Abstract
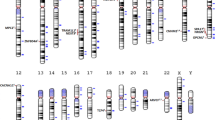
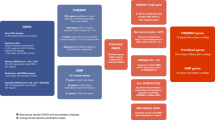
Schizophrenia is a debilitating psychiatric condition often associated with poor quality of life and decreased life expectancy. Lack of progress in improving treatment outcomes has been attributed to limited knowledge of the underlying biology, although large-scale genomic studies have begun to provide insights. We report a new genome-wide association study of schizophrenia (11,260 cases and 24,542 controls), and through meta-analysis with existing data we identify 50 novel associated loci and 145 loci in total. Through integrating genomic fine-mapping with brain expression and chromosome conformation data, we identify candidate causal genes within 33 loci. We also show for the first time that the common variant association signal is highly enriched among genes that are under strong selective pressures. These findings provide new insights into the biology and genetic architecture of schizophrenia, highlight the importance of mutation-intolerant genes and suggest a mechanism by which common risk variants persist in the population.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
03 June 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Owen, M. J., Sawa, A. & Mortensen, P. B. Schizophrenia. Lancet 388, 86–97 (2016).
Thornicroft, G. Physical health disparities and mental illness: the scandal of premature mortality. Br. J. Psychiatry 199, 441–442 (2011).
Olfson, M., Gerhard, T., Huang, C., Crystal, S. & Stroup, T. S. Premature mortality among adults with schizophrenia in the United States. JAMA Psychiatry 72, 1172–1181 (2015).
Morgan, C. et al. Reappraising the long-term course and outcome of psychotic disorders: the AESOP-10 study. Psychol. Med. 44, 2713–2726 (2014).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Singh, T. et al. Rare loss-of-function variants in SETD1A are associated with schizophrenia and developmental disorders. Nat. Neurosci. 19, 571–577 (2016).
Rees, E. et al. Analysis of copy number variations at 15 schizophrenia-associated loci. Br. J. Psychiatry 204, 108–114 (2014).
Purcell, S. M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014).
Power, R. A. et al. Fecundity of patients with schizophrenia, autism, bipolar disorder, depression, anorexia nervosa, or substance abuse vs their unaffected siblings. JAMA Psychiatry 70, 22–30 (2013).
Huxley, J., Mayr, E., Osmond, H. & Hoffer, A. Schizophrenia as a genetic morphism. Nature 204, 220–221 (1964).
Shaner, A., Miller, G. & Mintz, J. Schizophrenia as one extreme of a sexually selected fitness indicator. Schizophr. Res. 70, 101–109 (2004).
Srinivasan, S. et al. Genetic markers of human evolution are enriched in schizophrenia. Biol. Psychiatry 80, 284–292 (2016).
Uher, R. The role of genetic variation in the causation of mental illness: an evolution-informed framework. Mol. Psychiatry 14, 1072–1082 (2009).
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat. Genet. 45, 1150–1159 (2013).
Shi, Y. et al. Common variants on 8p12 and 1q24.2 confer risk of schizophrenia. Nat. Genet. 43, 1224–1227 (2011).
Deciphering Developmental Disorders Study. Prevalence and architecture of de novo mutations in developmental disorders. Nature 542, 433–438 (2017).
Kosmicki, J. A. et al. Refining the role of de novo protein-truncating variants in neurodevelopmental disorders by using population reference samples. Nat. Genet. 49, 504–510 (2017).
Samocha, K. E. et al. A framework for the interpretation of de novo mutation in human disease. Nat. Genet. 46, 944–950 (2014).
Genovese, G. et al. Increased burden of ultra-rare protein-altering variants among 4,877 individuals with schizophrenia. Nat. Neurosci. 19, 1433–1441 (2016).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Fagerberg, L. et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell. Proteomics 13, 397–406 (2014).
Smith, N. G. C. & Eyre-Walker, A. Human disease genes: patterns and predictions. Gene 318, 169–175 (2003).
Blekhman, R. et al. Natural selection on genes that underlie human disease susceptibility. Curr. Biol. 18, 883–889 (2008).
