Abstract
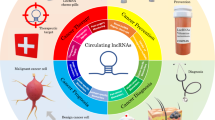
Over the past decade, the amount of research and the number of publications on associations between circulating small and long non-coding RNAs (ncRNAs) and cancer have grown exponentially. Particular focus has been placed on the development of diagnostic and prognostic biomarkers to enable efficient patient management — from early detection of cancer to monitoring for disease recurrence or progression after treatment. Owing to their high abundance and stability, circulating ncRNAs have potential utility as non-invasive, blood-based biomarkers that can provide information on tumour biology and the effects of treatments, such as targeted therapies and immunotherapies. Increasing evidence highlights the roles of ncRNAs in cell-to-cell communication, with a number of ncRNAs having the capacity to regulate gene expression outside of the cell of origin through extracellular vesicle-mediated transfer to recipient cells, with implications for cancer progression and therapy resistance. Moreover, ‘foreign’ microRNAs (miRNAs) encoded by non-human genomes (so-called xeno-miRNAs), such as viral miRNAs, have been shown to be present in human body fluids and can be used as biomarkers. Herein, we review the latest developments in the use of circulating ncRNAs as diagnostic and prognostic biomarkers and discuss their roles in cell-to-cell communication in the context of cancer. We provide a compendium of miRNAs and long ncRNAs that have been reported in the literature to be present in human body fluids and that have the potential to be used as diagnostic and prognostic cancer biomarkers.
Key points
-
Non-coding RNAs (ncRNAs) have important roles in regulating the expression of genes that control fundamental biological functions; altered levels of ncRNAs are frequently detected in cancer cells compared with nonmalignant cells.
-
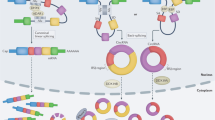
ncRNAs can be secreted into extracellular space as cell-free RNAs either encapsulated inside macrovesicles (such as extracellular vesicles) or in association with proteins (as RNA–protein complexes) or high-density lipoproteins (HDLs), and can subsequently be detected in blood samples as circulating ncRNAs.
-
Altered levels of circulating ncRNAs are found in body fluids, including serum, plasma, urine, and saliva, from patients with cancer compared with individuals without cancer and have potential clinical applications as diagnostic and prognostic blood-based biomarkers.
-
Circulating ncRNAs can mediate cell-to-cell communication and regulate gene expression in recipient cells, thereby acting similarly to hormones (mediating communication between cells within an individual) or as exogenous messengers (participating in inter-individual and cross-species molecular communication).
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Quail, D. F. & Joyce, J. A. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437 (2013).
Becker, A. et al. Extracellular vesicles in cancer: cell-to-cell mediators of metastasis. Cancer Cell 30, 836–848 (2016).
Redis, R. S., Calin, S., Yang, Y., You, M. J. & Calin, G. A. Cell-to-cell miRNA transfer: from body homeostasis to therapy. Pharmacol. Ther. 136, 169–174 (2012).
Pucci, M. et al. Extracellular vesicles as miRNA nano-shuttles: dual role in tumor progression. Target Oncol. 13, 175–187 (2018).
Berindan-Neagoe, I., Monroig Pdel, C., Pasculli, B. & Calin, G. A. MicroRNAome genome: a treasure for cancer diagnosis and therapy. CA Cancer J. Clin. 64, 311–336 (2014).
Cortez, M. A. et al. MicroRNAs in body fluids — the mix of hormones and biomarkers. Nat. Rev. Clin. Oncol. 8, 467–477 (2011).
Bejerano, G. et al. Ultraconserved elements in the human genome. Science 304, 1321–1325 (2004).
Calin, G. A. et al. Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 12, 215–229 (2007).
Rigoutsos, I. et al. N-BLR, a primate-specific non-coding transcript leads to colorectal cancer invasion and migration. Genome Biol. 18, 98 (2017).
Hansen, T. B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388 (2013).
Legnini, I. et al. Circ-ZNF609 is a circular rna that can be translated and functions in myogenesis. Mol. Cell 66, 22–37.e9 (2017).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659 (2007).
Conigliaro, A. et al. CD90+ liver cancer cells modulate endothelial cell phenotype through the release of exosomes containing H19 lncRNA. Mol. Cancer 14, 155 (2015).
