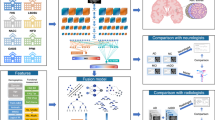
Abstract
Globally, there is a huge unmet need for effective treatments for neurodegenerative diseases. The complexity of the molecular mechanisms underlying neuronal degeneration and the heterogeneity of the patient population present massive challenges to the development of early diagnostic tools and effective treatments for these diseases. Machine learning, a subfield of artificial intelligence, is enabling scientists, clinicians and patients to address some of these challenges. In this Review, we discuss how machine learning can aid early diagnosis and interpretation of medical images as well as the discovery and development of new therapies. A unifying theme of the different applications of machine learning is the integration of multiple high-dimensional sources of data, which all provide a different view on disease, and the automated derivation of actionable insights.
Key points
-
Machine learning and natural language processing are forms of artificial intelligence that enable robust interrogation of multiple datasets to identify previously undiscovered patterns and relationships in the data.
-
Machine learning approaches have been applied to the study of neurodegenerative diseases and show promise in the areas of early diagnosis, prognosis and development of new therapies.
-
A substantial number of machine learning algorithms exist, and choosing the correct algorithm to apply to different types of data is crucial to obtain reliable results.
-
Neuroimaging was the first area of neurology to benefit from the application of machine learning approaches to improve diagnosis; more recently, application of machine learning methods to motor function and language feature analysis has shown promise in decreasing the time taken to perform clinical assessments.
-
The application of machine learning to longitudinal patient data collection and electronic health records has the potential to inform prognosis prediction and patient stratification.
-
Large collections of curated datasets and robust assessment of machine learning methods will be needed to achieve full integration of machine learning into diagnostic and prognostic neurology practice and the design of future therapeutics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
McCarthy, J. Basic questions. What is Artificial Intelligence? http://www-formal.stanford.edu/jmc/whatisai/node1.html (2007).
Agatonovic-Kustrin, S. & Beresford, R. Basic concepts of artificial neural network (ANN) modeling and its application in pharmaceutical research. J. Pharm. Biomed. Anal. 22, 717–727 (2000).
Yu, K. H., Beam, A. L. & Kohane, I. S. Artificial intelligence in healthcare. Nat. Biomed. Eng. 2, 719–731 (2018).
McDougall, R. J. Computer knows best? The need for value-flexibility in medical AI. J. Med. Ethics 45, 156–160 (2019).
McDougall, R. J. No we shouldn’t be afraid of medical AI; it involves risks and opportunities. J. Med. Ethics 45, 559 (2019).
Vellido, A. Societal issues concerning the application of artificial intelligence in medicine. Kidney Dis. 5, 11–17 (2019).
Di Nucci, E. Should we be afraid of medical AI? J. Med. Ethics 45, 556–558 (2019).
de Saint Laurent, C. In defence of machine learning: debunking the myths of artificial intelligence. Eur. J. Psychol. 14, 734–747 (2018).
Buch, V. H., Ahmed, I. & Maruthappu, M. Artificial intelligence in medicine: current trends and future possibilities. Br. J. Gen. Parctice 68, 143–144 (2018).
Denaxas, S. C. & Morley, K. I. Big biomedical data and cardiovascular disease research: opportunities and challenges. Eur. Heart. J. Qual. Care Clin. Outcomes 1, 9–16 (2015).
Weber, G., Mandl, K. & Kohane, I. Finding the missing link for big biomedical data. JAMA 311, 2479–2480 (2014).
Van Horn, J. & Toga, A. Human neuroimaging as a “big data” science. Brain Imaging Behav. 8, 323–331 (2014).
Zhou, L. & Verstreken, P. Reprogramming neurodegeneration in the big data era. Curr. Opin. Neurobiol. 48, 167–173 (2018).
Vallejos, C. A., Richardson, S. & Marioni, J. C. Beyond comparisons of means: understanding changes in gene expression at the single-cell level. Genome Biol. 17, 1–14 (2016).
Ritchie, M. D. et al. Multifactor-dimensionality reduction reveals high-order interactions among estrogen-metabolism genes in sporadic breast cancer. Am. J. Hum. Genet. 69, 138–147 (2001).
Xu, J., Zhang, Y., Qiu, C. & Cheng, F. Global and regional economic costs of dementia: a systematic review [abstract]. Lancet 390, S47 (2017).
Prince, M., Prina, M. & Guerchet, M. World Alzheimer’s Report 2013. The Journey of Caring: An Analysis of Long-Term Care for Dementia (Alzheimer’s Disease International, 2013).
Bishop, C. Pattern Recognition and Machine Learning (Springer, 2006).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015). This extensive review provides an elegant summary of deep learning methods and their application to images, video footage, speech recordings and written text.
Van Der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
Oyelade, J. et al. Clustering algorithms: their application to gene expression data. Bioinform. Biol. Insights 10, 237–253 (2016).
Chapelle, O., Schölkopf, B. & Zien, A. (eds) Semi-Supervised Learning (MIT Press, 2006).
Vapnik, V. Statistical Learning Theory (Wiley-Interscience, 1998).
Joachmis, T. in ICML ’99: Proceedings of the Sixteenth International Conference on Machine Learning (eds Bratko, I. & Dzeroski, S.) 200–209 (Morgan Kaufmann, 1999).
Watkins, C. J. C. H. Learning with Delayed Rewards. Thesis, King’s College, Cambridge (1989).
Mnih, V. et al. Human-level control through deep reinforcement learning. Nature 518, 529–533 (2015).
Popova, M., Isayev, O. & Tropsha, A. Deep reinforcement learning for de-novo drug design. Sci. Adv. 4, 1–14 (2017).
Raudys, Š. Statistical and Neural Classifiers: An Integrated Approach to Design (Springer, 2001).