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP–trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Takata, A., Ionita-Laza, I., Gogos, J. A., Xu, B. & Karayiorgou, M. De novo synonymous mutations in regulatory elements contribute to the genetic etiology of autism and schizophrenia. Neuron 89, 940–947 (2016).
Huber, C. D., DeGiorgio, M., Hellmann, I. & Nielsen, R. Detecting recent selective sweeps while controlling for mutation rate and background selection. Mol. Ecol. 25, 142–156 (2016).
McVicker, G., Gordon, D., Davis, C. & Green, P. Widespread genomic signatures of natural selection in hominid evolution. PLoS Genet. 5, e1000471 (2009).
Pocklington, A. J. et al. Novel findings from CNVs implicate inhibitory and excitatory signaling complexes in schizophrenia. Neuron 86, 1203–1214 (2015).
Fromer, M. et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 506, 179–184 (2014).
Darnell, J. C. et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 146, 247–261 (2011).
Iossifov, I. et al. De novo gene disruptions in children on the autistic spectrum. Neuron 74, 285–299 (2012).
Szatkiewicz, J. P. et al. Copy number variation in schizophrenia in Sweden. Mol. Psychiatry 19, 762–773 (2014).
Blake, J. A., Bult, C. J., Eppig, J. T., Kadin, J. A. & Richardson, J. E. The Mouse Genome Database: integration of and access to knowledge about the laboratory mouse. Nucleic Acids Res. 42, D810–D817 (2014).
Müller, C. S. et al. Quantitative proteomics of the Cav2 channel nano-environments in the mammalian brain. Proc. Natl. Acad. Sci. USA 107, 14950–14957 (2010).
Bécamel, C. et al. Synaptic multiprotein complexes associated with 5-HT2C receptors: a proteomic approach. EMBO J. 21, 2332–2342 (2002).
Liu, J. et al. Prediction of efficacy of vabicaserin, a 5-HT2C agonist, for the treatment of schizophrenia using a quantitative systems pharmacology model. CPT Pharmacometrics Syst. Pharmacol. 3, e111 (2014).
Benner, C. et al. FINEMAP: efficient variable selection using summary data from genome-wide association studies. Bioinformatics 32, 1493–1501 (2016).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164–e164 (2010).
Won, H. et al. Chromosome conformation elucidates regulatory relationships in developing human brain. Nature 538, 523–527 (2016).
Park, J. H. et al. SLC39A8 deficiency: a disorder of manganese transport and glycosylation. Am. J. Hum. Genet. 97, 894–903 (2015).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Charlesworth, B. The effects of deleterious mutations on evolution at linked sites. Genetics 190, 5–22 (2012).
Charlesworth, B., Betancourt, A. J., Kaiser, V. B. & Gordo, I. Genetic recombination and molecular evolution. Cold Spring Harb. Symp. Quant. Biol. 74, 177–186 (2009).
Comeron, J. M., Williford, A. & Kliman, R. M. The Hill–Robertson effect: evolutionary consequences of weak selection and linkage in finite populations. Heredity 100, 19–31 (2008).
Charlesworth, B. Background selection 20 years on: the Wilhelmine E. Key 2012 Invitational Lecture. J. Hered. 104, 161–171 (2013).
North, T. L. & Beaumont, M. A. Complex trait architecture: the pleiotropic model revisited. Sci. Rep. 5, 9351 (2015).
Rockman, M. V., Skrovanek, S. S. & Kruglyak, L. Selection at linked sites shapes heritable phenotypic variation in C. elegans. Science 330, 372–376 (2010).
Vitti, J. J., Grossman, S. R. & Sabeti, P. C. Detecting natural selection in genomic data. Annu. Rev. Genet. 47, 97–120 (2013).
Stephan, W. Signatures of positive selection: from selective sweeps at individual loci to subtle allele frequency changes in polygenic adaptation. Mol. Ecol. 25, 79–88 (2016).
Field, Y. et al. Detection of human adaptation during the past 2000 years. Science 354, 760–764 (2016).