Li, Y. et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res. 25, 981–984 (2015).
Arroyo, J. D. et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl Acad. Sci. USA 108, 5003–5008 (2011).
Turchinovich, A., Weiz, L., Langheinz, A. & Burwinkel, B. Characterization of extracellular circulating microRNA. Nucleic Acids Res. 39, 7223–7233 (2011).
Vickers, K. C., Palmisano, B. T., Shoucri, B. M., Shamburek, R. D. & Remaley, A. T. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat. Cell Biol. 13, 423–433 (2011).
Qi, P., Zhou, X. Y. & Du, X. Circulating long non-coding RNAs in cancer: current status and future perspectives. Mol. Cancer 15, 39 (2016).
Fabris, L. & Calin, G. A. Circulating free xeno-microRNAs — the new kids on the block. Mol. Oncol. 10, 503–508 (2016).
Mattick, J. S. & Makunin, I. V. Non-coding RNA. Hum. Mol. Genet. 15, R17–R29 (2006).
Volders, P. J. et al. LNCipedia: a database for annotated human lncRNA transcript sequences and structures. Nucleic Acids Res. 41, D246–D251 (2013).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Atianand, M. K., Caffrey, D. R. & Fitzgerald, K. A. Immunobiology of long noncoding RNAs. Annu. Rev. Immunol. 35, 177–198 (2017).
Marchese, F. P., Raimondi, I. & Huarte, M. The multidimensional mechanisms of long noncoding RNA function. Genome Biol. 18, 206 (2017).
Alvarez-Dominguez, J. R. & Lodish, H. F. Emerging mechanisms of long noncoding RNA function during normal and malignant hematopoiesis. Blood 130, 1965–1975 (2017).
Chen, Y. G., Satpathy, A. T. & Chang, H. Y. Gene regulation in the immune system by long noncoding RNAs. Nat. Immunol. 18, 962–972 (2017).
Esteller, M. Non-coding RNAs in human disease. Nat. Rev. Genet. 12, 861–874 (2011).
Morris, K. V. & Mattick, J. S. The rise of regulatory RNA. Nat. Rev. Genet. 15, 423–437 (2014).
Li, L. & Chang, H. Y. Physiological roles of long noncoding RNAs: insight from knockout mice. Trends Cell Biol. 24, 594–602 (2014).
Bartonicek, N., Maag, J. L. & Dinger, M. E. Long noncoding RNAs in cancer: mechanisms of action and technological advancements. Mol. Cancer 15, 43 (2016).
Spizzo, R., Almeida, M. I., Colombatti, A. & Calin, G. A. Long non-coding RNAs and cancer: a new frontier of translational research? Oncogene 31, 4577–4587 (2012).
Zhang, R. et al. Serum long non coding RNA MALAT-1 protected by exosomes is up-regulated and promotes cell proliferation and migration in non-small cell lung cancer. Biochem. Biophys. Res. Commun. 490, 406–414 (2017).
Liu, T. et al. Exosomal long noncoding RNA CRNDE-h as a novel serum-based biomarker for diagnosis and prognosis of colorectal cancer. Oncotarget 7, 85551–85563 (2016).
Isin, M. et al. Exosomal lncRNA-p21 levels may help to distinguish prostate cancer from benign disease. Front. Genet. 6, 168 (2015).
Nilsson, J. et al. Prostate cancer-derived urine exosomes: a novel approach to biomarkers for prostate cancer. Br. J. Cancer 100, 1603–1607 (2009).
Berrondo, C. et al. Expression of the long non-coding RNA HOTAIR correlates with disease progression in bladder cancer and is contained in bladder cancer patient urinary exosomes. PLoS ONE 11, e0147236 (2016).
Yuan, T. et al. Plasma extracellular RNA profiles in healthy and cancer patients. Sci. Rep. 6, 19413 (2016).
Bahn, J. H. et al. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin. Chem. 61, 221–230 (2015).
Sobala, A. & Hutvagner, G. Transfer RNA-derived fragments: origins, processing, and functions. Wiley Interdiscip. Rev. RNA 2, 853–862 (2011).
