Abstract
Within the past decade, the field of immunotherapy has revolutionized the treatment of many cancers with the development and regulatory approval of various immune-checkpoint inhibitors and chimeric antigen receptor T cell therapies in diverse indications. Another promising approach to cancer immunotherapy involves the use of personalized vaccines designed to trigger de novo T cell responses against neoantigens, which are highly specific to tumours of individual patients, in order to amplify and broaden the endogenous repertoire of tumour-specific T cells. Results from initial clinical studies of personalized neoantigen-based vaccines, enabled by the availability of rapid and cost-effective sequencing and bioinformatics technologies, have demonstrated robust tumour-specific immunogenicity and preliminary evidence of antitumour activity in patients with melanoma and other cancers. Herein, we provide an overview of the complex process that is necessary to generate a personalized neoantigen vaccine, review the types of vaccine-induced T cells that are found within tumours and outline strategies to enhance the T cell responses. In addition, we discuss the current status of clinical studies testing personalized neoantigen vaccines in patients with cancer and considerations for future clinical investigation of this novel, individualized approach to immunotherapy.
Key points
-
Personalized therapeutic cancer vaccines predicated on neoantigens have been shown to be feasible, safe and immunogenic in patients with melanoma and glioblastoma.
-
Different vaccine formats and delivery strategies are currently being tested in clinical studies involving patients with various tumour types.
-
Deeper evaluation of the phenotypes, functionality and long-lasting memory potential of vaccine-induced neoantigen-specific CD4+ and CD8+ T cells is warranted to improve understanding of their therapeutic activity and optimize vaccination strategies.
-
Neoantigen target discovery is continually being advanced to improve the identification of immunogenic neoepitopes that can be recognized by CD8+ T cells; algorithms for the more challenging task of predicting CD4+ T cell neoepitopes are also emerging.
-
Innovative vaccine delivery platforms and the most effective timing of combinatorial therapies should be further explored to reduce costs and time delays and increase clinical efficacy.
Similar content being viewed by others
Introduction
Vaccines have traditionally been used for the prevention of infectious diseases; however, the ability of such agents to elicit and amplify antigen-specific immune responses has long been recognized as a potentially valuable tool for the treatment of cancer. Early therapeutic vaccination strategies focused on self-antigens abnormally expressed or overexpressed in tumours, termed tumour-associated antigens (TAAs), were largely unsuccessful in generating clinically effective antitumour immune responses, probably owing to the TAA-specific T cells being subject to central and/or peripheral tolerance1. Such TAAs can also be expressed to some extent in nonmalignant tissues, which raises the risk of vaccine-induced autoimmune toxicities1. Thus, these early studies highlighted the lack of tumour specificity and poor immunogenicity as fundamental issues to overcome in developing cancer vaccines.
Mutations occurring in tumour cells can generate novel epitopes of self-antigens, which are referred to as neoepitopes or neoantigens. The advent of next-generation sequencing has provided opportunities to identify these tumour-specific mutations in individual patients in a timely and cost-effective manner and to explore therapies that target the mutated proteins in clinical studies. Additionally, the development of algorithms for the prediction of MHC class I (MHC I)-binding epitopes has paved the way for the identification of potentially immunogenic neoepitopes. Together, these scientific advances have enabled the generation of personalized therapeutic cancer vaccines that are tailored to the tumours of individual patients.
Vaccines predicated on neoantigens rather than traditionally used TAAs have several advantages. First, neoantigens are exclusively expressed by tumour cells and can, therefore, elicit truly tumour-specific T cell responses, thereby preventing ‘off-target’ damage to nonmalignant tissues. Second, neoantigens are de novo epitopes derived from somatic mutations, which presents the possibility to circumvent T cell central tolerance of self-epitopes and thus induce immune responses to tumours. Personalized neoantigen-based vaccines therefore afford the opportunity to boost tumour-specific immune responses and add an additional tool to the immunotherapy toolbox. Furthermore, the potential of these vaccine-boosted neoantigen-specific T cell responses to persist and provide post-treatment immunological memory presents the possibility of long-term protection against disease recurrence. Limitations of this personalized approach to immunotherapy include the high costs and time delays associated with the manufacture of individualized vaccines, uncertainty over the optimal neoantigen discovery platform and lack of consensus regarding the most suitable vaccine delivery platform.
In this Review, we discuss advances in the development of personalized neoantigen-based vaccines, current understanding of the T cell phenotypes and the immune responses elicited by these agents, and the clinical development landscape of this class of cancer therapy. We also consider avenues for improvement and future investigation.
Neoantigens and antitumour immunity
Neoantigens are targets of effective tumour-specific immune responses
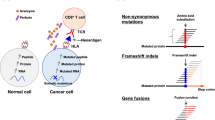
Neoantigens can arise through a variety of mutational events, with the numbers and types of mutation varying by cancer type2,3. These events include point mutations, insertions or deletions (indels)4 and gene fusions5. In tumours driven by viruses, viral proteins can be considered an alternative class of neoantigen, owing to their foreign nature6. Tumours with a high mutational burden are likely to have several candidate neoantigens for vaccine formulation. Fewer neoantigens are likely to be present in tumours with a low mutational burden, which might restrict the number of immunogenic epitopes7; however, the induction of neoantigen-specific immune responses using immunotherapy8 or therapeutic vaccination9,10 remains a possibility within this category of tumours.
Neoantigens can be recognized by tumour-infiltrating cytotoxic CD8+ T cells11,12, and increased immune cell infiltration and the related cytotoxicity signatures have been observed in tumours with a higher neoantigen load13,14,15. Accordingly, neoantigen presentation and load have been positively correlated with prognosis in patients with a variety of cancers6,14,16,17 and with benefit from immune-checkpoint inhibitors (ICIs) in patients with melanoma18,19, non-small-cell lung cancer (NSCLC)20,21 or colorectal cancer with mismatch-repair deficiency22. Together, these studies highlight the potential therapeutic benefit of developing immunotherapies that specifically ‘train’ the immune system to target neoantigens.
As reviewed previously23,24,25, tumour mutational burden alone is an imperfect predictor of response to immunotherapy: additional factors determine whether a given mutation has the ability to induce an effective antitumour immune response, including the type of mutation (for example, clonal versus subclonal and single-nucleotide variant versus frameshift), to what extent the mutation is expressed, the ability of the encoded neoepitope to be presented by tumour cells and/or antigen-presenting cells (APCs), trafficking of T cells into the tumour and the immunosuppressive mechanisms active in the tumour or periphery of the host. Thus, even when targetable alterations are present, their existence alone might not be sufficient for effective immunotherapy — other immunological parameters are key determinants of efficacy. For example, in patients with prostate cancer, tumour mutational burden was not associated with response following treatment with the anti-CTLA4 antibody ipilimumab; however, a response to immunotherapy with this ICI was associated with a higher abundance of tumour-infiltrating CD8+ T cells and IFNγ-response gene signatures in pretreatment tumours8. In addition, neoantigen-specific CD8+ T cells could be detected after therapy in blood from two of eight evaluated patients with favourable responses to therapy, whereas neoantigen-specific responses were not observed in four evaluated patients with unfavourable responses8. Moreover, CD8+ T cell responses and myeloid signatures have been associated with progression-free survival (PFS) in patients with renal cell carcinoma (RCC) treated with the anti-PD-L1 antibody atezolizumab plus the anti-VEGFA antibody bevacizumab26. In patients with resectable pancreatic cancer, neoantigen load alone was not correlated with overall survival (OS); however, correlations with OS were found when neoantigen load was combined with the abundance or diversity of tumour-infiltrating CD8+ T cells, or when neoantigen quality was considered; these correlations were independent of adjuvant chemotherapy6. Whether a mutation is clonal or subclonal might also influence response to immunotherapy; in patients with microsatellite-stable solid tumours, clonal mutations were found to be associated with a response to ICI therapy, whereas a greater abundance of subclonal mutations was more likely to result in progressive disease27. Furthermore, in this study, eight clonal neoantigens identified in driver genes (KRAS, BRAF and PIK3CA) were associated with response across more than one patient27. Notably, tumour mutational burden might correlate with OS but not with response to anti-PD-1 antibodies28. Taken together, these findings indicate that, although neoantigens are promising targets for immune-based therapies, consideration of additional factors that can affect neoantigen-specific T cell responses and antitumour activity following therapeutic vaccination is crucial.
Neoantigens and immunoediting
The importance of neoantigens in antitumour immunity is reflected by evidence of immunoediting. Immunoediting constitutes a dynamic process whereby tumour cells expressing immunogenic neoepitopes are recognized and eliminated by T cells, leading to a tumour phenotype with reduced recognition by the immune system and, ultimately, to tumour outgrowth29. Reduced expression of neoantigens has been detected in metastatic tumours arising in a patient with progression of pancreatic cancer following primary tumour resection6. Moreover, immunotherapy can shape the repertoire of neoantigens expressed by a variety of tumour types, including melanoma30, NSCLC31,32 and glioblastoma33, suggesting ongoing immunoediting.
Neoantigens and tumour heterogeneity
Tumours have a high degree of genetic heterogeneity, which has fundamental implications for the efficacy of cancer therapy, including immunotherapy34. The ability to identify tumour-specific mutations in individual patients is a key component of directing T cell responses to eliminate tumours through vaccination, thus warranting a personalized approach for therapeutic vaccine design. Tumour heterogeneity can also result in variable expression of neoantigens between or even within tumours present in an individual patient; therefore, elimination of tumour cells expressing a specific neoantigen can result in outgrowth of tumour cells not expressing that neoantigen34. Consequently, it will probably be necessary to target multiple neoantigens using a single vaccine to reduce the likelihood of immune evasion and effectively eradicate the disease in its entirety30.
