Abstract
In mammals, all somatic development originates from lineage segregation in early embryos. However, the dynamics of transcriptomes and epigenomes acting in concert with initial cell fate commitment remains poorly characterized. Here we report a comprehensive investigation of transcriptomes and base-resolution methylomes for early lineages in peri- and postimplantation mouse embryos. We found allele-specific and lineage-specific de novo methylation at CG and CH sites that led to differential methylation between embryonic and extraembryonic lineages at promoters of lineage regulators, gene bodies, and DNA-methylation valleys. By using Hi-C experiments to define chromatin architecture across the same developmental period, we demonstrated that both global demethylation and remethylation in early development correlate with chromatin compartments. Dynamic local methylation was evident during gastrulation, which enabled the identification of putative regulatory elements. Finally, we found that de novo methylation patterning does not strictly require implantation. These data reveal dynamic transcriptomes, DNA methylomes, and 3D chromatin landscapes during the earliest stages of mammalian lineage specification.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Rossant, J. & Tam, P. P. Emerging asymmetry and embryonic patterning in early mouse development. Dev. Cell 7, 155–164 (2004).
Zernicka-Goetz, M., Morris, S. A. & Bruce, A. W. Making a firm decision: multifaceted regulation of cell fate in the early mouse embryo. Nat. Rev. Genet. 10, 467–477 (2009).
Rossant, J. & Tam, P. P. Blastocyst lineage formation, early embryonic asymmetries and axis patterning in the mouse. Development 136, 701–713 (2009).
Bielinska, M., Narita, N. & Wilson, D. B. Distinct roles for visceral endoderm during embryonic mouse development. Int. J. Dev. Biol. 43, 183–205 (1999).
Arnold, S. J. & Robertson, E. J. Making a commitment: cell lineage allocation and axis patterning in the early mouse embryo. Nat. Rev. Mol. Cell Biol. 10, 91–103 (2009).
Lawson, K. A., Meneses, J. J. & Pedersen, R. A. Clonal analysis of epiblast fate during germ layer formation in the mouse embryo. Development 113, 891–911 (1991).
Smith, Z. D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Bourc’his, D., Xu, G. L., Lin, C. S., Bollman, B. & Bestor, T. H. Dnmt3L and the establishment of maternal genomic imprints. Science 294, 2536–2539 (2001).
Branco, M. R. et al. Maternal DNA methylation regulates early trophoblast development. Dev. Cell 36, 152–163 (2016).
McGraw, S. et al. Loss of DNMT1o disrupts imprinted X chromosome inactivation and accentuates placental defects in females. PLoS Genet. 9, e1003873 (2013).
Smith, Z. D. et al. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 484, 339–344 (2012).
Wang, L. et al. Programming and inheritance of parental DNA methylomes in mammals. Cell 157, 979–991 (2014).
Guo, H. et al. The DNA methylation landscape of human early embryos. Nature 511, 606–610 (2014).
Smith, Z. D. et al. DNA methylation dynamics of the human preimplantation embryo. Nature 511, 611–615 (2014).
Gao, F. et al. De novo DNA methylation during monkey pre-implantation embryogenesis. Cell Res. 27, 526–539 (2017).
Nagy, A., Gertsenstein, M., Vintersten, K. & Behringer, R. Separating postimplantation germ layers. CSH Protoc. http://dx.doi.org/10.1101/pdb.prot4368 (2006).
Beddington, R. S. P. Isolation, culture and manipulation of post-implantation mouse embryos. In: M. Monk ed. Mammalian Development: A Practical Approach (pp. 43–69. IRL Press, Oxford, UK, 1987).
Kwon, G. S., Viotti, M. & Hadjantonakis, A. K. The endoderm of the mouse embryo arises by dynamic widespread intercalation of embryonic and extraembryonic lineages. Dev. Cell 15, 509–520 (2008).