Dhahbi, J. M. et al. 5’ tRNA halves are present as abundant complexes in serum, concentrated in blood cells, and modulated by aging and calorie restriction. BMC Genomics 14, 298 (2013).
Dhahbi, J. M. et al. 5′-YRNA fragments derived by processing of transcripts from specific YRNA genes and pseudogenes are abundant in human serum and plasma. Physiol. Genom. 45, 990–998 (2013).
Christov, C. P., Trivier, E. & Krude, T. Noncoding human Y RNAs are overexpressed in tumours and required for cell proliferation. Br. J. Cancer 98, 981–988 (2008).
Martens-Uzunova, E. S. et al. Diagnostic and prognostic signatures from the small non-coding RNA transcriptome in prostate cancer. Oncogene 31, 978–991 (2012).
Martens-Uzunova, E. S., Olvedy, M. & Jenster, G. Beyond microRNA — novel RNAs derived from small non-coding RNA and their implication in cancer. Cancer Lett. 340, 201–211 (2013).
Dhahbi, J. M., Spindler, S. R., Atamna, H., Boffelli, D. & Martin, D. I. Deep sequencing of serum small RNAs identifies patterns of 5’ tRNA half and YRNA fragment expression associated with breast cancer. Biomark Cancer 6, 37–47 (2014).
Zhao, C. et al. 5′-tRNA halves are dysregulated in clear cell renal cell carcinoma. J. Urol. 199, 378–383 (2018).
Scholkmann, F., Fels, D. & Cifra, M. Non-chemical and non-contact cell-to-cell communication: a short review. Am. J. Transl Res. 5, 586–593 (2013).
Davis, D. M. & Sowinski, S. Membrane nanotubes: dynamic long-distance connections between animal cells. Nat. Rev. Mol. Cell Biol. 9, 431–436 (2008).
Karsenty, G. & Olson, E. N. Bone and muscle endocrine functions: unexpected paradigms of inter-organ communication. Cell 164, 1248–1256 (2016).
Skog, J. et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 10, 1470–1476 (2008).
Pitt, J. M., Kroemer, G. & Zitvogel, L. Extracellular vesicles: masters of intercellular communication and potential clinical interventions. J. Clin. Invest. 126, 1139–1143 (2016).
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell Biol. 200, 373–383 (2013).
Kowal, J., Tkach, M. & Thery, C. Biogenesis and secretion of exosomes. Curr. Opin. Cell Biol. 29, 116–125 (2014).
Colombo, M., Raposo, G. & Thery, C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu. Rev. Cell Dev. Biol. 30, 255–289 (2014).
Zoller, M. Tetraspanins: push and pull in suppressing and promoting metastasis. Nat. Rev. Cancer 9, 40–55 (2009).
Thery, C., Zitvogel, L. & Amigorena, S. Exosomes: composition, biogenesis and function. Nat. Rev. Immunol. 2, 569–579 (2002).
Zhang, H. et al. Exosome-delivered EGFR regulates liver microenvironment to promote gastric cancer liver metastasis. Nat. Commun. 8, 15016 (2017).
Phuyal, S., Hessvik, N. P., Skotland, T., Sandvig, K. & Llorente, A. Regulation of exosome release by glycosphingolipids and flotillins. FEBS J. 281, 2214–2227 (2014).
Thery, C., Ostrowski, M. & Segura, E. Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 9, 581–593 (2009).
Thakur, B. K. et al. Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell Res. 24, 766–769 (2014).
Bolukbasi, M. F. et al. miR-1289 and “zipcode”-like sequence enrich mRNAs in microvesicles. Mol. Ther. Nucleic Acids 1, e10 (2012).
Qu, L. et al. Exosome-transmitted lncARSR promotes sunitinib resistance in renal cancer by acting as a competing endogenous RNA. Cancer Cell 29, 653–668 (2016).
Ying, W. et al. Adipose tissue macrophage-derived exosomal miRNAs can modulate in vivo and in vitro insulin sensitivity. Cell 171, 372–384.e12 (2017).
Alexander, M. et al. Exosome-delivered microRNAs modulate the inflammatory response to endotoxin. Nat. Commun. 6, 7321 (2015).