Identifying immunogenic neoantigens
Neoantigen prediction in silico
The computational algorithm-based pipelines for neoantigen discovery and prioritization used in studies of therapeutic cancer vaccines to date have had several general commonalities9,10,35,36,37. Details of these workflows have been extensively reviewed25,38,39,40,41. As such, we only briefly outline the neoantigen prediction process here, with a focus on advances made over the past few years.
To identify tumour-specific somatic mutations, tumour biopsy specimens and nonmalignant tissue samples (usually peripheral blood mononuclear cells) are acquired from the patient in order to perform whole-exome sequencing for the comparison of tumour and germline DNA. Additional RNA sequencing provides information on the expression of the mutated genes and further confirmation of the mutation calls. A high number of tumour-specific mutations can often be identified, depending on the tumour type; however, not all mutations result in neoepitopes that are recognized by the immune system, owing to HLA restriction. More than 16,000 different alleles of the classical HLA class I genes (HLA-A, HLA-B and HLA-C) are known to exist;42 therefore, HLA typing is required to enable the prediction of potentially immunogenic epitopes.
MHC I-binding epitopes are predicted using computational approaches, and peptides with a predicted HLA-binding affinity in the moderate-to-strong range (IC50 < 150 nmol/l) are considered more likely to induce CD8+ T cell responses43. Various computational approaches for predicting MHC I-presented epitopes have been developed44, and historically these algorithms have been trained on peptide binding affinity data. Although these algorithms enabled the development of the first neoantigen-based vaccines, they are continuously being optimized by incorporating endogenous peptide processing and presentation rules to enable more precise prediction of neoepitopes with the highest likelihood of being presented on the cell surface via MHC I proteins. Epitope prediction based solely on binding affinity data inherently does not provide information about which endogenously processed peptides will be presented by MHC I proteins on the surface of tumour cells and/or APCs and will induce effective CD8+ T cell responses.
The use of mass spectrometry (MS) has enabled the development of improved prediction algorithms trained on peptides eluted from MHC proteins45. In particular, algorithms trained using peptides presented by specific mono-allelic HLA-expressing cell lines better incorporate endogenous antigen processing and presentation processes46,47. These MS-based algorithms can be further optimized by increasing the number of HLA alleles on which they are trained48. However, further studies are needed to improve understanding of the factors that can affect neoantigen expression, presentation and immunogenicity. For example, data suggest that incorporating information on the position of a mutated residue within the peptide-binding groove could improve neoantigen discovery49.
To date, epitope-prediction methods have largely focused on MHC I-binding epitopes; MHC II-binding epitope prediction is much less advanced. The MHC I peptide-binding groove has closed ends that define the positioning of peptide epitopes 8–11 amino acids long for presentation to CD8+ T cells. By contrast, the MHC II peptide-binding groove has open ends, which confers the ability to bind and present peptides of longer and more variable lengths; the peptide regions flanking the portion bound to the core of the MHC II molecule can influence the binding of the peptide, and in addition these peptides can bind to multiple different MHC II molecules50. Together, these features of MHC II peptide binding complicate the identification of immunogenic MHC II epitopes. MHC II epitopes have, however, been incorporated indirectly into vaccine peptide design. Long peptides (15–30-mers) that encompass predicted MHC I-binding CD8+ T cell epitopes are used in vaccination strategies to promote uptake and processing of the peptides by professional APCs and thereby enhance T cell stimulation51,52. In addition to the presentation of predicted CD8+ T cell epitopes on MHC I proteins, this long-peptide approach is compatible with the processing and presentation of CD4+ T cell epitopes bound to MHC II proteins. Indeed, the preferential induction of CD4+ T cell responses compared with CD8+ T cell responses has been observed following vaccination in several clinical trials7,9,10,35,36,53. This observation might at least partially reflect the potential for presentation of a greater variety of peptides in the open-ended MHC II peptide-binding groove than in the more stringent MHC I peptide-binding groove54. Other immunological factors might also contribute to the dominance of vaccine-mediated CD4+ T cell response, such as differences in the stimulation of dendritic cell (DC) subsets that promote CD4+ T cell activation or, via cross-presentation of antigen, the induction of CD8+ T cell responses55.
With the exception of one study36, specific MHC II-binding neoepitopes have not been incorporated into the neoantigen vaccines investigated in clinical trials to date. However, efforts have been made to develop better tools for the prediction of such epitopes, recognizing that CD4+ T cells can orchestrate antitumour immunity and, via their classical ‘helper’ functions, have a crucial role in generating and sustaining CD8+ T cell responses. A stride forward in HLA-II prediction was made in 2019 with the creation of a mono-allelic HLA expression system for profiling of the HLA ligandome, referred to as ‘mono-allelic purification with tagged allele constructs’ (MAPTAC)56. This system provided the ability to identify peptide-binding motifs of individual HLA alleles via MS, including >40 HLA class II alleles, which in turn enabled the training of a new machine learning-based prediction algorithm, neonmhc2, to identify MHC II-presented epitopes56. In another study published in 2019 (ref.57), a similar MS-based approach combined with motif deconvolution and annotation was used to train a machine learning prediction algorithm, MixMHC2pred. Moreover, the MHC II-binding epitope-prediction algorithm NetMHCIIpan, which was initially developed using peptide binding affinity data from the Immune Epitope Database (IEDB), was updated in 2020 to include the NNAlign_MA algorithm trained using MS-derived data on peptides eluted from MHC II proteins58. These efforts underscore a current focus on enhancing the induction of tumour-specific CD4+ T cell responses to improve the effectiveness of therapeutic cancer vaccines.
Although prediction algorithms are undergoing continual improvement, as stated earlier, additional factors can affect the ultimate immunogenicity of the predicted epitopes. These include the overall patterns of gene expression, RNA splicing, proteosomal processing and, crucially, peptide loading and presentation by MHCs25,40. Additionally, neoantigen sequences with similarity to pathogen-derived epitopes can exhibit enhanced immunogenicity probably because they can be better distinguished from self-epitopes by T cells6. Obtaining a better understanding of these factors will be important in optimizing the immunogenicity of neoantigen-based therapies, and incorporating such variables into computational algorithms will enable the design of more effective therapeutic T cell vaccines.
An alternative approach to inducing neoantigen-specific immune responses, beyond computational approaches, involves the use of tumour lysates59. Autologous APCs, typically DCs, can be isolated from the patient and exposed to tumour lysates ex vivo, before being injected back into the patient with the aim of stimulating an immune response to TAAs or neoantigens. This method circumvents the sequencing and computational analysis required for the identification of patient-specific neoantigens59. As mentioned earlier, however, TAAs are less likely to be immunogenic and, in addition, might reduce the ability of relevant neoepitopes to stimulate immune responses owing to the higher abundance of nonimmunogenic self-antigens.
Administration of tumour lysate-pulsed DCs to patients with recurrent ovarian cancer, either alone or in combination with bevacizumab with or without low-dose cyclophosphamide, was found to induce de novo neoantigen-specific CD8+ T cell responses and enhanced some neoantigen-specific responses that were detected prior to therapy60. Patients with ex vivo evidence of T cell responses to autologous tumour cells or DC-presented tumour antigens following therapy had improved PFS relative to those without such evidence of antitumour responses. Moreover, 2-year OS was 100% in those with antitumour T cell responses compared with 25% in non-responders. Addition of the vaccine to therapy also seemed to improve OS when compared with bevacizumab and low-dose cyclophosphamide combination therapy (2-year OS 78% versus 44% in a matched historical control group; P = 0.046). Low-dose cyclophosphamide significantly improved OS when given alongside bevacizumab and the vaccine (P = 0.012)60. These findings suggest that tumour-lysate-based vaccine therapies can induce antitumour T cell responses, including neoantigen-specific CD8+ T cell responses. Thus, the use of tumour-lysate-based approaches and/or antigen delivery via APCs merits further exploration. Additionally, tumour-lysate-based vaccines could potentially be used as priming vaccinations while a personalized neoantigen-based vaccine is being designed and manufactured59.
Monitoring of neoantigen vaccine-induced immune responses
Following vaccination, assessment of T cell responses to epitopes included in the vaccines can be accomplished in vitro using a variety of tools. One approach to validating T cell specificity involves the transfection of genetic constructs encoding multiple patient-specific mutated epitopes, termed tandem minigenes, into APCs, followed by co-culture of the APCs with T cells isolated from the blood or tumour of the patient. Any APC subpopulations that induce T cell responses can then be further investigated to identify immunogenic epitopes61. Neoantigen-specific T cells have been detected using this technology in several different cancers61,62,63,64, including following therapeutic vaccination10,35, and this technology has also enabled the identification of neoantigen-specific T cell receptors (TCRs)62.
Neoantigen vaccine-induced responses
Immunogenicity and immunological profiling analyses conducted in clinical trials of personalized neoantigen-based vaccines have provided insight into the magnitude, phenotype and functionality of vaccine-induced T cells (Table 1). The first such trial used an autologous DC-based vaccination strategy37, whereby three patients with stage III cutaneous melanoma previously treated with ipilimumab were vaccinated with HLA-A*02:01-specific peptides encoding patient-specific tumour neoepitopes and three peptides from the melanoma-associated antigen gp100. Vaccination induced CD8+ T cell responses and a diverse TCR repertoire37; thus, this study was the first to reveal that therapeutic vaccination with neoantigens can increase both the diversity and breadth of T cell responses. A subset of neoantigen-responsive T cells were detectable prior to vaccination37, indicating that existing neoantigen-specific T cell responses were boosted, in addition to induction of de novo responses.
