Abstract
Malformations of cortical development (MCDs) are neurodevelopmental disorders that result from abnormal development of the cerebral cortex in utero. MCDs place a substantial burden on affected individuals, their families and societies worldwide, as these individuals can experience lifelong drug-resistant epilepsy, cerebral palsy, feeding difficulties, intellectual disability and other neurological and behavioural anomalies. The diagnostic pathway for MCDs is complex owing to wide variations in presentation and aetiology, thereby hampering timely and adequate management. In this article, the international MCD network Neuro-MIG provides consensus recommendations to aid both expert and non-expert clinicians in the diagnostic work-up of MCDs with the aim of improving patient management worldwide. We reviewed the literature on clinical presentation, aetiology and diagnostic approaches for the main MCD subtypes and collected data on current practices and recommendations from clinicians and diagnostic laboratories within Neuro-MIG. We reached consensus by 42 professionals from 20 countries, using expert discussions and a Delphi consensus process. We present a diagnostic workflow that can be applied to any individual with MCD and a comprehensive list of MCD-related genes with their associated phenotypes. The workflow is designed to maximize the diagnostic yield and increase the number of patients receiving personalized care and counselling on prognosis and recurrence risk.
Similar content being viewed by others
Introduction
Abnormal formation of the cerebral cortex in utero leads to neurodevelopmental disorders known as malformations of cortical development (MCDs). Although individually rare, as a group MCDs represent a major cause of intellectual disability, autism, epilepsy and cerebral palsy1,2. The last update of the developmental and genetic classification for MCDs, which was published in 2012, includes 200 clinical entities and classifies them into three major groups: malformations secondary to abnormal neuronal and glial cell proliferation and apoptosis, including microcephaly and macrocephaly; neuronal migration disorders, represented by heterotopia, lissencephaly and cobblestone malformation (COB); and malformations of postmigrational cortical organization and connectivity, represented by conditions such as polymicrogyria, schizencephaly and focal cortical dysplasia (FCD)3.
Many MCDs are caused by an underlying genetic defect. Rapid advances in molecular genetics and neuroimaging techniques in recent years have substantially increased the number of recognized MCD forms and their associated genes, and have highlighted the considerable genetic heterogeneity associated with these disorders1. Next-generation sequencing (NGS) of a selection of genes related to a phenotype (gene panel), the coding exons of the human genes (exome sequencing) or the genome of an individual (genome sequencing) has enabled rapid sequencing of large numbers of genes.
Even following intensive diagnostic assessments, many individuals with an MCD remain without a molecular diagnosis4,5,6. The complex nature and high degree of clinical and genetic heterogeneity of MCDs demand highly specialized and multidisciplinary expertise. However, MCD experts usually work individually or in small multidisciplinary teams. Currently, comprehensive guidelines for diagnosis and management are lacking, adding to the variability in the diagnostic approach between different centres. The disease course and long-term clinical outcome are often difficult to predict at an early stage, and medical management is rarely evidence-based. These challenges highlight the need for an expert-driven multidisciplinary effort to better understand these disorders. The availability of carefully curated MCD gene panels to the wider medical community will enable accurate molecular diagnosis in a larger number of patients without long delays or unnecessary investigations.
We established the international multidisciplinary network Neuro-MIG with the aim of disseminating knowledge to the broad medical community, improving the diagnosis and management of MCDs and accelerating research into MCDs7. In this article, we first review the clinical presentation and aetiology of the main MCD types. On the basis of a critical review of the literature, expert surveys and discussions, we then present a consensus statement on the clinical and molecular investigations in patients with MCDs, including specific recommendations on clinical work-up, molecular diagnostic methods and alternative strategies in undiagnosed patients.
Methods
This article represents a consensus document based on three face-to-face expert meetings within the Neuro-MIG network that were held in St Julians, Malta, from 21 to 23 February 2018, in Lisbon, Portugal, on 13 and 14 September 2018, and in Rehovot, Israel, on 17 March 2019. The meetings were funded by the European Cooperation in Science & Technology (COST Action CA16118). Two Neuro-MIG working groups, WG1 and WG3, took the lead in preparing the draft, although a larger group within the network was invited to participate in the Delphi consensus procedure and comment on the second draft. The final version of the consensus document was reviewed by the drafting team and circulated among all COST network members before submission.
PubMed was systematically queried for phenotypes, genes and mutation rates associated with MCDs, using the key words “microcephaly”, “megalencephaly”, “lissencephaly”, “polymicrogyria”, “schizencephaly”, “cobblestone malformation”, “focal cortical dysplasia” and “heterotopia”. The most recent search was performed on 31 October 2019.
From the MCD expert laboratories within the Neuro-MIG network, headed by M.W., K.S., U.H., E.P. and N.D.D., we collected data regarding gene panels, enrichment strategies and diagnostic yield. Using the data obtained as described above, we compiled lists of genes associated with the various MCD subtypes and defined a diagnostic strategy for patients with MCDs. The gene list was curated — that is, checked, corrected and completed — by all authors on the basis of long-standing personal experience gained through molecular diagnostics in patients with MCDs. The first draft was finalized before the second meeting. During the first round of voting, 21 of the authors voted on 101 recommendation statements. Agreement (>90% positive votes) was reached for 89 statements, and the remaining 12 were revised according to the reasons provided for disagreement. The second round of voting involved 42 experts. At the end of the process, 94 recommendations found >90% consensus. In addition, five statements were agreed on by 80–90%, two statements by 75–80% and one statement by 70–75% of the participants (Supplementary Table 1). Recommendations with consensus <80% were excluded from the recommendations section below. Unless specified otherwise, we report on recommendation statements with >90% consensus.
Clinical presentation of MCDs
MCDs can be isolated or associated with a wide variety of neurological and extra-neurological features, including other birth defects and facial dysmorphism. The age at clinical referral and the severity of neurological deficits vary substantially between affected individuals. The most common presenting features are epilepsy, developmental delay and/or motor abnormalities of tone, movement and posture1. These features are listed in relation to the typical ages of presentation in Box 1.
Main MCD types
In this section, we provide an overview of the most common types of MCD and their aetiologies. Different descriptions have been introduced in the literature over the years depending on the study design and the medical background of the research group (for example, neurologists, radiologists, geneticists or pathologists). Table 1 summarizes the consensus definitions that were agreed on by our working group. These definitions are used throughout the text, and brain imaging examples are provided in Fig. 1. The descriptions are specific to each term and do not consider the presence of abnormalities of other brain structures, which often coexist with MCD. Each MCD type can be further classified on the basis of morphology, topography, severity gradient and involvement of other brain structures1. A detailed paper on the MCD neuroimaging features has been published separately by representatives from the Neuro-MIG network8.
a | Normal brain on T1-weighted images. b | Normal brain on T2-weighted images. c | Primary microcephaly with a small brain. d | Abnormally large brain (megalencephaly) with abnormal appearance of the perisylvian cortex (arrows point to small gyri suggestive of polymicrogyria). e | Bilateral nodular heterotopia (arrows) situated along the ventricular walls. f | Lissencephaly spectrum with agyria–severe pachygyria (arrows). g | Lissencephaly spectrum with subcortical band heterotopia visible as a thick band isointense to the cortex (asterisks). h | Generalized thickened cortex with broad gyri and white matter abnormalities consistent with cobblestone complex (arrows). i | Bilateral frontoparietal polymicrogyria with abnormally small gyri and shallow sulci (arrows). j | Coronal scan showing schizencephaly, characterized by a cleft lined by grey matter extending from the cortex to the ventricle (arrow). k | Abnormally oriented sulci of varying depth with normal cortical thickness (arrows). l | Focal cortical dysplasia with blurring of the grey–white matter boundary and hyperintensity of the white matter on T2-weighted imaging (arrow).
Microcephaly
Microcephaly is defined as a significant reduction in the occipitofrontal circumference (OFC) compared with controls matched for age and sex. Microcephaly is the most common MCD and is present in 15% of children referred for evaluation of developmental disabilities9. The relevant degree of reduction differs throughout the literature, being set at 2–3 s.d. below the mean9,10,11,12. Strictly speaking, microcephaly is a clinical finding rather than a disease; however, it provides a reliable estimation of the brain volume10. The final brain size is the result of a complex process of neural stem cell proliferation, migration, and ongoing organization, synaptogenesis and apoptosis11. Microcephaly is classed as congenital if present at birth (primary microcephaly) or postnatal if it develops after birth (secondary microcephaly)10,13,14. These two groups also have different molecular aetiologies11. Microcephaly can present with a normal or simplified gyral pattern, or with additional, more complex brain abnormalities11. The clinical outcome cannot be predicted by head size alone and largely depends on the underlying cause and the appearance of the brain on MRI.
Macrocephaly and megalencephaly
Macrocephaly is defined as an OFC ≥2 s.d. above the mean, whereas megalencephaly refers to an abnormally large brain size1. Macrocephaly has a wide variety of causes besides megalencephaly, including hydrocephalus and increased skull thickness. Mild megalencephaly (2–3 s.d. above the mean) with an otherwise structurally normal brain can be seen in typically developing children, often in the setting of benign familial macrocephaly15. However, megalencephaly can point to an underlying neurodevelopmental or generalized overgrowth disorder.
Periventricular nodular heterotopia
The term neuronal heterotopia refers to groups of neurons in an abnormal location, and periventricular nodular heterotopia (PVNH) describes nodular masses of grey matter located along the ventricular walls protruding into the ventricle1. PVNH can occur in isolation or together with other brain or body malformations and is not rare: in one study, PVNH was observed in 0.48% of the general paediatric population16. The nodules can occur unilaterally or bilaterally, and should be further defined according to their number and location (for example, involving the frontal or temporal and/or occipital horns of the lateral ventricles).
PVNH is associated with numerous different copy number variations (CNVs) and single gene variants, and can be part of a complex syndromic disorder.
Lissencephaly spectrum
The lissencephaly spectrum encompasses agyria, pachygyria and subcortical band heterotopia (SBH)17. Agyria and pachygyria are characterized by an abnormal gyral pattern with absent gyri (agyria) or broad gyri (pachygyria) in combination with an abnormally thick cortex18. SBH describes a band of grey matter separated from the cortex and lateral ventricles by zones of white matter18. In rare cases, pachygyria and SBH can co-occur in the same brain, with a typical pattern of frontal pachygyria and posterior SBH19. Microlissencephaly represents a separate subgroup and is defined as a combination of lissencephaly (usually in the form of agyria or pachygyria) with severe congenital microcephaly (OFC at birth ≥3 s.d. below the mean)20.
Subcortical heterotopia
Subcortical heterotopia (SUBH) refers to brain malformations with clusters of neurons located within the white matter, between the cortex and lateral ventricles21. The well-recognized and aforementioned PVNH and SBH have distinct imaging patterns and are classified separately. Multiple terms have been used to describe this type of malformation, including giant, curvilinear, nodular, focal and massive heterotopias21. In 2019, a group within the Neuro-MIG network provided the first framework for an imaging classification of SUBH that encompasses five groups further subdivided into specific entities21.
Cobblestone malformation
COB is recognized as an undersulcated, irregular and ‘pebbled’ cerebral surface, with a moderately thick cortex22,23. This malformation is caused by defects of the pial limiting membrane with resulting neuronal overmigration from the cortical plate into the leptomeninges3,24. COB often co-occurs with eye, muscle and additional brain malformations within the spectrum of the α-dystroglycanopathies, with Walker–Warburg syndrome at the most severe end25.