Bala, S. et al. Biodistribution and function of extracellular miRNA-155 in mice. Sci. Rep. 5, 10721 (2015).
Fabbri, M. et al. MicroRNAs bind to Toll-like receptors to induce prometastatic inflammatory response. Proc. Natl Acad. Sci. USA 109, E2110–E2116 (2012).
Liu, Y. et al. Tumor exosomal RNAs promote lung pre-metastatic niche formation by activating alveolar epithelial TLR3 to recruit neutrophils. Cancer Cell 30, 243–256 (2016).
Brannan, C. I., Dees, E. C., Ingram, R. S. & Tilghman, S. M. The product of the H19 gene may function as an RNA. Mol. Cell. Biol. 10, 28–36 (1990).
Matouk, I. J. et al. The H19 non-coding RNA is essential for human tumor growth. PLoS ONE 2, e845 (2007).
Iizuka, N. et al. Comparison of gene expression profiles between hepatitis B virus- and hepatitis C virus-infected hepatocellular carcinoma by oligonucleotide microarray data on the basis of a supervised learning method. Cancer Res. 62, 3939–3944 (2002).
Barsyte-Lovejoy, D. et al. The c-Myc oncogene directly induces the H19 noncoding RNA by allele-specific binding to potentiate tumorigenesis. Cancer Res. 66, 5330–5337 (2006).
Adriaenssens, E. et al. Cross-talk between mesenchyme and epithelium increases H19 gene expression during scattering and morphogenesis of epithelial cells. Exp. Cell Res. 275, 215–229 (2002).
Matouk, I. J. et al. Oncofetal H19 RNA promotes tumor metastasis. Biochim. Biophys. Acta 1843, 1414–1426 (2014).
Zhang, L. et al. Epigenetic activation of the MiR-200 family contributes to H19-mediated metastasis suppression in hepatocellular carcinoma. Carcinogenesis 34, 577–586 (2013).
Schultheiss, C. et al. The long non-coding RNA H19 suppresses carcinogenesis and chemoresistance in hepatocellular carcinoma. Cell Stress 1, 37–54 (2017).
Kogure, T., Yan, I. K., Lin, W. L. & Patel, T. Extracellular vesicle-mediated transfer of a novel long noncoding RNA TUC339: a mechanism of intercellular signaling in human hepatocellular cancer. Genes Cancer 4, 261–272 (2013).
Vannini, I. et al. Transcribed ultraconserved region 339 promotes carcinogenesis by modulating tumor suppressor microRNAs. Nat. Commun. 8, 1801 (2017).
Gezer, U., Ozgur, E., Cetinkaya, M., Isin, M. & Dalay, N. Long non-coding RNAs with low expression levels in cells are enriched in secreted exosomes. Cell Biol. Int. 38, 1076–1079 (2014).
Takahashi, K., Yan, I. K., Kogure, T., Haga, H. & Patel, T. Extracellular vesicle-mediated transfer of long non-coding RNA ROR modulates chemosensitivity in human hepatocellular cancer. FEBS Open Bio 4, 458–467 (2014).
Takahashi, K., Yan, I. K., Haga, H. & Patel, T. Modulation of hypoxia-signaling pathways by extracellular linc-RoR. J. Cell Sci. 127, 1585–1594 (2014).
Wang, Y. et al. Endogenous miRNA sponge lincRNA-RoR regulates Oct4, Nanog, and Sox2 in human embryonic stem cell self-renewal. Dev. Cell 25, 69–80 (2013).
Xu, Q. et al. MiR-145 directly targets p70S6K1 in cancer cells to inhibit tumor growth and angiogenesis. Nucleic Acids Res. 40, 761–774 (2012).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013).
Chen, X., Liang, H., Zhang, J., Zen, K. & Zhang, C. Y. Secreted microRNAs: a new form of intercellular communication. Trends Cell Biol. 22, 125–132 (2012).
Kawano, Y. et al. Plasma viral microRNA profiles reveal potential biomarkers for chronic active Epstein-Barr virus infection. J. Infect. Dis. 208, 771–779 (2013).
Fuentes-Mattei, E. et al. Plasma viral miRNAs indicate a high prevalence of occult viral infections. EBioMedicine 20, 182–192 (2017).