Summers, M. J. et al. Deep machine learning application to the detection of preclinical neurodegenerative diseases of aging. Sci. J. Digit. Cult. 2, 9–24 (2017).
Ho, T. K. Random decision forests perceptron training. in ICDAR ’95: Proceedings of the Third International Conference on Document Analysis and Recognition 278–282 (IEEE Computer Society, 1995).
Hothorn, T. & Jung, H. H. RandomForest4Life: a random forest for predicting ALS disease progression. Amyotroph. Lateral Scler. Front. Degener. 15, 444–452 (2014).
Cortes, C. & Vapnik, V. Support-vector networks. Mach. Learn. 297, 273–297 (1995).
Rosenblatt, F. The Perceptron – A Perceiving and Rocognizing Automation (Cornell Aeronautical Laboratory, 1957).
McCulloch, W. S. & Pitts, W. A logical calculus of the idea immanent in nervous activity. Bull. Math. Biophys. 5, 115–133 (1943). This paper describes the first steps towards mathematical modelling of neuronal function, which eventually resulted in the development of artificial neural networks.
Fukushima, K. Neocognitron: a self-organizing neural network model for a mechanism of pattern recognition unaffected by shift in position. Biol. Cybern. 36, 193–202 (1980).
LeCun, Y., Bottou, L., Bengio, Y. & Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 86, 2278–2324 (1998).
LeCun, Y., Hafner, P., Bottou, L. & Bengio, Y. in Shape, Contour and Grouping in Computer Vision. Lecture Notes in Computer Science Vol 1681 (eds Forsyth, D. A., Mundy, J. L., di Gesú, V. & Cipolla, R.) 319–345 (Springer, 1999).
Burt, J. R. et al. Deep learning beyond cats and dogs: recent advances in diagnosing breast cancer with deep neural networks. Br. J. Radiol. 91, 2–11 (2018).
Hopfield, J. J. Neural networks and physical systems with emergent collective computational abilities. Proc. Natl Acad. Sci. USA 79, 2554–2558 (1982).
Hochreiter, S. & Schmidhuber, J. Long short-term memory. Neural Comput. 9, 1735–1780 (1997).
Cho, K. et al. in Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing (EMNLP) 1724–1734 (Association for Computational Linguistics, 2014).
Cawley, G. C. & Talbot, N. L. C. On over-fitting in model selection and subsequent selection bias in performance evaluation. J. Mach. Learn. Res. 11, 2079–2107 (2010).
Chicco, D. Ten quick tips for machine learning in computational biology. BioData Min. 10, 1–17 (2017).
Neumaier, A. Solving ill-conditioned and singular linear systems: a tutorial on regularization. SIAM Rev. 40, 636–666 (1998).
Michel, P. P., Hirsch, E. C. & Hunot, S. Understanding dopaminergic cell death pathways in Parkinson disease. Neuron 90, 675–691 (2016).
Donev, R., Kolev, M., Millet, B. & Thome, J. Neuronal death in Alzheimer’s disease and therapeutic opportunities. J. Cell. Mol. Med. 13, 4329–4348 (2009).
Fischer, L. R. et al. Amyotrophic lateral sclerosis is a distal axonopathy: evidence in mice and man. Exp. Neurol. 182, 232–240 (2004).
Hainc, N. et al. The bright, artificial intelligence-augmented future of neuroimaging reading. Front. Neurol. 8, 10–12 (2017).
Grenander, U., Chow, Y. & Keenan, D. HANDS: a Pattern Theoretic Study of Biological Shapes (Springer, 1990).
Evans, A. C., Marrett, S., Torrescorzo, J., Ku, S. & Collins, L. MRI-PET correlation in three dimensions using a volume-of-interest (VOI) atlas. J. Cereb. Blood Flow. Metab. 11, A69–A78 (1991).
Woods, R. P., Mazziotta, J. C. & Cherry, S. R. MRI-PET registration with automated algorithm. J. Comput. Assist. Tomogr. 17, 536–546 (1993).
Joshi, S. C. et al. Hierarchical brain mapping via a generalized dirichlet solution for mapping brain manifolds. in Proceedings of the SPIE’s 1995 international symposium on optical science, engineering, and instrumentation. Vision geometry IV Vol. 2573 (eds Melter, R. A., Wu, A. Y., Bookstein, F. L. & Green, W. D. K.) 278–289 (SPIE, 1995).
Grady, C. L. et al. Subgroups in dementia of the Alzheimer type identified using positron emission tomography. J. Neuropsychiatry Clin. Neurosci. 2, 373–384 (1990).
DeFigueiredo, R. J. P. et al. Neural-network-based classification of cognitively normal, demented, Alzheimer disease and vascular dementia from single photon emission with computed tomography image data from brain. Proc. Natl Acad. Sci. USA 92, 5530–5534 (1995). This study is one of the first to have used an artificial neural network algorithm to automate the identification of normal ageing, AD and vascular dementia from SPECT data.
Wang, S. et al. Pathological brain detection by artificial intelligence in magnetic resonance imaging scanning. Prog. Electromagn. Res. 156, 105–133 (2016).
Haller, J. W. et al. Hippocampal MR imaging morphometry by means of general pattern matching. Radiology 199, 787–791 (1996).
Davatzikos, C. et al. A computerized approach for morphological analysis of the corpus callosum. J. Comput. Assist. Tomogr. 20, 88–97 (1996).
Gur, R. C. et al. Sex differences in brain gray and white matter in healthy young adults: correlations with cognitive performance. J. Neurosci. 19, 4065–4072 (1999).
Mega, M. S. et al. Cerebral correlates of psychotic symptoms in Alzheimer’s disease. J. Neurol. Neurosurg. Psychiatry 69, 167–171 (2000).