Key, F. M., Fu, Q., Romagné, F., Lachmann, M. & Andrés, A. M. Human adaptation and population differentiation in the light of ancient genomes. Nat. Commun. 7, 10775 (2016).
Harris, K. & Nielsen, R. The genetic cost of Neanderthal introgression. Genetics 203, 881–891 (2016).
Peloso, G. M. & Lunetta, K. L. Choice of population structure informative principal components for adjustment in a case–control study. BMC Genet. 12, 64 (2011).
Pirinen, M., Donnelly, P. & Spencer, C. C. A. Including known covariates can reduce power to detect genetic effects in case–control studies. Nat. Genet. 44, 848–851 (2012).
Bouaziz, M., Ambroise, C. & Guedj, M. Accounting for population stratification in practice: a comparison of the main strategies dedicated to genome-wide association studies. PLoS One 6, e28845 (2011).
Tucker, G., Price, A. L. & Berger, B. Improving the power of GWAS and avoiding confounding from population stratification with PC-Select. Genetics 197, 1045–1049 (2014).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Perälä, J. et al. Lifetime prevalence of psychotic and bipolar I disorders in a general population. Arch. Gen. Psychiatry 64, 19–28 (2007).
McGrath, J., Saha, S., Chant, D. & Welham, J. Schizophrenia: a concise overview of incidence, prevalence, and mortality. Epidemiol. Rev. 30, 67–76 (2008).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Schizophrenia Psychiatric Genome-Wide Association Study (GWAS) Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 43, 969–976 (2011).
Tansey, K. E. et al. Common alleles contribute to schizophrenia in CNV carriers. Mol. Psychiatry 21, 1085–1089 (2015).
Dudbridge, F. Polygenic epidemiology. Genet. Epidemiol. 40, 268–272 (2016).
Lee, S. H., Goddard, M. E., Wray, N. R. & Visscher, P. M. A better coefficient of determination for genetic profile analysis. Genet. Epidemiol. 36, 214–224 (2012).
Maston, G. A., Evans, S. K. & Green, M. R. Transcriptional regulatory elements in the human genome. Annu. Rev. Genomics Hum. Genet. 7, 29–59 (2006).
Network and Pathway Analysis Subgroup of Psychiatric Genomics Consortium. Psychiatric genome-wide association study analyses implicate neuronal, immune and histone pathways. Nat. Neurosci. 18, 199–209 (2015).
Cox, D. D. & Lee, J. S. Pointwise testing with functional data using the Westfall–Young randomization method. Biometrika 95, 621–634 (2008).
Batada, N. N., Hurst, L. D. & Tyers, M. Evolutionary and physiological importance of hub proteins. PLoS Comput. Biol. 2, e88 (2006).
Parikshak, N. N., Gandal, M. J. & Geschwind, D. H. Systems biology and gene networks in neurodevelopmental and neurodegenerative disorders. Nat. Rev. Genet. 16, 441–458 (2015).
Pocklington, A. J., Cumiskey, M., Armstrong, J. D. & Grant, S. G. N. The proteomes of neurotransmitter receptor complexes form modular networks with distributed functionality underlying plasticity and behaviour. Mol. Syst. Biol. 2, 2006.0023 (2006).
Fernández, E. et al. Targeted tandem affinity purification of PSD-95 recovers core postsynaptic complexes and schizophrenia susceptibility proteins. Mol. Syst. Biol. 5, 269 (2009).
Kirov, G. et al. De novo CNV analysis implicates specific abnormalities of postsynaptic signalling complexes in the pathogenesis of schizophrenia. Mol. Psychiatry 17, 142–153 (2012).
Pers, T. H. et al. Comprehensive analysis of schizophrenia-associated loci highlights ion channel pathways and biologically plausible candidate causal genes. Hum. Mol. Genet. 25, 1247–1254 (2016).
Gene Ontology Consortium. Gene Ontology Consortium: going forward. Nucleic Acids Res. 43, D1049–D1056 (2015).