Lagatie, O., Van Loy, T., Tritsmans, L. & Stuyver, L. J. Viral miRNAs in plasma and urine divulge JC polyomavirus infection. Virol. J. 11, 158 (2014).
Tudor, S. et al. Cellular and Kaposi’s sarcoma-associated herpes virus microRNAs in sepsis and surgical trauma. Cell Death Dis. 5, e1559 (2014).
Pinsky, M. R. et al. Serum cytokine levels in human septic shock. Relation to multiple-system organ failure and mortality. Chest 103, 565–575 (1993).
Giza, D. E. et al. Cellular and viral microRNAs in sepsis: mechanisms of action and clinical applications. Cell Death Differ. 23, 1906–1918 (2016).
Ferrajoli, A. et al. Epstein-Barr virus microRNAs are expressed in patients with chronic lymphocytic leukemia and correlate with overall survival. EBioMedicine 2, 572–582 (2015).
Zhang, G. et al. Circulating Epstein-Barr virus microRNAs miR-BART7 and miR-BART13 as biomarkers for nasopharyngeal carcinoma diagnosis and treatment. Int. J. Cancer 136, E301–E312 (2015).
Hoshina, S. et al. Profile of exosomal and intracellular microRNA in gamma-herpesvirus-infected lymphoma cell lines. PLoS ONE 11, e0162574 (2016).
Gourzones, C. et al. Extra-cellular release and blood diffusion of BART viral micro-RNAs produced by EBV-infected nasopharyngeal carcinoma cells. Virol. J. 7, 271 (2010).
Chugh, P. E. et al. Systemically circulating viral and tumor-derived microRNAs in KSHV-associated malignancies. PLoS Pathog. 9, e1003484 (2013).
Pegtel, D. M. et al. Functional delivery of viral miRNAs via exosomes. Proc. Natl Acad. Sci. USA 107, 6328–6333 (2010).
Liu, S. et al. The host shapes the gut microbiota via fecal microRNA. Cell Host Microbe 19, 32–43 (2016).
Sivan, A. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350, 1084–1089 (2015).
Vetizou, M. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084 (2015).
Schwarzenbach, H., Hoon, D. S. & Pantel, K. Cell-free nucleic acids as biomarkers in cancer patients. Nat. Rev. Cancer 11, 426–437 (2011).
Jarry, J., Schadendorf, D., Greenwood, C., Spatz, A. & van Kempen, L. C. The validity of circulating microRNAs in oncology: five years of challenges and contradictions. Mol. Oncol. 8, 819–829 (2014).
Becker, N. & Lockwood, C. M. Pre-analytical variables in miRNA analysis. Clin. Biochem. 46, 861–868 (2013).
Fernandez-Mercado, M. et al. The circulating transcriptome as a source of non-invasive cancer biomarkers: concepts and controversies of non-coding and coding RNA in body fluids. J. Cell. Mol. Med. 19, 2307–2323 (2015).
Weber, J. A. et al. The microRNA spectrum in 12 body fluids. Clin. Chem. 56, 1733–1741 (2010).
Khan, J., Lieberman, J. A. & Lockwood, C. M. Variability in, variability out: best practice recommendations to standardize pre-analytical variables in the detection of circulating and tissue microRNAs. Clin. Chem. Lab Med. 55, 608–621 (2017).
Glinge, C. et al. Stability of circulating blood-based microRNAs — pre-analytic methodological considerations. PLoS ONE 12, e0167969 (2017).
Mitchell, A. J. et al. Platelets confound the measurement of extracellular miRNA in archived plasma. Sci. Rep. 6, 32651 (2016).
Brunet-Vega, A. et al. Variability in microRNA recovery from plasma: comparison of five commercial kits. Anal. Biochem. 488, 28–35 (2015).
Hantzsch, M. et al. Comparison of whole blood RNA preservation tubes and novel generation RNA extraction kits for analysis of mRNA and miRNA profiles. PLoS ONE 9, e113298 (2014).
Sultan, M. et al. Influence of RNA extraction methods and library selection schemes on RNA-seq data. BMC Genomics 15, 675 (2014).