In another phase I trial evaluating personalized neoantigen-based vaccination, a peptide-based vaccine (NeoVax) was administered to four patients with previously high-risk stage III and two with stage IV melanoma following initial curative-intent surgery35. The NeoVax vaccines comprised up to 20 different long peptides (15–30-mers), formulated with the adjuvant poly-ICLC (a TLR3 agonist consisting of carboxymethylcellulose, polyinosinic–polycytidylic acid and poly-l-lysine double-stranded RNA). Previously undetectable neoantigen-specific CD4+ and CD8+ T cells were induced post-vaccination, with a greater fraction of the response consisting of CD4+ T cells35. These T cell populations were polyfunctional, and neoantigen-specific CD4+ T cell transcriptional profiles post-vaccination revealed T helper 1 (TH1), effector and memory programmes35. The four patients with stage III disease remained disease-free at a median follow-up duration of 25 months (range 20–32 months) after vaccination. The two patients with stage IV disease had disease recurrence within a few months after the last vaccination and subsequently received pembrolizumab (an anti-PD-1 antibody), which resulted in complete regression of metastatic tumours in both patients and broadening of antitumour T cell responses35. This observation highlights the potential of combination therapies to improve vaccine-induced T cell responses.
In a third phase I study36, a vaccine platform consisting of mRNAs encoding shared melanoma antigens (NY-ESO-1 and/or tyrosinase) and personalized neoantigen peptides was tested in 13 patients with stage III or IV melanoma. Profiling of T cells post-vaccination revealed that neoantigen-specific cytokine-producing CD8+ T cells present in blood comprised central memory (TCM) and effector memory (TEM) populations36. Similar to the responses observed with NeoVax35, CD4+ T cell responses were greater in magnitude than CD8+ T cell responses36. Neoantigen-specific tumour-infiltrating lymphocytes were found in resected tumours from two patients. Together, these findings from studies in patients with melanoma provided proof of concept that personalized neoantigen-based vaccines can induce tumour-specific T cell responses in a therapeutic setting.
More recently, neoantigen-based vaccines have been studied in patients with glioblastoma, a cancer type that typically has a low mutational burden3,65 and that is generally considered to constitute an immunologically ‘cold’ tumour. In a phase I/Ib trial involving ten patients with MGMT promoter-unmethylated glioblastoma, personalized long-peptide-based vaccines formulated with poly-ICLC (NeoVax) were administered following standard-of-care surgical resection and radiotherapy10. Six of the eight vaccinated patients also received dexamethasone for the treatment of brain oedema, which impeded vaccine immunogenicity10. In the two vaccinated patients who did not receive dexamethasone, however, neoantigen-specific CD4+ T cell and CD8+ T cell responses were detected in the peripheral blood, and increased numbers of T cells were observed in intracranial tumours post-vaccination10. Transcriptomic analysis of tumour-associated T cells from one patient revealed varying degrees of expression of inhibitory receptors (including TIM3, LAG3, TIGIT and CTLA4), in addition to cytotoxicity signatures in both CD4+ T cells and CD8+ T cells10. TCR reconstruction revealed four neoantigen-specific CD4+ T cell and two neoantigen-specific CD8+ T cell clonotypes that were present in the post-treatment intracranial tumour.
In the phase I GAPVAC-101 trial9, patients with glioblastoma similarly received peptide-based vaccination (with GM-CSF and poly-ICLC as adjuvants) following surgical resection and chemoradiotherapy with temozolomide, in parallel with temozolomide maintenance therapy. The vaccination regimen consisted of two parts, the first component (APVAC1) involving vaccination with up to seven MHC I-binding peptides and MHC II-binding peptides derived from unmutated glioblastoma-associated antigens, and the second component (APVAC2) involving patient-specific neoantigens (typically two)9. The unmutated glioblastoma-associated antigens included in APVAC1 induced CD8+ T cell responses across multiple cell subsets, including those with a central memory phenotype9. Neoantigens in APVAC2 induced predominantly CD4+ T cell responses that were multifunctional and TH1 polarized9. CD4+ T cells specific for glioblastoma-associated antigen incorporated in the APVAC1 vaccine were found in resected tumour tissue obtained after recurrence from one patient, which suggests that vaccine-induced T cells have the ability to traffic into tumour tissue.
No clear signals of clinical benefit that could be attributed to the vaccines were seen in either of the studies involving patients with glioblastoma. Nevertheless, evidence from both studies provides proof of concept that neoantigen-based vaccines can generate tumour-infiltrating T cell responses in immunologically cold tumours with a low mutational burden, which supports further investigations aiming to optimize neoantigen-based therapeutic vaccines for such tumours.
Altogether, these initial studies provided important knowledge of the immunogenicity and therapeutic potential of neoantigen-based personalized cancer vaccines. However, the extent of the induction and persistence of long-lived neoantigen-specific memory T cells in long-term survivors remains to be determined. Deeper phenotypic and functional analysis of these memory T cell populations could provide insights into the importance of subsets of memory T cells induced post-vaccination, particularly their potential to eliminate residual tumour cells.
Optimally inducing antitumour responses
Vaccine-induced immune responses have been extensively studied in the context of preventive vaccines for infectious disease, in order to understand the immunological mechanisms of vaccination and their implications for vaccine-mediated protection against infectious organisms. In the context of cancer, the stimulation of tumour-specific immune responses faces unique challenges that can limit the efficacy of therapeutic vaccines.
Vaccine development and optimization strategies typically involve testing the suitability of different delivery platforms, adjuvants, routes of administration, prime–boost strategies and incorporated antigens, all of which can affect the nature of the induced immune responses (as previously reviewed38). A key factor in the choice of a therapeutic cancer vaccine platform relates to how rapidly the vaccine can be designed, manufactured, administered and elicit antitumour T cell responses. Candidate vaccine platforms include peptides66, RNA67, DNA68, DCs69 and viral vectors70. Indeed, multiple platforms hold promise for use in the development of personalized cancer vaccines and can be built upon for improvements.
Priming of T cell responses
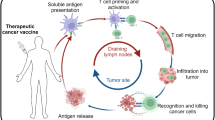
The immunological cascade elicited following vaccination commences rapidly with the induction of innate immune responses and uptake of antigens by APCs, which leads to priming of T cell responses in the draining lymph nodes56. Antigen-specific T cell responses then undergo an expansion phase, followed after antigen clearance by a contraction phase that culminates in the generation of small populations of long-lived memory T cells71 (Fig. 1).
Following vaccination, both neoantigen-specific CD4+ T cells and CD8+ T cells are induced either de novo or through boosting of existing neoantigen-specific T cell responses. These T cells proliferate and kill tumour cells expressing the neoantigen. As tumour cells are eliminated, the release of tumour antigens might contribute to epitope spreading and thus an increased breadth of the tumour-specific T cell response. Following tumour eradication, the responding T cell populations contract, although subpopulations persist as components of the memory T cell pool. Indeed, vaccine-induced neoantigen-specific T cells have the potential to generate long-lived central memory T (TCM) cells and effector memory T (TEM) cells. Whether neoantigen-based vaccination can generate tissue-resident memory T (TRM) cells or peripheral memory T (TPM) cells remains unknown.
Priming immunization can be crucial given that the phenotypes and magnitudes of initial T cell responses elicited can determine the subsequent ‘boostability’ of vaccine-elicited memory T cell populations72. This consideration is relevant because boosting immunizations will probably be needed to ensure adequate induction of neoantigen-specific T cell responses. Indeed, repetitive antigen stimulation has the potential to continually affect the gene-expression and functional profiles of memory CD8+ T cells72,73,74. Similar to preventive vaccines75, the timing of the administration of priming and boosting immunizations with therapeutic vaccines should be considered carefully, as should the timing of any combination therapies, given that these factors can all affect the resulting T cell phenotypes. Additionally, different adjunctive treatments might be needed for various tumour types to account for their distinct immune microenvironments.
The quality and magnitude of CD8+ T cell responses can be influenced by numerous factors71,76,77, resulting in variable memory T cell populations with distinct anatomical distributions and functional capabilities78. In the context of cancer, induction of effective neoantigen-specific T cell responses can be compromised by peripheral tolerance mechanisms, a result of either regulatory T (Treg) cell-mediated suppression79 or tolerogenic T cell priming by DCs80. Theoretically, memory T cells induced through therapeutic vaccination should persist in the long term following tumour clearance and might, therefore, contribute to protection against future tumour recurrence (Fig. 1). Thus, the memory T cell populations that develop after therapeutic vaccination need to be investigated in preclinical models and, ideally, in patients with cancer78,81.
Multiple memory T cell subsets exist, including TCM, TEM, resident memory T (TRM) and peripheral memory T (TPM) populations. TCM cells are most associated with localization to secondary lymphoid tissues, albeit with some degree of circulation in the blood81. TCM cells have a vast proliferative capacity, which enables them to mount vigorous recall responses77. TEM cells mostly circulate in the blood and periphery, and can, therefore, rapidly respond to antigenic challenges82. TEM cells have a greater capacity for cytotoxicity and cytokine production than TCM cells, but have a lesser proliferative capacity than other memory T cell populations77.
TRM cells localize within tissues to provide rapid responses upon antigen detection, including the release of cytokines and activation of other immune cell populations83,84,85,86,87, and could potentially contribute to the immunological response following vaccination87,88. High numbers of intratumoural TRM cells are correlated with favourable OS in patients with lung cancer89,90 or head and neck cancer91. Moreover, a high abundance of tumour-resident TRM cells expressing TIM3 and PD-1 has been associated with a response to anti-PD-1 therapy in patients with lung cancer92. In a mouse model of head and neck cancer, TRM cells induced by preventive vaccination with a DC-targeted human papillomavirus 16 (HPV16)-derived E7 peptide contributed to tumour control, together with other memory T cell populations89. TRM cells patrol a wide variety of organs85,93; therefore, inducing neoantigen-specific TRM cells through therapeutic vaccination might provide long-term immunosurveillance and protection against disease recurrence94.
