Abstract
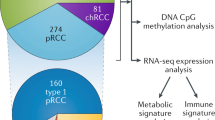
The molecular features that define clear cell renal cell carcinoma (ccRCC) initiation and progression are being increasingly defined. The TRACERx Renal studies and others that have described the interaction between tumour genomics and remodelling of the tumour microenvironment provide important new insights into the molecular drivers underlying ccRCC ontogeny and progression. Our understanding of common genomic and chromosomal copy number abnormalities in ccRCC, including chromosome 3p loss, provides a mechanistic framework with which to organize these abnormalities into those that drive tumour initiation events, those that drive tumour progression and those that confer lethality. Truncal mutations in ccRCC, including those in VHL, SET2, PBRM1 and BAP1, may engender genomic instability and promote defects in DNA repair pathways. The molecular features that arise from these defects enable categorization of ccRCC into clinically and therapeutically relevant subtypes. Consideration of the interaction of these subtypes with the tumour microenvironment reveals that specific mutations seem to modulate immune cell populations in ccRCC tumours. These findings present opportunities for disease prevention, early detection, prognostication and treatment.
Key points
-
Chromosome 3p loss is an almost universal finding in both hereditary and sporadic clear cell renal cell carcinoma (ccRCC).
-
The near ubiquitous loss of a second copy of VHL seems to provide a selective advantage for cells, as well as leading to defects in DNA repair and an increase in genomic instability.
-
Secondarily mutated genes in ccRCC, including PBRM1, SETD2 and BAP1, as well as copy number changes in chromosomes 9p and 14q, are associated with prognostically important molecular and phenotypic characteristics that can be used to classify tumours into subgroups.
-
Tumour genomic features are associated with distinct immune phenotypes; for example, PBRM1 mutations are associated with reduced infiltration of T cells.
-
Efforts are underway to link genomic features to specific therapeutic strategies for patients with ccRCC.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Jonasch, E., Gao, J. & Rathmell, W. K. Renal cell carcinoma. BMJ 349, g4797 (2014).
Hagemeijer, A., Hoehn, W. & Smit, E. M. Cytogenetic analysis of human renal carcinoma cell lines of common origin (NC 65). Cancer Res. 39, 4662–4667 (1979).
Szucs, S., Muller-Brechlin, R., DeRiese, W. & Kovacs, G. Deletion 3p: the only chromosome loss in a primary renal cell carcinoma. Cancer Genet. Cytogenet. 26, 369–373 (1987).
Yoshida, M. A. et al. Rearrangement of chromosome 3 in renal cell carcinoma. Cancer Genet. Cytogenet. 19, 351–354 (1986).
Latif, F. et al. Identification of the von Hippel-Lindau disease tumor suppressor gene. Science 260, 1317–1320 (1993).
Varela, I. et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469, 539–542 (2011).
Pena-Llopis, S. et al. BAP1 loss defines a new class of renal cell carcinoma. Nat. Genet. 44, 751–759 (2012).
Dalgliesh, G. L. et al. Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature 463, 360–363 (2010).
The Cancer Genome Atlas Research Network. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499, 43–49 (2013).
Klatte, T. et al. Cytogenetic profile predicts prognosis of patients with clear cell renal cell carcinoma. J. Clin. Oncol. 27, 746–753 (2009).
La Rochelle, J. et al. Chromosome 9p deletions identify an aggressive phenotype of clear cell renal cell carcinoma. Cancer 116, 4696–4702 (2010).
Monzon, F. A. et al. Chromosome 14q loss defines a molecular subtype of clear-cell renal cell carcinoma associated with poor prognosis. Mod. Pathol. 24, 1470–1479 (2011).
Shen, C. et al. Genetic and functional studies implicate HIF1alpha as a 14q kidney cancer suppressor gene. Cancer Discov. 1, 222–235 (2011).
Jiang, F. et al. Construction of evolutionary tree models for renal cell carcinoma from comparative genomic hybridization data. Cancer Res. 60, 6503–6509 (2000).
Brannon, A. R. et al. Molecular stratification of clear cell renal cell carcinoma by consensus clustering reveals distinct subtypes and survival patterns. Genes Cancer 1, 152–163 (2010).
Rini, B. et al. A 16-gene assay to predict recurrence after surgery in localised renal cell carcinoma: development and validation studies. Lancet Oncol. 16, 676–685 (2015).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Turajlic, S. et al. Deterministic evolutionary trajectories influence primary tumor growth: TRACERx Renal. Cell 173, 595–610.e11 (2018).
Mitchell, T. J. et al. Timing the landmark events in the evolution of clear cell renal cell cancer: TRACERx Renal. Cell 173, 611–623.e17 (2018).
Turajlic, S. et al. Tracking cancer evolution reveals constrained routes to metastases: TRACERx Renal. Cell 173, 581–594.e12 (2018).
Monzon, F. A. et al. Whole genome SNP arrays as a potential diagnostic tool for the detection of characteristic chromosomal aberrations in renal epithelial tumors. Mod. Pathol. 21, 599–608 (2008).
Wang, L. et al. Whole-exome sequencing of human pancreatic cancers and characterization of genomic instability caused by MLH1 haploinsufficiency and complete deficiency. Genome Res. 22, 208–219 (2012).
Chiang, Y. C. et al. SETD2 Haploinsufficiency for microtubule methylation is an early driver of genomic instability in renal cell carcinoma. Cancer Res. 78, 3135–3146 (2018).
Fei, S. S. et al. Patient-specific factors influence somatic variation patterns in von Hippel-Lindau disease renal tumours. Nat. Commun. 7, 11588 (2016).
Bakhoum, S. F. & Cantley, L. C. The multifaceted role of chromosomal instability in cancer and its microenvironment. Cell 174, 1347–1360 (2018).
Kalsbeek, D. & Golsteyn, R. M. G2/M-Phase checkpoint adaptation and micronuclei formation as mechanisms that contribute to genomic instability in human cells. Int. J. Mol. Sci. 18, 2344 (2017).
Podrimaj-Bytyqi, A. et al. The frequencies of micronuclei, nucleoplasmic bridges and nuclear buds as biomarkers of genomic instability in patients with urothelial cell carcinoma. Sci. Rep. 8, 17873 (2018).