COB was originally described as lissencephaly type 2 but this term has now been abandoned26. In addition, COB is often confused with polymicrogyria27. The strict differentiation of COB-related and polymicrogyria-related genes in the literature remains difficult, as several conditions characterized by COB were reported as polymicrogyria-associated disorders (for example, GPR56-associated frontoparietal ‘polymicrogyria’ and CHIME syndrome).
Polymicrogyria
Polymicrogyria is one of the most frequent types of MCD and is also one of the most heterogeneous in aetiology1. Polymicrogyria is defined as an excessive number of abnormally small cerebral gyri with cortical overfolding, an irregular, pebbled cortical surface and a stippled grey–white matter boundary28.
As highlighted in the previous section, polymicrogyria can be difficult to differentiate from COB, and might also be confused with dysgyria or pachygyria. High-resolution imaging can aid the differentiation of these conditions, as it can show microgyri, microsulci and stippling of the grey–white matter junction — a specific feature of polymicrogyria that is not seen in other MCDs1. Of note, the Sylvian fissures, which are best viewed on sagittal imaging, should be closely scrutinized as polymicrogyria often affects these areas preferentially, with abnormal posterior extension and sulcal branching being observed28. Polymicrogyria is frequently seen in association with many other brain malformations and is sporadically described in various syndromic disorders. Polymicrogyria has been classified into six topographic patterns that are further divided into 13 morphological subtypes28. Moreover, at least six polymicrogyria syndromes have been defined on the basis of radiological and clinical features29.
Dysgyria
Dysgyria translates as abnormal gyration and can therefore be applied to almost every type of MCD. However, this term was introduced to describe cortical malformations that do not meet classic features of any of the abovementioned well-established MCD types. Dysgyria describes a cortex of variable thickness and an abnormal gyral pattern characterized by abnormalities of sulcal depth or orientation (for example, obliquely oriented sulci directed radially towards the centre of the cerebrum and narrow gyri separated by abnormally deep or shallow sulci)30,31. In the vast majority of cases, the term dysgyria describes an abnormal non-lissencephaly, non-polymicrogyria cortex within the spectrum of tubulinopathies.
FCD and hemimegalencephaly
FCD is identified on brain imaging by focal irregularities of cortical morphology and thickness, blurring of the grey–white matter boundary, and white matter T2 hyperintensity. Depending on the size of the lesion and the resolution of the brain imaging, FCD can be missed on MRI. Smaller lesions are often only identified on neuropathological studies after surgery for epilepsy. FCD type II is characterized by the presence of dysplastic, megalocytic neurons, a feature that is also present in hemimegalencephaly. Balloon cells are also observed in FCD IIB and hemimegalencephaly32. The size of the lesion varies from submicroscopic involvement of one or several sulci (FCD) to a larger area involving a lobe (partial hemimegalencephaly) or involvement of an entire cerebral hemisphere (classic hemimegalencephaly)32. In the latter condition, the affected hemisphere is visibly enlarged. In hemimegalencephaly, the lesion can extend to non-brain tissue, and clinicians should look out for skin abnormalities and localized overgrowth of one or several body parts.
Molecular testing: current practice
Chromosomal testing
MCDs have been linked to a wide range of CNVs, as detected by chromosomal microarray analysis (CMA)1,33,34. Several CNVs are consistently associated with MCD, the most common of which are the 22q11 and 1p36 deletions associated with polymicrogyria, the 17p13.3 deletion (encompassing LIS1 (also known as PAFAH1B1), YWHAE and other genes) that causes Miller–Dieker syndrome and isolated lissencephaly, and 6qter deletions associated with various brain malformations including polymicrogyria and PVNH33,35,36. A study published in 2019 reported a diagnostic yield of 36% when CMA was used in patients who had PVNH with or without other malformations, and 9% in a group with polymicrogyria only37. Another study did not show an increased burden of rare CNVs in people with polymicrogyria compared with healthy controls38. In patients with microcephaly, the yield was ~5–7%13,39. In a large cohort of patients with lissencephaly (n = 811), Miller–Dieker syndrome was diagnosed in 9% of cases40. Several MCD-related genes frequently harbour intragenic deletions or duplications, which might be identified by standard microarrays41,42,43.
Single gene testing
Single gene testing is being superseded by NGS gene panels, and we were only able to identify systematic studies for a small number of MCD types. The yield of single gene testing varies greatly depending on the MCD type and extension of the malformation. For SBH, the yield of molecular testing is high, with pathogenic variants in DCX or LIS1 being found in 79% of patients (123 of 155)40. Pathogenic variants in FLNA are important aetiological factors for PVNH. The highest frequency is found in women with bilateral frontocentral PVNH, especially in combination with cerebellar hypoplasia and/or mega cisterna magna, with a positive family history of PVNH44,45. The yield varies from 80–100% in female familial cases to 9–26% in sporadic cases44,45,46.
In a cohort of 113 patients with MCDs, a molecular diagnosis was established in 21 patients (19%) by targeted testing of one or more genes selected on the basis of the phenotype4. In a more recent study consisting of an Argentinian cohort of 38 patients with lissencephaly, SBH or PVNH, pathogenic variants were identified in 36% of cases46.
Pathogenic variants of ASPM are the most common genetic cause of primary microcephaly, with a mutation rate of 10–40% depending on ethnicity and the presence or absence of consanguinity47,48. Among consanguineous families, alterations in ASPM and WDR62 accounted for >50% of cases of primary microcephaly49,50.
For COB, mutation detection rates vary considerably, depending on the age at diagnosis and clinical inclusion criteria. For the most severe prenatal manifestations, the detection rate was usually >60% when the six genes most commonly linked to dystroglycanopathy were analysed25.
Gene panels
Despite multiple publications reporting on the yield of gene panels in cohorts of patients with neurodevelopmental disorders51,52,53, similar studies for MCDs are scarce. The only study that we identified reported on testing of a small gene panel (ten genes) in 158 individuals with brain malformations, including 30 individuals with SBH, 20 with megalencephaly, 61 with PVNH and 47 with pachygyria. Causal pathogenic variants were found in 27 individuals (17%, range 10–30% depending on the phenotype)54.
Several genes encoding components of the PI3K–AKT–mTOR pathway have been implicated in FCD, and targeted testing of PI3K–AKT–mTOR pathway genes, using highly sensitive sequencing methods that allowed detection of low-frequency brain somatic variants, produced diagnostic yields ranging from 12% to 40%55,56,57. In a different cohort, a targeted NGS panel that included the most commonly mutated PI3K–AKT–mTOR pathway genes uncovered PIK3CA pathogenic variants in 50 of 131 individuals (174 samples) with the megalencephaly–capillary malformation syndrome58.
Exome sequencing
One study investigated the yield of exome sequencing, combined with CMA, in 54 patients with various MCD types5. This approach yielded a definitive (9/16) or presumptive (7/16) molecular diagnosis in 16 of 54 enrolled individuals (30%). Another study of 62 patients with microcephaly followed a similar approach and identified causative variants in 48% of the individuals39.
Neuro-MIG laboratories
We have also analysed the yield from the diagnostic laboratories within the Neuro-MIG network. Targeted gene panels resulted in diagnostic yields of 15–37%, although wide variability was observed among the different clinical subtypes (Table 2). The combination of expert evaluation of MRI scans followed by targeted analysis of the most plausible causative variants can considerably increase the diagnostic yield. Substantiating this point, the availability of MRI scans resulted in an improved mutation detection rate of 37% in a mixed cohort of 117 patients with MCDs, compared with only 18% in a cohort of 784 patients analysed without previous expert re-evaluation of MRI scans at the Human Genetics Center Regensburg (U.H., unpublished work). In the former cohort, the testing strategy was selected by the laboratory depending on the MRI pattern, and the approaches included single gene, panel and exome sequencing. A similar trend was noted in the Department of Clinical Genetics, Erasmus MC University Medical Center, Rotterdam, where the diagnostic yields from in-house requests accompanied by expert MRI review by G.M.S.M. were almost double those from the tests ordered from other medical specialists outside the university hospital (M.W., unpublished work).
In utero infections
Prenatal infections can cause extensive damage to the fetal brain, including the cerebral cortex59,60,61. Cytomegalovirus (CMV) is one of the most frequent non-genetic causes of MCDs and is specifically associated with polymicrogyria, intracranial calcifications, white matter abnormalities and microcephaly1. In a cohort of 26 patients with bilateral polymicrogyria, six (31%) tested positive for CMV; however, it was unclear whether these patients were infected prenatally or postnatally62. In a larger group of 50 patients with polymicrogyria, six (12%) tested positive on Guthrie cards (W.B.D., unpublished work).
In one study of 41 newborn babies with symptomatic CMV, eight (19.5%) presented with microcephaly63. Not all CMV-infected individuals are symptomatic at birth, and neurological sequelae can develop later in life64. Other infectious agents, including rubella virus65, varicella zoster66 and herpes simplex virus61,67, can also cause microcephaly. In recent years, Zika virus has been associated with primary microcephaly and a spectrum of brain malformations68,69,70,71,72,73,74.
New recommendations
The Neuro-MIG network recommends that a concerted effort be made to reach an aetiological diagnosis in every individual with an MCD. The diagnosis serves several functions. First, it explains the cause of the malformation, ends the diagnostic odyssey and prevents further unnecessary investigations. Second, it provides information on prognosis and recurrence risk for the patient and family members4. Third, it aids the prediction of treatment outcomes; for example, the success rate for epilepsy surgery depends on the underlying genetic cause75. Fourth, it directs patient management (for example, antiviral treatment and screening for progressive hearing loss in infants with congenital CMV infection76, cardiovascular surveillance in FLNA-related and ARFGEF2-related PNVH77,78 or mTORC1 inhibition in patients with tuberous sclerosis complex (TSC))79. Fifth, it enables natural history studies80,81 and targeted research into personalized therapy and prevention82,83.
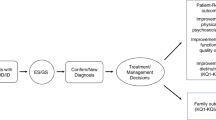
Imaging findings, such as generalized versus focal and bilateral versus unilateral malformations, cannot reliably distinguish genetic from non-genetic causes, and the diagnostic yield of targeted testing is determined to a large extent by the availability of a multidisciplinary expert evaluation. However, such an ideal setting can rarely be met in practice. Therefore, we have formulated a general diagnostic workflow that can be applied in most clinics to any individual with an MCD (Fig. 2). Lists of currently known MCD-associated genes are presented in Supplementary Tables 2 and 3. These lists can assist variant interpretation and guide targeted testing if exome (or genome) sequencing is not available. These general recommendations should minimize the chance of missing a known causative variant. The workflow can be started when a person is first diagnosed with an MCD, although clinicians should check whether any of the investigations have already been performed.
This step-by-step diagnostic approach was formulated by Neuro-MIG. The main diagnostic steps are in purple-lined boxes. aSeek expert review. bIncluding prenatal and perinatal history. cIncludes construction of a pedigree and enquiry for consanguinity. dBased on additional features (for example, sick infant, abnormal liver function tests, retinal scarring or hearing loss), perinatal history (for example, maternal rash or fever) and/or imaging abnormalities (for example, calcifications, white matter injury or cysts). eOffer genetic counselling and segregation analysis to the patient and family members. fAffected brain tissue (if available), fibroblasts or saliva. CMV, cytomegalovirus; GOUS, gene of uncertain significance; MCD, malformation of cortical development; NGS, next-generation sequencing; OFC, occipitofrontal circumference, VOUS, variant of uncertain significance.