TPM cells are a more-recently described subset of CD8+ T cells and have been observed to participate in immunosurveillance of peripheral tissues in viral infection models82. TPM cells can traffic through peripheral tissues, have self-renewal capacity and can differentiate into the highly proliferative TCM cells82, which underscores the potential benefits of harnessing this cell population in vaccination strategies. Although a better understanding of this novel T cell subset is needed, the immunosurveillance function of TPM cells might prove particularly useful in preventing metastatic tumour recurrences.
Circumventing CD8+ T cell exhaustion
T cell exhaustion is one of the major challenges in antitumour immunity95. CD8+ T cell exhaustion is a gradual process96,97, which eventually leads to an irreversible dysfunctional programme owing to epigenetic changes98,99,100,101,102. The personalized neoantigen-based vaccines tested in patients with cancer to date induce both CD4+ T cells and CD8+ T cells expressing several inhibitory receptors9,10,35,36, suggesting that functional blockade of these receptors will be necessary to produce effective neoantigen-specific T cell responses. This hypothesis is supported by data from the trial of the melanoma vaccine NeoVax, in which two patients who had recurrent tumours had complete tumour regression following treatment with pembrolizumab35. Findings from a number of studies demonstrate that TCF1-expressing ‘progenitor exhausted’ CD8+ T cells are capable of self-renewal or differentiation into terminally exhausted effector T cells and are important for the efficacy of ICIs103,104,105,106 and therapeutic vaccines106.
The importance of CD4+ T cells
CD4+ T cells have crucial roles in supporting the development of CD8+ T cell responses through licensing of DCs that prime CD8+ T cells, in addition to the provision of cytokines such as IL-2 (ref.107) This ‘help’ provided by CD4+ T cells is important for developing antitumour CD8+ T cell responses108,109,110,111,112, including in the context of immune-checkpoint inhibition111. CD4+ T cells also produce cytokines that mediate the recruitment and activation of immune cell populations109,111,113,114,115,116 (Fig. 2). Furthermore, IFNγ produced by CD4+ T cells can indirectly result in the elimination of tumour cells117. Alternatively, immunosuppressive CD4+ Treg cells can dampen immune responses and are often found within the tumour microenvironment (TME)79.
CD4+ T cells have many roles in the vaccine-induced antitumour immune response. By interacting with dendritic cells and enhancing antigen cross-presentation, CD4+ T cells perform the well-described roles of providing ‘help’ in the generation of the antitumour cytotoxic CD8+ T cell responses. CD4+ T cells can also produce many cytokines, including IFNγ and TNF, that stimulate and modulate immune responses. In addition, other types of immune cell with potential antitumour activity, such as macrophages, can be recruited as a result of CD4+ T cell-related responses. Interestingly, cytotoxic gene-expression signatures have been observed in neoantigen-specific CD4+ T cells induced following therapeutic vaccination, which indicates that these cells might contribute to the elimination of tumours with upregulation of MHC class II (MHC II) proteins (dashed arrow); however, whether vaccine-induced neoantigen-specific CD4+ T cells do indeed have this capacity needs to be investigated further.
CD4+ T cells are also important in inducing systemic immune responses, which are required for effective tumour control following immunotherapy in various mouse models118. Notably, neoantigen-specific CD4+ T cells have been found in tumour tissue from immunotherapy-naive patients with melanoma119. Adoptive transfer of tumour-specific autologous CD4+ T cells has also been shown to result in tumour regression in patients with metastatic melanoma120 and in a patient with metastatic cholangiocarcinoma121. Given the variety of crucial immunological roles of CD4+ T cells and their potential to shape vaccine-elicited immune responses, understanding the antitumour functions of this cell population (Fig. 2) and how they could be harnessed for neoantigen vaccines will be important122,123.
An early example suggesting a potential role of CD4+ T cells in therapeutic vaccination is provided by a study of a DC-based vaccine involving patients with stage III or IV melanoma, wherein inclusion of MHC II epitopes for TAAs in the vaccine resulted in increased induction of both TAA-specific CD4+ T cells and CD8+ T cells (in skin and/or blood) in some patients124. In mouse models of therapeutic cancer vaccines, the induction of tumour-specifc CD4+ T cells resulted in CD8+ T cells with improved effector functions111,125, a memory phenotype111, reduced expression of inhibitory markers (such as PD-1) and enhanced migration to and infiltration of tumours, resulting in improved tumour control125. Indeed, the inclusion of MHC II neoantigens in vaccines contributed substantially to tumour eradication in multiple mouse models, with evidence of TME reshaping and CD8+ T cell epitope spreading observed53. These findings indicate the importance of eliciting CD4+ T cell responses using therapeutic vaccination to optimize CD8+ T cell responses and tumour control.
In addition to supporting CD8+ T cell responses, data from a mouse model of sarcoma have shown that expression of antigen that is MHC II restricted by tumours has been shown to be crucial for the efficacy of ICI therapy or a tumour vaccine even when the tumour cells themselves do not express MHC II111. This study similarly revealed that expression of an MHC II epitope and CD4+ T cell help was important for the induction of CD8+ T cell responses, and expression of an MHC II epitope by the tumour resulted in increased frequency of both intratumoural CD4+ T cells and CD8+ T cells111. Moreover, expression of MHC II antigens was associated with greater accumulation of inducible nitric oxide synthase (iNOS)-positive macrophages111. Thus, CD4+ T cell assistance in recruiting innate immune cells might have a role in vaccine efficacy. A potential role of recruited innate immunity is supported by findings of cell-depletion studies in a mouse model of therapeutic vaccination in combination with recombinant IL-2, an anti-PD-1 antibody and a TAA-targeting antibody126. Thus, the effects of vaccine-induced neoantigen-specific T cells on the landscape of innate immune cell populations in tumours should be investigated in patients with cancer.
Similar to CD8+ T cells, the substantial diversity of CD4+ T cell subsets is well established127. Single-cell transcriptomic analyses of CD4+ T cells in a mouse model of colon adenocarcinoma and in human liver cancers demonstrated heterogeneity among the CD4+ T cells infiltrating tumours and tumour-draining lymph nodes, and a particular subset had a defined gene signature that was enriched in patients with intrinsic resistance to ICIs128. Furthermore, transcriptional profiling of neoantigen-vaccine-induced CD4+ T cells in patients with glioblastoma revealed cytotoxic signatures10. Most tumours are MHC II negative, although emerging evidence indicates that some tumours express MHC II129. Moreover, IFNγ produced by tumour-specific CD4+ T cells can stimulate upregulation of tumoural expression of MHC II, which resulted in direct CD4+ T cell-mediated tumour elimination in a mouse model of melanoma130. Tumour-infiltrating cytotoxic CD4+ T cells have also been found in human bladder cancers and to have MHC II-dependent cytotoxic activity against the autologous tumour cells131. Whether cytotoxic CD4+ T cells induced by a therapeutic vaccine could mediate antitumour activity directly remains unclear and should be further investigated. This question is particularly important given that the neoantigen-based vaccines evaluated in the initial clinical studies induced a greater degree of CD4+ T cell response than CD8+ T cell response.
Although antitumour CD4+ T cell responses have clearly been demonstrated, emerging evidence indicates that neoantigen-specific CD4+ Treg cells also exist. Tumour antigen-specific Treg cells have been observed in both peripheral blood and tumour specimens from patients with metastatic melanoma or gastrointestinal or ovarian cancers; the expansion of these specific Treg cell clonotypes probably occurs in the TME, in a neoantigen-selective manner132. Neoantigen-specific Treg cells might have important implications for therapeutic vaccines. Theoretically, immunosuppressive Treg cells induced by vaccination could dampen other vaccine-induced antitumour immune responses.
In line with potential inhibitory effects of vaccination, the ATLAS epitope-selection platform has been designed to account for ‘inhibitory’ neoantigens during vaccine formulation133. The exact mechanism of these inhibitory neoantigens has not been defined; however, using the ATLAS platform, immunostimulatory and inhibitory epitopes can be identified through high-throughput ex vivo screening assays involving autologous T cells and APCs134. Together, these observations highlight the need to obtain a better understanding of the CD4+ T cell phenotypes induced by therapeutic vaccination.
Designing effective vaccine-induced immune responses
The question of how to elicit specific beneficial T cell phenotypes is crucial to the development of effective therapeutic cancer vaccines. Data indicate the importance of sequencing of combination therapies as well as roles for CD4+ T cells and novel CD8+ T cell subsets. Methodical testing of vaccination regimens will provide insights into how various vaccine strategies influence the magnitude and phenotype of neoantigen-specific CD4+ and CD8+ T cell responses. Thus, detailed immunophenotyping should be conducted to clearly define T cell phenotypes and responses. Together, this information could contribute to the development of improved neoantigen-based vaccination regimens.
Clinical neoantigen vaccine development
As discussed previously, the initial clinical trials testing personalized neoantigen-based vaccines were conducted largely in disease settings in which all tumours had been surgically removed and no additional standard therapy was indicated, enabling the administration of an investigational vaccine alone. Given that ICIs targeting PD-1 or PD-L1 have broad clinical efficacy across cancers and that antitumour activity seems to depend on the presence of pre-existing T cell responses in the tumour (that is, an immunologically ‘hot’ TME), testing personalized vaccines in combination with PD-1 or PD-L1 inhibition is a logical next step in the clinical development of these therapies (Fig. 3). The observation of complete responses to PD-1 inhibitors in patients with melanoma who had disease progression during or after therapeutic neoantigen-based vaccination35 has been interpreted as potentially indicative of synergy between ICIs and vaccination, although complete responses can sporadically occur with PD-1 inhibition alone in patients with metastatic melanoma. In line with these considerations, ongoing clinical trials are combining personalized neoantigen-based vaccines with PD-1, PD-L1 and/or CTLA4 inhibitors in various tumour types (Table 2).