Lonser, R. R. et al. Von Hippel-Lindau disease. Lancet 361, 2059–2067 (2003).
Ho, T. H. & Jonasch, E. Genetic kidney cancer syndromes. J. Natl Compr. Canc. Netw. 12, 1347–1355 (2014).
Nickerson, M. L. et al. Improved identification of von Hippel-Lindau gene alterations in clear cell renal tumors. Clin. Cancer Res. 14, 4726–4734 (2008).
Maxwell, P. H. et al. Hypoxia-inducible factor-1 modulates gene expression in solid tumors and influences both angiogenesis and tumor growth. Proc. Natl Acad. Sci. USA 94, 8104–8109 (1997).
Flamme, I., Krieg, M. & Plate, K. H. Up-regulation of vascular endothelial growth factor in stromal cells of hemangioblastomas is correlated with up-regulation of the transcription factor HRF/HIF-2alpha. Am. J. Pathol. 153, 25–29 (1998).
Krieg, M. et al. Up-regulation of hypoxia-inducible factors HIF-1alpha and HIF-2alpha under normoxic conditions in renal carcinoma cells by von Hippel-Lindau tumor suppressor gene loss of function. Oncogene 19, 5435–5443 (2000).
Maxwell, P. H. et al. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 399, 271–275 (1999).
Keith, B., Johnson, R. S. & Simon, M. C. HIF1alpha and HIF2alpha: sibling rivalry in hypoxic tumour growth and progression. Nat. Rev. Cancer 12, 9–22 (2011).
Takahashi, A. et al. Markedly increased amounts of messenger RNAs for vascular endothelial growth factor and placenta growth factor in renal cell carcinoma associated with angiogenesis. Cancer Res. 54, 4233–4237 (1994).
Hu, C. J., Wang, L. Y., Chodosh, L. A., Keith, B. & Simon, M. C. Differential roles of hypoxia-inducible factor 1alpha (HIF-1alpha) and HIF-2alpha in hypoxic gene regulation. Mol. Cell Biol. 23, 9361–9374 (2003).
Raval, R. R. et al. Contrasting properties of hypoxia-inducible factor 1 (HIF-1) and HIF-2 in von Hippel-Lindau-associated renal cell carcinoma. Mol. Cell Biol. 25, 5675–5686 (2005).
Carroll, V. A. & Ashcroft, M. Role of hypoxia-inducible factor (HIF)-1alpha versus HIF-2alpha in the regulation of HIF target genes in response to hypoxia, insulin-like growth factor-I, or loss of von Hippel-Lindau function: implications for targeting the HIF pathway. Cancer Res. 66, 6264–6270 (2006).
Kim, J. W., Tchernyshyov, I., Semenza, G. L. & Dang, C. V. HIF-1-mediated expression of pyruvate dehydrogenase kinase: a metabolic switch required for cellular adaptation to hypoxia. Cell Metab. 3, 177–185 (2006).
Semenza, G. L. et al. Hypoxia response elements in the aldolase A, enolase 1, and lactate dehydrogenase A gene promoters contain essential binding sites for hypoxia-inducible factor 1. J. Biol. Chem. 271, 32529–32537 (1996).
Zhang, H. et al. Mitochondrial autophagy is an HIF-1-dependent adaptive metabolic response to hypoxia. J. Biol. Chem. 283, 10892–10903 (2008).
Bellot, G. et al. Hypoxia-induced autophagy is mediated through hypoxia-inducible factor induction of BNIP3 and BNIP3L via their BH3 domains. Mol. Cell Biol. 29, 2570–2581 (2009).
O’Bryan, J. P. et al. axl, a transforming gene isolated from primary human myeloid leukemia cells, encodes a novel receptor tyrosine kinase. Mol. Cell Biol. 11, 5016–5031 (1991).
Lemke, G. Biology of the TAM receptors. Cold Spring Harb. Perspect. Biol. 5, a009076 (2013).
Lai, C., Gore, M. & Lemke, G. Structure, expression, and activity of Tyro 3, a neural adhesion-related receptor tyrosine kinase. Oncogene 9, 2567–2578 (1994).
Rankin, E. B. et al. Direct regulation of GAS6/AXL signaling by HIF promotes renal metastasis through SRC and MET. Proc. Natl Acad. Sci. USA 111, 13373–13378 (2014).
Zhou, L. et al. Targeting MET and AXL overcomes resistance to sunitinib therapy in renal cell carcinoma. Oncogene 35, 2687–2697 (2015).
Rankin, E. B. et al. Inactivation of the arylhydrocarbon receptor nuclear translocator (Arnt) suppresses von Hippel-Lindau disease-associated vascular tumors in mice. Mol. Cell Biol. 25, 3163–3172 (2005).
Fu, L., Wang, G., Shevchuk, M. M., Nanus, D. M. & Gudas, L. J. Generation of a mouse model of Von Hippel-Lindau kidney disease leading to renal cancers by expression of a constitutively active mutant of HIF1alpha. Cancer Res. 71, 6848–6856 (2011).
Fu, L., Wang, G., Shevchuk, M. M., Nanus, D. M. & Gudas, L. J. Activation of HIF2alpha in kidney proximal tubule cells causes abnormal glycogen deposition but not tumorigenesis. Cancer Res. 73, 2916–2925 (2013).
Schietke, R. E. et al. Renal tubular HIF-2alpha expression requires VHL inactivation and causes fibrosis and cysts. PLoS One 7, e31034 (2012).
Schonenberger, D. et al. Formation of renal cysts and tumors in vhl/trp53-deficient mice requires HIF1alpha and HIF2alpha. Cancer Res. 76, 2025–2036 (2016).
Zhang, J. & Zhang, Q. VHL and hypoxia signaling: beyond HIF in cancer. Biomedicines 6, 35 (2018).
Guo, J. et al. pVHL suppresses kinase activity of Akt in a proline-hydroxylation-dependent manner. Science 353, 929–932 (2016).
An, J. & Rettig, M. B. Mechanism of von Hippel-Lindau protein-mediated suppression of nuclear factor kappa B activity. Mol. Cell Biol. 25, 7546–7556 (2005).
Yang, H. et al. pVHL acts as an adaptor to promote the inhibitory phosphorylation of the NF-kappaB agonist Card9 by CK2. Mol. Cell 28, 15–27 (2007).