Fischl, B. & Dale, A. Measuring the thickness of the human cerebral cortex from magnetic resonance images. Proc. Natl Acad. Sci. USA 97, 11050–11055 (2000).
Ashburner, J. & Friston, K. Voxel-based morphometry–the methods. Neuroimage 11, 805–821 (2000).
Haller, J. W. et al. Three-dimensional hippocampal volumetry by high dimensional transformation of a neuroanatomical atlas. Radiology 202, 504–510 (1997).
Maldijan, J. A., Laurienti, P. J., Kraft, R. A. & Burdette, J. H. An automated method for neuroanatomic and cytoarchitectonic atlas-based interrogation of fMRI data sets. Neuroimage 19, 1233–1239 (2003). The paper presents the first automated analysis system based on a digital brain atlas to show robust application to fMRI data, without the need for pre-definied region of interest masks.
Lao, Z. et al. Morphological classification of brains via high-dimensional shape transformations and machine learning methods. Neuroimage 21, 46–57 (2004). This paper presents an early application of SVM to MR image analysis and highlights the importance of analysing all voxels simultaneously, rather than focusing on a pre-defined region of interest.
Mourão-Miranda, J., Bokde, A. L. W., Born, C., Hampel, H. & Stetter, M. Classifying brain states and determining the discriminating activation patterns: support vector machine on functional MRI data. Neuroimage 28, 980–995 (2005). This study is an early demonstration of the superior performance of SVM over traditional statistical methods for MRI analysis and highlights the ability of SVM to select the brain regions from which the most accurate classification can be drawn.
Mitchell, T. M. et al. Learning to decode cognitive states from brain images. Mach. Learn. 57, 145–175 (2004). In this study multiple machine learning algorithms, including SVM, are used on functional MR images to assess the feasibility of detecting patients’ transient cognitive states during a single time interval.
Reczko, M., Karras, D. A., Mertzios, B. G., Graveron-Demilly, D. & Van Ormondt, D. Improved MR image reconstruction from sparsely sampled scans based on neural networks. Pattern Recognit. Lett. 22, 35–46 (2001).
Zhu, G. et al. Applications of deep learning to neuro-imaging techniques. Front. Neurol. 10, 1–13 (2019).
He, K., Zhang, X., Ren, S. & Sun, J. in Proceedings of the 29th IEEE Conference on Computer Vision and Pattern Recognition (CVPR 2016) 770–778 (IEEE, 2016).
Gray, K. R. et al. Random forest-based similarity measures for multi-modal classification of Alzheimer’s disease. Neuroimage 65, 167–175 (2013).
Korolev, S., Safiullin, A., Belyaev, M. & Dodonova, Y. in Proceedings of the 14th International Symposium on Biomedical Imaging 835–838 (IEEE, 2017).
Choi, H., Kang, H. & Lee, D. S. Predicting aging of brain metabolic topography using variational autoencoder. Front. Aging Neurosci. 10, 212 (2018).
Lundervold, A. S. & Lundervold, A. An overview of deep learning in medical imaging focusing on MRI. Z. Med. Phys. 29, 102–127 (2019).
Klöppel, S. et al. Automatic classification of MR scans in Alzheimer’s disease. Brain 131, 681–689 (2008). This study shows that an SVM can use MR scans to successfully distinguish between individuals with AD and individuals with FTLD as well as between individuals with AD and healthy individuals.
Bron, E. E., Smits, M., Niessen, W. J. & Klein, S. Feature selection based on the SVM weight vector for classification of dementia. IEEE J. Biomed. Heal. Inform. 19, 1617–1626 (2015).
Moradi, E., Pepe, A., Gaser, C., Huttunen, H. & Tohka, J. Machine learning framework for early MRI-based Alzheimer’s conversion prediction in MCI subjects. Neuroimage 104, 398–412 (2015).
Magnin, B. et al. Support vector machine-based classification of Alzheimer’s disease from whole-brain anatomical MRI. Neuroradiology 51, 78–83 (2009).
Gerardin, E. et al. Multidimensional classification of hippocampal shape features discriminates Alzheimer’s disease and mild cognitive impairment from normal aging. Neuroimage 47, 1476–1486 (2009).
Li, S. et al. Hippocampal shape analysis of Alzheimer disease based on machine learning methods. Am. J. Neuroradiol. 28, 1339–1345 (2007).
Amoroso, N. et al. Alzheimer’s disease diagnosis based on the hippocampal unified multi-atlas network (HUMAN) algorithm. Biomed. Eng. Online 17, 1–16 (2018).
De Marco, M., Beltrachini, L., Biancardi, A., Frangi, A. F. & Venneri, A. Machine-learning support to individual diagnosis of mild cognitive impairment using multimodal MRI and cognitive assessments. Alzheimer Dis. Assoc. Disord. 31, 278–286 (2017).
Ahn, W., Krawitz, A. & Kim, W. A model-based fMRI analysis with hierarchical Bayesian parameter estimation. J. Neurosci. Psychol. Econ. 4, 95–110 (2011).
Rehme, A. K. et al. Identifying neuroimaging markers of motor disability in acute stroke by machine learning techniques. Cereb. Cortex 25, 3046–3056 (2014).
Weygandt, M. et al. MRI pattern recognition in multiple sclerosis normal-appearing brain areas. PLoS One 6, e21138 (2011).
Duchesne, S., Rolland, Y. & Vérin, M. Automated computer differential classification in Parkinsonian syndromes via pattern analysis on MRI. Acad. Radiol. 16, 61–70 (2009).
Chen, L. et al. Rapid automated quantification of cerebral leukoaraiosis on CT images: a multicenter validation study. Radiology 288, 573–581 (2018).
Prevedello, L. M., Little, K. J., Qian, S. & White, R. D. Automated critical test findings identification and online notification system using artificial intelligence in imaging. Radiology 285, 923–931 (2017).