For some MCD subtypes, the most cost-effective strategy would be targeted gene analysis, but the success of this approach depends greatly on accurate pattern recognition. The relevant subtype-specific patterns and aetiologies are outlined in the section ‘Phenotype-specific considerations’ below.
The correct interpretation of genetic test results requires detailed phenotypic analysis, including re-evaluation of the brain MRI, to confirm that the identified single nucleotide variant (SNV) or CNV fully explains the phenotype. In the case of a negative result, the re-evaluation should help determine whether the malformation was correctly classified, whether additional diagnostic testing, such as deep sequencing or analysis of a different tissue, might be helpful, and whether a non-genetic cause is more likely.
We recommend that a final clinical interpretation is done by a qualified medical geneticist, preferably after an interdisciplinary discussion with a molecular geneticist, neuroradiologist and/or neurologist. Unusual cases can be presented at an expert review session. Selected case reports demonstrating the importance of phenotype-guided interpretation of the test results are summarized in Supplementary Box 1.
Strategy if no diagnosis is reached
If no diagnosis has been reached after the general workflow has been applied, several strategies can be considered.
Patients with an MCD pattern that is known to be highly specific for one or a few genes could benefit from visual inspection of NGS reads and/or alternative targeted sequencing methods such as Sanger sequencing complemented by deletion/duplication testing of genes of interest84, as outlined in the section ‘Phenotype-specific considerations’ below. Review of NGS data might reveal inadequate coverage of the genes of interest, or the filtering out of potentially relevant splice site or flanking intronic sequences.
If not performed previously, karyotype analysis should be considered in undiagnosed patients with MCDs (86% consensus from the Neuro-MIG network). Balanced translocations and ring chromosome abnormalities are a rare cause of MCDs but have occasionally been described35,85.
Patients from consanguineous pedigrees and families with multiple affected siblings might benefit from a single nucleotide polymorphism microarray analysis to identify regions of homozygosity. If a homozygous region contains a known MCD-related gene that is compatible with the phenotype, special attention must be given to the known deep intronic variants86,87,88,89 (listed in Supplementary Table 3).
Metabolic investigations should be considered in patients with microcephaly, polymicrogyria or COB, as a broad range of metabolic diseases, including peroxisomal disorders, glutaric aciduria, fumarase deficiency and D-bifunctional protein deficiency, can manifest with cortical malformations resembling these MCD patterns1.
In patients with unexplained MCDs and muscle weakness and/or elevated creatine kinase, a muscle biopsy might be considered to allow specific analysis for dystroglycanopathies and mitochondrial disorders. The results of muscle biopsy allied to characteristic brain imaging findings in the CNS may help to indicate the affected gene90.
Some patients might benefit from repeat brain imaging, especially if the first MRI scan was performed before completion of myelination (3 months to 2.5 years of age) or was of low quality (for example, low resolution, or inadequate exploration of the brain according to the axial, coronal and sagittal plan and/or inadequate sequences). Occasionally, brain MRI scans of the parents can identify a previously unrecognized familial malformation syndrome41,91,92.
Autopsy represents an important final procedure in deceased patients with unexplained MCDs as it can provide additional information that cannot be obtained during life93. Also, after brain surgery, DNA can be extracted from affected brain tissue to identify somatic pathogenic variants. Specific protocols are recommended for the evaluation of perinatal and postnatal brain tissue, including both frozen and fixed tissue samples from key brain regions (that is, regions that are vulnerable to epilepsy-related damage) to identify specific structural abnormalities and rule out other pathologies94.
Finally, patients without a diagnosis should be considered for trio-based whole-genome sequencing and RNA sequencing, preferably within a large collaborative research network to allow rapid discovery of novel causative variants, non-coding variants in regulatory elements and epigenetic variations95,96,97.
Recurrence risk and genetic counselling
Only when the cause of the MCD is known can an accurate recurrence risk be provided to the patient and their family. When the cause is unknown, an attempt should be made to provide an empirical risk figure. This figure depends on the type of malformation, clinical presentation and the causes that have been reliably excluded (Table 3). We should point out that empirical risk counselling requires very high confidence in correct MRI interpretation and recognition of the specific phenotype.
Phenotype-specific considerations
Microcephaly
The aetiology of microcephaly is heterogeneous and includes both genetic and non-genetic factors. Non-genetic causes, including intrauterine teratogen exposure (for example, alcohol or drugs), congenital infections and perinatal and postnatal brain injuries (placental insufficiency, birth complications, postnatal infarcts and concussions), account for almost 30% of microcephaly cases. Recognized genetic causes include chromosomal aneuploidies, CNVs, some of which are submicroscopic, and a rapidly growing number of single gene disorders (reviewed by Pirozzi et al.11). Accurate perinatal history-taking aids the identification of teratogen exposure and infections, although a negative history can never reliably rule out these causes. Brain scans should be scrutinized for signs of fetal injury, including gliosis, cysts and calcifications. Clinicians should be aware that cortical malformations, especially polymicrogyria, can also be caused by fetal injury (see also below). Recurrence in the family, dysmorphic features and congenital abnormalities outside the CNS can be indicative of a genetic cause.
Ophthalmological abnormalities are found in up to 48% of patients with microcephaly98,99, including chorioretinal lacunae in Aicardi syndrome, chorioretinopathy in KIF11-related microcephaly, microphthalmia and cataract in Warburg Micro syndrome and cerebro-oculo-facio-skeletal syndrome, chorioretinitis after in utero CMV or toxoplasmosis infection, and a wide spectrum of abnormalities of the macula, retina and optic nerve after in utero Zika virus infection. Therefore, a detailed eye examination should be routinely performed in every individual with microcephaly so that appropriate support and diagnostics can be implemented.
Megalencephaly
Examination of an individual with megalencephaly should include an assessment of whether the malformation is confined to the brain or whether it is associated with a generalized or segmental overgrowth syndrome. Careful assessment of serial height, weight and OFC measurements is helpful, as is examining the body for any asymmetries and skin abnormalities. Overgrowth usually manifests within the first 2 years of life100. Currently, >20 generalized overgrowth syndromes are known (reviewed elsewhere100,101). Distinctive facial features can also aid identification of the underlying syndrome.
Generalized overgrowth syndromes are most often caused by germline gene mutations or CNVs, which can be identified with the standardized workflow. By contrast, segmental overgrowth syndromes and some isolated megalencephaly syndromes are caused by somatic mutations that might elude detection by standard workflows. To increase the chance of identifying the disease-causing variant, it might be necessary to sequence DNA derived from affected tissue (for example, skin or brain specimens) instead of blood. Further details of this approach are provided in the section ‘Detecting mosaic variants’ below. Several overgrowth syndromes, as well as the PTEN hamartoma tumour syndrome, are associated with an increased risk of malignancies.
An increasing number of defects in genes involved in cell growth and proliferation pathways are being identified in megalencephaly. The affected pathways and molecules include the PI3K–AKT–mTOR and RAS–MAPK–ERK pathways, DNA methyltransferases, transcription initiation regulators and receptor tyrosine kinases11,102,103. In our experience, PI3K–AKT–mTOR pathway-associated megalencephaly is often ≥3 s.d. above the mean. Mutations in this pathway can cause either isolated or syndromal megalencephaly, with other features including somatic (body) overgrowth and/or other MCDs, including polymicrogyria104,105. Given the high prevalence of mosaicism in these disorders, a tailored approach is recommended (see below).
Lissencephaly spectrum
The lissencephaly imaging classification was updated in 2017 and now includes 21 patterns17. Lissencephaly is considered to be an exclusively genetic disorder40, with 28 genes currently known to be associated with this condition (Supplementary Table 3). Four lissencephaly patterns are highly specific for pathogenic variants in one or two genes, with diagnostic yields >90%40. The first pattern is diffuse agyria with cortical thickness >10 mm, which is caused by LIS1 and DCX variants. The main cause in this group is a microdeletion at chromosome 17p13.3, the LIS1 locus, which can cause isolated lissencephaly, or Miller–Dieker syndrome in the case of a larger deletion40. The second specific pattern is occipital agyria combined with frontal pachygyria, which is primarily associated with deletions and pathogenic variants in LIS1, but also in rare cases with TUBG1 variants and TUBA1A variants affecting codon Arg402. The third pattern is pachygyria with a cortical thickness of 5–10 mm, most prominent over the temporal lobes, combined with complete agenesis of the corpus callosum and severe hypomyelination. This pattern is caused by ARX pathogenic variants. Note that pathogenic variants in DYNC1H1 have been linked to a similar lissencephaly pattern but without hypomyelination. The fourth pattern, diffuse SBH with a band thickness >5 mm, is a pathognomonic pattern strongly associated with pathogenic variants in DCX in both women and men40. Posterior-predominant SBH is associated with mild or mosaic LIS1 mutations40.
No other genes have been associated with these patterns. Therefore, a negative test result for those genes in a patient with a specific phenotype should prompt an offer to the family to participate in a research project focusing on gene discovery.
Periventricular nodular heterotopia
PVNH is associated with numerous CNVs and single gene mutations and can be part of a complex syndromic disorder, such as van Maldergem syndrome, Donnai–Barrow syndrome, Au–Kline syndrome or Noonan-like syndrome with loose anagen hair37. Proteins encoded by the genes associated with PVNH are involved in several cellular and molecular mechanisms, including the formation of the radial glial scaffold, cell–cell adhesion and vesicle trafficking. In addition, dysregulation of PI3K–AKT–mTOR or SMAD2/3 signalling pathways, RNA processing or transcriptional regulation has been reported in people with PVNH106,107,108. At least 20 genes have been associated with this condition (Supplementary Table 2).
FLNA mutations are an important monogenic cause of PVNH and, owing to a substantial risk of cardiovascular and other organ complications, identification of FLNA-related disorders is of great clinical importance77,109. Although no single feature is pathognomonic, several features should raise suspicion of an FLNA mutation, including female sex, with or without a positive family history that follows an X-linked dominant pattern; absence of overt intellectual disability, although learning difficulties, dyslexia and/or psychiatric problems can be present110,111; bilateral clusters of confluent nodules extending along the walls of the frontocentral lateral ventricles (classic PVNH)44; and the presence of a retrocerebellar cyst or mega cisterna magna44,110. Less frequently, corpus callosum hypoplasia, inward rotated anterior ventricular horns, white matter abnormalities and/or focal cortical abnormalities can be observed77,110. Systemic involvement is not an obligatory feature but can be present, leading to cardiovascular abnormalities such as patent ductus arteriosus, aortic aneurysm and cardiac valvular dystrophy; obstructive lung disease; constipation; coagulopathy; joint hypermobility; and other connective tissue abnormalities77,109,110.
In individuals with one or two single nodules, normal cognitive functioning and no other congenital abnormalities, the yield of genetic testing is low. However, these individuals can harbour mosaic FLNA mutations that might be passed on through the germline to their offspring44.
Posterior-predominant PVNH is a common pattern that is often associated with overlying polymicrogyria and/or subcortical heterotopia, as well as abnormalities of the fossa posterior, corpus callosum and/or hippocampus112. This pattern can be caused by a microdeletion of chromosome 6q27, but has also been associated with fetal brain injury36,113.