Peri, S., Devarajan, K., Yang, D. H., Knudson, A. G. & Balachandran, S. Meta-analysis identifies NF-kappaB as a therapeutic target in renal cancer. PLoS One 8, e76746 (2013).
Hu, L. et al. TBK1 is a synthetic lethal target in cancer with VHL loss. Cancer Discov. 10, 460–475 (2020).
Metcalf, J. L. et al. K63-ubiquitylation of VHL by SOCS1 mediates DNA double-strand break repair. Oncogene 33, 1055–1065 (2014).
Scanlon, S. E., Hegan, D. C., Sulkowski, P. L. & Glazer, P. M. Suppression of homology-dependent DNA double-strand break repair induces PARP inhibitor sensitivity in VHL-deficient human renal cell carcinoma. Oncotarget 9, 4647–4660 (2018).
Espana-Agusti, J., Warren, A., Chew, S. K., Adams, D. J. & Matakidou, A. Loss of PBRM1 rescues VHL dependent replication stress to promote renal carcinogenesis. Nat. Commun. 8, 2026 (2017).
Zhou, L. et al. Abstract: Tip60 dependent DNA homologous recombination repair is impaired in VHL-deficient clear cell renal cell carcinoma. Cancer Res. 78, 1364 (2018).
Zhang, J. et al. VHL substrate transcription factor ZHX2 as an oncogenic driver in clear cell renal cell carcinoma. Science 361, 290–295 (2018).
Calzada, M. J. et al. Von Hippel-Lindau tumor suppressor protein regulates the assembly of intercellular junctions in renal cancer cells through hypoxia-inducible factor-independent mechanisms. Cancer Res. 66, 1553–1560 (2006).
Stickle, N. H. et al. pVHL modification by NEDD8 is required for fibronectin matrix assembly and suppression of tumor development. Mol. Cell Biol. 24, 3251–3261 (2004).
Ohh, M. et al. The von Hippel-Lindau tumor suppressor protein is required for proper assembly of an extracellular fibronectin matrix. Mol. Cell 1, 959–968 (1998).
Kurban, G. et al. Collagen matrix assembly is driven by the interaction of von Hippel-Lindau tumor suppressor protein with hydroxylated collagen IV alpha 2. Oncogene 27, 1004–1012 (2008).
Kurban, G., Hudon, V., Duplan, E., Ohh, M. & Pause, A. Characterization of a von Hippel Lindau pathway involved in extracellular matrix remodeling, cell invasion, and angiogenesis. Cancer Res. 66, 1313–1319 (2006).
Clifford, S. C. & Maher, E. R. Von Hippel-Lindau disease: clinical and molecular perspectives. Adv. Cancer Res. 82, 85–105 (2001).
Hoffman, M. A. et al. Von Hippel-Lindau protein mutants linked to type 2C VHL disease preserve the ability to downregulate HIF. Hum. Mol. Genet. 10, 1019–1027 (2001).
Tang, N., Mack, F., Haase, V. H., Simon, M. C. & Johnson, R. S. pVHL function is essential for endothelial extracellular matrix deposition. Mol. Cell Biol. 26, 2519–2530 (2006).
Lolkema, M. P. et al. Tumor suppression by the von Hippel-Lindau protein requires phosphorylation of the acidic domain. J. Biol. Chem. 280, 22205–22211 (2005).
German, P. et al. Phosphorylation-dependent cleavage regulates von Hippel Lindau proteostasis and function. Oncogene 35, 4973–4980 (2016).
Hergovich, A., Lisztwan, J., Barry, R., Ballschmieter, P. & Krek, W. Regulation of microtubule stability by the von Hippel-Lindau tumour suppressor protein pVHL. Nat. Cell Biol. 5, 64–70 (2003).
Schermer, B. et al. The von Hippel-Lindau tumor suppressor protein controls ciliogenesis by orienting microtubule growth. J. Cell Biol. 175, 547–554 (2006).
Esteban, M. A., Harten, S. K., Tran, M. G. & Maxwell, P. H. Formation of primary cilia in the renal epithelium is regulated by the von Hippel-Lindau tumor suppressor protein. J. Am. Soc. Nephrol. 17, 1801–1806 (2006).
Lutz, M. S. & Burk, R. D. Primary cilium formation requires von Hippel-Lindau gene function in renal-derived cells. Cancer Res. 66, 6903–6907 (2006).
Devlin, L. A. & Sayer, J. A. Renal ciliopathies. Curr. Opin. Genet. Dev. 56, 49–60 (2019).
Hasanov, E. et al. Ubiquitination and regulation of AURKA identifies a hypoxia-independent E3 ligase activity of VHL. Oncogene 36, 3450–3463 (2017).
Thoma, C. R. et al. VHL loss causes spindle misorientation and chromosome instability. Nat. Cell Biol. 11, 994–1001 (2009).
Hell, M. P., Duda, M., Weber, T. C., Moch, H. & Krek, W. Tumor suppressor VHL functions in the control of mitotic fidelity. Cancer Res. 74, 2422–2431 (2014).
Hell, M. P. et al. miR-28-5p promotes chromosomal instability in VHL-associated cancers by inhibiting Mad2 translation. Cancer Res. 74, 2432–2443 (2014).
Janku, F., Yap, T. A. & Meric-Bernstam, F. Targeting the PI3K pathway in cancer: are we making headway? Nat. Rev. Clin. Oncol. 15, 273–291 (2018).
Horiguchi, A., Oya, M., Uchida, A., Marumo, K. & Murai, M. Elevated Akt activation and its impact on clinicopathological features of renal cell carcinoma. J. Urol. 169, 710–713 (2003).
Sourbier, C. et al. The phosphoinositide 3-kinase/Akt pathway: a new target in human renal cell carcinoma therapy. Cancer Res. 66, 5130–5142 (2006).
Yang, J. et al. Targeting PI3K in cancer: mechanisms and advances in clinical trials. Mol. Cancer 18, 26 (2019).
Kim, J. et al. Cytoplasmic sequestration of p27 via AKT phosphorylation in renal cell carcinoma. Clin. Cancer Res. 15, 81–90 (2009).
Haase, V. H., Glickman, J. N., Socolovsky, M. & Jaenisch, R. Vascular tumors in livers with targeted inactivation of the von Hippel-Lindau tumor suppressor. Proc. Natl Acad. Sci. USA 98, 1583–1588 (2001).