Fabregat, A. et al. The Reactome Pathway Knowledgebase. Nucleic Acids Res. 44 (D1), D481–D487 (2016).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44 (D1), D457–D462 (2016).
Amberger, J. S., Bocchini, C. A., Schiettecatte, F., Scott, A. F. & Hamosh, A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 43, D789–D798 (2015).
de Leeuw, C. A., Neale, B. M., Heskes, T. & Posthuma, D. The statistical properties of gene-set analysis. Nat. Rev. Genet. 17, 353–364 (2016).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44 (D1), D733–D745 (2016).
van Dongen, J. & Boomsma, D. I. The evolutionary paradox and the missing heritability of schizophrenia. Am. J. Med. Genet. B Neuropsychiatr. Genet. 162B, 122–136 (2013).
Xu, K., Schadt, E. E., Pollard, K. S., Roussos, P. & Dudley, J. T. Genomic and network patterns of schizophrenia genetic variation in human evolutionary accelerated regions. Mol. Biol. Evol. 32, 1148–1160 (2015).
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J. K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Sabeti, P. C. et al. Genome-wide detection and characterization of positive selection in human populations. Nature 449, 913–918 (2007).
Grossman, S. R. et al. Identifying recent adaptations in large-scale genomic data. Cell 152, 703–713 (2013).
Sankararaman, S. et al. The genomic landscape of Neanderthal ancestry in present-day humans. Nature 507, 354–357 (2014).
Sabeti, P. C. et al. Positive natural selection in the human lineage. Science 312, 1614–1620 (2006).
Ronen, R., Udpa, N., Halperin, E. & Bafna, V. Learning natural selection from the site frequency spectrum. Genetics 195, 181–193 (2013).
Nordborg, M., Charlesworth, B. & Charlesworth, D. The effect of recombination on background selection. Genet. Res. 67, 159–174 (1996).
Fu, W. & Akey, J. M. Selection and adaptation in the human genome. Annu. Rev. Genomics Hum. Genet. 14, 467–489 (2013).
Zhao, H. et al. CrossMap: a versatile tool for coordinate conversion between genome assemblies. Bioinformatics 30, 1006–1007 (2014).
Cunningham, F. et al. Ensembl 2015. Nucleic Acids Res. 43, D662–D669 (2015).
Altshuler, D. M. et al. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
Maclean, C. A., Chue Hong, N. P. & Prendergast, J. G. hapbin: an efficient program for performing haplotype-based scans for positive selection in large genomic datasets. Mol. Biol. Evol. 32, 3027–3029 (2015).
Hussin, J. G. et al. Recombination affects accumulation of damaging and disease-associated mutations in human populations. Nat. Genet. 47, 400–404 (2015).
Ptok, U., Barkow, K. & Heun, R. Fertility and number of children in patients with Alzheimer’s disease. Arch. Womens Ment. Health 5, 83–86 (2002).
Whitworth, K. W., Baird, D. D., Stene, L. C., Skjaerven, R. & Longnecker, M. P. Fecundability among women with type 1 and type 2 diabetes in the Norwegian Mother and Child Cohort Study. Diabetologia 54, 516–522 (2011).
Jokela, M. Birth-cohort effects in the association between personality and fertility. Psychol. Sci. 23, 835–841 (2012).
Zheng, J. et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics 33, 272–279 (2017).
Lambert, J.-C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Smith, D. J. et al. Genome-wide analysis of over 106 000 individuals identifies 9 neuroticism-associated loci. Mol. Psychiatry 21, 749–757 (2016).
Mahajan, A. et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat. Genet. 46, 234–244 (2014).