Titano, J. J. et al. Automated deep-neural-network surveillance of cranial images for acute neurologic events. Nat. Med. 24, 1337–1341 (2018).
Chilamkurthy, S. et al. Deep learning algorithms for detection of critical findings in head CT scans: a retrospective study. Lancet 392, 2388–2396 (2018).
Davatzikos, C., Fan, Y., Wu, X., Shen, D. & Resnick, S. M. Detection of prodromal Alzheimer’s disease via pattern classification of magnetic resonance imaging. Neurobiol. Aging 29, 514–523 (2008).
Fan, Y., Resnick, S. M., Wu, X. & Davatzikos, C. Structural and functional biomarkers of prodromal Alzheimer’s disease. Neuroimage 41, 277–285 (2008).
Fan, Y., Batmanghelich, N. K., Clark, C. M. & Davatzikos, C., Alzeimer’s Disease Neuroimaging Initiative. Spatial patterns of brain atrophy in MCI patients, identified via high-dimensional pattern classification, predict subsequent cognitive decline. Neuroimage 39, 1731–1743 (2008).
Kivipelto, M. et al. The Finnish geriatric intervention study to prevent cognitive impairment and disability (FINGER): study design and progress. Alzheimer’s Dement. 9, 657–665 (2013).
Zhang, Y. C. & Kagen, A. C. Machine learning interface for medical image analysis. J. Digit. Imaging 30, 615–621 (2017).
Mufford, M. S. et al. Neuroimaging genomics in psychiatry–a translational approach. Genome Med. 9, 1–12 (2017).
Bookheimer, S. Y. et al. Patterns of brain activation in people at risk for Alzheimer’s disease. N. Engl. J. Med. 343, 450–456 (2000).
Heinz, A. et al. Genotype influences in vivo dopamine transporter availability in human striatum. Neuropsychopharmacology 22, 133–139 (2000).
Liang, Z. & Lauterbur, P. Principles of Magnetic Resonance Imaging: a Signal Processing Approach (IEEE, 2000).
Hibar, D. P. et al. Common genetic variants influence human subcortical brain structures. Nature 520, 224–229 (2015).
Czigler, B. et al. Quantitative EEG in early Alzheimer’s disease patients – power spectrum and complexity features. Int. J. Psychophysiol. 68, 75–80 (2008).
Lee, H., Brekelmans, G. J. F. & Roks, G. The EEG as a diagnostic tool in distinguishing between dementia with Lewy bodies and Alzheimer’s disease. Clin. Neurophysiol. 126, 1735–1739 (2015).
Barcelon, E. A. et al. Grand total EEG score can differentiate Parkinson’s disease from Parkinson-related disorders. Front. Neurol. 10, 1–11 (2019).
Buscema, M. et al. An improved I-FAST system for the diagnosis of Alzheimer’s disease from unprocessed electroencephalograms by using robust invariant features. Artif. Intell. Med. 64, 59–74 (2015).
Bosco, D. A., LaVoie, M. J., Petsko, G. A. & Ringe, D. Proteostasis and movement disorders: Parkinson’s disease and amyotrophic lateral sclerosis. Cold Spring Harb. Perspect. Biol. 3, 1–24 (2011).
Ross, C. A. & Tabrizi, S. J. Huntington’s disease: from molecular pathogenesis to clinical treatment. Lancet Neurol. 10, 83–98 (2011).
[No authors listed.]. The amyotrophic lateral sclerosis functional rating scale: assessment of activities of daily living in patients with amyotrophic lateral sclerosis. Arch. Neurol. 53, 141–147 (1996).
[No authors listed.]. Unified Huntington’s disease rating scale: reliability and consistency. Mov. Disord. 11, 136–142 (1996).
Fahn, S., Elton, R. & Members of the UPDRS Development Committee. in Recent Developments in Parkinson’s Disease Vol. 2 (eds. Fahn, S., Marsden, C. D., Calne, D. B. & Goldstein, M.) 153–163, 293–304 (Macmillan Health Care Information, 1987).
Davenport, T. & Kalakota, R. The potential for artificial intelligence in healthcare. Futur. Healthc. J. 6, 94–98 (2019).
Rosenblum, S., Samuel, M., Zlotnik, S., Erikh, I. & Schlesinger, I. Handwriting as an objective tool for Parkinson’s disease diagnosis. J. Neurol. 260, 2357–2361 (2013).
Alty, J., Cosgrove, J., Thorpe, D. & Kempster, P. How to use pen and paper tasks to aid tremor diagnosis in the clinic. Pract. Neurol. 17, 456–463 (2017).
McLennan, J., Nakano, K., Tyler, H. & Schwab, R. Micrographia in Parkinson’s disease. J. Neurol. Sci. 15, 141–152 (1972).
Kotsavasiloglou, C., Kostikis, N., Hristu-Varsakelis, D. & Arnaoutoglou, M. Machine learning-based classification of simple drawing movements in Parkinson’s disease. Biomed. Signal. Process. Control. 31, 174–180 (2017). This is the first study to have used a combination of simple line drawings and machine learning algorithms to aid PD diagnosis.
Westin, J. et al. A new computer method for assessing drawing impairment in Parkinson’s disease. J. Neurosci. Methods 190, 143–148 (2010).
Griffiths, R. I., Kotschet, K., Arfon, S., Ming, Z. & Johnson, W. Automated assessment of bradykinesia and dyskinesia in Parkinson’s disease. J. Parkinson’s Dis. 2, 47–55 (2012).
Giuffrida, J. P., Riley, D. E., Maddux, B. N. & Heldman, D. A. Clinically deployable KinesiaTM technology for automated tremor assessment. Mov. Disord. 24, 723–730 (2009).