Peng, X. et al. TELP, a sensitive and versatile library construction method for next-generation sequencing. Nucleic Acids Res. 43, e35 (2015).
Hon, G. C. et al. Epigenetic memory at embryonic enhancers identified in DNA methylation maps from adult mouse tissues. Nat. Genet. 45, 1198–1206 (2013).
Habibi, E. et al. Whole-genome bisulfite sequencing of two distinct interconvertible DNA methylomes of mouse embryonic stem cells. Cell Stem Cell 13, 360–369 (2013).
Hu, Y. G. et al. Regulation of DNA methylation activity through Dnmt3L promoter methylation by Dnmt3 enzymes in embryonic development. Hum. Mol. Genet. 17, 2654–2664 (2008).
He, Y. & Ecker, J. R. Non-CG methylation in the human genome. Annu. Rev. Genomics Hum. Genet. 16, 55–77 (2015).
Pastor, W. A. et al. Naive human pluripotent cells feature a methylation landscape devoid of blastocyst or germline memory. Cell Stem Cell 18, 323–329 (2016).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Jeong, M. et al. Large conserved domains of low DNA methylation maintained by Dnmt3a. Nat. Genet. 46, 17–23 (2014).
Auclair, G., Guibert, S., Bender, A. & Weber, M. Ontogeny of CpG island methylation and specificity of DNMT3 methyltransferases during embryonic development in the mouse. Genome Biol. 15, 545 (2014).
Schroeder, D. I. et al. Early developmental and evolutionary origins of gene body DNA methylation patterns in mammalian placentas. PLoS Genet. 11, e1005442 (2015).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Du, Z. et al. Allelic reprogramming of 3D chromatin architecture during early mammalian development. Nature 547, 232–235 (2017).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Gu, T. P. et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature 477, 606–610 (2011).
Peat, J. R. et al. Genome-wide bisulfite sequencing in zygotes identifies demethylation targets and maps the contribution of TET3 oxidation. Cell Reports 9, 1990–2000 (2014).
Amouroux, R. et al. De novo DNA methylation drives 5hmC accumulation in mouse zygotes. Nat. Cell Biol. 18, 225–233 (2016).
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011).
Burger, L., Gaidatzis, D., Schübeler, D. & Stadler, M. B. Identification of active regulatory regions from DNA methylation data. Nucleic Acids Res. 41, e155 (2013).
Vierstra, J. et al. Mouse regulatory DNA landscapes reveal global principles of cis-regulatory evolution. Science 346, 1007–1012 (2014).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Schöler, H. R., Dressler, G. R., Balling, R., Rohdewohld, H. & Gruss, P. Oct-4: a germline-specific transcription factor mapping to the mouse t-complex. EMBO J. 9, 2185–2195 (1990).
DeVeale, B. et al. Oct4 is required ~E7.5 for proliferation in the primitive streak. PLoS Genet. 9, e1003957 (2013).
Iwafuchi-Doi, M. et al. Transcriptional regulatory networks in epiblast cells and during anterior neural plate development as modeled in epiblast stem cells. Development 139, 3926–3937 (2012).
Ang, S. L. et al. The formation and maintenance of the definitive endoderm lineage in the mouse: involvement of HNF3/forkhead proteins. Development 119, 1301–1315 (1993).
Bossard, P. & Zaret, K. S. GATA transcription factors as potentiators of gut endoderm differentiation. Development 125, 4909–4917 (1998).
Kuo, C. T. et al. GATA4 transcription factor is required for ventral morphogenesis and heart tube formation. Genes Dev. 11, 1048–1060 (1997).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Libbus, B. L. & Hsu, Y. C. Sequential development and tissue organization in whole mouse embryos cultured from blastocyst to early somite stage. Anat. Rec. 197, 317–329 (1980).
Morris, S. A. et al. Dynamics of anterior-posterior axis formation in the developing mouse embryo. Nat. Commun. 3, 673 (2012).