El-Khoury, V., Pierson, S., Kaoma, T., Bernardin, F. & Berchem, G. Assessing cellular and circulating miRNA recovery: the impact of the RNA isolation method and the quantity of input material. Sci. Rep. 6, 19529 (2016).
Benes, V. & Castoldi, M. Expression profiling of microRNA using real-time quantitative PCR, how to use it and what is available. Methods 50, 244–249 (2010).
Shi, J. et al. Circulating lncRNAs associated with occurrence of colorectal cancer progression. Am. J. Cancer Res. 5, 2258–2265 (2015).
Chapin, S. C., Appleyard, D. C., Pregibon, D. C. & Doyle, P. S. Rapid microRNA profiling on encoded gel microparticles. Angew. Chem. Int. Ed. 50, 2289–2293 (2011).
Chapin, S. C. & Doyle, P. S. Ultrasensitive multiplexed microRNA quantification on encoded gel microparticles using rolling circle amplification. Anal. Chem. 83, 7179–7185 (2011).
Le, M. T. et al. miR-200-containing extracellular vesicles promote breast cancer cell metastasis. J. Clin. Invest. 124, 5109–5128 (2014).
Moldovan, L. et al. Methodological challenges in utilizing miRNAs as circulating biomarkers. J. Cell. Mol. Med. 18, 371–390 (2014).
Shore, S. et al. Small RNA library preparation method for next-generation sequencing using chemical modifications to prevent adapter dimer formation. PLoS ONE 11, e0167009 (2016).
Kashi, K., Henderson, L., Bonetti, A. & Carninci, P. Discovery and functional analysis of lncRNAs: methodologies to investigate an uncharacterized transcriptome. Biochim. Biophys. Acta 1859, 3–15 (2016).
Clark, M. B. et al. Quantitative gene profiling of long noncoding RNAs with targeted RNA sequencing. Nat. Methods 12, 339–342 (2015).
Cohen, L., Hartman, M. R., Amardey-Wellington, A. & Walt, D. R. Digital direct detection of microRNAs using single molecule arrays. Nucleic Acids Res. 45, e137 (2017).
Bustin, S. et al. Variability of the reverse transcription step: practical implications. Clin. Chem. 61, 202–212 (2015).
Lu, N. et al. CMOS-compatible silicon nanowire field-effect transistors for ultrasensitive and label-free microRNAs sensing. Small 10, 2022–2028 (2014).
Husale, S., Persson, H. H. & Sahin, O. DNA nanomechanics allows direct digital detection of complementary DNA and microRNA targets. Nature 462, 1075–1078 (2009).
Yang, C. et al. In situ DNA-templated synthesis of silver nanoclusters for ultrasensitive and label-free electrochemical detection of microRNA. ACS Appl. Mater. Interfaces 7, 1188–1193 (2015).
Androvic, P., Valihrach, L., Elling, J., Sjoback, R. & Kubista, M. Two-tailed RT-qPCR: a novel method for highly accurate miRNA quantification. Nucleic Acids Res. 45, e144 (2017).
Schwarzenbach, H., da Silva, A. M., Calin, G. & Pantel, K. Data normalization strategies for microRNA quantification. Clin. Chem. 61, 1333–1342 (2015).
Benz, F. et al. U6 is unsuitable for normalization of serum miRNA levels in patients with sepsis or liver fibrosis. Exp. Mol. Med. 45, e42 (2013).
Schmidt, B., Rehbein, G. & Fleischhacker, M. Liquid profiling in lung cancer — quantification of extracellular miRNAs in bronchial lavage. Adv. Exp. Med. Biol. 924, 33–37 (2016).
Muller, V. et al. Changes in serum levels of miR-21, miR-210, and miR-373 in HER2-positive breast cancer patients undergoing neoadjuvant therapy: a translational research project within the Geparquinto trial. Breast Cancer Res. Treat. 147, 61–68 (2014).
McDermott, A. M., Kerin, M. J. & Miller, N. Identification and validation of miRNAs as endogenous controls for RQ-PCR in blood specimens for breast cancer studies. PLoS ONE 8, e83718 (2013).
Ge, W. et al. Expression of serum miR-16, let-7f, and miR-21 in patients with hepatocellular carcinoma and their clinical significances. Clin. Lab 60, 427–434 (2014).