Hickey, M. M., Lam, J. C., Bezman, N. A., Rathmell, W. K. & Simon, M. C. Von Hippel-Lindau mutation in mice recapitulates Chuvash polycythemia via hypoxia-inducible factor-2alpha signaling and splenic erythropoiesis. J. Clin. Invest. 117, 3879–3889 (2007).
Lee, C. M. et al. VHL Type 2B gene mutation moderates HIF dosage in vitro and in vivo. Oncogene 28, 1694–1705 (2009).
Gnarra, J. R. et al. Defective placental vasculogenesis causes embryonic lethality in VHL-deficient mice. Proc. Natl Acad. Sci. USA 94, 9102–9107 (1997).
Frew, I. J. & Moch, H. A clearer view of the molecular complexity of clear cell renal cell carcinoma. Annu. Rev. Pathol. 10, 263–289 (2015).
Rankin, E. B., Tomaszewski, J. E. & Haase, V. H. Renal cyst development in mice with conditional inactivation of the von Hippel-Lindau tumor suppressor. Cancer Res. 66, 2576–2583 (2006).
Mathia, S. et al. Action of hypoxia-inducible factor in liver and kidney from mice with Pax8-rtTA-based deletion of von Hippel-Lindau protein. Acta Physiol. 207, 565–576 (2013).
Schley, G. et al. Hypoxia-inducible transcription factors stabilization in the thick ascending limb protects against ischemic acute kidney injury. J. Am. Soc. Nephrol. 22, 2004–2015 (2011).
Lubensky, I. A. et al. Allelic deletions of the VHL gene detected in multiple microscopic clear cell renal lesions in von Hippel-Lindau disease patients. Am. J. Pathol. 149, 2089–2094 (1996).
Paraf, F. et al. Renal lesions in von Hippel-Lindau disease: immunohistochemical expression of nephron differentiation molecules, adhesion molecules and apoptosis proteins. Histopathology 36, 457–465 (2000).
Albers, J. et al. Combined mutation of Vhl and Trp53 causes renal cysts and tumours in mice. EMBO Mol. Med. 5, 949–964 (2013).
Frew, I. J. et al. pVHL and PTEN tumour suppressor proteins cooperatively suppress kidney cyst formation. EMBO J. 27, 1747–1757 (2008).
Harlander, S. et al. Combined mutation in Vhl, Trp53 and Rb1 causes clear cell renal cell carcinoma in mice. Nat. Med. 23, 869–877 (2017).
Gu, Y. F. et al. Modeling renal cell carcinoma in mice: Bap1 and Pbrm1 inactivation drive tumor grade. Cancer Discov. 7, 900–917 (2017).
Nargund, A. M. et al. The SWI/SNF protein PBRM1 restrains VHL-loss-driven clear cell renal cell carcinoma. Cell Rep. 18, 2893–2906 (2017).
Lee, H. et al. BAF180 regulates cellular senescence and hematopoietic stem cell homeostasis through p21. Oncotarget 7, 19134–19146 (2016).
de Cubas, A. A. & Rathmell, W. K. Epigenetic modifiers: activities in renal cell carcinoma. Nat. Rev. Urol. 15, 599–614 (2018).
Thompson, M. Polybromo-1: the chromatin targeting subunit of the PBAF complex. Biochimie 91, 309–319 (2009).
Pulice, J. L. & Kadoch, C. Composition and function of mammalian SWI/SNF Chromatin remodeling complexes in human disease. Cold Spring Harb. Symp. Quant. Biol. 81, 53–60 (2016).
Kim, S. H. et al. The prognostic value of BAP1, PBRM1, pS6, PTEN, TGase2, PD-L1, CA9, PSMA, and Ki-67 tissue markers in localized renal cell carcinoma: a retrospective study of tissue microarrays using immunohistochemistry. PLoS One 12, e0179610 (2017).
Wang, Z. et al. Sarcomatoid renal cell carcinoma has a distinct molecular pathogenesis, driver mutation profile, and transcriptional landscape. Clin. Cancer Res. 23, 6686–6696 (2017).
Gao, W., Li, W., Xiao, T., Liu, X. S. & Kaelin, W. G. Jr. Inactivation of the PBRM1 tumor suppressor gene amplifies the HIF-response in VHL−/− clear cell renal carcinoma. Proc. Natl Acad. Sci. USA 114, 1027–1032 (2017).
Liao, L. et al. Multiple tumor suppressors regulate a HIF-dependent negative feedback loop via ISGF3 in human clear cell renal cancer. eLife 7, e37925 (2018).
Liu, X. D. et al. PBRM1 loss defines a nonimmunogenic tumor phenotype associated with checkpoint inhibitor resistance in renal carcinoma. Nat. Commun. 11, 2135 (2020).
Cai, W. et al. PBRM1 acts as a p53 lysine-acetylation reader to suppress renal tumor growth. Nat. Commun. 10, 5800 (2019).
Xue, Y. et al. The human SWI/SNF-B chromatin-remodeling complex is related to yeast rsc and localizes at kinetochores of mitotic chromosomes. Proc. Natl Acad. Sci. USA 97, 13015–13020 (2000).
Huang, J., Hsu, J. M. & Laurent, B. C. The RSC nucleosome-remodeling complex is required for Cohesin’s association with chromosome arms. Mol. Cell 13, 739–750 (2004).
Baetz, K. K., Krogan, N. J., Emili, A., Greenblatt, J. & Hieter, P. The ctf13-30/CTF13 genomic haploinsufficiency modifier screen identifies the yeast chromatin remodeling complex RSC, which is required for the establishment of sister chromatid cohesion. Mol. Cell Biol. 24, 1232–1244 (2004).
Brownlee, P. M., Chambers, A. L., Cloney, R., Bianchi, A. & Downs, J. A. BAF180 promotes cohesion and prevents genome instability and aneuploidy. Cell Rep. 6, 973–981 (2014).
Clark, D. J. et al. Integrated proteogenomic characterization of clear cell renal cell carcinoma. Cell 179, 964–983.e31 (2019).
Hakimi, A. A. et al. Transcriptomic profiling of the tumor microenvironment reveals distinct subgroups of clear cell renal cell cancer — data from a randomized phase III trial. Cancer Discov. 9, 510–525 (2019).