Kalkan, T. & Smith, A. Mapping the route from naive pluripotency to lineage specification. Philos. Trans. R. Soc. Lond. B Biol. Sci. 369, 20130540 (2014).
Baubec, T. et al. Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520, 243–247 (2015).
Berman, B. P. et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina-associated domains. Nat. Genet. 44, 40–46 (2011).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Smith, Z. D. et al. Epigenetic restriction of extraembryonic lineages mirrors the somatic transition to cancer. Nature 549, 543–547 (2017).
Zylicz, J. J. et al. Chromatin dynamics and the role of G9a in gene regulation and enhancer silencing during early mouse development. eLife 4, e09571 (2015).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Solter, D. & Knowles, B. B. Immunosurgery of mouse blastocyst. Proc. Natl. Acad. Sci. USA 72, 5099–5102 (1975).
Ohnishi, Y. et al. Cell-to-cell expression variability followed by signal reinforcement progressively segregates early mouse lineages. Nat. Cell Biol. 16, 27–37 (2014).
Harrison, S. M., Dunwoodie, S. L., Arkell, R. M., Lehrach, H. & Beddington, R. S. Isolation of novel tissue-specific genes from cDNA libraries representing the individual tissue constituents of the gastrulating mouse embryo. Development 121, 2479–2489 (1995).
Libbus, B. L. & Hsu, Y. C. Changes in S-phase associated with differentiation of mouse embryos in culture from blastocyst to early somite stage. Anat. Embryol. (Berl.) 159, 235–244 (1980).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Guo, W. et al. BS-Seeker2: a versatile aligning pipeline for bisulfite sequencing data. BMC Genomics 14, 774 (2013).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Akdemir, K. C. & Chin, L. HiCPlotter integrates genomic data with interaction matrices. Genome Biol. 16, 198 (2015).
de Hoon, M. J., Imoto, S., Nolan, J. & Miyano, S. Open source clustering software. Bioinformatics 20, 1453–1454 (2004).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Huang, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Bell, R. E. et al. Enhancer methylation dynamics contribute to cancer plasticity and patient mortality. Genome Res. 26, 601–611 (2016).
Acknowledgements
We are grateful to members of the Xie laboratory for helpful comments during preparation of the manuscript. We thank J. Na for critical reading of the manuscript. This work was supported by the National Key R&D Program of China (2016YFC0900301 to W. Xie; 2017YFC1001401 to L.L.), the National Basic Research Program of China (2015CB856201 to W. Xie), the National Natural Science Foundation of China (31422031 to W. Xie), the THU-PKU Center for Life Sciences (W. Xie), Beijing Advanced Innovation Center for Structural Biology (W. Xie), and the Biomedical Research Council of A*STAR (Agency for Science, Technology and Research), Singapore (F.X.). W. Xie is a Howard Hughes Medical Institute (HHMI) International Research Scholar. J.W. was funded by grants from the NIH (R01GM095942 and R21HD087722) and the Empire State Stem Cell Fund through the New York State Department of Health (NYSTEM) (C028103 and C028121), and is a recipient of an Irma T. Hirschl and Weill-Caulier Trusts Career Scientist Award.
Author information
Authors and Affiliations
Contributions
Y.Z. and Q.Y. developed and conducted STEM-seq experiments. Y.X. dissected mouse tissues from embryos in vivo, carried out in vitro culture of embryos, and conducted RNA-seq. Z.D. conducted Hi-C experiments and related analysis. Z.Z. and L.L. advised on embryo lineage dissection. X.P. and F.X. advised on the development of STEM-seq. Y.L. and Q.W. conducted high-throughput sequencing. Y.Z. and Y.X. carried out data analysis. Q.W., W.Z., and W. Xia helped with the generation of Tet1/2 double-knockout mice. J.M., M.F., and J.W. helped with various experiments and/or advised the project. Y.Z. and W. Xie wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
A patent for STEM-seq has been filed (2014104662612 China and PCT/CN2015/088680).