Jeon, H., Lee, W. & Park, H. High-accuracy automatic classification of parkinsonian tremor severity using machine learning method. Physiol. Meas. 38, 1980–1999 (2017).
Zhao, A., Qi, L., Dong, J. & Yu, H. Dual channel LSTM based multi-feature extraction in gait for diagnosis of neurodegenerative diseases. Knowl. Syst. 145, 91–97 (2018).
Pushparani, M. & Athisakthi, A. Detection of movement disorders using multi SVM. Glob. J. Comput. Sci. Technol. 13, 23–25 (2013).
Sacco, G. et al. Detection of activities of daily living impairment in Alzheimer’s disease and mild cognitive impairment using information and communication technology. Clin. Interv. Ageing 7, 539–549 (2012).
Ji, S., Xu, W., Yang, M. & Yu, K. 3D convolutional neural networks for human action recognition. IEEE Trans. Pattern Anal. Mach. Intell. 35, 221–231 (2013).
Raya, Z. et al. in Proceedings of SPIE: Applications of Machine Learning Vol. 11139 (eds Zelinski, M. E., Taha, T. M., Howe, J., Awwal, A. A. S. & Iftekharuddin, K. M.) 1113909 (SPIE, 2019).
Brand, D., DiGennaro Reed, F. D., Morley, M. D., Erath, T. G. & Novak, M. D. A survey assessing privacy concerns of smart-home services provided to individuals with disabilities. Behav. Anal. Pract. 13, 11–21 (2020).
Riboni, D., Bettini, C., Civitarese, G., Janjua, Z. H. & Helaoui, R. SmartFABER: Recognizing fine-grained abnormal behaviors for early detection of mild cognitive impairment. Artif. Intell. Med. 67, 57–74 (2016).
Ordóñez, F. J. & Roggen, D. Deep convolutional and LSTM recurrent activity recognition. Sensors 16, 115–140 (2016).
Alam, R., Homdee, N., Wolfe, S., Hayes, J. & Lach, J. In IoTDI 2019: Proceedings of the International Conference on Internet of Things Design and Implementation 281–282 (Association for Computing Machinery, 2019).
Rankin, K. P., Baldwin, E., Pace-Savitsky, C., Kramer, J. H. & BL, M. Self awareness and personality change in dementia. J. Neurol. Neurosurg. Psychiatry 76, 632–639 (2005).
Sollberger, M. et al. Neural basis of interpersonal traits in neurodegenerative diseases. Neuropsychologia 47, 2812–2827 (2009).
Christidi, F., Migliaccio, R., Santamaría-García, H., Santangelo, G. & Trojsi, F. Social cognition dysfunctions in neurodegenerative diseases: neuroanatomical correlates and clinical implications. Behav. Neurol. 2018, 18 (2018).
Orimaye, S., Wong, J. & Golden, K. in Proceedings of the Workshop on Computational Linguistics and Clinical Psychology: From Linguistic Signal to Clinical Reality 78–87 (Association for Computational Linguistics, 2014).
Wankerl, S., Nöth, E. & Evert, S. in Proceedings of the Annual Conference of the International Speech Communication Association, INTERSPEECH 2017 3162–3166 (Association for Computational Linguistics, 2017).
Weizenbaum, J. ELIZA — a computer program for the study of natural language communication between man and machine. Commun. ACM 26, 23–28 (1983). This article describes the first question-and-answer computer program, which paved the way for AI-driven avatars as we know them today.
Ireland, D. et al. Hello Harlie: enabling speech monitoring through chat-bot conversations. Stud. Health Technol. Inform. 227, 55–60 (2016).
Tanaka, H. et al. Detecting dementia through interactive computer avatars. IEEE J. Transl. Eng. Heal. Med. 5, 1–11 (2017).
Blackburn, D. et al. An avatar aid in memory clinic [abstract PO029]. J. Neurol. Neurosurg. Psychiatry 88, A19–A20 (2017).
Schmidtke, K., Pohlmann, S. & Metternich, B. The syndrome of functional memory disorder: definition, etiology, and natural course. Am. J. Geriatr. Psychiatry 16, 981–988 (2008).
Mahley, R. W., Weisgraber, K. H. & Huang, Y. Apolipoprotein E4: a causative factor and therapeutic target in neuropathology, including Alzheimer’s disease. Proc. Natl Acad. Sci. USA 103, 5644–5651 (2006).
Van Cauwenberghe, C., Van Broeckhoven, C. & Sleegers, K. The genetic landscape of Alzheimer disease: clinical implications and perspectives. Genet. Med. 18, 421–430 (2016).
Huang, X. et al. Revealing Alzheimer’s disease genes spectrum in the whole-genome by machine learning. BMC Neurol. 18, 1–8 (2018).
Maj, C. et al. Integration of machine learning methods to dissect genetically imputed transcriptomic profiles in Alzheimer’s disease. Front. Genet. 10, 1–16 (2019).
Lopez, C., Tucker, S., Salameh, T. & Tucker, C. An unsupervised machine learning method for discovering patient clusters based on genetic signatures. J. Biomed. Inform. 85, 30–39 (2018).
Ray, S. et al. Classification and prediction of clinical Alzheimer’s diagnosis based on plasma signaling proteins. Nat. Med. 13, 1359–1362 (2007).
Agarwal, S., Ghanty, P. & Pal, N. R. Identification of a small set of plasma signalling proteins using neural network for prediction of Alzheimer’s disease. Bioinformatics 31, 2505–2513 (2015).
Andersen, S. L. et al. Metabolome-based signature of disease pathology in MS. Mult. Scler. Relat. Disord. 31, 12–21 (2019).