Subcortical heterotopia
Several rare, mostly symmetrical bilateral forms of SUBH have a genetic origin, usually with an autosomal recessive mode of inheritance. Extensive brain involvement is seen in the mesial parasagittal form associated with Chudley–McCullough syndrome, which results from biallelic variants in GPSM2, and ribbon-like heterotopia, in combination with agenesis of the corpus callosum and megalencephaly, is observed in individuals with biallelic EML1 variants114,115. Another rare subtype affecting the peritrigonal regions has been observed in patients with variants in genes encoding a microtubule component (TUBB), a microtubule-severing protein that localizes to the centrosome and mitotic spindle during cell division (KATNB1), or a centrosomal protein with tubulin-dimer binding activity (CENPJ)21.
In parallel with the diverse morphology of SUBH, the aetiology of this condition is also very heterogeneous, and for certain subtypes is largely unknown. For example, no genetic cause has been identified for curvilinear heterotopia, which is often asymmetric and can extend from the cortex to the ependyma21,116. However, a vascular disruptive cause has been suggested in several patients on the basis of a prenatal history of twinning, near miscarriage or trauma117,118,119,120, and some cases are hypothesized to result from postzygotic mutations21.
Polymicrogyria
The aetiology of polymicrogyria can be either genetic or disruptive27, and our new clinical workflow has been designed to make the physician aware of potential pitfalls. Despite extensive work-up, including genomic testing, the underlying aetiology of polymicrogyria often remains unknown.
In a substantial proportion of patients, polymicrogyria has a genetic aetiology. Various CNVs, in particular, 22q11.2 and 1p36 deletions, have been linked to this condition, along with a rapidly growing number of monogenic causes, including several metabolic disorders (Supplementary Table 2). Dozens of genes implicated in different pathways or groups of related disorders, including the mTORopathies (affecting the PI3K–AKT–mTOR pathway), the tubulinopathies and the RABopathies, have been associated with polymicrogyria121.
A common cause of polymicrogyria is a congenital CMV infection, which is thought to account for 12–30% of cases, or even more among patients with specific white matter changes62,64. Congenital CMV infection should be suspected if polymicrogyria is observed in the presence of clinical features such as microcephaly and congenital sensorineural hearing loss. Imaging features suggestive of congenital CMV, besides polymicrogyria, include white matter hyperintensities and intracranial calcifications62,64,122. Toxoplasmosis, syphilis, varicella zoster virus and Zika virus have also been associated with polymicrogyria27,60. Additional non-genetic causes include vascular disruptive events during pregnancy and, according to a few reports, maternal ergotamine use123. Twinning is also a risk factor for polymicrogyria, particularly in the case of death of a monozygotic co-twin, and in some cases of twin-to-twin transfusion syndrome, in which the donor twin is most commonly affected124. The association with twinning is proposed to be related to vascular disturbance and/or hypoperfusion125.
Dysmorphic features, multiple congenital abnormalities, megalencephaly and microcephaly are all indicative of a genetic cause, although the latter condition can also be associated with congenital infection. Evaluation of head circumference is an essential part of the clinical work-up and could assist with variant interpretation, as several genes are specifically associated with microcephaly or megalencephaly121. The best-known gene associated with polymicrogyria and microcephaly is WDR62, and germline or somatic variants in genes encoding components of the mTOR pathway, such as PIK3CA and PIK3R2, are usually associated with megalencephaly, often with other abnormalities such as vascular skin lesions and digital anomalies121. Calcifications on brain imaging are indicative of fetal brain injury (dystrophic calcification). However, COL4A1 and COL4A2 pathogenic variants can genetically predispose to fetal vascular injuries, and the pseudo-TORCH syndrome mimics congenital infection126,127.
Polymicrogyria can be associated with peroxisomal disorders such as Zellweger syndrome or D-bifunctional protein deficiency, and is reported in up to 65% of patients with the latter condition128. A peroxisomal disorder should be suspected if a child with polymicrogyria is unusually sick for an individual with a static brain malformation, particularly in the neonatal period or early infancy. Additional abnormalities might be found, including dysmorphic features, hepatomegaly and profound hypotonia. In addition to polymicrogyria, brain MRI will usually show severe leukoencephalopathy129. If a peroxisomal disorder is suspected, plasma levels of very long chain fatty acids (VLCFAs) should be checked, and further investigations such as skin fibroblast enzymatic analysis or genomic testing should be initiated.
The work-up of a patient with polymicrogyria first requires astute clinical assessment and review of the brain MRI scan. If CMV is suspected, attempts should be made to retrieve the Guthrie neonatal blood spot for CMV PCR. VLCFA analysis should be requested if a peroxisomal disorder is suspected. CMA remains the first tier of genomic analysis. Although many genes have been associated with polymicrogyria, the yield of standard genomic testing is generally ~20% (unpublished work from Neuro-MIG laboratories). Deep sequencing might be required to identify mosaic variants, especially in patients with megalencephaly. However, patients with mosaic PIK3R2 mutations and normal OFC have been reported.
Cobblestone malformation
All currently known COB syndromes are genetic and inherited in an autosomal recessive mode. A major group is the dystroglycanopathies, which are linked to various genes required for O-glycosylation of α-dystroglycan (Supplementary Table 1). Patients often have muscular dystrophy with markedly elevated serum creatine kinase levels. Moreover, eye involvement, such as severe myopia or structural malformations, is frequently observed. Recurrent biallelic microdeletions at the ISPD locus are the most common cause of dystroglycanopathies. Other COB syndromes include laminopathies, congenital disorders of glycosylation and basement membrane transmigration disorders (reviewed by Dobyns et al.27). At the imaging level, COB can be difficult to distinguish from polymicrogyria27, but creatine kinase analysis and/or an ophthalmological examination can potentially guide the clinical diagnosis25.
Differentiation of COB syndromes from polymicrogyria might be especially challenging on low-resolution images and at a young age when myelination is still ongoing (from 3 months to 2 years of age). Useful distinguishing characteristics include the intracortical striations that appear at regular intervals vertical and perpendicular to the grey–white matter border in COB and that differ from the chaotic striations seen in polymicrogyria27. Other structural malformations that can co-occur with COB include hydrocephalus, brainstem hypoplasia and cerebellar cysts. The white matter might show an abnormal MRI signal and small cysts. However, what clearly appears as polymicrogyria on MRI can present as typical neuronal overmigration on microscopic examination, suggesting that COB and polymicrogyria have a common pathogenesis130.
Tubulinopathies
Tubulinopathy is caused by heterozygous missense variants in any one of six tubulin-encoding genes, TUBA1A, TUBB2A, TUBB2B, TUBB3, TUBB and TUBG1. The variants probably exert dominant-negative effects on microtubule assembly and/or function. Although several pathogenic variants are recurrent, many patients harbour a unique variant, which can be difficult to confidently classify as pathogenic without functional studies131.
The tubulinopathies present with highly heterogeneous yet very recognizable patterns of brain malformations. The presence of a typical tubulinopathy pattern can be helpful in the interpretation of variants of uncertain significance (VOUS)131. Abnormalities of the cortex can be obvious or subtle, and the range encompasses microlissencephaly, pachygyria with a cortical thickness >10 mm, pachygyria with a 5–10 mm thick cortex (often more prominent in the perisylvian regions), polymicrogyria, dysgyria and a simplified gyral pattern17,30,131,132. The basal ganglia are usually dysmorphic, including an enlarged caudate and absent or diminutive anterior limb of the internal capsule (dividing the caudate from the putamen), resulting in a fused striatum that in turn gives the frontal horns of the lateral ventricles a characteristic ‘hooked’ appearance. Callosal abnormalities (partial or complete agenesis of the corpus callosum), ventriculomegaly, vermian dysplasia with ‘diagonal’ folia (folia crossing the midline at an oblique angle), cerebellar hypoplasia and asymmetric hypoplasia of the brainstem might also be seen30,31,131,133. TUBB3 pathogenic variants can cause an ocular motility disorder, known as congenital fibrosis of the extra-ocular muscles type 3, with or without MCD or axonal polyneuropathy132.
Pathogenic variants in DYNC1H1 and KIF2A, which encode microtubule-associated motor proteins, also lead to a spectrum of MCDs, ranging from pachygyria to dysgyria. Similar to the tubulinopathy spectrum, most individuals demonstrate a large caudate and vermian hypoplasia. DYNC1H1 variants can be associated with peripheral nerve disease ranging from fetal akinesia to spinal muscular atrophy with lower extremity predominance134.
FCD and hemimegalencephaly
Somatic and/or germline variants in numerous PI3K–AKT–mTOR pathway genes, including TSC2, TSC1, MTOR, PIK3CA, AKT3, RHEB, DEPDC5, NPRL3 and NPRL2, are known to be associated with malformations within the FCD–hemimegalencephaly spectrum55,135,136,137,138,139. TSC encompasses a wide spectrum of severity and clinical presentation, including FCD, and the diagnosis has consequences for surveillance and treatment79. In people who present with FCD, the skin and MRI should be checked for manifestations such as hypomelanotic macules, shagreen patch, additional FCD foci and subependymal nodules. If any of these features are present, a full diagnostic work-up including TSC1/TSC2 testing is recommended140. Germline pathogenic variants in the GATOR1 complex genes DEPDC5, NPRL2 and NPRL3 are associated with focal onset seizures with or without FCD on imaging. In families with epilepsy in particular, these genes should be carefully checked for SNVs and CNVs that segregate in an autosomal dominant pattern with reduced penetrance141,142,143. Two-hit models involving germline plus somatic variants in TSC2 and DEPDC5 have been proposed to explain the aetiology of TSC-associated FCD and isolated FCD type IIA141,142,144. In recent years, somatic mutations in SLC35A2, which encodes an enzyme involved in glycosylation, have been found in focal epilepsy specimens and seem to be specific to FCD type I137,145,146. Analysis of resected brain tissue using deep sequencing and single-cell techniques might be required for detection of somatic mutations.
Cerebrovascular disorders associated with MCDs
Prenatal and postnatal cerebrovascular events can lead to ischaemic and disruptive brain malformations, including schizencephaly, polymicrogyria, intracranial calcifications, cysts and porencephaly. Disorders with a vascular and/or inflammatory basis, such as familial stroke, pseudo-TORCH syndrome, Aicardi–Goutières syndrome, leukoencephalopathy with cortical cysts, and cerebral microangiopathy syndromes with calcifications and cysts, can cause damage to the developing brain. A case series of 119 individuals with intracranial calcifications revealed a specific diagnosis in 50% of the cases147. Of these, 33 had Aicardi–Goutières syndrome, 6 had OCLN-related pseudo-TORCH syndrome and 3 had a COL4A1-related disease. Pathogenic variants in USP18 have been associated with cerebral haemorrhage in utero, leading to polymicrogyria148. However, polymicrogyria is a rare feature in cerebrovascular disorders.
Several reports have shown porencephaly, schizencephaly, polymicrogyria and PVNH associated with COL4A1 pathogenic variants, which cause imbalance or structural distortion of the collagen IV triple helix126,149,150. Evidence for a link between COL4A2 and MCDs is weaker, although, considering the functional interactions between the two collagen IV proteins, COL4A1 and COL4A2 should be tested together149. Despite reports of EMX2 as a ‘schizencephaly gene’, evidence of a role for EMX2 mutations in schizencephaly is lacking151,152.
A list of genes that have been associated with early-onset and often severe cerebrovascular phenotypes is provided in Supplementary Table 4.
Laboratory requirements
Chromosomal microarray analysis
A survey within the Neuro-MIG network, which was conducted in preparation for this Consensus Statement, indicated that multiple different microarray platforms can be used, with no specific technology showing a clear advantage.