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated Supplementary Information
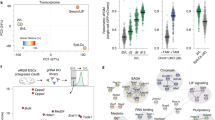
Supplementary Figure 1 Transcriptome profiling for early lineages during peri- and postimplantation development.
a) Schematic showing tissue dissection of early embryos from E3.5 to E7.5 (see Methods). b) The correlation of gene expression levels across the genome between biological replicates for RNA-seq samples generated in this study. c) The expression of various lineage marker genes is shown for dissected tissues at each developmental stage as determined by RNA-seq. Error bars denote the standard deviation of FPKM values from biological replicates. d) Heatmap showing the expression levels of various marker genes in each tissue isolated from E3.5 to E7.5 embryos based on RNA-seq. Due to wide distributions of gene expression for different genes, each gene expression was relatively normalized by setting the FPKM of the highest lineage as 10. e) Hierarchical clustering analysis of global gene expression levels (with replicates) based on the RNA-seq data for tissues isolated from early embryos
Supplementary Figure 2 Development and validation of STEM-seq.
a Schematic of STEM-seq procedure. Genomic DNA (or lysed cells) is first subjected to bisulfite conversion, followed by sequencing library preparation using TELP, a highly sensitive single-strand DNA amplification and library preparation method. Briefly, the purified converted DNA is tailed by poly C, followed by extension with a biotinylated poly G containing primer. The extension product is ligated to an adaptor followed by PCR amplification for sequencing library preparation (See Methods for details). b A UCSC genome browser snapshot shows comparison of mESC methylomes determined by STEM-seq and MethylC-seq using various amounts of DNA or cells in two mESC lines (TT2 and R1) near the Hoxa gene cluster. CpG islands (CGIs) in this region are also shown. c Scatterplots show the comparison of mESC methylation levels between those determined by STEM-seq and MethylC-seq (2kb bin), or between replicates of STEM-seq data across the whole genome. Pearson’s correlation coefficients are also shown. d Comparison of average CG methylation levels in mESCs at different genomic elements between those determined by STEM-seq using 10ng or 100ng DNA and MethylC-seq. e A similar plot as d for STEM-seq using 500 mESCs with two replicates. Please note that different mESC lines were unintentionally used in d (TT2) and e (R1)
Supplementary Figure 3 The landscapes of DNA methylomes in early lineages.
a Scatterplots comparing biological replicates (2kb bin, across the whole genome) for lineage methylomes. Dashed lines indicate mCG difference = 0.2. b The plot showing the percentages of CG sites covered by various numbers of STEM-seq reads for each lineage. c The percentages of CG sites (≥5x) across various types of genomic elements for methylome datasets in this study. d The promoter methylation and gene expression levels for Hnf4a (VE marker) and Oct4, Tdgf1 (epiblast markers). e The promoter methylation and gene expression levels for E3.5 ICM and E3.5 TE specific expressed genes. f The dynamics of average DNA methylation levels across different classes of genomic elements for lineages from E3.5 to E7.5. g Barcharts showing the expression levels of Dnmts in early embryos from E3.5 to E7.5. h The promoter methylation and expression levels for Dnmt3l in development are shown
Supplementary Figure 4 Allelic DNA-methylome reprogramming during early lineage specification.
a Barcharts showing the percentages of reads that were assigned to the maternal or the paternal genome in each tissue. Only reads that contain SNPs were counted. b Heatmaps showing allelic DNA methylation levels at imprinting control regions. Only regions that are covered by sufficient SNPs were included for analysis. Gray (marked by asterisks) indicates stages with no or insufficient allelic reads
Supplementary Figure 5 Dynamic DNA methylation at promoters and DNA methylation valleys during lineage specification.