Sapkota, S. et al. Alzheimer’s biomarkers from multiple modalities selectively discriminate clinical status: relative importance of salivary metabolomics panels, genetic, lifestyle, cognitive, functional health and demographic risk markers. Front. Aging Neurosci. 10, 1–13 (2018).
Tavares, J. & Oliveira, T. Electronic health record portal adoption: a cross country analysis. BMC Med. Inform. Decis. Mak. 17, 1–17 (2017).
Stone, C. P. A glimpse at EHR implementation around the world: the lessons the US can learn. e-healthpolicy.org https://www.e-healthpolicy.org/sites/e-healthpolicy.org/files/A_Glimpse_at_EHR_Implementation_Around_the_World1_ChrisStone.pdf (2014).
Chen, Y. et al. Applying active learning to high-throughput phenotyping algorithms for electronic health records data. J. Am. Med. Inform. Assoc. 20, 253–259 (2013).
Schank, R. C. & Tesler, L. in Proceedings of the 1969 Conference on Computational linguistics 1–3 (Association for Computational Linguistics, 1969).
Winograd, T. Procedures as a representation for data in a computer program for understanding natural language (Massachusetts Institute of Technology, 1971).
Schank, R. C. Computer understanding of natural language. Behav. Res. Methods Instrum. 10, 132–138 (1978).
Devlin, J., Chang, M.-W., Lee, K. & Toutanova, K. BERT: pre-training of deep bidirectional transformers for language understanding. Preprint at arXiv https://arxiv.org/abs/1810.04805 (2018).
Manning, C. et al. in Proceedings of 52nd Annual Meeting of the Association for Computational Linguistics: System Demonstrations 55–60 (Association for Computational Linguistics, 2014).
Honnibal, M. & Johnson, M. in Proceedings of the 2015 Conference on Empirical Methods in Natural Language Processing 1373–1378 (Association for Computational Linguistics, 2015).
Petrov, S. Announcing syntaxnet: the world’s most accurate parser goes open source. Google AI Blog https://ai.googleblog.com/2016/05/announcing-syntaxnet-worlds-most.html (2016).
Ford, E., Carroll, J. A., Smith, H. E., Scott, D. & Cassell, J. Extracting information from the text of electronic medical records to improve case detection: a systematic review. J. Am. Med. Inform. Assoc. 23, 1007–1015 (2016).
Weissenbacher, D. et al. in Proceedings of the 2016 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies 1198–1207 (Association for Computational Linguistics, 2016).
Grassi, M. et al. A novel ensemble-based machine learning algorithm to predict the conversion from mild cognitive impairment to Alzheimer’s disease using socio-demographic characteristics, clinical information, and neuropsychological measures. Front. Neurol. 10, 1–15 (2019).
Gordon, P. H. & Meininger, V. How can we improve clinical trials in amyotrophic lateral sclerosis? Nat. Rev. Neurol. 7, 650–654 (2011).
Moura, M. C., Casulari, L. A., Rita, M. & Garbi, C. A predictive model for prognosis in motor neuron disease. J. Neurol. Disord. 4, 4–10 (2016).
Westeneng, H.-J. et al. Prognosis for patients with amyotrophic lateral sclerosis: development and validation of a personalised prediction model. Lancet Neurol. 17, 423–433 (2018). This study shows how the application of machine learning to large clinical datasets from various clinical centres enables the prediction of disease prognosis in individuals with amyotrophic lateral sclerosis.
Latourelle, J. C. et al. Large-scale identification of clinical and genetic predictors of motor progression in patients with newly diagnosed Parkinson’s disease: a longitudinal cohort study and validation. Lancet Neurol. 16, 908–916 (2017). This study exemplifies how integration of large clinical, molecular and genetic longitudinal datasets can be used to provide information on disease progression in PD.
Wang, T., Qiu, R. G. & Yu, M. Predictive modeling of the progression of Alzheimer’s disease with recurrent neural networks. Sci. Rep. 8, 1–12 (2018).
Che, C. et al. in Proceedings of the 2017 SIAM International Conference on Data Mining 198–206 (SIAM, 2017).
Rajkomar, A. et al. Scalable and accurate deep learning for electronic health records. NPJ Digit. Med. 1, 1–10 (2018).
Fernandes, A. C. et al. Development and evaluation of a de-identification procedure for a case register sourced from mental health electronic records. BMC Med. Inform. Decis. Mak. 13, 1–14 (2013).
[No authors listed]. Stimulus package. Nat. Med. 24, 247 (2018).
Zwierzyna, M., Davies, M., Hingorani, A. D. & Hunter, J. Clinical trial design and dissemination: comprehensive analysis of clinicaltrials.gov and PubMed data since 2005. BMJ 361, 1–11 (2018).
Hay, M., Thomas, D. W., Craighead, J. L., Economides, C. & Rosenthal, J. Clinical development success rates for investigational drugs. Nat. Biotechnol. 32, 40–51 (2014).
Cummings, J. Lessons learned from Alzheimer disease: clinical trials with negative outcomes. Clin. Transl. Sci. 11, 147–152 (2018).
Cummings, J. L., Morstorf, T. & Zhong, K. Alzheimer’s disease drug-development pipeline: few candidates, frequent failures. Alzheimers. Res. Ther. 6, 1–7 (2014).
Mitsumoto, H., Brooks, B. R. & Silani, V. Clinical trials in amyotrophic lateral sclerosis: why so many negative trials and how can trials be improved? Lancet Neurol. 13, 1127–1138 (2014).
Ferraiuolo, L., Kirby, J., Grierson, A. J., Sendtner, M. & Shaw, P. J. Molecular pathways of motor neuron injury in amyotrophic lateral sclerosis. Nat. Rev. Neurol. 7, 616–630 (2011).
