding stage after Pspc1, Tet2 or control knock-down. Data are representative from 3 independent experiments. Statistical analysis: Two-tailed Student´s t-test; ns, not significant. c-d, Expression of 2C genes at 1.5 dpc and 2.5 dpc, relative to Hprt and a non-targeting siRNA, in presence of the indicated siRNAs. e, Relative expression of Pspc1 and Tet2 at the blastocyst stage after knock-down of respective gene (siPspc1 or siTet2) compared to a non-targeting control (siNT) during in vitro embryo development. Expression was calculated relative to β-actin (Actb). f, Relative expression of PSPC1 and TET2-regulated 2C genes Zfp352 and Zscan4 at the blastocyst stage after Pspc1, Tet2 or a control (siNT) knock-down. Data in c-f are shown as mean ± s.e.m. (n=3 independent experiments). Statistical analysis: Two-tailed Student's t-test; ns, not significant.
Supplementary Figure 12 Analysis of DNA 5mC and 5hmC modification at ERV loci.
a, Line plots of mean hMeDIP-seq and MeDIP-seq density enrichment of 5hmC and 5mC, respectively, at chromatin in non-repetitive (n=10343) and LTR-containing (n=2493) DNA sequences centered in PSPC1 CLIP-seq regions. b, 5hmC DNA IP and RT-qPCR analysis of the indicated loci. Representative data from 2 independent experiments are shown. Data are presented as mean ± s.e.m. (n=4 technical replicates). c, Relative expression of Tet2 after rescue of Tet TKO cells with a wild-type (+TET2WT) or catalytic mutant (+TET2Mut) TET2 compared to a wild-type cell line (WT). β-actin was used as a calibrator. Data are shown as mean ±s.e.m. (n=3 technical replicates). d, Relative expression of IAP and MusD in Tet TKO cells and the indicated rescue lines. Data are shown as mean ± s.e.m. (n=3 independent experiments). Statistical analysis: Two-tailed Student’s t-test; ns, not significant.
Supplementary Figure 13 Determination of 5mC and 5hmC levels in ERV RNAs.
a, Relative enrichment of MERVL, IAP, MusD and U6 negative control among anti-5hmC immunoprecipitated RNAs compared to control IgG immunoprecipitation. b, (Left) Scheme of the unfragmented RIP-qPCR used to assess MERVL abundance among 5mC and 5hmC modified RNAs (center and right, respectively), relative to U6 negative control. Percentage was calculated as the ratio of immunoprecipitated (IPed) RNA over total RNA (as the sum of IPed and unbound (flow-through, FT) RNAs). c, Overlap of PSPC1 CLIP-seq peaks and previously reported chimeric transcripts10. d, Percentages of input recovery of the indicated chimeric RNAs after immunoprecipitation of 5hmC compared to IgG control. Representative data from 2 independent experiments are shown in a, b and d. Data are presented as mean ± s.e.m. (n=3 technical replicates). e, (Left) RNA EMSA experiment of biotin-conjugated and methylated (5mC) or hydroxymethylated (5hmC) probes incubated with MycPSPC1 and 3xFLTET2-immunopurified complexes (by anti-FLAG IP) and developed with streptavidin antibody. A representative blot from three independent experiments (with similar results) is shown. (Right) Quantification of the percentage of each probe bound by the PSPC1-TE2 protein complex relative to total probe. Statistical test: Two-tailed Student´s t-test. The p-values indicate significant higher affinity of 5mC probes than 5hmC probes in binding to the PSPC1-TET2 complex. f, A putative model for PSPC1-TET2 mediated MERVL degradation in pluripotent ESCs. MERVL RNA is readily detectable for 5mC and 5hmC modification in ESCs (see panel b). The higher affinity of the PSPC1-TET2 protein complex for 5mC-modified than 5hmC-modified MERVL transcripts (data shown in e) suggests the tendency of oxidation of 5mC-modified MERVL RNAs and subsequent release of PSPC1-TET2 from MERVL transcripts. The 5hmC may attract unknown "readers" for RNA degradation in pluripotent stem cells leading to their destabilization (dotted line). "X" denotes additional factor(s) that may associate with PSPC1 and contribute to the targeting specificity of PSPC1-TET2 in RNA hydroxymethylation.