Various factors should be considered during the design of therapeutic vaccination regimens. Following sample collection, the time required to generate the personalized vaccine is a crucial factor, particularly in the metastatic disease setting. The manufacturing time is dependent on the choice of vaccine platform, as indicated for the various platforms listed along the red arrow. However, while the personalized vaccine is being designed and manufactured, combinatorial therapies can be administered to the patient with the aim of fostering a favourable immunological milieu. Adjunctive therapies can also be given either at the time of or following vaccination to enhance the immune response. Additional variables include the route of administration of the vaccine and any combination therapies, as well as the number of booster vaccinations. In the case of disease recurrence, tumour DNA sequencing can be repeated (for example, to understand why the vaccine was ineffective for long-term tumour control and/or to identify potential alternative neoantigens), and vaccine-induced T cell responses can be evaluated using both blood and tumour samples to inform decisions regarding subsequent therapy. FLT3L, Fms-related tyrosine kinase 3 ligand; GM-CSF, granulocyte–macrophage colony-stimulating factor.
A personalized long-peptide vaccine formulated with poly-ICLC (NEO-PV-01) has been assessed in combination with nivolumab in the phase Ib NT-001 trial involving patients with advanced-stage melanoma (n = 34), NSCLC (n = 27) or urothelial carcinoma (n = 21)135. Patients received nivolumab monotherapy while the vaccines were being manufactured, as well as during the vaccination and post-vaccination periods. T cells responsive to neoantigens included in the vaccine were detected in all vaccinated patients (n = 60) using ex vivo IFNγ ELISpot assays of peripheral blood135. Vaccine-specific TCRs were detected in post-vaccination tumour samples from two patients with melanoma, one of whom had a radiographic response, and one who had stable disease following treatment135, indicating that the vaccines induced trafficking of neoantigen-specific T cells into metastatic tumours. Clinical benefit observed after vaccination could not be directly attributed to the vaccine owing to the use of nivolumab and the nonrandomized design of the study, although longitudinal assessments of biopsy samples revealed that major pathological responses occurred after vaccination in nine patients with melanoma who did not have such a response to initial pre-vaccination treatment with nivolumab, three of whom had a residual tumour burden of >80% prior to vaccination135. Moreover, evidence of epitope spreading of the T cell responses to neoantigens not contained in the vaccines (suggesting vaccine-mediated tumour cell killing) was observed. Notably, epitope spreading was associated with a lack of disease progression at 6 months after initiation of therapy in the urothelial carcinoma group and at 9 months in the NSCLC and melanoma groups (P = 0.03 across groups), translating into improved PFS in the three groups overall (HR 0.23, 95% CI 0.06–0.83; P = 0.01)135. These findings suggest vaccine-mediated clinical benefit. The combination of nivolumab and NEO-PV-01 was well tolerated, with mild injection-site reaction and transient flu-like symptoms observed in 52% and 35% of patients, respectively.
GEN-009 is a personalized neoantigen-based vaccine comprising 4–20 synthetic long peptides selected using the aforementioned ATLAS epitope-discovery platform and formulated with poly-ICLC. In an ongoing multicentre phase I/IIa study (NCT03633110), eight patients with solid tumours with a high risk of recurrence after completion of curative-intent treatment received GEN-009 and tolerated the vaccine well, with only local injection-site discomfort reported136. All patients were found to have peripheral CD4+ T cell and CD8+responses against at least one neoantigen, and 99% of all peptides stimulated T cell responses (74% induced CD8+ T cell responses and 92% induced CD4+ T cell responses)137. Of note, T cell responses were often sustained for more than 12 months136. Further data from this trial, which also includes a cohort of patients with advanced-stage disease who are receiving the vaccine in combination with PD-1 inhibitors (Table 2), are awaited.
In a phase Ib study137, a personalized RNA–lipoplex neoantigen-based vaccine encoding up to 20 neoantigens (RO7198457) was tested in combination with the anti-PD-L1 antibody atezolizumab in 132 patients with advanced-stage solid tumours (NCT03289962) (Table 2). Circulating T cell responses to a median of 2.6 neoantigens were detected ex vivo in 77% of patients. Vaccine-specific CD8+ T cells were detected at frequencies >5% in the peripheral blood and had a TEM phenotype with high levels of PD-1 expression137. Vaccine-specific TCRs were also detected in post-vaccination but not in pre-vaccination tumour specimens. The antitumour activity of RO7198475 in combination with atezolizumab was modest, with two objective responses in 28 patients (7%) with various advanced-stage solid tumours who were enrolled in the dose-escalation part of the study and objective response rates (ORRs) between 4% and 30% (8% overall) in the expansion cohorts, which comprised patients with triple-negative breast cancer, urothelial carcinoma, RCC, melanoma or NSCLC137. The ORRs were not higher than those expected with PD-L1 inhibition alone in these patient groups. Treatment-related adverse events were primarily systemic reactions (low-grade cytokine-release syndrome, infusion-related reactions or flu-like symptoms).
mRNA-4157, another lipid-encapsulated RNA-based neoantigen-based vaccine, has been tested as monotherapy in 13 patients with high-risk resectable solid tumours and in combination with pembrolizumab in 20 patients with unresectable advanced-stage solid tumours, with the latter group including 12 patients who had disease progression on prior ICI therapy (NCT03313778)138 (Table 2). No dose-limiting toxicities or grade 3–4 adverse events were observed138. Six clinical responses were observed in the 20 patients (ORR 30%) treated with the combination, including two (17%) of the 12 patients who had previously received ICIs.
Do personalized neoantigen-based vaccines mediate antitumour activity in patients with cancer?
Preliminary signals of clinical activity have been obtained using personalized neoantigen-based vaccines in several proof-of-concept studies (conducted primarily in patients with resected melanoma who were at high risk of disease recurrence)35,36. However, initial clinical trials in patients with advanced-stage solid tumours treated with personalized vaccines in combination with ICI have not demonstrated ORRs that are clearly superior to those achieved with ICI monotherapy135,137,138. In the NT-001 study135, the number of patients who converted from initial disease progression during the ICI monotherapy period to a response after vaccination with NEO-PV-01 was modest (13%). The possibility that combination therapy with personalized vaccines and ICI could prolong anti-PD-1-mediated responses remains; however, the personalized neoantigen vaccines used in the studies to date have been mostly ineffective in addressing the issue of primary resistance to PD-1 ICIs, at least when given after the initiation of and/or concurrently with PD-1 inhibition. Randomized trials will be necessary to provide conclusive evidence of improved antitumour activity of PD-1 or PD-L1 ICIs in combination with personalized vaccines and are ongoing in the settings of first-line therapy for advanced-stage melanoma (NCT03815058) and adjuvant therapy for high-risk NSCLC (NCT04267237) or melanoma (NCT03897881) (Table 2).
How can clinical efficacy be improved?
Despite the apparently modest levels of clinical benefit shown to date, correlative analyses of data from the initial trials of neoantigen-based vaccines provided evidence of robust immunogenicity and on-target tumour cell killing. Neoepitope-specific responses of both CD4+ T cells and CD8+ T cells were seen in all vaccinated patients in the phase Ib studies of peptide plus poly-ICLC vaccine NEO-PV-01 and the RNA–lipoplex vaccine RO7198457 (refs135,137). Furthermore, trafficking of vaccine-induced neoantigen-specific T cells to metastatic tumours was observed in both studies135,137. In the trial of NEO-PV-01, epitope spreading and post-vaccination pathological responses correlated with prolonged PFS135. Nevertheless, T cell responses were not consistently detected in ex vivo assays, and a relatively large proportion of the vaccine neoepitopes did not stimulate T cell responses135,137. Furthermore, although large numbers of T cells targeting certain neoepitopes were detected, many vaccine neoepitopes induced only low-frequency responses135,137. As we have described, one challenge in the field of cancer vaccine research relates to improving our understanding of how to induce maximum activation and expansion of T cell responses and of CD8+ T cells in particular. To accomplish this aim, complementary therapies that promote APC function and optimal priming of T cells in the lymph nodes will probably be necessary (Fig. 3). Therapeutic agents with the potential to achieve this include ICIs, agonists of co-stimulatory receptors (such as CD40), TLR agonists and growth factors supporting DC development and/or function (such as granulocyte–macrophage colony-stimulating factor (GM-CSF) and Fms-related tyrosine kinase 3 ligand (FLT3L). We are conducting phase I trials in patients with completely resected advanced-stage RCC (NCT02950766) or unresectable melanoma (NCT03929029), in which personalized neoantigen-based vaccines are being coadministered with ipilimumab, an ICI targeting CTLA4 (Table 2). In preclinical models of lymphoma, in situ vaccination approaches consisting of intumoural injections of FLT3L and poly-ICLC plus local radiotherapy led to the recruitment of cross-priming DCs and improved the efficacy of PD-1 inhibition139. The in situ vaccination strategy alone induced objective responses, including some abscopal responses, in patients with treatment-resistant non-Hodgkin lymphomas; furthermore, non-responders had increases in the abundance of phenotypically exhausted, PD-1+CD8+ T cells in peripheral blood following treatment, supporting the combination of this approach with PD-1 inhibition139.