Neil, D. et al. Interpretable graph convolutional neural networks for inference on noisy knowledge graphs. Preprint at arXiv https://arxiv.org/abs/1812.00279 (2018).
Zitnik, M., Agrawal, M. & Leskovec, J. Modeling polypharmacy side effects with graph convolutional networks. Bioinformatics 34, i457–i466 (2018).
Duvenaud, D. et al. in NIPS’15: Proceedings of the 28th International Conference on Neural Information Processing Systems Vol. 2 2224–2232 (Neural Information Processing Systems Foundation, 2015).
Kipf, T. N. & Welling, M. Semi-supervised classification with graph convolutional networks. Preprint at arXiv https://arxiv.org/abs/1609.02907 (2017).
Palop, J. J., Chin, J. & Mucke, L. A network dysfunction perspective on neurodegenerative diseases. Nature 443, 768–773 (2006).
Zakeri, P., Simm, J., Arany, A., Elshal, S. & Moreau, Y. Gene prioritization using Bayesian matrix factorization with genomic and phenotypic side information. Bioinformatics 34, i447–i456 (2018).
Bakkar, N. et al. Artificial intelligence in neurodegenerative disease research: use of IBM Watson to identify additional RNA-binding proteins altered in amyotrophic lateral sclerosis. Acta Neuropathol. 135, 227–247 (2018).
Zhang, B. et al. Resource integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 153, 707–720 (2013). This study exemplifies how machine learning approaches applied to omics data can lead to identification of new therapeutic targets.
Haure-Mirande, J. V. et al. Deficiency of TYROBP, an adapter protein for TREM2 and CR3 receptors, is neuroprotective in a mouse model of early Alzheimer’s pathology. Acta Neuropathol. 134, 769–788 (2017).
Haure-Mirande, J. V. et al. Integrative approach to sporadic Alzheimer’s disease: deficiency of TYROBP in cerebral Aβ amyloidosis mouse normalizes clinical phenotype and complement subnetwork molecular pathology without reducing Aβ burden. Mol. Psychiatry 24, 431–446 (2019).
Wauters, E. et al. Neurobiology of aging clinical variability and onset age modifiers in an extended Belgian GRN founder family. Neurobiol. Aging 67, 84–94 (2018).
Grollemund, V. et al. Machine learning in amyotrophic lateral sclerosis: achievements, pitfalls, and future directions. Front. Neurosci. 13, 1–28 (2019).
Maudsley, S., Devanarayan, V., Martin, B. & Geerts, H. Intelligent and effective informatic deconvolution of ‘big data’ and its future impact on the quantitative nature of neurodegenerative disease therapy. Alzheimer’s Dement. 14, 961–975 (2018).
Meyer, S. et al. Optimizing ADAS-Cog worksheets: a survey of clinical trial raters’ perceptions. Curr. Alzheimer Res. 14, 1008–1016 (2017).
McDermott, J. E. et al. Challenges in biomarker discovery: combining expert insights with statistical analysis of complex omics data. Expert. Opin. Med. Diagn. 7, 37–51 (2013).
Popejoy, A. & Fullerton, S. Genomics is failing on diversity. Nature 538, 161–164 (2016).
Cohn, D. A., Ghahramani, Z. & Jordan, M. I. Active learning with statistical models. J. Artif. Intell. Res. 4, 129–145 (1996).
Sellwood, M. A., Ahmed, M., Segler, M. H. S. & Brown, N. Artificial intelligence in drug discovery. Future Med. Chem. 10, 2025–2028 (2018).
Gupta, A., Ayhan, M. S. & Maida, A. S. Natural image bases to represent neuroimaging data. PMLR 28, 987–994 (2013).
Xu, Y., Raj, A. & Victor, J. D. Systematic differences between perceptually relevant image statistics of brain MRI and natural images. Front. Neuroinform. 13, 1–15 (2019).
Marinescu, R. V. et al. in Medical Image Computing and Computer Assisted Intervention – MICCAI 2019. Lecture Notes in Computer Science Vol. 11765 (eds Shen, D. et al.) 860–868 (Springer, 2019).
Ganchev, P., Malehorn, D., Bigbee, W. L. & Gopalakrishnan, V. Transfer learning of classification rules for biomarker discovery and verification from molecular profiling studies. J. Biomed. Inform. 44, S17–S23 (2011).
Young, J. et al. Accurate multimodal probabilistic prediction of conversion to Alzheimer’s disease in patients with mild cognitive impairment. NeuroImage Clin. 19, 735–745 (2013).
Cheng, B. et al. Multi-domain transfer learning for early diagnosis of Alzheimer’s disease. Neuroinformatics 15, 115–132 (2017).
Hon, M. & Khan, N. Towards Alzheimer’s disease classification through transfer learning. Preprint at arXiv https://arxiv.org/abs/1711.11117 (2017).
Goodfellow, I. J. et al. Generative adversarial nets. Neural Inf. Process. Syst. 27, 1–9 (2014).
Huang, H., Yu, P. S. & Wang, C. An introduction to image synthesis with generative adversarial nets. Preprint at arXiv https://arxiv.org/abs/1803.04469 (2018).
Kazuhiro, K. et al. Generative adversarial networks for the creation of realistic artificial brain magnetic resonance images. Tomography 4, 159–163 (2018).
Palacio-Niño, J.-O. & Berzal, F. Evaluation metrics for unsupervised learning algorithms. Preprint at arXiv https://arxiv.org/abs/1905.05667 (2019).
Lötsch, J., Lerch, F., Djaldetti, R., Tegder, I. & Ultsch, A. Identification of disease-distinct complex biomarker patterns by means of unsupervised machine-learning using an interactive R toolbox (Umatrix). Big Data Anal. 3, 1–17 (2018).
Ravi, D. et al. Deep learning for health informatics. IEEE J. Biomed. Heal. Inform. 21, 4–21 (2017).