When choosing CMA platforms for MCD diagnostics, special attention should be paid to the exon-level resolution of genes in which single-exon aberrations have been described (Supplementary Table 3). Single nucleotide polymorphism arrays have the advantage of detecting regions of homozygosity, thereby facilitating diagnostics in consanguineous families. Mosaic CNVs showing as little as 15–20% chromosomal mosaicism were successfully detected in patients with neurodevelopmental disorders153. We anticipate that CMA will become redundant in the future as NGS costs further decrease and algorithms for CNV analysis from NGS data become more robust.
High-throughput sequencing
As MCDs constitute a genetically heterogeneous group of disorders and the number of known disease-associated genes is rapidly increasing, we strongly recommend genome-wide testing approaches combined with targeted evaluation of genes that are currently implicated in MCDs (the ‘slice approach’). If the results of these tests are negative, the strategy can be expanded to a full trio exome analysis after appropriate genetic counselling. Neuro-MIG network laboratories are applying various exome enrichment strategies with comparable efficiency across the platforms and compliance with published NGS guidelines154,155. Most current exome sequencing enrichment kits provide sufficient coverage to offer an MCD panel as a type A or type B test154. The terms type A and type B refer to the definitions from the current guidelines for diagnostic NGS from the European Society of Human Genetics (ESHG), whereby the laboratory guarantees >99% reliable reference or variant calls of the target regions (type A) or describes exactly which regions are sequenced at >99% reliable reference or variant calls (type B)154.
Variant calling and prioritization
Our experience shows that an average per base coverage of 100 reads with a minimum coverage of 30 reads is sufficient for reliable calls within coding and flanking intronic regions. Neuro-MIG network members preferentially use a variant calling threshold of 20% of the non-reference (alternative) reads and variant calling is performed within exons and 10 bp of the flanking intronic sequence (80% consensus). However, deep intronic variants affecting splicing have already been described in several MCD-associated genes (Supplementary Table 5). Such variants need to be considered in patients with highly suggestive phenotypes, but might require genome or targeted sequencing.
The described approach is applicable for the identification of constitutional (germline) and high-grade mosaic variants (>30% of cells). Special considerations regarding detection and validation of low-grade mosaic variants are summarized in the section ‘Detecting mosaic variants’ below.
Supplementary Table 2 provides a curated list of the core MCD-associated genes, including information on the observed mutational spectrum and associated phenotypes. Supplementary Table 3 summarizes selected genes associated with syndromic, often postnatal microcephaly. Microcephaly is a frequent accompanying feature of these conditions but is not a key manifestation. Genes associated with disorders that always present with microcephaly are listed in Supplementary Table 2. Taking into account the number of novel disease-associated genes that are emerging, we strongly suggest updating the gene lists according to the current literature every 6 months.
Variant interpretation follows the general recommendations of EuroGentest, the ESHG and the American College of Medical Genetics and Genomics (ACMG)154,156.
As all MCD entities are rare disorders, we recommend classifying a variant as benign if the allele frequency is >1% in the Genome Aggregation Database (gnomAD), which differs from the ACMG stand-alone evidence of benign impact with an allele frequency of >5%157. As the Neuro cohort of gnomAD includes individuals with neuropsychiatric disorders, which represent a rare manifestation of MCDs, one should consider excluding variants from this cohort when estimating gnomAD allele frequency, as pathogenic MCD-associated variants might be present. The presence of a variant as a homozygous allele in multiple (at least five) individuals in gnomAD strongly suggests its benign impact and irrelevance for the phenotype. However, one should be careful to check that the variant is truly homozygous and not hemizygous, combined with a deletion of the second allele. The impact of a homozygous SNV might differ substantially from the impact of deletion of one allele and the same SNV on the remaining allele158.
Pitfalls in variant prioritization
In-house variant databases, which contain data from a single institution, are another important source to distinguish benign from potentially causative variants. However, some MCD-relevant genes, especially those encoding tubulin, which are prone to read-alignment errors, might have high false-positive in-house frequencies. One TUBB2B pathogenic variant, Ala248Val159, was listed in gnomAD with an allele frequency of 3% but is currently flagged as failed — that is, probably an artefact — by random forest filters. However, when in-house data are analysed, this variant might erroneously show up in control samples in up to 30% of the reads (K.S., unpublished work) and might, therefore, be filtered out as a ‘frequent’ in-house variant, despite being pathogenic. On the basis of this example, we suggest that manual curation of in-house variants in the tubulin-encoding genes should include consideration of mapping quality and comparison of in-house frequencies with the curated gnomAD dataset. Sanger sequencing of TUBB2B could be considered in undiagnosed patients with an MCD pattern highly suggestive of a tubulinopathy. In the near future, such misalignment errors should be solved through high-resolution mapping and application of long-read DNA sequencing platforms160.
The presence of highly homologous pseudogenes also complicates accurate variant calling for a number of MCD-relevant target genes161 (Supplementary Table 2).
Penetrance of MCD-associated variants
With the exception of X-chromosomal genes such as ARX and DCX, and gene encoding components of the GATOR1 complex, variants in other MCD-associated genes seem to be fully penetrant, as carrier probands always show characteristic structural changes in the brain. However, individuals with these variants might be clinically asymptomatic and therefore never undergo brain MRI. In the case of inheritance of likely pathogenic variants from apparently unaffected parents, parental brain imaging is essential for accurate variant interpretation91,110. Female carriers of the X-chromosomal variants might be clinically unaffected and have normal brain scans43,162. Incomplete and/or age-related penetrance were reported for variants mainly associated with a seizure phenotype (for example, GATOR1 complex genes163); therefore, variants inherited from unaffected parents might be considered causative.
Clinical laboratory report
The final laboratory report, including reporting of incidental findings, should follow the general requirements published by EuroGentest, the ESHG and the ACMG, as well as country-specific guidelines for genetic laboratory reports.
If the review board includes a medical professional with sufficient expertise in MRI interpretation, we recommend that MRI scans should be presented together with the clinical information and relevant variants. Relevant clinical information and brain imaging are important for accurate interpretation of the variants and should be actively requested.
If parents and similarly affected siblings (if applicable) were not analysed together with the index patient, segregation analysis must be strongly recommended in the final report.
All pathogenic and likely pathogenic variants (class 5 and class 4 variants, respectively) must be included in the final report. The final report should also contain all VOUS (class 3 variants) in MCD-associated genes. The laboratory should consider including protein-altering de novo, homozygous or compound heterozygous rare variants in potentially relevant genes of uncertain significance in the final report. The relevance for the MCD phenotype might be determined on the basis of the expression pattern of the gene or its potential importance for human brain evolution (human-specific genes or transcripts). Despite the fact that most MCD-associated genes are evolutionarily conserved, primate-specific genes and isoforms should not be ignored as they can be linked to neurodevelopmental disorders164. The relevance of such variants must be continually re-evaluated over time.
High-resolution, single-exon-level CNV analysis is essential to complement the sequencing report. CNV analysis can be provided with different methods including CNV calling from NGS data if robustly established and validated, multiplex ligation-dependent probe amplification, quantitative PCR or customized high-resolution microarrays.
The final report must specify whether CNV analysis has been performed, including information about the genes analysed and methods used for the analysis. If no copy number analysis has been carried out, the report must contain information about the genes that require copy number tests.
If (likely) pathogenic variants or VOUS have been identified, patients and/or their families should be referred to a clinical geneticist for return of results and counselling on their clinical and prognostic implications.
We recommend sharing VOUS in the available databases, such as ClinVar and the Leiden Open Variation Database. Depending on the local ethical and legal regulations, some laboratories might choose to use different country-specific databases.
Additional considerations
Detecting mosaic variants
Mosaic (postzygotic somatic) mutations, including mutations in PI3K–AKT–mTOR pathway genes, as well as in DCX, LIS1, FLNA and TUBB2B, have been described in a wide range of MCDs54. Mosaic mutation variant detection requires dedicated deep sequencing and bioinformatics tools, as these variants are likely to be missed by standard-coverage exon sequencing, especially in blood-derived DNA58. When available, affected brain tissue is the recommended tissue for genetic testing. If this tissue is not available, the use of ‘proxies’ such as saliva or skin-derived fibroblasts is recommended over lymphocytes when a mosaic disorder is suspected58,165. Ideally, multiple tissues from the same individual should be examined.
Reliable testing requires a targeted approach to specific loci, using a customized gene panel with ultra-deep sequencing (for example, >1,000-times coverage). A gene panel for PI3K–AKT–mTOR-related syndromes is provided in Supplementary Table 6. As a general rule, hybridization-based assays offer superior performance over amplicon assays166. However, amplicon protocols with unique molecular identifiers during library preparation have also proved effective for detecting somatic mutations167. The variant-calling algorithm (percentage of non-reference allele reads) must be adapted for detecting low-grade mosaic SNVs.
As false-positive mosaic mutation calls can arise from many different sources, we strongly suggest confirmation of every low-grade mosaic variant using an orthogonal technology such as droplet digital PCR or a second independent round of ultra-deep sequencing168,169,170.
Neuropathological work-up
Detailed neuropathological examination, biobanking and genetic testing are required after epilepsy surgery or autopsy in patients with MCDs, and also after sudden unexpected death in epilepsy, as individuals who die as a result of sudden unexpected death in epilepsy might have a previously undiagnosed MCD.
In 2016, the task force of neuropathology from the International League Against Epilepsy (ILAE) Commission on Diagnostic Methods published a consensus standard operational procedure for collection and processing of cortical samples from patients with MCDs such as FCDs94. Whenever feasible, anatomically intact surgical neocortical samples should be obtained to allow systematic analysis to identify the affected area. Correct orientation of the cortical sample and determination of its relationship to neurophysiologically aberrant sites and MRI findings requires an interdisciplinary diagnostic approach with good communication between pathology, neurology, radiology and neurosurgical teams. Representative tissue should be apportioned for histology and biobanking. Brain tissue-derived DNA is often required for genetic diagnosis in FCD and hemimegalencephaly; thus, highly standardized tissue processing is recommended. A neuropathologist should be involved in the interpretation of the brain pathology, and molecular biologists (or pathologists) and geneticists should participate in the set-up and analysis of the sequencing results171. A consensus protocol with details of how to best process resected brain specimens for somatic mutational analysis to detect mosaic variants for hemimegalencephaly, FCD types I and II, and other MCDs is under development by a task force of the ILAE (E.A., unpublished work).
Analysis of low-level mosaic mutations, such as those reported in FCDs57,136,142,172, requires careful selection of brain regions and cells to ensure enrichment of the mutated cells, followed by deep sequencing136,138. A study published in 2019 used resected brain tissue from a large cohort of patients after epilepsy surgery to explore the possibility of detecting low-level somatic mutations in unmatched formalin-fixed paraffin-embedded (FFPE) brain tissue samples (that is, brain samples without a blood sample from the same patient). FFPE samples often represent the most relevant samples in the standard neuropathological diagnostic approach to MCDs146,173. The researchers showed that deep sequencing, even when applied to unmatched FFPE brain tissues, can be used to accurately and efficiently detect low-level somatic mutations.
Conclusions
In this Consensus Statement, we present a diagnostic work-up for individuals affected by brain malformations within the MCD spectrum, encompassing current best practices and recommendations based on the consensus of a multidisciplinary group of international experts within the Neuro-MIG network. With this approach, we aim to increase diagnostic yield, thereby improving patient care and management worldwide and facilitating the development of targeted therapeutic approaches in the long term.