Supplementary Figure 14 PSPC1 and TET2 destabilize MERVL RNAs.
a, MERVL enrichment among 5hmC-modified RNAs in Tet TKO ESCs (+EV) or Tet TKO cells rescued with each of the three TET family members. U6 RNA is a negative control. Representative data from 2 independent experiments are shown. Data are presented as mean ± s.e.m. (n=3 technical replicates). b, Relative abundance of MERVL RNAs after transcriptional inhibition with α-amanitin in wild-type (WT), Pspc1 knock-out (Pspc1KO) and Tet2 knock-out (Tet2KO) ESCs compared to cells treated with Milli-Q water (Vehicle). 18S rRNA was used as an internal calibrator. c, (Top) Schematic representation of the 5-ethynyl uridine (EU) pulse given to ESCs to measure RNA stability. (Bottom) MERVL RNA stability measurement in absence of PSPC1 or TET2, assessed by a EU-incorporation pulse and measurement of RNA abundance in the three specified cell lines. Data in a and b are presented as mean ± s.e.m. (n=3 independent experiments). Statistical test: Two-tailed Student´s t-test. d, De novo motif analysis using the Homer’s find Motifs Genome program FindMotifsGenome.pl. The top five P-value ranking sequence motifs identified after analysis of PSPC1 CLIP-seq targets are displayed. e, (Top) Schematic depiction of pMotif-d2EGFP reporter construct containing each of the top five consensus motifs of PSPC1 CLIP-seq upstream of the destabilized EGFP protein (d2EGFP). (Bottom) Sequences of the motifs cloned in the reporter system. Cytosines were mutated to Adenines to generate the mutated motifs. f, Mean fluorescence intensity of EGFP of cells transfected with the indicated reporters and treated with α-Amanitin to inhibit transcription, relative to untreated (Veh.). Data are shown as mean ± s.e.m. (n=3 independent experiments). Statistical analysis: Two-tailed Student´s t-test; ns, not significant.
Supplementary Figure 15 Cooperation of HDAC1/2 and associated activities with PSPC1 and TET2 in transcriptional repression of MERVL and MERVL-regulated genes.
a-b, PSPC1 physically associates with HDAC1 and HDAC2. Co-immunoprecipitation of exogenously expressed MycPSPC1 and 3xFLHDAC1 (a) or 3xFLHDAC2 (b) in HEK293T cells was performed. Representative images of two independent experimental replicates are shown. c, Relative enrichment of HDAC1 and HDAC2 at MERVL genomic loci in Pspc1 knock-out (Pspc1 KO+3xFLEV) and rescued cell lines (Pspc1 KO+3xFLPSPC1WT and Pspc1 KO+3xFLPSPC1Mut) determined by ChIP-qPCR. Enrichment is calculated relative to a negative genomic region in chromosome 6. Data are shown as mean± s.e.m. (n=3 independent experiments). Statistical test: Two-tailed Student's t-test. d, Gene expression of MERVL, Zfp352 and Oct4 in cells treated with an HDAC inhibitor (Valproic acid, VPA) or DMSO control (Veh.) relative to β-actin as a calibrator. e, Relative gene expression of MERVL, Zfp352 and Oct4 in ESCs with HDAC1 or HDAC2 knock-down (shHdac1 and shHdac2, respectively) compared to a control knock-down (shEV). Expression was normalized relative to β-actin as a calibrator. Representative data from 2 independent experiments are shown in d and e. Data points are shown as mean ± s.e.m. (n=3 technical replicates). f, Relative expression of MERVL in Pspc1 KO ESCs rescued with EV, PSPC1WT or PSPC1Mut (left) or Tet TKO ESCs rescued with EV, TET2WT or TET2Mut (Right), in the presence (VPA) or absence (Veh.) of HDAC inhibition. g, Percentages of input recovery of MERVL and U6 negative control upon ChIP of H3K9me2 in Pspc1 KO and Tet2 KO ESCs compared to WT cells. Data in f and g are shown as mean ± s.e.m. (n=3 independent experiments). Statistical analysis: Two-tailed Student´s t-test; ns, not significant.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15
Supplementary Table 1
List of TET2-interacting proteins ranked by their enrichment in two complementary AP-MS techniques, related to Fig. 1a
Supplementary Table 2
PSPC1 CLIP-seq peaks (piranha-annotated), related to Fig. 3a
Supplementary Table 3
RNA-seq of PSPC1-knockdown ESCs. List of deregulated genes obtained from biological duplicates, related to Supplementary Fig. 7b
Supplementary Table 4
shRNAs, RT-qPCR, CLIP-seq, RIP-qPCR and ChIPqPCR oligos used in this study
Rights and permissions
About this article
Cite this article
Guallar, D., Bi, X., Pardavila, J.A. et al. RNA-dependent chromatin targeting of TET2 for endogenous retrovirus control in pluripotent stem cells. Nat Genet 50, 443–451 (2018). https://doi.org/10.1038/s41588-018-0060-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0060-9
This article is cited by
-
A multiomics dataset for the study of RNA modifications in human macrophage differentiation and polarisation
Scientific Data (2024)
-
K235 acetylation couples with PSPC1 to regulate the m6A demethylation activity of ALKBH5 and tumorigenesis
Nature Communications (2023)
-
Nuclear localization of TET2 requires β-catenin activation and correlates with favourable prognosis in colorectal cancer
Cell Death & Disease (2023)
-
TET (Ten-eleven translocation) family proteins: structure, biological functions and applications
Signal Transduction and Targeted Therapy (2023)
-
GID complex regulates the differentiation of neural stem cells by destabilizing TET2
Frontiers of Medicine (2023)