Another challenge relates to the identification of a vaccine delivery system that enables rapid, cost-effective manufacturing and thus timely deployment of the vaccines (Fig. 3). As reviewed previously140, different vaccine formats — comprising peptides, RNA, DNA, viral constructs or DCs — each have advantages and disadvantages; however, head-to-head comparisons of these different approaches in patients are lacking. This situation most probably reflects the current reality of the clinical drug development landscape, in which vaccine formats are typically chosen based on the available technology, in-house experience and intellectual property, among other criteria. Conceivably, cooperative groups could perform such comparative studies, although systematic comparisons of different personalized vaccine platforms are unlikely to be undertaken in the near term given the current high costs and complexity of this therapeutic approach.
The timing of ICI administration and therapeutic vaccination is another important consideration (Fig. 3). This point was highlighted in a preclinical study in which concurrent administration of a PD-1 inhibitor and a therapeutic cancer vaccine improved both vaccine-induced immune responses and tumour control in a mouse model141. Conversely, PD-1 inhibition before vaccination resulted in a population of PD-1+CD38hiCD8+ T cells that were dysfunctional in the same model, a finding that was attributed to a suboptimal priming environment for induction of CD8+ T cell responses141. Indeed, this PD-1+CD38hiCD8+ T cell population was shown to contribute to immunotherapy failure in mice, and patients with intrinsic resistance to PD-1 ICIs were found to have higher numbers of these dysfunctional T cells in blood and tumour samples than responders141. Thus, combining ICIs with therapeutic vaccination regimens can be beneficial, but will require careful consideration of the most appropriate sequencing of the component therapies.
Conclusions
The ability to rapidly and comprehensively identify tumour-specific mutations has provided the cancer vaccine field with long-elusive tumour-specific targets. Initial studies have shown that personalized neoantigen-based vaccines can be generated and administered safely to individual patients with cancer. In-depth profiling of vaccine-induced T cell responses, including the use of newly available immunogenomic tools, has demonstrated that all vaccinated patients develop T cell responses to subsets of the immunizing epitopes and that vaccine-induced T cells can traffic into metastatic tumours. Neoantigen-prediction algorithms have largely focused on MHC I epitopes, but a predominance of CD4+ T cell responses has been observed. Despite emerging evidence of crucial roles of CD4+ T cells, developing strategies that induce maximal cytotoxic CD8+ T cell responses remains important, in addition to the robust neoantigen-specific CD4+ T cell responses elicited in the initial trials to date (Fig. 4). Numerous studies utilizing a spectrum of different vaccine delivery platforms and combination therapies are underway, with the ultimate goal of stimulating effective, long-lasting, tumour-specific immunity in patients with cancer.
Many areas of advancement might improve the efficacy of personalized vaccines, as indicated in this figure. For example, the identification of targetable neoantigens could be improved through development of better prediction algorithms. Various vaccine modalities and dosing schedules could be explored to generate maximally effective antitumour immune responses. Importantly, detailed assessments of the characteristics of the immune responses induced by vaccination could be used to inform iterative improvements in the approach to vaccine design and administration. These include evaluating the magnitude of neoantigen-specific responses (by ELISpot) and in-depth phenotyping via single-cell RNA sequencing (scRNA-seq) analyses. MHC II, MHC class II; MS, mass spectrometry; scTCR, single-cell T cell receptor; SFC, spot-forming cell; SNV, single-nucleotide variant.
References
Hollingsworth, R. E. & Jansen, K. Turning the corner on therapeutic cancer vaccines. NPJ Vaccines 4, 7–10 (2019).
Vormehr, M., Türeci, Ö. & Sahin, U. Harnessing tumor mutations for truly individualized cancer vaccines. Annu. Rev. Med. 70, 395–407 (2019).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
MD, S. T. et al. Insertion-and-deletion-derived tumour-specific neoantigens and the immunogenic phenotype: a pan-cancer analysis. Lancet Oncol. 18, 1009–1021 (2017).
Yang, W. et al. Immunogenic neoantigens derived from gene fusions stimulate T cell responses. Nat. Med. 25, 767–775 (2019).
Balachandran, V. P. et al. Identification of unique neoantigen qualities in long-term survivors of pancreatic cancer. Nature 108, 914–934 (2017).
Martin, S. D. et al. Low mutation burden in ovarian cancer may limit the utility of neoantigen-targeted vaccines. PLoS ONE 11, e0155189 (2016).
Subudhi, S. K. et al. Neoantigen responses, immune correlates, and favorable outcomes after ipilimumab treatment of patients with prostate cancer. Sci. Transl Med. 12, eaaz3577 (2020).
Hilf, N. et al. Actively personalized vaccination trial for newly diagnosed glioblastoma. Nature 565, 240–245 (2019).
Keskin, D. B. et al. Neoantigen vaccine generates intratumoral T cell responses in phase Ib glioblastoma trial. Nature 565, 234–239 (2019).
van Rooij, N. et al. Tumor exome analysis reveals neoantigen-specific T-cell reactivity in an ipilimumab-responsive melanoma. J. Clin. Oncol. 31, e439–e442 (2013).
Tran, E. et al. Immunogenicity of somatic mutations in human gastrointestinal cancers. Science 350, 1387–1390 (2015).
Giannakis, M. et al. Genomic correlates of immune-cell infiltrates in colorectal carcinoma. Cell Rep. 15, 857–865 (2016).
Brown, S. D. et al. Neo-antigens predicted by tumor genome meta-analysis correlate with increased patient survival. Genome Res. 24, 743–750 (2014).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Strickland, K. C. et al. Association and prognostic significance of BRCA1/2-mutation status with neoantigen load, number of tumor-infiltrating lymphocytes and expression of PD-1/PD-L1 in high grade serous ovarian cancer. Oncotarget 7, 13587–13598 (2016).
Matsushita, H. et al. Neoantigen load, antigen presentation machinery, and immune signatures determine prognosis in clear cell renal cell carcinoma. Cancer Immunol. Res. 4, 463–471 (2016).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
Rizvi, N. A. et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Chan, T. A. et al. Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann. Oncol. 30, 44–56 (2019).
Keenan, T. E., Burke, K. P. & Van Allen, E. M. Genomic correlates of response to immune checkpoint blockade. Nat. Med. 25, 389–402 (2019).
Schumacher, T. N., Scheper, W. & Kvistborg, P. Cancer neoantigens. Annu. Rev. Immunol. 37, 173–200 (2019).
McDermott, D. F. et al. Clinical activity and molecular correlates of response to atezolizumab alone or in combination with bevacizumab versus sunitinib in renal cell carcinoma. Nat. Med. 24, 749–757 (2018).
Miao, D. et al. Genomic correlates of response to immune checkpoint blockade in microsatellite-stable solid tumors. Nat. Genet. 50, 1271–1281 (2018).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Matsushita, H. et al. Cancer exome analysis reveals a T-cell-dependent mechanism of cancer immunoediting. Nature 482, 400–404 (2012).
Verdegaal, E. M. E. et al. Neoantigen landscape dynamics during human melanoma–T cell interactions. Nature 536, 91–95 (2016).
Anagnostou, V. et al. Evolution of neoantigen landscape during immune checkpoint blockade in non-small cell lung cancer. Cancer Discov. 7, 264–276 (2017).
Rosenthal, R. et al. Neoantigen-directed immune escape in lung cancer evolution. Nature 567, 479–485 (2019).
Zhao, J. et al. Immune and genomic correlates of response to anti-PD-1 immunotherapy in glioblastoma. Nat. Med. 25, 462–469 (2019).
Dagogo-Jack, I. & Shaw, A. T. Tumour heterogeneity and resistance to cancer therapies. Nat. Rev. Clin. Oncol. 15, 81–94 (2018).
Ott, P. A. et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 547, 217–221 (2017).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Carreno, B. M. et al. A dendritic cell vaccine increases the breadth and diversity of melanoma neoantigen-specific T cells. Science 348, 803–808 (2015).
Hu, Z., Ott, P. A. & Wu, C. J. Towards personalized, tumour-specific, therapeutic vaccines for cancer. Nat. Rev. Immunol. 18, 168–182 (2018).
Lee, C.-H., Yelensky, R., Jooss, K. & Chan, T. A. Update on tumor neoantigens and their utility: why it is good to be different. Trends Immunol. 39, 536–548 (2018).
Roudko, V., Greenbaum, B. & Bhardwaj, N. Computational prediction and validation of tumor-associated neoantigens. Front. Immunol. 11, 609–611 (2020).
Türeci, Ö. et al. Challenges towards the realization of individualized cancer vaccines. Nat. Biomed. Eng. 2, 566–569 (2018).
Robinson, J. et al. IPD-IMGT/HLA database. Nucleic Acids Res. 55, 280–288 (2019).
Fritsch, E. F. et al. HLA-binding properties of tumor neoepitopes in humans. Cancer Immunol. Res. 2, 522–529 (2014).
Peters, B., Nielsen, M. & Sette, A. T cell epitope predictions. Annu. Rev. Immunol. 38, 123–145 (2020).
Jurtz, V. et al. NetMHCpan-4.0: improved peptide-MHC class I interaction predictions integrating eluted ligand and peptide binding affinity data. J. Immunol. 199, 3360–3368 (2017).
Abelin, J. G. et al. Mass spectrometry profiling of HLA-associated peptidomes in mono-allelic cells enables more accurate epitope prediction. Immunity 46, 315–326 (2017).
Bulik-Sullivan, B. et al. Deep learning using tumor HLA peptide mass spectrometry datasets improves neoantigen identification. Nat. Biotechnol. 37, 55–63 (2019).
Sarkizova, S. et al. A large peptidome dataset improves HLA class I epitope prediction across most of the human population. Nat. Biotechnol. 38, 199–209 (2020).