References
Guerrini, R. & Dobyns, W. B. Malformations of cortical development: clinical features and genetic causes. Lancet Neurol. 13, 710–726 (2014).
Coryell, J. et al. Neuroimaging of early life epilepsy. Pediatrics 142, e20180672 (2018).
Barkovich, A. J., Guerrini, R., Kuzniecky, R. I., Jackson, G. D. & Dobyns, W. B. A developmental and genetic classification for malformations of cortical development: update 2012. Brain 135, 1348–1369 (2012). A comprehensive classification of around 200 MCD subtypes according to developmental mechanism.
de Wit, M. C. et al. Cortical brain malformations: effect of clinical, neuroradiological, and modern genetic classification. Arch. Neurol. 65, 358–366 (2008).
Wiszniewski, W. et al. Comprehensive genomic analysis of patients with disorders of cerebral cortical development. Eur. J. Hum. Genet. 26, 1121–1131 (2018).
van Nimwegen, K. J. et al. The diagnostic pathway in complex paediatric neurology: a cost analysis. Eur. J. Paediatr. Neurol. 19, 233–239 (2015).
Mancini, G. M. S., Neuro-MIG Network (COST Action CA16118). Neuro-MIG: a European network on brain malformations. Eur. J. Med. Genet. 61, 741–743 (2018). An introduction to the Neuro-MIG network.
Severino, M. et al. Definitions and classification of malformations of cortical development: practical guidelines. Brain https://doi.org/10.1093/brain/awaa174 (2020).
Ashwal, S. et al. Practice parameter: evaluation of the child with microcephaly (an evidence-based review): report of the quality standards subcommittee of the American Academy of Neurology and the practice committee of the Child Neurology Society. Neurology 73, 887–897 (2009).
Woods, C. G. & Parker, A. Investigating microcephaly. Arch. Dis. Child. 98, 707–713 (2013).
Pirozzi, F., Nelson, B. & Mirzaa, G. From microcephaly to megalencephaly: determinants of brain size. Dialogues Clin. Neurosci. 20, 267–282 (2018).
Jayaraman, D., Bae, B. I. & Walsh, C. A. The genetics of primary microcephaly. Annu. Rev. Genomics Hum. Genet. 19, 177–200 (2018). A review on the molecular mechanisms underlying primary microcephaly.
von der Hagen, M. et al. Diagnostic approach to microcephaly in childhood: a two-center study and review of the literature. Dev. Med. Child. Neurol. 56, 732–741 (2014).
Alcantara, D. & O’Driscoll, M. Congenital microcephaly. Am. J. Med. Genet. C. Semin. Med. Genet. 166C, 124–139 (2014).
DeMyer, W. Megalencephaly: types, clinical syndromes, and management. Pediatr. Neurol. 2, 321–328 (1986).
Jansen, P. R. et al. Incidental findings on brain imaging in the general pediatric population. N. Engl. J. Med. 377, 1593–1595 (2017).
Di Donato, N. et al. Lissencephaly: expanded imaging and clinical classification. Am. J. Med. Genet. A 173, 1473–1488 (2017). A classification of the lissencephaly spectrum malformations: agyria, pachygyria and subcortical band heterotopia.
Dobyns, W. B. The clinical patterns and molecular genetics of lissencephaly and subcortical band heterotopia. Epilepsia 51 (Suppl. 1), 5–9 (2010).
Kato, M. & Dobyns, W. B. Lissencephaly and the molecular basis of neuronal migration. Hum. Mol. Genet. 2, Spec. No. 1 R89–R96 (2003).
Basel-Vanagaite, L. & Dobyns, W. B. Clinical and brain imaging heterogeneity of severe microcephaly. Pediatr. Neurol. 43, 7–16 (2010).
Oegema, R., Barkovich, A. J., Mancini, G. M. S., Guerrini, R. & Dobyns, W. B. Subcortical heterotopic gray matter brain malformations: classification study of 107 individuals. Neurology 93, e1360–e1373 (2019). Imaging patterns associated with subcortical MCDs.
van der Knaap, M. S. et al. Magnetic resonance imaging in classification of congenital muscular dystrophies with brain abnormalities. Ann. Neurol. 42, 50–59 (1997).
Brun, B. N. et al. Comparison of brain MRI findings with language and motor function in the dystroglycanopathies. Neurology 88, 623–629 (2017).
Luo, R. et al. G protein-coupled receptor 56 and collagen III, a receptor-ligand pair, regulates cortical development and lamination. Proc. Natl Acad. Sci. USA 108, 12925–12930 (2011).
Devisme, L. et al. Cobblestone lissencephaly: neuropathological subtypes and correlations with genes of dystroglycanopathies. Brain 135, 469–482 (2012).
Walker, E. Lissencephaly. Arch. Neurol. Psychiatry 48, 13–29 (1942).
Dobyns, W. B., Leventer, R. J. & Guerrini, R. In Swaiman’s Pediatric Neurology, 6th edn (eds Swaiman, K. F. et al.) part V. 29 (Elsevier, 2018).
Leventer, R. J. et al. Clinical and imaging heterogeneity of polymicrogyria: a study of 328 patients. Brain 133, 1415–1427 (2010). A classification and review of polymicrogyria subtypes.
Leventer, R. J., Guerrini, R. & Dobyns, W. B. Malformations of cortical development and epilepsy. Dialogues Clin. Neurosci. 10, 47–62 (2008).
Oegema, R. et al. Recognizable cerebellar dysplasia associated with mutations in multiple tubulin genes. Hum. Mol. Genet. 24, 5313–5325 (2015).
Mutch, C. A. et al. Disorders of microtubule function in neurons: imaging correlates. Am. J. Neuroradiol. 37, 528–535 (2016).
Najm, I. M., Sarnat, H. B. & Blumcke, I. Review: the international consensus classification of focal cortical dysplasia — a critical update 2018. Neuropathol. Appl. Neurobiol. 44, 18–31 (2018).
Dobyns, W. B. et al. Consistent chromosome abnormalities identify novel polymicrogyria loci in 1p36.3, 2p16.1-p23.1, 4q21.21-q22.1, 6q26-q27, and 21q2. Am. J. Med. Genet. A 146, 1637–1654 (2008).
van Kogelenberg, M. et al. Periventricular heterotopia in common microdeletion syndromes. Mol. Syndromol. 1, 35–41 (2010).
Dobyns, W. B. et al. Miller–Dieker syndrome: lissencephaly and monosomy 17p. J. Pediatr. 102, 552–558 (1983).
Peddibhotla, S. et al. Delineation of candidate genes responsible for structural brain abnormalities in patients with terminal deletions of chromosome 6q27. Eur. J. Hum. Genet. 23, 54–60 (2015).
Cellini, E. et al. Multiple genomic copy number variants associated with periventricular nodular heterotopia indicate extreme genetic heterogeneity. Eur. J. Hum. Genet. 27, 909–918 (2019).
Sajan, S. A. et al. Both rare and de novo copy number variants are prevalent in agenesis of the corpus callosum but not in cerebellar hypoplasia or polymicrogyria. PLoS Genet. 9, e1003823 (2013).
Boonsawat, P. et al. Elucidation of the phenotypic spectrum and genetic landscape in primary and secondary microcephaly. Genet. Med. 21, 2043–2058 (2019).
Di Donato, N. et al. Analysis of 17 genes detects mutations in 81% of 811 patients with lissencephaly. Genet. Med. 20, 1354–1364 (2018).
Clapham, K. R. et al. FLNA genomic rearrangements cause periventricular nodular heterotopia. Neurology 78, 269–278 (2012).
Dobyns, W. B. & Das, S. PAFAH1B1-associated lissencephaly/subcortical band heterotopia. GeneReviews (2014).
Hehr, U., Uyanik, G., Aigner, L., Couillard-Despres, S. & Winkler, J. DCX-related disorders. GeneReviews (2019).
Parrini, E. et al. Periventricular heterotopia: phenotypic heterogeneity and correlation with filamin A mutations. Brain 129, 1892–1906 (2006).
Sheen, V. L. et al. Mutations in the X-linked filamin 1 gene cause periventricular nodular heterotopia in males as well as in females. Hum. Mol. Genet. 10, 1775–1783 (2001).
Gonzalez-Moron, D. et al. Germline and somatic mutations in cortical malformations: molecular defects in Argentinean patients with neuronal migration disorders. PLoS ONE 12, e0185103 (2017).
Nicholas, A. K. et al. The molecular landscape of ASPM mutations in primary microcephaly. J. Med. Genet. 46, 249–253 (2009).
Tan, C. A. et al. Analysis of ASPM in an ethnically diverse cohort of 400 patient samples: perspectives of the molecular diagnostic laboratory. Clin. Genet. 85, 353–358 (2014).
Letard, P. et al. Autosomal recessive primary microcephaly due to ASPM mutations: an update. Hum. Mutat. 39, 319–332 (2018).
Mahmood, S., Ahmad, W. & Hassan, M. J. Autosomal recessive primary microcephaly (MCPH): clinical manifestations, genetic heterogeneity and mutation continuum. Orphanet J. Rare Dis. 6, 39 (2011).
de Ligt, J. et al. Diagnostic exome sequencing in persons with severe intellectual disability. N. Engl. J. Med. 367, 1921–1929 (2012).
Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature 519, 223–228 (2015).
O’Roak, B. J. et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science 338, 1619–1622 (2012).
Jamuar, S. S. et al. Somatic mutations in cerebral cortical malformations. N. Engl. J. Med. 371, 733–743 (2014).
Jansen, L. A. et al. PI3K/AKT pathway mutations cause a spectrum of brain malformations from megalencephaly to focal cortical dysplasia. Brain 138, 1613–1628 (2015).
Marsan, E. & Baulac, S. Review: mechanistic target of rapamycin (mTOR) pathway, focal cortical dysplasia and epilepsy. Neuropathol. Appl. Neurobiol. 44, 6–17 (2018).
Lim, J. S. et al. Brain somatic mutations in MTOR cause focal cortical dysplasia type II leading to intractable epilepsy. Nat. Med. 21, 395–400 (2015).
Mirzaa, G. et al. PIK3CA-associated developmental disorders exhibit distinct classes of mutations with variable expression and tissue distribution. JCI Insight 1, e87623 (2016).
Barkovich, A. J. in Magnetic Resonance of Epilepsy Ch. 7 (eds Kuzniecky, R. I. & Jackson, G. D.) 221–248 (Elsevier, 2005).
de Vries, L. S. Viral infections and the neonatal brain. Semin. Pediatr. Neurol. 32, 100769 (2019).
Barkovich, A. J. & Raybaud, C. Pediatric Neuroimaging 5th edn (Lippincott Williams and Wilkins, 2011).
Mavili, E. et al. Polymicrogyria: correlation of magnetic resonance imaging and clinical findings. Childs Nerv. Syst. 28, 905–909 (2012).
Noyola, D. E. et al. Early predictors of neurodevelopmental outcome in symptomatic congenital cytomegalovirus infection. J. Pediatr. 138, 325–331 (2001).
van der Knaap, M. S. et al. Pattern of white matter abnormalities at MR imaging: use of polymerase chain reaction testing of Guthrie cards to link pattern with congenital cytomegalovirus infection. Radiology 230, 529–536 (2004).