Jensen, D. E. et al. BAP1: a novel ubiquitin hydrolase which binds to the BRCA1 RING finger and enhances BRCA1-mediated cell growth suppression. Oncogene 16, 1097–1112 (1998).
Kapur, P. et al. Effects on survival of BAP1 and PBRM1 mutations in sporadic clear-cell renal-cell carcinoma: a retrospective analysis with independent validation. Lancet Oncol. 14, 159–167 (2013).
Scheuermann, J. C. et al. Histone H2A deubiquitinase activity of the Polycomb repressive complex PR-DUB. Nature 465, 243–247 (2010).
Endoh, M. et al. Histone H2A mono-ubiquitination is a crucial step to mediate PRC1-dependent repression of developmental genes to maintain ES cell identity. PLoS Genet. 8, e1002774 (2012).
Dey, A. et al. Loss of the tumor suppressor BAP1 causes myeloid transformation. Science 337, 1541–1546 (2012).
Cejas, P. et al. Enhancer signatures stratify and predict outcomes of non-functional pancreatic neuroendocrine tumors. Nat. Med. 25, 1260–1265 (2019).
Peng, J. et al. Stabilization of MCRS1 by BAP1 prevents chromosome instability in renal cell carcinoma. Cancer Lett. 369, 167–174 (2015).
Kobayashi, A. et al. Six2 defines and regulates a multipotent self-renewing nephron progenitor population throughout mammalian kidney development. Cell Stem Cell 3, 169–181 (2008).
Wang, S. S. et al. Bap1 is essential for kidney function and cooperates with Vhl in renal tumorigenesis. Proc. Natl Acad. Sci. USA 111, 16538–16543 (2014).
Joseph, R. W. et al. Loss of BAP1 protein expression is an independent marker of poor prognosis in patients with low-risk clear cell renal cell carcinoma. Cancer 120, 1059–1067 (2014).
Hsieh, J. J. et al. Genomic biomarkers of a randomized trial comparing first-line everolimus and sunitinib in patients with metastatic renal cell carcinoma. Eur. Urol. 71, 405–414 (2017).
Farley, M. N. et al. A novel germline mutation in BAP1 predisposes to familial clear-cell renal cell carcinoma. Mol. Cancer Res. 11, 1061–1071 (2013).
Lickwar, C. R. et al. The Set2/Rpd3S pathway suppresses cryptic transcription without regard to gene length or transcription frequency. PLoS One 4, e4886 (2009).
Hacker, K. E. et al. Structure/function analysis of recurrent mutations in SETD2 protein reveals a critical and conserved role for a SET domain residue in maintaining protein stability and histone H3 Lys-36 trimethylation. J. Biol. Chem. 291, 21283–21295 (2016).
Fahey, C. C. & Davis, I. J. SETting the stage for cancer development: SETD2 and the consequences of lost methylation. Cold Spring Harb. Perspect. Med. 7, a026468 (2017).
Park, I. Y. et al. Dual chromatin and cytoskeletal remodeling by SETD2. Cell 166, 950–962 (2016).
Giaccia, A. J. A new chromatin-cytoskeleton link in cancer. Mol. Cancer Res. 14, 1173–1175 (2016).
Seervai, R. N. H. et al. SETD2 is an actin lysine methyltransferase. Preprint at bioRxiv https://doi.org/10.1101/2020.04.13.034629 (2020).
Karki, M. et al. A cytoskeletal function for PBRM1 reading methylated microtubules. Preprint at bioRxiv https://doi.org/10.1101/2020.04.21.053942 (2020).
Pilie, P. G., Tang, C., Mills, G. B. & Yap, T. A. State-of-the-art strategies for targeting the DNA damage response in cancer. Nat. Rev. Clin. Oncol. 16, 81–104 (2019).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Dere, R., Perkins, A. L., Bawa-Khalfe, T., Jonasch, D. & Walker, C. L. β-catenin links von Hippel-Lindau to aurora kinase A and loss of primary cilia in renal cell carcinoma. J. Am. Soc. Nephrol. 26, 553–564 (2014).
Zhang, M. et al. HDAC6 deacetylates and ubiquitinates MSH2 to maintain proper levels of MutSalpha. Mol. Cell 55, 31–46 (2014).
Li, F. et al. The histone mark H3K36me3 regulates human DNA mismatch repair through its interaction with MutSalpha. Cell 153, 590–600 (2013).
Naqvi, R. A. et al. Hypermethylation analysis of mismatch repair genes (hmlh1 and hmsh2) in locally advanced breast cancers in Indian women. Hum. Pathol. 39, 672–680 (2008).
Altavilla, G., Fassan, M., Busatto, G., Orsolan, M. & Giacomelli, L. Microsatellite instability and hMLH1 and hMSH2 expression in renal tumors. Oncol. Rep. 24, 927–932 (2010).
Jasin, M. & Rothstein, R. Repair of strand breaks by homologous recombination. Cold Spring Harb. Perspect. Biol. 5, a012740 (2013).
Hall, J. M. et al. Linkage of early-onset familial breast cancer to chromosome 17q21. Science 250, 1684–1689 (1990).
Hall, J. M. et al. Closing in on a breast cancer gene on chromosome 17q. Am. J. Hum. Genet. 50, 1235–1242 (1992).
Wooster, R. et al. Identification of the breast cancer susceptibility gene BRCA2. Nature 378, 789–792 (1995).
Antoniou, A. C. et al. Breast-cancer risk in families with mutations in PALB2. N. Engl. J. Med. 371, 497–506 (2014).
Swift, M., Morrell, D., Massey, R. B. & Chase, C. L. Incidence of cancer in 161 families affected by ataxia-telangiectasia. N. Engl. J. Med. 325, 1831–1836 (1991).
Bindra, R. S. et al. Down-regulation of Rad51 and decreased homologous recombination in hypoxic cancer cells. Mol. Cell Biol. 24, 8504–8518 (2004).
Bindra, R. S. et al. Hypoxia-induced down-regulation of BRCA1 expression by E2Fs. Cancer Res. 65, 11597–11604 (2005).
Scanlon, S. E. & Glazer, P. M. Hypoxic stress facilitates acute activation and chronic downregulation of fanconi anemia proteins. Mol. Cancer Res. 12, 1016–1028 (2014).