Capietto, A.-H. et al. Mutation position is an important determinant for predicting cancer neoantigens. J. Exp. Med. 217, 315–322 (2020).
Andreatta, M. et al. Accurate pan-specific prediction of peptide-MHC class II binding affinity with improved binding core identification. Immunogenetics 67, 641–650 (2015).
Bijker, M. S. et al. CD8+ CTL priming by exact peptide epitopes in incomplete freund’s adjuvant induces a vanishing CTL response, whereas long peptides induce sustained CTL reactivity. J. Immunol. 179, 5033–5040 (2007).
Kenter, G. G. et al. Vaccination against HPV-16 oncoproteins for vulvar intraepithelial neoplasia. N. Engl. J. Med. 361, 1838–1847 (2009).
Kreiter, S. et al. Mutant MHC class II epitopes drive therapeutic immune responses to cancer. Nature 520, 692–696 (2015).
Rossjohn, J. et al. T cell antigen receptor recognition of antigen-presenting molecules. Annu. Rev. Immunol. 33, 169–200 (2015).
Mildner, A. & Jung, S. Development and function of dendritic cell subsets. Immunity 40, 642–656 (2014).
Abelin, J. G. et al. Defining HLA-II ligand processing and binding rules with mass spectrometry enhances cancer epitope prediction. Immunity 51, 766–779.e17 (2019).
Racle, J. et al. Robust prediction of HLA class II epitopes by deep motif deconvolution of immunopeptidomes. Nat. Biotechnol. 37, 1283–1286 (2019).
Reynisson, B. et al. Improved prediction of MHC II antigen presentation through integration and motif deconvolution of mass spectrometry MHC eluted ligand data. J. Proteome Res. 19, 2304–2315 (2020).
Harari, A., Graciotti, M., Bassani-Sternberg, M. & Kandalaft, L. E. Antitumour dendritic cell vaccination in a priming and boosting approach. Nat. Rev. Drug Discov. 19, 635–652 (2020).
Tanyi, J. L. et al. Personalized cancer vaccine effectively mobilizes antitumor T cell immunity in ovarian cancer. Sci. Transl Med. 10, eaao5931 (2018).
Lu, Y.-C. et al. Efficient identification of mutated cancer antigens recognized by T cells associated with durable tumor regressions. Clin. Cancer Res. 20, 3401–3410 (2014).
Lu, Y.-C. et al. An efficient single-cell RNA-Seq approach to identify neoantigen-specific T cell receptors. Mol. Ther. 26, 379–389 (2018).
Cafri, G. et al. Memory T cells targeting oncogenic mutations detected in peripheral blood of epithelial cancer patients. Nat. Commun. 10, 449 (2019).
Leko, V. et al. Identification of neoantigen-reactive tumor-infiltrating lymphocytes in primary bladder cancer. J. Immunol. 202, 3458–3467 (2019).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Malonis, R. J., Lai, J. R. & Vergnolle, O. Peptide-based vaccines: current progress and future challenges. Chem. Rev. 120, 3210–3229 (2020).
Pardi, N., Hogan, M. J., Porter, F. W. & Weissman, D. mRNA vaccines — a new era in vaccinology. Nat. Rev. Clin. Oncol. 17, 261–279 (2018).
Gary, E. N. & Weiner, D. B. DNA vaccines: prime time is now. Curr. Opin. Immunol. 65, 21–27 (2020).
Saxena, M. & Bhardwaj, N. Re-emergence of dendritic cell vaccines for cancer treatment. Trends Cancer 4, 119–137 (2018).
Humphreys, I. R. & Sebastian, S. Novel viral vectors in infectious diseases. Immunology 153, 1–9 (2017).
Harty, J. T. & Badovinac, V. P. Shaping and reshaping CD8+ T-cell memory. Nat. Rev. Immunol. 8, 107–119 (2008).
Fraser, K. A., Schenkel, J. M., Jameson, S. C., Vezys, V. & Masopust, D. Preexisting high frequencies of memory CD8+ T cells favor rapid memory differentiation and preservation of proliferative potential upon boosting. Immunity 39, 171–183 (2013).
Wirth, T. C. et al. Repetitive antigen stimulation induces stepwise transcriptome diversification but preserves a core signature of memory CD8+ T cell differentiation. Immunity 33, 128–140 (2010).
Masopust, D., Ha, S.-J., Vezys, V. & Ahmed, R. Stimulation history dictates memory CD8 T cell phenotype: implications for prime-boost vaccination. J. Immunol. 177, 831–839 (2006).
Sallusto, F., Lanzavecchia, A., Araki, K. & Ahmed, R. From vaccines to memory and back. Immunity 33, 451–463 (2010).
Tscharke, D. C., Croft, N. P., Doherty, P. C. & La Gruta, N. L. Sizing up the key determinants of the CD8 + T cell response. Nat. Rev. Immunol. 15, 705–716 (2015).
Kaech, S. M. & Cui, W. Transcriptional control of effector and memory CD8+ T cell differentiation. Nat. Rev. Immunol. 12, 749–761 (2012).
Martin, M. D. & Badovinac, V. P. Defining memory CD8 T cell. Front. Immunol. 9, 2692 (2018).
Plitas, G. & Rudensky, A. Y. Regulatory T cells in cancer. Annu. Rev. Cancer Biol. 4, 459–477 (2020).
Wculek, S. K. et al. Dendritic cells in cancer immunology and immunotherapy. Nat. Rev. Immunol. 20, 7–24 (2019).
Jameson, S. C. & Masopust, D. Understanding subset diversity in T cell memory. Immunity 48, 214–226 (2018).
Gerlach, C. et al. The chemokine receptor CX3CR1 defines three antigen-experienced CD8 T cell subsets with distinct roles in immune surveillance and homeostasis. Immunity 45, 1270–1284 (2016).
Schenkel, J. M. et al. Resident memory CD8 T cells trigger protective innate and adaptive immune responses. Science 346, 98–101 (2014).
Mueller, S. N. & Mackay, L. K. Tissue-resident memory T cells: local specialists in immune defence. Nat. Rev. Immunol. 16, 79–89 (2015).
Szabo, P. A., Miron, M. & Farber, D. L. Location, location, location: tissue resident memory T cells in mice and humans. Sci. Immunol. 4, eaas9673 (2019).
Gebhardt, T. et al. Different patterns of peripheral migration by memory CD4+ and CD8+ T cells. Nature 477, 216–219 (2011).
Park, S. L., Gebhardt, T. & Mackay, L. K. Tissue-resident memory T cells in cancer immunosurveillance. Trends Immunol. 40, 735–747 (2019).
Blanc, C. et al. Targeting resident memory T cells for cancer immunotherapy. Front. Immunol. 9, 524–527 (2018).
Nizard, M. et al. Induction of resident memory T cells enhances the efficacy of cancer vaccine. Nat. Commun. 8, 1–11 (2017).
Ganesan, A.-P. et al. Tissue-resident memory features are linked to the magnitude of cytotoxic T cell responses in human lung cancer. Nat. Immunol. 18, 940–950 (2017).
Duhen, T. et al. Co-expression of CD39 and CD103 identifies tumor-reactive CD8 T cells in human solid tumors. Nat. Commun. 9, 2724 (2018).
Clarke, J. et al. Single-cell transcriptomic analysis of tissue-resident memory T cells in human lung cancer. J. Exp. Med. 216, 2128–2149 (2019).
Dijkgraaf, F. E. et al. Tissue patrol by resident memory CD8+ T cells in human skin. Nat. Immunol. 20, 756–764 (2019).
Steinert, E. M. et al. Quantifying memory CD8 T cells reveals regionalization of immunosurveillance. Cell 161, 737–749 (2015).
McLane, L. M., Abdel-Hakeem, M. S. & Wherry, E. J. CD8 T cell exhaustion during chronic viral infection and cancer. Annu. Rev. Immunol. 37, 457–495 (2019).
Wherry, E. J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007).
Angelosanto, J. M., Blackburn, S. D., Crawford, A. & Wherry, E. J. Progressive loss of memory T cell potential and commitment to exhaustion during chronic viral infection. J. Virol. 86, 8161–8170 (2012).
Pauken, K. E. et al. Epigenetic stability of exhausted T cells limits durability of reinvigoration by PD-1 blockade. Science 354, 1160–1165 (2016).
Philip, M. et al. Chromatin states define tumour-specific T cell dysfunction and reprogramming. Nature 545, 452–456 (2017).
Sen, D. R. et al. The epigenetic landscape of T cell exhaustion. Science 354, 1165–1169 (2016).
Ghoneim, H. E. et al. De novo epigenetic programs inhibit PD-1 blockade- mediated T cell rejuvenation. Cell 170, 142–157.e19 (2017).
Scott-Browne, J. P. et al. Dynamic changes in chromatin accessibility occur in CD8. Immunity 45, 1327–1340 (2016).
Jansen, C. S. et al. An intra-tumoral niche maintains and differentiates stem-like CD8 T cells. Nature 476, 465–470 (2019).
Miller, B. C. et al. Subsets of exhausted CD8+ T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Brummelman, J. et al. High-dimensional single cell analysis identifies stem-like cytotoxic CD8+ T cells infiltrating human tumors. J. Exp. Med. 215, 2520–2535 (2018).
Siddiqui, I. et al. Intratumoral Tcf1+PD-1+CD8+T cells with stem-like properties promote tumor control in response to vaccination and checkpoint blockade immunotherapy. Immunity 50, 195–211.e10 (2019).
Laidlaw, B. J., Craft, J. E. & Kaech, S. M. The multifaceted role of CD4+T cells in CD8+T cell memory. Nat. Rev. Immunol. 16, 102–111 (2016).