Plotkin, S. A., Reef, S. E., Cooper, L. Z. & Alford, C. A. In Infectious Diseases of the Fetus and Newborn Infant 7th edn (eds Remington, J. S. et al.) 861–889 (Elsevier Saunders, 2011).
Koren, G. Congenital varicella syndrome in the third trimester. Lancet 366, 1591–1592 (2005).
Marquez, L., Levy, M. L., Munoz, F. M. & Palazzi, D. L. A report of three cases and review of intrauterine herpes simplex virus infection. Pediatr. Infect. Dis. J. 30, 153–157 (2011).
Honein, M. A. et al. Birth defects among fetuses and infants of US women with evidence of possible Zika virus infection during pregnancy. JAMA 317, 59–68 (2017).
WHO. Zika virus. https://www.who.int/news-room/fact-sheets/detail/zika-virus (2020).
Melo, A. S. et al. Congenital Zika virus infection: beyond neonatal microcephaly. JAMA Neurol. 73, 1407–1416 (2016).
Miranda-Filho Dde, B. et al. Initial description of the presumed congenital Zika syndrome. Am. J. Public Health 106, 598–600 (2016).
Moore, C. A. et al. Characterizing the pattern of anomalies in congenital Zika syndrome for pediatric clinicians. JAMA Pediatr. 171, 288–295 (2017).
Parra-Saavedra, M. et al. Serial head and brain imaging of 17 fetuses with confirmed Zika virus infection in Colombia, South America. Obstet. Gynecol. 130, 207–212 (2017).
Strafela, P. et al. Zika virus-associated microcephaly: a thorough description of neuropathologic findings in the fetal central nervous system. Arch. Pathol. Lab. Med. 141, 73–81 (2017).
Stevelink, R. et al. Epilepsy surgery for patients with genetic refractory epilepsy: a systematic review. Epileptic Disord. 20, 99–115 (2018).
Lim, Y. & Lyall, H. Congenital cytomegalovirus — who, when, what-with and why to treat? J. Infect. 74, S89–S94 (2017).
Lange, M. et al. 47 patients with FLNA associated periventricular nodular heterotopia. Orphanet J. Rare Dis. 10, 134 (2015). The key features in the heterogeneous clinical spectrum of FLNA-associated disorder.
Yilmaz, S. et al. The expanding phenotypic spectrum of ARFGEF2 gene mutation: cardiomyopathy and movement disorder. Brain Dev. 38, 124–127 (2016).
Krueger, D. A. & Northrup, H. & International Tuberous Sclerosis Complex Consensus Group. Tuberous sclerosis complex surveillance and management: recommendations of the 2012 International Tuberous Sclerosis Complex Consensus Conference. Pediatr. Neurol. 49, 255–265 (2013). An evidence-based, standardized approach for optimal clinical care provided for individuals with TSC.
Geis, T. et al. Clinical long-time course, novel mutations and genotype–phenotype correlation in a cohort of 27 families with POMT1-related disorders. Orphanet J. Rare Dis. 14, 179 (2019).
Herbst, S. M. et al. LIS1-associated classic lissencephaly: a retrospective, multicenter survey of the epileptogenic phenotype and response to antiepileptic drugs. Brain Dev. 38, 399–406 (2016).
Venot, Q. et al. Targeted therapy in patients with PIK3CA-related overgrowth syndrome. Nature 558, 540–546 (2018).
Kim, J. K. et al. Brain somatic mutations in MTOR reveal translational dysregulations underlying intractable focal epilepsy. J. Clin. Invest. 129, 4207–4223 (2019).
Pena, L. D. M. et al. Looking beyond the exome: a phenotype-first approach to molecular diagnostic resolution in rare and undiagnosed diseases. Genet. Med. 20, 464–469 (2018).
Zaki, M. et al. Identification of a novel recessive RELN mutation using a homozygous balanced reciprocal translocation. Am. J. Med. Genet. A 143A, 939–944 (2007).
Kobayashi, K. et al. Deep-intronic variant of fukutin is the most prevalent point mutation of Fukuyama congenital muscular dystrophy in Japan. J. Hum. Genet. 62, 945–948 (2017).
Lax, N. Z. et al. Neuropathologic characterization of pontocerebellar hypoplasia type 6 associated with cardiomyopathy and hydrops fetalis and severe multisystem respiratory chain deficiency due to novel RARS2 mutations. J. Neuropathol. Exp. Neurol. 74, 688–703 (2015).
Mancini, C. et al. Megalencephalic leukoencephalopathy with subcortical cysts type 1 (MLC1) due to a homozygous deep intronic splicing mutation (c.895-226T>G) abrogated in vitro using an antisense morpholino oligonucleotide. Neurogenetics 13, 205–214 (2012).
Vandervore, L. V. et al. Heterogeneous clinical phenotypes and cerebral malformations reflected by rotatin cellular dynamics. Brain 142, 867–884 (2019).
Bonnemann, C. G. et al. Diagnostic approach to the congenital muscular dystrophies. Neuromuscul. Disord. 24, 289–311 (2014).
Harel, T. et al. Homozygous null variant in CRADD, encoding an adaptor protein that mediates apoptosis, is associated with lissencephaly. Am. J. Med. Genet. A 173, 2539–2544 (2017).
Zillhardt, J. L. et al. Mosaic parental germline mutations causing recurrent forms of malformations of cortical development. Eur. J. Hum. Genet. 24, 611–614 (2016).
Aminoff, M. J., Boller, F., Swaab, D. F., Sarnat, H. B. & Curatolo, P. Handbook of Clinical Neurology: Malformations of the Nervous System (Elsevier Science & Technology 2007).
Blumcke, I. et al. International recommendation for a comprehensive neuropathologic workup of epilepsy surgery brain tissue: a consensus Task Force report from the ILAE Commission on Diagnostic Methods. Epilepsia 57, 348–358 (2016). A standardized protocol for neuropathological work-up of epilepsy surgery specimens.
Barbosa, M. et al. Identification of rare de novo epigenetic variations in congenital disorders. Nat. Commun. 9, 2064 (2018).
Kremer, L. S. et al. Genetic diagnosis of Mendelian disorders via RNA sequencing. Nat. Commun. 8, 15824 (2017).
Perenthaler, E., Yousefi, S., Niggl, E. & Barakat, T. S. Beyond the exome: the non-coding genome and enhancers in neurodevelopmental disorders and malformations of cortical development. Front. Cell Neurosci. 13, 352 (2019).
Stoll, C., Alembik, Y., Dott, B. & Roth, M. P. Congenital eye malformations in 212,479 consecutive births. Ann. Genet. 40, 122–128 (1997).
Krauss, M. J., Morrissey, A. E., Winn, H. N., Amon, E. & Leet, T. L. Microcephaly: an epidemiologic analysis. Am. J. Obstet. Gynecol. 188, 1484–1489 (2003).
Burkardt, D. D., Tatton-Brown, K., Dobyns, W. & Graham, J. M. Jr. Approach to overgrowth syndromes in the genome era. Am. J. Med. Genet. C. Semin. Med. Genet. 181, 483–490 (2019).
Kamien, B. et al. A clinical review of generalized overgrowth syndromes in the era of massively parallel sequencing. Mol. Syndromol. 9, 70–82 (2018).
Mirzaa, G. M., Riviere, J. B. & Dobyns, W. B. Megalencephaly syndromes and activating mutations in the PI3K-AKT pathway: MPPH and MCAP. Am. J. Med. Genet. C Semin. Med. Genet. 163, 122–130 (2013).
Tidyman, W. E. & Rauen, K. A. The RASopathies: developmental syndromes of Ras/MAPK pathway dysregulation. Curr. Opin. Genet. Dev. 19, 230–236 (2009).
Mirzaa, G. M. et al. Megalencephaly-capillary malformation (MCAP) and megalencephaly-polydactyly-polymicrogyria-hydrocephalus (MPPH) syndromes: two closely related disorders of brain overgrowth and abnormal brain and body morphogenesis. Am. J. Med. Genet. A 158A, 269–291 (2012).
Riviere, J. B. et al. De novo germline and postzygotic mutations in AKT3, PIK3R2 and PIK3CA cause a spectrum of related megalencephaly syndromes. Nat. Genet. 44, 934–940 (2012).
Ferland, R. J. et al. Disruption of neural progenitors along the ventricular and subventricular zones in periventricular heterotopia. Hum. Mol. Genet. 18, 497–516 (2009).
Broix, L. et al. Mutations in the HECT domain of NEDD4L lead to AKT-mTOR pathway deregulation and cause periventricular nodular heterotopia. Nat. Genet. 48, 1349–1358 (2016).
Oegema, R. et al. Human mutations in integrator complex subunits link transcriptome integrity to brain development. PLoS Genet. 13, e1006809 (2017).
Chen, M. H. et al. Thoracic aortic aneurysm in patients with loss of function filamin A mutations: clinical characterization, genetics, and recommendations. Am. J. Med. Genet. A 176, 337–350 (2018).
Chen, M. H. & Walsh, C. A. FLNA-related periventricular nodular heterotopia. GeneReviews (2015).
Fry, A. E. et al. Neuropsychiatric disease in patients with periventricular heterotopia. J. Neuropsychiatry Clin. Neurosci. 25, 26–31 (2013).
Mandelstam, S. A. et al. Bilateral posterior periventricular nodular heterotopia: a recognizable cortical malformation with a spectrum of associated brain abnormalities. Am. J. Neuroradiol. 34, 432–438 (2013).
Aldinger, K. A. et al. Redefining the etiologic landscape of cerebellar malformations. Am. J. Hum. Genet. 105, 606–615 (2019).
Doherty, D. et al. GPSM2 mutations cause the brain malformations and hearing loss in Chudley–McCullough syndrome. Am. J. Hum. Genet. 90, 1088–1093 (2012).
Oegema, R. et al. EML1-associated brain overgrowth syndrome with ribbon-like heterotopia. Am. J. Med. Genet. C Semin. Med. Genet. 181, 627–637 (2019).
Barkovich, A. J. Morphologic characteristics of subcortical heterotopia: MR imaging study. Am. J. Neuroradiol. 21, 290–295 (2000).
Dubeau, F. et al. Periventricular and subcortical nodular heterotopia. A study of 33 patients. Brain 118, 1273–1287 (1995).
Kuzniecky, R., Gilliam, F. & Faught, E. Discordant occurrence of cerebral unilateral heterotopia and epilepsy in monozygotic twins. Epilepsia 36, 1155–1157 (1995).
Novegno, F. et al. Giant subcortical heterotopia involving the temporo-parieto-occipital region: a challenging cause of drug-resistant epilepsy. Epilepsy Res. 87, 88–94 (2009).
Preul, M. C. et al. Function and organization in dysgenic cortex. Case report. J. Neurosurg. 87, 113–121 (1997).
Stutterd, C. A., Dobyns, W. B., Jansen, A., Mirzaa, G. & Leventer, R. J. Polymicrogyria overview. GeneReviews (2018).
Barkovich, A. J. & Lindan, C. E. Congenital cytomegalovirus infection of the brain: imaging analysis and embryologic considerations. Am. J. Neuroradiol. 15, 703–715 (1994).
Barkovich, A. J., Rowley, H. & Bollen, A. Correlation of prenatal events with the development of polymicrogyria. Am. J. Neuroradiol. 16, 822–827 (1995).
Ascherl, R. et al. Severe gyration and migration disorder in fetofetal transfusion syndrome: two case reports and a review of the literature on the neurological outcome of children with lesions on neuroimaging. Childs Nerv. Syst. 34, 155–163 (2018).