Roe, J. S. et al. Phosphorylation of von Hippel-Lindau protein by checkpoint kinase 2 regulates p53 transactivation. Cell Cycle 10, 3920–3928 (2011).
Tang, J. et al. Acetylation limits 53BP1 association with damaged chromatin to promote homologous recombination. Nat. Struct. Mol. Biol. 20, 317–325 (2013).
Sun, Y., Jiang, X., Chen, S., Fernandes, N. & Price, B. D. A role for the Tip60 histone acetyltransferase in the acetylation and activation of ATM. Proc. Natl Acad. Sci. USA 102, 13182–13187 (2005).
Pfister, S. X. et al. SETD2-dependent histone H3K36 trimethylation is required for homologous recombination repair and genome stability. Cell Rep. 7, 2006–2018 (2014).
Carvalho, S. et al. SETD2 is required for DNA double-strand break repair and activation of the p53-mediated checkpoint. eLife 3, e02482 (2014).
Aymard, F. et al. Transcriptionally active chromatin recruits homologous recombination at DNA double-strand breaks. Nat. Struct. Mol. Biol. 21, 366–374 (2014).
Kanu, N. et al. SETD2 loss-of-function promotes renal cancer branched evolution through replication stress and impaired DNA repair. Oncogene 34, 5699–5708 (2015).
Li, L. & Wang, Y. Cross-talk between the H3K36me3 and H4K16ac histone epigenetic marks in DNA double-strand break repair. J. Biol. Chem. 292, 11951–11959 (2017).
Ismail, I. H. et al. Germline mutations in BAP1 impair its function in DNA double-strand break repair. Cancer Res. 74, 4282–4294 (2014).
Peng, G. et al. Genome-wide transcriptome profiling of homologous recombination DNA repair. Nat. Commun. 5, 3361 (2014).
Pilie, P. G. et al. Homologous repair deficiency in VHL-mutated clear cell renal cell carcinoma. J. Clin. Oncol. 36, 585 (2018).
Carter, S. L., Eklund, A. C., Kohane, I. S., Harris, L. N. & Szallasi, Z. A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat. Genet. 38, 1043–1048 (2006).
McGranahan, N., Burrell, R. A., Endesfelder, D., Novelli, M. R. & Swanton, C. Cancer chromosomal instability: therapeutic and diagnostic challenges. EMBO Rep. 13, 528–538 (2012).
Gordan, J. D. et al. HIF-alpha effects on c-Myc distinguish two subtypes of sporadic VHL-deficient clear cell renal carcinoma. Cancer Cell 14, 435–446 (2008).
Wolfer, A. et al. MYC regulation of a “poor-prognosis” metastatic cancer cell state. Proc. Natl Acad. Sci. USA 107, 3698–3703 (2010).
Jamal-Hanjani, M., Quezada, S. A., Larkin, J. & Swanton, C. Translational implications of tumor heterogeneity. Clin. Cancer Res. 21, 1258–1266 (2015).
Malouf, G. G. et al. Genomic characterization of renal cell carcinoma with sarcomatoid dedifferentiation pinpoints recurrent genomic alterations. Eur. Urol. 70, 348–357 (2016).
Bi, M. et al. Genomic characterization of sarcomatoid transformation in clear cell renal cell carcinoma. Proc. Natl Acad. Sci. USA 113, 2170–2175 (2016).
Hinshaw, D. C. & Shevde, L. A. The tumor microenvironment innately modulates cancer progression. Cancer Res. 79, 4557–4566 (2019).
Chevrier, S. et al. An immune atlas of clear cell renal cell carcinoma. Cell 169, 736–749 e718 (2017).
Senbabaoglu, Y. et al. Tumor immune microenvironment characterization in clear cell renal cell carcinoma identifies prognostic and immunotherapeutically relevant messenger RNA signatures. Genome Biol. 17, 231 (2016).
Ko, J. S. et al. Sunitinib mediates reversal of myeloid-derived suppressor cell accumulation in renal cell carcinoma patients. Clin. Cancer Res. 15, 2148–2157 (2009).
Wang, T. et al. An empirical approach leveraging tumorgrafts to dissect the tumor microenvironment in renal cell carcinoma identifies missing link to prognostic inflammatory factors. Cancer Discov. 8, 1142–1155 (2018).
Miao, D. et al. Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science 359, 801–806 (2018).
McDermott, D. F. et al. Clinical activity and molecular correlates of response to atezolizumab alone or in combination with bevacizumab versus sunitinib in renal cell carcinoma. Nat. Med. 24, 749–757 (2018).
Bindea, G. et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39, 782–795 (2013).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Winslow, S., Leandersson, K., Edsjo, A. & Larsson, C. Prognostic stromal gene signatures in breast cancer. Breast Cancer Res. 17, 23 (2015).
Heng, D. Y. et al. Prognostic factors for overall survival in patients with metastatic renal cell carcinoma treated with vascular endothelial growth factor-targeted agents: results from a large, multicenter study. J. Clin. Oncol. 27, 5794–5799 (2009).
Seung, S. K., Shu, H. K., McDermott, M. W., Sneed, P. K. & Larson, D. A. Stereotactic radiosurgery for malignant melanoma to the brain. Surg. Clin. North Am. 76, 1399–1411 (1996).
Braun, D. A. et al. Clinical validation of PBRM1 alterations as a marker of immune checkpoint inhibitor response in renal cell carcinoma. JAMA Oncol. 5, 1631–1633 (2019).
Zisman, A. et al. Improved prognostication of renal cell carcinoma using an integrated staging system. J. Clin. Oncol. 19, 1649–1657 (2001).
Frank, I. et al. An outcome prediction model for patients with clear cell renal cell carcinoma treated with radical nephrectomy based on tumor stage, size, grade and necrosis: the SSIGN score. J. Urol. 168, 2395–2400 (2002).
Kattan, M. W., Reuter, V., Motzer, R. J., Katz, J. & Russo, P. A postoperative prognostic nomogram for renal cell carcinoma. J. Urol. 166, 63–67 (2001).
Bettegowda, C. et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl. Med. 6, 224ra224 (2014).
Cohen, J. D. et al. Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science 359, 926–930 (2018).