a Heatmap showing the promoter methylation (left) and expression (right) levels for genes that are differentially methylated between E6.5 VE and E6.5 Epi but are silenced in both lineages. b The GO analysis result for all genes located in DMVs identified in E6.5 Epi. c Barcharts showing the percentages of DMVs identified in early embryo (combining five lineages) that are marked by H3K27me3 (using data from a panel of somatic cells). d Barcharts showing the enrichment (logratio of observed/expected) of E5.5 Epi hypermethylated regions (vs. E6.5 Epi) in various classes of genomic elements. A set of random regions with equal lengths of individual hypermethylated regions were used as controls. e The boxplot showing the methylation levels in all CGIs and the non-CGI regions in DMVs in E5.5 and E6.5 embryos. f Barcharts showing the percentages of hypermethylated CGIs in E6.5 VE that fall into DMVs identified in early embryos (combining five lineages). The percentage of all CGIs that are located in DMVs (background) is also shown. g The barplot shows the expression of all Tet genes in early development
Supplementary Figure 6 Hi-C analysis for early lineages during peri- and postimplantation development.
a The Directionality Index (DI) tracks for E3.5 ICM, E6.5 epiblast, E6.5 VE and E7.5 ectoderm are shown. b The scatterplots comparing the DI values between E6.5 epiblast and other lineages, or between E6.5 epiblast replicates. The correlation coefficients (Spearman) are also shown. c The P(s) curves (chromatin contract frequency vs. genomic distances) for mESC and early lineages are shown
Supplementary Figure 7 Both de novo methylation and demethylation are correlated with chromatin compartments.
a Barcharts showing the average methylation levels gained from E4.0 ICM to E5.5 Epi or E5.5 VE (left) in active gene bodies (+), inactive gene bodies (-), intergenic regions (i), and the whole compartment (w), for either compartment A or compartment B (left). b A chromosome-wide view of CH methylation levels (1Mb bin) is shown for E5.5 Epi and E5.5 VE (top). Chromatin compartments in E3.5 ICM, E6.5 Epi, E6.5 VE and gene-dense regions are also shown (bottom). Arrows indicate hypomethylated regions that overlap with compartment B. c Scatterplots comparing CG and CH methylation levels (1Mb bin) between epiblast and VE at E5.5 and E6.5. The Pearson correlation coefficients are indicated. d The average allelic methylation levels near active (green) or silenced (black) genes for E3.5 ICM are shown, before (top) or after (bottom) TAD-based normalization (subtracting TAD background methylation levels for each gene). The background methylation level for each TAD was calculated by averaging DNA methylation levels across the TAD (excluding gene bodies and regulatory elements such as promoters and putative enhancers). e Barcharts showing the average methylation differences between wild type and Tet3 knockout zygotes in active gene bodies, inactive gene bodies, intergenic regions and the whole compartment for either compartment A or B. f A chromosome-wide view of DNA methylation levels (1Mb bin) is shown for mESC (serum and 2i). Chromatin compartments and gene-dense regions are also shown
Supplementary Figure 8 Dynamic DNA methylation at UMRs/LMRs during early lineage commitment.
a Heatmap showing the pairwise overlap of UMRs or LMRs among individual lineages. Tissues with global hypomethylation were excluded from the analysis. The percentages of UMRs that overlap with annotated TSSs are also shown. b Venn diagram showing the overlap of identified LMRs/UMRs in E6.5 Epi with DHS sites in mESC identified by ENCODE. c Heatmap showing the overlap of tissue-specific LMRs (tsLMRs) and putative enhancers previously defined in various tissues using histone modification signatures. The enrichment was calculated as logratio of observed overlap divided by expected overlap using a random set of regions with equal lengths of individual tsLMRs. d Heatmap showing the average methylation levels of early embryo-specific UMRs/LMRs (left) and associated gene expression (right) between early embryonic tissues (average of E6.5 Epi, Ect, PS, Mes, End) and somatic tissues (average of 11 tissues). Representative genes associated with UMRs/LMRs are shown on the right. e The snapshot showing methylation levels near Dnmt3b in oocyte, early embryos, somatic tissues, and mESCs (left). The expression for Dnmt3b in each cell type is also shown as heatmap (right). The shade indicates early embryo specific UMRs/LMRs. The DNaseI hypersensitive sites in mESCs are also shown
Supplementary Figure 9 Lineage-specific gene expression in embryos cultured in vitro.