Vial, A. et al. The role of deep learning and radiomic feature extraction in cancer-specific predictive modelling: a review. Transl. Cancer Res. 7, 803–816 (2018).
Gulshan, V. et al. Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. JAMA 316, 2402–2410 (2016).
Gilpin, L. H. et al. Explaining explanations: an overview of interpretability of machine learning. Preprint at arXiv https://arxiv.org/abs/1806.00069 (2018).
Sarwar, S. et al. Physician perspectives on integration of artificial intelligence into diagnostic pathology. NPJ Digit. Med. 2, 28 (2019). This article reports the perspective of pathologists towards the integration of artificial intelligence into diagnostic pathology.
Fan, Y., Shen, D. & Davatzikos, C. in Lecture Notes in Computer Science, Vol. 3749 (eds Duncan, J. S. & Gerig, G.) 1–8 (Springer, 2005).
Shi, B., Chen, Y., Zhang, P., Smith, C. D. & Liu, J. Nonlinear feature transformation and deep fusion for Alzheimer’s disease staging analysis. Pattern Recognit. 63, 487–498 (2017).
Author information
Authors and Affiliations
Contributions
L.F. researched data for the article, made a substantial contribution to the discussion of article content, wrote the article, and reviewed and edited the manuscript before submission. M.A.M. researched data for the article, made a substantial contribution to the discussion of article content and wrote the article. P.N.O. and J.D.H. made a substantial contribution to discussion of article content, and reviewed and edited the manuscript before submission. A.M.B.L. and D.N. researched data for the article, and reviewed and edited the manuscript before submission. A.S., R.M. and G.M.H. reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
M.A.M. is funded by BenevolentAI. P.N.O., A.M.B.L., D.N., A.S. and J.D.H. work for BenevolentAI. R.M. and L.F. have a project in collaboration with BenevolentAI.
Additional information
Peer review information
Nature Reviews Neurology thanks S. Baranzini, D. Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Allen Brain Atlas: http://portal.brain-map.org/
Alzheimer’s Disease Neuroimaging Initiative: http://adni.loni.usc.edu/
Amazon Comprehend Medical initiative: https://aws.amazon.com/comprehend/medical/
Enhancing Neuro Imaging Genetics through Meta-Analysis (ENIGMA) consortium: http://enigma.ini.usc.edu/
European Alzheimer’s Disease Consortium Impact of Cholinergic Treatment Use (EADC-ICTUS): http://www.eadc.info/sito/pagine/d_01.php?nav=d
Google TensorFlow: https://www.tensorflow.org/
Parkinson’s Progression Markers Initiative: http://www.ppmi-info.org/
The Finnish Geriatric Intervention Study to Prevent Cognitive Impairment and Disability (FINGER): http://www.alz.org/wwfingers/overview.asp
UK Biobank: http://www.ukbiobank.ac.uk/
Glossary
- Omics
-
A collective term for a field within biological research concerned with the study of ‘omes’; for example, the genome, transcriptome or proteome.
- Endotypes
-
Clusters of individuals within a disease population that share functional and pathological traits.
- Molecular signatures
-
A collection of proteins, genes and their variants that can be used as hallmarks for a given phenotype.
- Regularizing
-
The technique of adding constraints or knowledge within the training process in order to prevent overfitting.
- Data leakage
-
An undesirable process whereby information is accidentally shared between the training data and the test data, resulting in test evaluation scores that are not representative of real-world unseen data.
- Sample-to-feature ratio
-
(SFR). The number of data points divided by the number of features; for example, gene expression data comprising tens of patients with thousands of gene expression levels would have an SFR of <1.
- Overfitting
-
When an algorithm learns the patterns within the training dataset as opposed to the patterns representative of all data.
- Classifier
-
A type of model used to identify the correct category for a data point.
- Hyperparameter
-
A parameter the value of which is set before training; for example, the attributes of the model architecture.
- Cross-validation
-
A training and evaluation procedure that consists of splitting the data into subsets and alternately holding out one subset for evaluation until all subsets have been evaluated.
- Metadata
-
Data about other data; for example, information about an experimental protocol or the time and date of sample collection.
- Genome-wide association study
-
(GWAS). An observational method of studying genetic variants across a population in search for associations between genetic changes and traits such as diseases.
- Next-generation sequencing
-
High-throughput, deep sequencing of DNA and RNA; this technique utilizes sequencing technologies that are capable of processing multiple DNA or RNA sequences in parallel.
- Metabolomics
-
A study of metabolites, that is, the small molecule substrates, intermediates and products of cellular metabolism, and their interactions within living organisms.
- Bayesian inference
-
A method of statistical inference that uses Bayes’ theorem to calculate the probability of a hypothesis being true on the basis of observed data and prior information.
Rights and permissions
About this article
Cite this article
Myszczynska, M.A., Ojamies, P.N., Lacoste, A.M.B. et al. Applications of machine learning to diagnosis and treatment of neurodegenerative diseases. Nat Rev Neurol 16, 440–456 (2020). https://doi.org/10.1038/s41582-020-0377-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41582-020-0377-8
This article is cited by
-
Comprehensive analysis of mitochondria-related genes indicates that PPP2R2B is a novel biomarker and promotes the progression of bladder cancer via Wnt signaling pathway
Biology Direct (2024)
-
Immunological aspects of central neurodegeneration
Cell Discovery (2024)
-
Prediction of xerostomia in elderly based on clinical characteristics and salivary flow rate with machine learning
Scientific Reports (2024)
-
A sustainable approach to universal metabolic cancer diagnosis
Nature Sustainability (2024)
-
Discovery of potent inhibitors of α-synuclein aggregation using structure-based iterative learning
Nature Chemical Biology (2024)