Pal, S. K. et al. Evolution of circulating tumor DNA profile from first-line to subsequent therapy in metastatic renal cell carcinoma. Eur. Urol. 72, 557–564 (2017).
Merker, J. D. et al. Circulating tumor DNA analysis in patients with cancer: American Society of Clinical Oncology and College of American Pathologists joint review. Arch. Pathol. Lab. Med. 142, 1242–1253 (2018).
Rouzier, R. et al. Breast cancer molecular subtypes respond differently to preoperative chemotherapy. Clin. Cancer Res. 11, 5678–5685 (2005).
Travis, W. D. et al. The 2015 World Health Organization classification of lung tumors: impact of genetic, clinical and radiologic advances since the 2004 classification. J. Thorac. Oncol. 10, 1243–1260 (2015).
Turcotte, S. et al. A molecule targeting VHL-deficient renal cell carcinoma that induces autophagy. Cancer Cell 14, 90–102 (2008).
Ding, Z. et al. Genetic and pharmacological strategies to refunctionalize the von Hippel Lindau R167Q mutant protein. Cancer Res. 74, 3127–3136 (2014).
Ding, Z. et al. Agents that stabilize mutated von Hippel-Lindau (VHL) protein: results of a high-throughput screen to identify compounds that modulate VHL proteostasis. J. Biomol. Screen. 17, 572–580 (2012).
Gameiro, P. A. et al. In vivo HIF-mediated reductive carboxylation is regulated by citrate levels and sensitizes VHL-deficient cells to glutamine deprivation. Cell Metab. 17, 372–385 (2013).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03428217 (2019).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03163667 (2019).
Courtney, K. D. et al. Phase I dose-escalation trial of PT2385, a first-in-class hypoxia-inducible factor-2alpha antagonist in patients with previously treated advanced clear cell renal cell carcinoma. J. Clin. Oncol. 36, 867–874 (2018).
Chen, W. et al. Targeting renal cell carcinoma with a HIF-2 antagonist. Nature 539, 112–117 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03634540 (2020).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03401788 (2020).
Jonasch, E. et al. An open-label phase II study to evaluate PT2977 for the treatment of von Hippel-Lindau disease-associated renal cell carcinoma. J. Clin. Oncol. 37, TPS680 (2019).
Jonasch, E. et al. Phase II study of the oral HIF-2α inhibitor MK-6482 for Von Hippel-Lindau disease-associated renal cell carcinoma. J. Clin. Oncol. 38, 5003–5003 (2020).
LaFave, L. M. et al. Loss of BAP1 function leads to EZH2-dependent transformation. Nat. Med. 21, 1344–1349 (2015).
Schoumacher, M. et al. Uveal melanoma cells are resistant to EZH2 inhibition regardless of BAP1 status. Nat. Med. 22, 577–578 (2016).
Pfister, S. X. et al. Inhibiting WEE1 selectively kills histone H3K36me3-deficient cancers by dNTP starvation. Cancer Cell 28, 557–568 (2015).
Terzo, E. A. et al. SETD2 loss sensitizes cells to PI3Kbeta and AKT inhibition. Oncotarget 10, 647–659 (2019).
Motzer, R. J. et al. Efficacy of everolimus in advanced renal cell carcinoma: a double-blind, randomised, placebo-controlled phase III trial. Lancet 372, 449–456 (2008).
Hudes, G. et al. Temsirolimus, interferon alfa, or both for advanced renal-cell carcinoma. N. Engl. J. Med. 356, 2271–2281 (2007).
Jonasch, E. et al. A randomized phase 2 study of MK-2206 versus everolimus in refractory renal cell carcinoma. Ann. Oncol. 28, 804–808 (2017).
Ibrahim, Y. H. et al. PI3K inhibition impairs BRCA1/2 expression and sensitizes BRCA-proficient triple-negative breast cancer to PARP inhibition. Cancer Discov. 2, 1036–1047 (2012).
Juvekar, A. et al. Combining a PI3K inhibitor with a PARP inhibitor provides an effective therapy for BRCA1-related breast cancer. Cancer Discov. 2, 1048–1063 (2012).
Escudier, B. et al. A randomized, controlled, double-blind phase III study (AVOREN) of bevacizumab/interferon-α2a vs placebo/interferon-α2a as first-line therapy in metastatic renal cell carcinoma. J. Clin. Oncol. 25, 3 (2007).
Rini, B. I. et al. Bevacizumab plus interferon alfa compared with interferon alfa monotherapy in patients with metastatic renal cell carcinoma: CALGB 90206. J. Clin. Oncol. 26, 5422–5428 (2008).
Rini, B. I. et al. Comparative effectiveness of axitinib versus sorafenib in advanced renal cell carcinoma (AXIS): a randomised phase 3 trial. Lancet 378, 1931–1939 (2012).
Motzer, R. J. et al. Sunitinib versus interferon alfa in metastatic renal-cell carcinoma. N. Engl. J. Med. 356, 115–124 (2007).
Choueiri, T. K. et al. Cabozantinib versus everolimus in advanced renal-cell carcinoma. N. Engl. J. Med. 373, 1814–1823 (2015).
Escudier, B. et al. Sorafenib in advanced clear-cell renal-cell carcinoma. N. Engl. J. Med. 356, 125–134 (2007).
Sternberg, C. N. et al. Pazopanib in locally advanced or metastatic renal cell carcinoma: results of a randomized phase III trial. J. Clin. Oncol. 28, 1061–1068 (2010).
Motzer, R. J. et al. Pazopanib versus sunitinib in metastatic renal-cell carcinoma. N. Engl. J. Med. 369, 722–731 (2013).
Jonasch, E. et al. Phase II study of two weeks on, one week off sunitinib scheduling in patients with metastatic renal cell carcinoma. J. Clin. Oncol. 36, 1588–1593 (2018).
Jonasch, E. et al. Pilot trial of sunitinib therapy in patients with von Hippel-Lindau disease. Ann. Oncol. 22, 2661–2666 (2011).
Kawakami, F. et al. Programmed cell death ligand 1 and tumor-infiltrating lymphocyte status in patients with renal cell carcinoma and sarcomatoid dedifferentiation. Cancer 123, 4823–4831 (2017).
Joseph, R. W. et al. PD-1 and PD-L1 expression in renal cell carcinoma with sarcomatoid differentiation. Cancer Immunol. Res. 3, 1303–1307 (2015).