a The embryos were collected at E4.0 and were cultured in vitro for 4 days. Epiblast-like and VE-like tissues were dissected from in vitro cultured (IVC) embryos. b Barcharts showing the expression of marker genes for epiblast, VE/endoderm, ectoderm, PS, and mesoderm for lineages isolated from in vivo (red) and in vitro cultured (IVC) embryos (blue). As some germ layer markers are also expressed at earlier stages including epiblast (in vivo), only those that are exclusively expressed during gastrulation were examined. c Hierarchical clustering analysis of RNA-seq data for tissues isolated from in vivo embryos and IVC embryos. d Barcharts showing the expression levels of Dnmts in tissues isolated from IVC embryos and in vivo embryos
Supplementary Figure 10 Lineage-specific de novo methylation in embryos cultured in vitro.
a Chromosome-wide view of CG methylation for tissues isolated from in vivo E6.5 embryos (red) and IVC embryos (blue, replicate 1). The second replicate of IVC embryos showed similar patterns (data not shown). b The average methylation levels near active (green, FPKM≥10) or silenced (black, FPKM≤1) genes for IVC epiblast and VE (replicate 1). Similar observations were made for replicate 2 (data not shown). c The enrichment (logratio of observed/expected) of various types of genomic elements for regions hypermethylated in IVC epiblast compared to E5.5 or E6.5 epiblast in the genome. d A model of allele and lineage-specific DNA methylation reprogramming in mouse early embryos. After fertilization, the maternal allele inherits gene body DNA methylation pattern from oocyte to blastocyst. Gene body methylation then occurs in postimplantation embryos on both alleles during de novo methylation. Such pattern is retained in extraembryonic tissues but is gradually diminished in embryonic tissues. On the other hand, the paternal allele in preimplantation embryos undergoes mega-base chromatin compartment A-specific demethylation. During de novo methylation, VE preferentially gains DNA methylation in compartment A while epiblast shows even DNA methylation in both compartments. The differences between epiblast and VE are likely in part contributed by the differential expression of Dnmts
Supplementary information
Supplementary Figures
Supplementary Figures 1–10.
Supplementary Table 1
Lists of lineage-specific genes from E3.5 to E7.5.
Supplementary Table 2
Sequencing summary.
Supplementary Table 3
Differentially methylated genes.
Supplementary Table 4
Lists of all UMRs and LMRs.
Supplementary Table 5
Early-embryo-specific UMRs and LMRs.
Rights and permissions
About this article
Cite this article
Zhang, Y., Xiang, Y., Yin, Q. et al. Dynamic epigenomic landscapes during early lineage specification in mouse embryos. Nat Genet 50, 96–105 (2018). https://doi.org/10.1038/s41588-017-0003-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-017-0003-x
This article is cited by
-
Integrative analysis of transcriptomic and epigenomic data reveals distinct patterns for developmental and housekeeping gene regulation
BMC Biology (2024)
-
MiR-124-3p negatively impacts embryo implantation via suppressing uterine receptivity formation and embryo development
Reproductive Biology and Endocrinology (2024)
-
DNA methylation restricts coordinated germline and neural fates in embryonic stem cell differentiation
Nature Structural & Molecular Biology (2024)
-
The impact of DNA methylation on CTCF-mediated 3D genome organization
Nature Structural & Molecular Biology (2024)
-
3D Enhancer–promoter networks provide predictive features for gene expression and coregulation in early embryonic lineages
Nature Structural & Molecular Biology (2024)