Acknowledgements
General. This project has received funding from the European Union’s Seventh Framework Programme for research, technological development and demonstration under grant agreement 279227 (CRESTAR Consortium). The work at Cardiff University was funded by the Medical Research Council (MRC) Centre (MR/L010305/1), a program grant (G0800509) and a project grant (MR/L011794/1) and by the European Community’s Seventh Framework Programme HEALTH-F2-2010-241909 (project EU-GEI). U.D. received funding from the German Research Foundation (DFG, grant FOR2107 DA1151/5-1; SFB-TRR58, project C09) and the Interdisciplinary Center for Clinical Research (IZKF) of the medical faculty of Münster (grant Dan3/012/17). E.M.B. and N.R.W. received salary funding from the National Health and Medical Research Council (NHMRC; 1078901, 105363). E. Santiago and A.C. received funding from the Agencia Estatal de Investigación (AEI; CGL2016-75904-C2-1-P), Xunta de Galicia (ED431C 2016-037) and Fondo Europeo de Desarrollo Regional (FEDER). The iPSYCH and GEMS2 teams acknowledge funding from the Lundbeck Foundation (grants R102-A9118 and R155-2014-1724), the Stanley Medical Research Institute, an advanced grant from the European Research Council (project 294838), the Danish Strategic Research Council and grants from Aarhus University to the iSEQ and CIRRAU centers.
Case data. We thank the participants and clinicians who took part in the CardiffCOGS study. For the CLOZUK2 sample, we thank Leyden Delta for supporting the sample collection, anonymization and data preparation (particularly M. Helthuis, J. Jansen, K. Jollie and A. Colson), Magna Laboratories, UK (A. Walker) and, for CLOZUK1, Novartis and the Doctor’s Laboratory staff for their guidance and cooperation. We acknowledge L. Bates, C. Bresner and L. Hopkins, at Cardiff University, for laboratory sample management. We acknowledge W. Lawrence and M. Einon, at Cardiff University, for support with the use and setup of computational infrastructures.
Control data. A full list of the investigators who contributed to the generation of the Wellcome Trust Case Control Consortium (WTCCC) data is available from its website. Funding for the project was provided by the Wellcome Trust under award 076113. The UK10K project was funded by Wellcome Trust award WT091310. Venous blood collection for the 1958 Birth Cohort (NCDS) was funded by UK MRC grant G0000934, peripheral blood lymphocyte preparation was funded by the Juvenile Diabetes Research Foundation (JDRF) and the Wellcome Trust, and cell line production, DNA extraction and processing were funded by Wellcome Trust grant 06854/Z/02/Z. Genotyping was supported by the Wellcome Trust (083270) and the European Union (ENGAGE: HEALTH-F4-2007-201413). The UK Blood Services Common Controls (UKBS-CC collection) was funded by the Wellcome Trust (076113/C/04/Z) and by a National Institute for Health Research (NIHR) programme grant to the NHS Blood and Transplant authority (NHSBT; RP-PG-0310-1002). NHSBT also made possible the recruitment of the Cardiff Controls, from participants who provided informed consent. Generation Scotland (GS) received core funding from the Chief Scientist Office of the Scottish government Health Directorates (CZD/16/6) and the Scottish Funding Council (HR03006). Genotyping of the GS:SFHS samples was carried out by the Genetics Core Laboratory at the Wellcome Trust Clinical Research Facility, Edinburgh, Scotland, and was funded by the MRC and Wellcome Trust (grant 10436/Z/14/Z). The Type 1 Diabetes Genetics Consortium (T1DGC; EGA dataset EGAS00000000038) is a collaborative clinical study sponsored by the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK), the National Institute of Allergy and Infectious Diseases (NIAID), the National Human Genome Research Institute (NHGRI), the National Institute of Child Health and Human Development (NICHD) and JDRF. The People of the British Isles project (POBI) is supported by the Wellcome Trust (088262/Z/09/Z). TwinsUK is funded by the Wellcome Trust, MRC, European Union, NIHR-funded BioResource, Clinical Research Facility and Biomedical Research Centre based at Guy’s and St Thomas’ NHS Foundation Trust in partnership with King’s College London. Funding for the QIMR samples was provided by the Australian NHMRC (241944, 339462, 389875, 389891, 389892, 389927, 389938, 442915, 442981, 496675, 496739, 552485, 552498, 613602, 613608, 613674, 619667), the Australian Research Council (FT0991360, FT0991022), the FP-5 GenomEUtwin Project (QLG2-CT-2002-01254) and the US National Institutes of Health (NIH; AA07535, AA10248, AA13320, AA13321, AA13326, AA14041, MH66206, DA12854, DA019951) and the Center for Inherited Disease Research (Baltimore, MD, USA). TEDS is supported by a program grant from the MRC (G0901245-G0500079), with additional support from the NIH (HD044454, HD059215). In the GERAD1 Consortium, Cardiff University was supported by the Wellcome Trust, the MRC, Alzheimer’s Research UK (ARUK) and the Welsh government. King’s College London acknowledges support from the MRC. The University of Belfast acknowledges support from ARUK, the Alzheimer’s Society, Ulster Garden Villages, the Northern Ireland R&D Office and the Royal College of Physicians/Dunhill Medical Trust. Washington University was funded by NIH grants, the Barnes Jewish Foundation, and the Charles and Joanne Knight Alzheimer’s Research Initiative. The Bonn group was supported by the German Federal Ministry of Education and Research (BMBF), Competence Network Dementia and Competence Network Degenerative Dementia and by the Alfried Krupp von Bohlen und Halbach-Stiftung.