Raychaudhuri, R. et al. Immune check point inhibition in sarcomatoid renal cell carcinoma: a new treatment paradigm. Clin. Genitourin. Cancer 15, e897–e901 (2017).
Paolino, M. et al. The E3 ligase Cbl-b and TAM receptors regulate cancer metastasis via natural killer cells. Nature 507, 508–512 (2014).
Paolino, M. & Penninger, J. M. The role of TAM family receptors in immune cell function: implications for cancer therapy. Cancers 8, 97 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03141177 (2020).
Heemskerk, B., Kvistborg, P. & Schumacher, T. N. The cancer antigenome. EMBO J. 32, 194–203 (2013).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Motzer, R. J. et al. Nivolumab versus everolimus in advanced renal-cell carcinoma. N. Engl. J. Med. 373, 1803–1813 (2015).
Motzer, R. J. et al. Nivolumab plus ipilimumab versus sunitinib in advanced renal-cell carcinoma. N. Engl. J. Med. 378, 1277–1290 (2018).
Woo, S. R., Corrales, L. & Gajewski, T. F. The STING pathway and the T cell-inflamed tumor microenvironment. Trends Immunol. 36, 250–256 (2015).
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
E.J. has received research funding from Exelixis, Novartis, Peloton and Pfizer and consulting fees from Eisai, Exelixis, Genentech, Ipsen, Novartis, Peloton, Pfizer and Roche. The other authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Nephrology thanks O. Iliopoulos, S. Turajlic and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Haploinsufficient
-
A situation in which one copy of a gene is inactivated; for such haploinsufficient genes, gene function is thus altered in heterozygotes, as the remaining (functional) copy does not produce sufficient gene product for normal function.
- Von Hippel–Lindau (VHL) disease
-
A hereditary condition associated with the development of cysts and tumours in multiple organs.
- Chromothripsis
-
A phenomenon by which tens to thousands of chromosomal rearrangements occur; chromothripsis may result from a single catastrophic event during the life history of a cell.
- Micronuclei
-
The small nucleus that forms when a chromosome or a fragment of a chromosome is not incorporated into one of the daughter nuclei during mitosis. It is usually a sign of genotoxic events and chromosomal instability.
- Aneugens
-
Substances that cause a daughter cell to have an abnormal number of chromosomes.
- Clastogens
-
Mutagenic agents that give rise to or induce disruption or breakages of chromosomes.
- Chromosome bridges
-
Attachments of the telomeric ends of sister chromatids during mitosis, specifically during anaphase, preventing segregation into respective daughter cells.
- Cytokinesis
-
The physical process of cell division, which divides the cytoplasm of a parental cell into two daughter cells.
- Aneuploidy
-
An abnormal number of chromosomes in a cell.
- Replication stress
-
Any condition that compromises the faithful duplication of the genome once per cell cycle.
- Acetyl-lysine reader
-
A protein that recognizes acetylated lysine residues in promoters and enhancers and has an active role in gene transcription.
- Kinetochore
-
A large protein complex that assembles at the centromere of a chromosome and functions to connect the chromosome to microtubules in the mitotic spindle.
- Genome instability index
-
An estimate of the proportion of the genome with aberrant copy number compared with the median ploidy, weighted on a per chromosome basis.
- Intratumoural heterogeneity
-
(ITH). The presence of distinct tumour cell populations within the same tumour specimen.
- Cryptic transcription
-
Transcripts initiated from intragenic promoters that are usually not accessible to transcriptional machinery.
- DNA mismatch repair
-
A process for recognizing and repairing erroneous insertion, deletion and misincorporation of bases that can arise during DNA replication and recombination, or through DNA damage.
- Microsatellite instability
-
Genetic predisposition to mutation that results from impaired DNA mismatch repair.
- Lynch syndrome
-
An inherited disorder that arises from germline genetic defects in mismatch repair components and increases the risk of many types of cancer, particularly cancers of the colon.
- Non-homologous end joining
-
(NHEJ). A pathway that repairs double-strand breaks in DNA without the need for a homologous template.
- Homologous recombination repair
-
(HRR). A DNA repair process in which an intact DNA molecule is used as a template for the repair process, in particular during the S phase.
- Cell cycle checkpoint
-
Control mechanisms in the eukaryotic cell cycle that ensure its proper progression.
- Microhomology repair
-
An error-prone, alternative form of non-homologous end joining that involves alignment of microhomologous sequences internal to the broken ends before joining.
- PARP inhibitors
-
A group of pharmacological inhibitors of the enzyme poly(ADP-ribose) polymerase (PARP).
- Somatic copy number alterations
-
(SCNAs). Somatic changes to chromosome structure that result in gain or loss of copies of sections of DNA.
- Circulating tumour DNA
-
(ctDNA). DNA in the bloodstream that comes from cancerous cells and tumours.
- Synthetic lethality
-
The concept whereby the simultaneous mutation of two genes leads to the lethality of a cell, but the mutation of one alone is viable.
- cGAS–STING pathway
-
A component of the innate immune system that functions to detect the presence of cytosolic DNA and triggers the expression of inflammatory genes.
Rights and permissions
About this article
Cite this article
Jonasch, E., Walker, C.L. & Rathmell, W.K. Clear cell renal cell carcinoma ontogeny and mechanisms of lethality. Nat Rev Nephrol 17, 245–261 (2021). https://doi.org/10.1038/s41581-020-00359-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41581-020-00359-2
This article is cited by
-
Metabolic heterogeneity in clear cell renal cell carcinoma revealed by single-cell RNA sequencing and spatial transcriptomics
Journal of Translational Medicine (2024)
-
Single-cell combined bioinformatics analysis: construction of immune cluster and risk prognostic model in kidney renal clear cells based on CD8+ T cell-associated genes
European Journal of Medical Research (2024)
-
Cellular senescence and metabolic reprogramming model based on bulk/single-cell RNA sequencing reveals PTGER4 as a therapeutic target for ccRCC
BMC Cancer (2024)
-
Targeting STING elicits GSDMD-dependent pyroptosis and boosts anti-tumor immunity in renal cell carcinoma
Oncogene (2024)
-
Integration of basement membrane-related genes in a risk signature for prognosis in clear cell renal cell carcinoma
Scientific Reports (2024)