Baxevanis, C. N. et al. Tumor-specific CD4+T lymphocytes from cancer patients are required for optimal induction of cytotoxic T cells against the autologous tumor. J. Immunol. 164, 3902–3912 (2000).
Marzo, A. L. et al. Tumor-specific CD4+ T cells have a major ‘post-licensing’ role in CTL mediated anti-tumor immunity. J. Immunol. 165, 6047–6055 (2000).
Ossendorp, F., Mengedé, E., Camps, M., Filius, R. & Melief, C. J. Specific T helper cell requirement for optimal induction of cytotoxic T lymphocytes against major histocompatibility complex class II negative tumors. J. Exp. Med. 187, 693–702 (1998).
Alspach, E. et al. MHC-II neoantigens shape tumour immunity and response to immunotherapy. Nature 574, 696–701 (2019).
Zander, R. et al. CD4+ T cell help is required for the formation of a cytolytic CD8+T cell subset that protects against chronic infection and cancer. Immunity 51, 1028–1042.e4 (2019).
Ahrends, T. & Borst, J. The opposing roles of CD4+ T cells in anti-tumour immunity. Immunology 154, 582–592 (2018).
Bos, R. & Sherman, L. A. CD4+ T-cell help in the tumor milieu is required for recruitment and cytolytic function of CD8+ T lymphocytes. Cancer Res. 70, 8368–8377 (2010).
Corthay, A. et al. Primary antitumor immune response mediated by CD4+ T cells. Immunity 22, 371–383 (2005).
Wong, S. B. J., Bos, R. & Sherman, L. A. Tumor-specific CD4+ T cells render the tumor environment permissive for infiltration by low-avidity CD8+ T cells. J. Immunol. 180, 3122–3131 (2008).
Mumberg, D. et al. CD4+ T cells eliminate MHC class II-negative cancer cells in vivo by indirect effects of IFN-gamma. Proc. Natl Acad. Sci. USA 96, 8633–8638 (1999).
Spitzer, M. H. et al. Systemic immunity is required for effective cancer immunotherapy. Cell 168, 487–502.e15 (2017).
Linnemann, C. et al. High-throughput epitope discovery reveals frequent recognition of neo-antigens by CD4+ T cells in human melanoma. Nat. Med. 21, 81–85 (2014).
Hunder, N. N. et al. Treatment of metastatic melanoma with autologous CD4+T cells against NY-ESO-1. N. Engl. J. Med. 358, 2698–2703 (2008).
Tran, E. et al. Cancer immunotherapy based on mutation-specific CD4+T cells in a patient with epithelial cancer. Science 344, 641–645 (2014).
Melssen, M. & Slingluff, C. L. Jr. Vaccines targeting helper T cells for cancer immunotherapy. Curr. Opin. Immunol. 47, 85–92 (2017).
Wilk, M. M. & Mills, K. H. G. CD4 TRM cells following infection and immunization: implications for more effective vaccine design. Front. Immunol. 9, 2413–2418 (2018).
Aarntzen, E. H. J. G. et al. Targeting CD4+T-helper cells improves the induction of antitumor responses in dendritic cell–based vaccination. Cancer Res. 73, 19–29 (2013).
Ahrends, T. et al. CD4+T cell help confers a cytotoxic T cell effector program including coinhibitory receptor downregulation and increased tissue invasiveness. Immunity 47, 848–861.e5 (2017).
Moynihan, K. D. et al. Eradication of large established tumors in mice by combination immunotherapy that engages innate and adaptive immune responses. Nat. Med. 22, 1402–1410 (2016).
Jaigirdar, S. A. & MacLeod, M. K. L. Development and function of protective and pathologic memory CD4 T cells. Front. Immunol. 6, 456 (2015).
Magen, A. et al. Single-cell profiling defines transcriptomic signatures specific to tumor-reactive versus virus-responsive CD4+T cells. Cell Rep. 29, 3019–3032.e6 (2019).
Axelrod, M. L., Cook, R. S., Johnson, D. B. & Balko, J. M. Biological consequences of MHC-II expression by tumor cells in cancer. Clin. Cancer Res. 25, 2392–2402 (2019).
Quezada, S. A. et al. Tumor-reactive CD4+T cells develop cytotoxic activity and eradicate large established melanoma after transfer into lymphopenic hosts. J. Exp. Med. 207, 637–650 (2010).
Oh, D. Y. et al. Intratumoral CD4+ T cells mediate anti-tumor cytotoxicity in human bladder cancer. Cell 181, 1612–1625 (2020).
Ahmadzadeh, M. et al. Tumor-infiltrating human CD4+ regulatory T cells display a distinct TCR repertoire and exhibit tumor and neoantigen reactivity. Sci. Immunol. 4, eaao4310 (2019).
Cohen, R. B. et al. GEN-009, a neoantigen vaccine containing ATLAS selected neoantigens, to generate broad sustained immunity against immunogenic tumor mutations and avoid inhibitory peptides. J. Clin. Oncol. 38, 3107–3107 (2020).
Nogueira, C., Kaufmann, J. K., Lam, H. & Flechtner, J. B. Improving cancer immunotherapies through empirical neoantigen selection. Trends Cancer 4, 97–100 (2018).
Ott, P. A. et al. A phase Ib trial of personalized neoantigen therapy plus anti-PD-1 in patients with advanced melanoma, non-small cell lung cancer, bladder cancer. Cell 183, 347–362.e24 (2020).
Cohen, R. B. et al. Broad immunogenicity from GEN-009, a neoantigen vaccine using ATLASTM, an autologous immune assay, to identify immunogenic and inhibitory tumor neoantigens. J. Immunother. Cancer 7 (Suppl. 1), P420 (2019).
Lopez, J. S. et al. A phase Ib study to evaluate RO7198457, an individualized neoantigen specific immunotherapy (iNeST), in combination with atezolizumab in patients with locally advanced or metastatic solid tumors. Cancer Res. 80, CT301 (2020).
Burris, H. A. et al. A phase I multicenter study to assess the safety, tolerability, and immunogenicity of mRNA-4157 alone in patients with resected solid tumors and in combination with pembrolizumab in patients with unresectable solid tumors. J. Clin. Oncol. 37 (Suppl. 15), 2523 (2019).
Hammerich, L. et al. Systemic clinical tumor regressions and potentiation of PD1 blockade with in situ vaccination. Nat. Med. 25, 814–824 (2019).
Sahin, U. & Türeci, Ö. Personalized vaccines for cancer immunotherapy. Science 359, 1355–1360 (2018).
Verma, V. et al. PD-1 blockade in subprimed CD8 cells induces dysfunctional PD-1+ CD38hi cells and anti-PD-1 resistance. Nat. Immunol. 20, 1231–1243 (2019).
Rajasagi, M. et al. Systematic identification of personal tumor-specific neoantigens in chronic lymphocytic leukemia. Blood 124, 453–462 (2014).
Blazquez, A. et al. A phase I study of the safety and immunogenicity of a multi-peptide personalized genomic vaccine in the adjuvant treatment of solid tumors and hematological malignancies. J. Clin. Oncol. 37 (Suppl. 15), e14307 (2019).
Rubinsteyn, A. et al. Computational pipeline for the PGV-001 neoantigen vaccine trial. Front. Immunol. 8, 1807 (2017).
Wesolowski R. et al. A phase 1 study of safety and tolerability of AutoSynVax™ vaccine in patients with advanced cancer. Presented at the Society for the Immunotherapy of Cancer Annual Meeting (2018).
Krauss J. et al. Preliminary safety, efficacy and immunogenicity results from a phase 1/2a study (DIRECT-01) of cancer neoantigen DNA vaccine VB10.NEO in patients with locally advanced or metastatic solid tumors. Presented at the Society for the Immunotherapy of Cancer Annual Meeting (2019).
Acknowledgements
The work of P.A.O. is supported by the National Institute for Health Research (NIHR) Efficacy and Mechanism Evaluation Programme (National Cancer Institute grant 1R01CA229261-01), a Team Science Award from the Melanoma Research Alliance, the Francis and Adele Kittredge Family Immuno-Oncology and Melanoma Research Fund, the Faircloth Family Research Fund, the Bender Family Research Fund, and the Dana-Farber Cancer Institute (DFCI) Center for Cancer Immunotherapy Research fellowship.
Author information
Authors and Affiliations
Contributions
Both authors contributed to all stages of the preparation of this manuscript for publication.
Corresponding author
Ethics declarations
Competing interests
P.A.O. has received research funding from and has been an adviser of Amgen, Armo BioSciences, Array, AstraZeneca/MedImmune, Bristol-Myers Squibb, Celldex, CytomX, Merck, Neon Therapeutics, Novartis, Pfizer and Roche/Genentech. E.B. declares no competing interests.
Additional information
Peer review information
Nature Reviews Clinical Oncology thanks Craig Slingluff, Lana Kandalaft and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Blass, E., Ott, P.A. Advances in the development of personalized neoantigen-based therapeutic cancer vaccines. Nat Rev Clin Oncol 18, 215–229 (2021). https://doi.org/10.1038/s41571-020-00460-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41571-020-00460-2
This article is cited by
-
Immunosuppressive tumor microenvironment and immunotherapy of hepatocellular carcinoma: current status and prospectives
Journal of Hematology & Oncology (2024)
-
The potential and promise for clinical application of adoptive T cell therapy in cancer
Journal of Translational Medicine (2024)
-
An engineered influenza virus to deliver antigens for lung cancer vaccination
Nature Biotechnology (2024)
-
ERK pathway agonism for cancer therapy: evidence, insights, and a target discovery framework
npj Precision Oncology (2024)
-
Systematic pan-cancer analyses of the potential function of the Golgi scaffold protein PAQR3
Scientific Reports (2024)