Jansen, A. & Andermann, E. Genetics of the polymicrogyria syndromes. J. Med. Genet. 42, 369–378 (2005).
Cavallin, M. et al. Further refinement of COL4A1 and COL4A2 related cortical malformations. Eur. J. Med. Genet. 61, 765–772 (2018).
O’Driscoll, M. C. et al. Recessive mutations in the gene encoding the tight junction protein occludin cause band-like calcification with simplified gyration and polymicrogyria. Am. J. Hum. Genet. 87, 354–364 (2010).
Ferdinandusse, S. et al. Clinical and biochemical spectrum of D-bifunctional protein deficiency. Ann. Neurol. 59, 92–104 (2006).
Barkovich, A. J. & Peck, W. W. MR of Zellweger syndrome. Am. J. Neuroradiol. 18, 1163–1170 (1997).
Vandervore, L. V. et al. TMX2 is a crucial regulator of cellular redox state, and its dysfunction causes severe brain developmental abnormalities. Am. J. Hum. Genet. 105, 1126–1147 (2019).
Bahi-Buisson, N. et al. The wide spectrum of tubulinopathies: what are the key features for the diagnosis? Brain 137, 1676–1700 (2014).
Romaniello, R. et al. Tubulin genes and malformations of cortical development. Eur. J. Med. Genet. 61, 744–754 (2018). Together with Bahi-Buisson et al. (2014), this article provides aids to recognizing the brain imaging abnormalities associated with the tubulinopathies.
Bahi-Buisson, N. & Cavallin, M. Tubulinopathies overview. GeneReviews (2016).
Poirier, K. et al. Mutations in TUBG1, DYNC1H1, KIF5C and KIF2A cause malformations of cortical development and microcephaly. Nat. Genet. 45, 639–647 (2013).
Hoelz, H. et al. Childhood-onset epileptic encephalopathy associated with isolated focal cortical dysplasia and a novel TSC1 germline mutation. Clin. EEG Neurosci. 49, 187–191 (2018).
Lim, J. S. et al. Somatic mutations in TSC1 and TSC2 cause focal cortical dysplasia. Am. J. Hum. Genet. 100, 454–472 (2017).
Baldassari, S. et al. Dissecting the genetic basis of focal cortical dysplasia: a large cohort study. Acta Neuropathol. 138, 885–900 (2019).
D’Gama, A. M. et al. Somatic mutations activating the mTOR pathway in dorsal telencephalic progenitors cause a continuum of cortical dysplasias. Cell Rep. 21, 3754–3766 (2017). References 137 and 138 report the spectrum of genetic variants associated with cortical dysplasia.
Pelorosso, C. et al. Somatic double-hit in MTOR and RPS6 in hemimegalencephaly with intractable epilepsy. Hum. Mol. Genet. 28, 3755–3765 (2019).
Northrup, H. & Krueger, D. A. & International Tuberous Sclerosis Complex Consensus Group. Tuberous sclerosis complex diagnostic criteria update: recommendations of the 2012 International Tuberous Sclerosis Complex Consensus Conference. Pediatr. Neurol. 49, 243–254 (2013).
Baulac, S. et al. Familial focal epilepsy with focal cortical dysplasia due to DEPDC5 mutations. Ann. Neurol. 77, 675–683 (2015).
Ribierre, T. et al. Second-hit mosaic mutation in mTORC1 repressor DEPDC5 causes focal cortical dysplasia-associated epilepsy. J. Clin. Invest. 128, 2452–2458 (2018).
Sim, J. C. et al. Familial cortical dysplasia caused by mutation in the mammalian target of rapamycin regulator NPRL3. Ann. Neurol. 79, 132–137 (2016).
Martin, K. R. et al. The genomic landscape of tuberous sclerosis complex. Nat. Commun. 8, 15816 (2017).
Winawer, M. R. et al. Somatic SLC35A2 variants in the brain are associated with intractable neocortical epilepsy. Ann. Neurol. 83, 1133–1146 (2018).
Sim, N. S. et al. Brain somatic mutations in SLC35A2 cause intractable epilepsy with aberrant N-glycosylation. Neurol. Genet. 4, e294 (2018).
Livingston, J. H., Stivaros, S., van der Knaap, M. S. & Crow, Y. J. Recognizable phenotypes associated with intracranial calcification. Dev. Med. Child. Neurol. 55, 46–57 (2013).
Meuwissen, M. E. et al. Human USP18 deficiency underlies type 1 interferonopathy leading to severe pseudo-TORCH syndrome. J. Exp. Med. 213, 1163–1174 (2016).
Plaisier, E. & Ronco, P. COL4A1-related disorders. GeneReviews (2016).
Zagaglia, S. et al. Neurologic phenotypes associated with COL4A1/2 mutations: expanding the spectrum of disease. Neurology 91, e2078–e2088 (2018).
Granata, T. et al. Schizencephaly: clinical spectrum, epilepsy, and pathogenesis. J. Child. Neurol. 20, 313–318 (2005).
Merello, E. et al. No major role for the EMX2 gene in schizencephaly. Am. J. Med. Genet. A 146A, 1142–1150 (2008).
Oneda, B. et al. Low-level chromosomal mosaicism in neurodevelopmental disorders. Mol. Syndromol. 8, 266–271 (2017).
Matthijs, G. et al. Guidelines for diagnostic next-generation sequencing. Eur. J. Hum. Genet. 24, 2–5 (2016).
Vermeesch, J. R. et al. Guidelines for molecular karyotyping in constitutional genetic diagnosis. Eur. J. Hum. Genet. 15, 1105–1114 (2007).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015). Helpful and widely used guideline for classification of genomic variants.
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Albers, C. A. et al. Compound inheritance of a low-frequency regulatory SNP and a rare null mutation in exon-junction complex subunit RBM8A causes TAR syndrome. Nat. Genet. 44, 435–439 (2012).
Amrom, D. et al. Polymicrogyria with dysmorphic basal ganglia? Think tubulin! Clin. Genet. 85, 178–183 (2014).
van Dijk, E. L., Jaszczyszyn, Y., Naquin, D. & Thermes, C. The third revolution in sequencing technology. Trends Genet. 34, 666–681 (2018).
Jenkinson, E. M. et al. Comprehensive molecular screening strategy of OCLN in band-like calcification with simplified gyration and polymicrogyria. Clin. Genet. 93, 228–234 (2018).
Kato, M. et al. Mutations of ARX are associated with striking pleiotropy and consistent genotype–phenotype correlation. Hum. Mutat. 23, 147–159 (2004).
Baulac, S. & Weckhuysen, S. DEPDC5-related epilepsy. GeneReviews (2016).
O’Neill, A. C. et al. A primate-specific isoform of PLEKHG6 regulates neurogenesis and neuronal migration. Cell Rep. 25, 2729–2741.e6 (2018).
Kuentz, P. et al. Molecular diagnosis of PIK3CA-related overgrowth spectrum (PROS) in 162 patients and recommendations for genetic testing. Genet. Med. 19, 989–997 (2017).
Samorodnitsky, E. et al. Evaluation of hybridization capture versus amplicon-based methods for whole-exome sequencing. Hum. Mutat. 36, 903–914 (2015).
Xu, C. A review of somatic single nucleotide variant calling algorithms for next-generation sequencing data. Comput. Struct. Biotechnol. J. 16, 15–24 (2018).
Campbell, I. M., Shaw, C. A., Stankiewicz, P. & Lupski, J. R. Somatic mosaicism: implications for disease and transmission genetics. Trends Genet. 31, 382–392 (2015).
Sims, D., Sudbery, I., Ilott, N. E., Heger, A. & Ponting, C. P. Sequencing depth and coverage: key considerations in genomic analyses. Nat. Rev. Genet. 15, 121–132 (2014).
Hindson, C. M. et al. Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat. Methods 10, 1003–1005 (2013).
Blumcke, I. et al. The clinicopathologic spectrum of focal cortical dysplasias: a consensus classification proposed by an ad hoc Task Force of the ILAE Diagnostic Methods Commission. Epilepsia 52, 158–174 (2011).
Mirzaa, G. M. et al. Association of MTOR mutations with developmental brain disorders, including megalencephaly, focal cortical dysplasia, and pigmentary mosaicism. JAMA Neurol. 73, 836–845 (2016). This article shows the importance of skin examination in patients with brain malformations.
Sim, N. S. et al. Precise detection of low-level somatic mutation in resected epilepsy brain tissue. Acta Neuropathol. 138, 901–912 (2019).
Byrd, S. E., Osborn, R. E., Bohan, T. P. & Naidich, T. P. The CT and MR evaluation of migrational disorders of the brain. Part II. Schizencephaly, heterotopia and polymicrogyria. Pediatr. Radiol. 19, 219–222 (1989).
Schaffer, A. E. et al. Biallelic loss of human CTNNA2, encoding alphaN-catenin, leads to ARP2/3 complex overactivity and disordered cortical neuronal migration. Nat. Genet. 50, 1093–1101 (2018).
Lee, S. et al. Bi-allelic loss of human APC2, encoding adenomatous polyposis coli protein 2, leads to lissencephaly, subcortical heterotopia, and global developmental delay. Am. J. Hum. Genet. 105, 844–853 (2019).
Acknowledgements
This article represents a consensus document based on three Neuro-MIG network meetings, which were supported by COST Action CA16118. The authors would like to thank all members of the network for their participation and feedback.
Author information
Authors and Affiliations
Contributions
R.O., T.S.B., M.W., K.S., D.A., E.A., N.B.-B., V.C., T.G., D.G.A., E.P., I.P., E.S., D.S., L.M.V., M.S.Z., G.M., O.R., U.H., A.J., G.M.S.M. and N.D.D. researched data for the article. R.O., M.W., E.A., N.B.-B., A.E.F., G.M., W.B.D., O.R., R.G., D.T.P., U.H., R.J.L., A.J., G.M.S.M. and N.D.D. made substantial contributions to discussion of the content. R.O., T.S.B. and N.D.D. wrote the article. All authors reviewed and edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
ClinVar: https://www.ncbi.nlm.nih.gov/clinvar/
COST Action CA16118: https://www.cost.eu/actions/CA16118/#tabs
Leiden Open Variation Database: http://www.lovd.nl/3.0/home
Neuro-MIG: https://www.neuro-mig.org
WG1: https://www.neuro-mig.org/content/wg1-standardization-clinical-phenotyping-and-medical-management
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Oegema, R., Barakat, T.S., Wilke, M. et al. International consensus recommendations on the diagnostic work-up for malformations of cortical development. Nat Rev Neurol 16, 618–635 (2020). https://doi.org/10.1038/s41582-020-0395-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41582-020-0395-6
This article is cited by
-
Exploring unsolved cases of lissencephaly spectrum: integrating exome and genome sequencing for higher diagnostic yield
Journal of Human Genetics (2024)
-
Angeborene Hirnfehlbildungen
Die Radiologie (2024)
-
Focally Enlarged Perivascular Spaces in Pediatric and Adolescent Patients with Polymicrogyria—an MRI Study
Clinical Neuroradiology (2024)
-
A new variant confirms GNAI2 as a rare cause of periventricular nodular heterotopia
Neurological Sciences (2024)
-
EANM practice guidelines for an appropriate use of PET and SPECT for patients with epilepsy
European Journal of Nuclear Medicine and Molecular Imaging (2024)