Author information
Authors and Affiliations
Consortia
Contributions
A.F.P. curated and processed genetic data, performed statistical analyses, contributed to the interpretation of results and participated in the primary drafting of the manuscript. P.H., A.J.P., V.E.-P., A.C. and E. Santiago performed statistical analyses, contributed to the interpretation of results and participated in the primary drafting of the manuscript. S.R. curated and processed genetic data and participated in the primary drafting of the manuscript. N.C. and M.L.H. contributed to the interpretation of results and participated in the primary drafting of the manuscript. S.E.L., S.B. and A.L. participated in the recruitment of participants for the study and curated and managed their phenotypic information. D.C., J.H., L.H., E.R. and G.K. contributed and curated data used in the statistical analyses. K.M. managed the laboratory and genotyping procedures at Cardiff University. J.H.M., D.A.C. and D.R. supervised the recruitment of the participants for the study. S.A.M. managed the genotyping of samples for the study. N.R.W. contributed genotypes of control samples and participated in the primary drafting of the manuscript. Control data were obtained from the GERAD1 Consortium; as such, the investigators within the GERAD1 Consortium contributed to the design and implementation of GERAD1 and/or provided control data but did not participate in analysis or writing of this report. D.H.G., L.M.H., D.M.R., P.S., E.A.S. and H.W. performed statistical analyses and contributed to the interpretation of results. M.J.O. and M.C.O’D. conceived and supervised the project, contributed to the interpretation of results and participated in the primary drafting of the manuscript. J.T.R.W. conceived and supervised the project, led the recruitment of the participants and sample acquisition for the study, performed statistical analysis, contributed to the interpretation of results and participated in the primary drafting of the manuscript. All other authors contributed genotypes of control samples or summary statistics of replication samples. All authors had the opportunity to review and comment on the manuscript, and all approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.A.C. is a full-time employee and stockholder of Eli Lilly and Company. The remaining authors declare no conflicts of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Note
Supplementary Tables
Supplementary Tables 1–15
Supplementary Data
Gene sets that survive conditional analysis
Rights and permissions
About this article
Cite this article
Pardiñas, A.F., Holmans, P., Pocklington, A.J. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat Genet 50, 381–389 (2018). https://doi.org/10.1038/s41588-018-0059-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0059-2
This article is cited by
-
Genetic effects of sequence-conserved enhancer-like elements on human complex traits
Genome Biology (2024)
-
A preliminary metabolomics study of the database for biological samples of schizophrenia among Chinese ethnic minorities
BMC Psychiatry (2024)
-
Incorporating functional annotation with bilevel continuous shrinkage for polygenic risk prediction
BMC Bioinformatics (2024)
-
Temporal changes of gene expression in health, schizophrenia, bipolar disorder, and major depressive disorder
Schizophrenia (2024)
-
Genetic evidence for causal effects of immune dysfunction in psychiatric disorders: where are we?
Translational Psychiatry (2024)