Li, S. et al. Direct targeting of HGF by miR-16 regulates proliferation and migration in gastric cancer. Tumour Biol. 37, 15175–15183 (2016).
Guo, S. et al. Serum miR-16: a potential biomarker for predicting melanoma prognosis. J. Invest. Dermatol. 136, 985–993 (2016).
Filkova, M. et al. Association of circulating miR-223 and miR-16 with disease activity in patients with early rheumatoid arthritis. Ann. Rheum. Dis. 73, 1898–1904 (2014).
Figueredo Dde, S., Gitai, D. L. & Andrade, T. G. Daily variations in the expression of miR-16 and miR-181a in human leukocytes. Blood Cells Mol. Dis. 54, 364–368 (2015).
Kok, M. G. et al. Normalization panels for the reliable quantification of circulating microRNAs by RT-qPCR. FASEB J. 29, 3853–3862 (2015).
Yang, J. et al. Expression analysis of microRNA as prognostic biomarkers in colorectal cancer. Oncotarget 8, 52403–52412 (2017).
Song, J. et al. Identification of suitable reference genes for qPCR analysis of serum microRNA in gastric cancer patients. Dig. Dis. Sci. 57, 897–904 (2012).
Mahn, R. et al. Circulating microRNAs (miRNA) in serum of patients with prostate cancer. Urology 77, 1265.e9–1265.e16 (2011).
Bae, I. S. et al. Identification of reference genes for relative quantification of circulating microRNAs in bovine serum. PLoS ONE 10, e0122554 (2015).
Kirschner, M. B. et al. Haemolysis during sample preparation alters microRNA content of plasma. PLoS ONE 6, e24145 (2011).
Bignotti, E. et al. Identification of stably expressed reference small non-coding RNAs for microRNA quantification in high-grade serous ovarian carcinoma tissues. J. Cell. Mol. Med. 20, 2341–2348 (2016).
Sourvinou, I. S., Markou, A. & Lianidou, E. S. Quantification of circulating miRNAs in plasma: effect of preanalytical and analytical parameters on their isolation and stability. J. Mol. Diagn. 15, 827–834 (2013).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, research0034.1–research0034.11 (2002).
De Spiegelaere, W. et al. Reference gene validation for RT-qPCR, a note on different available software packages. PLoS ONE 10, e0122515 (2015).
Andersen, C. L., Jensen, J. L. & Orntoft, T. F. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 64, 5245–5250 (2004).
Ram, T. P. et al. The diagnostic sensitivity of circulating cardio-enriched microRNAs is increased after normalization of high-density lipoprotein levels. Int. J. Cardiol. 236, 498–500 (2017).
Jin, C. et al. Long non-coding RNA HULC as a novel serum biomarker for diagnosis and prognosis prediction of gastric cancer. Oncotarget 7, 51763–51772 (2016).
Zhang, L. et al. Circulating DNA of HOTAIR in serum is a novel biomarker for breast cancer. Breast Cancer Res. Treat. 152, 199–208 (2015).
Tang, Q. et al. Three circulating long non-coding RNAs act as biomarkers for predicting NSCLC. Cell Physiol. Biochem. 37, 1002–1009 (2015).
Li, N. et al. Identification of circulating long noncoding RNA HOTAIR as a novel biomarker for diagnosis and monitoring of non-small cell lung cancer. Technol. Cancer Res. Treat. 16, 1060–1066 (2017).
Dong, L. et al. Circulating CUDR, LSINCT-5 and PTENP1 long noncoding RNAs in sera distinguish patients with gastric cancer from healthy controls. Int. J. Cancer 137, 1128–1135 (2015).
Zhou, X., Yin, C., Dang, Y., Ye, F. & Zhang, G. Identification of the long non-coding RNA H19 in plasma as a novel biomarker for diagnosis of gastric cancer. Sci. Rep. 5, 11516 (2015).
Baek, R., Sondergaard, E. K., Varming, K. & Jorgensen, M. M. The impact of various preanalytical treatments on the phenotype of small extracellular vesicles in blood analyzed by protein microarray. J. Immunol. Methods 438, 11–20 (2016).
Lacroix, R. et al. Impact of pre-analytical parameters on the measurement of circulating microparticles: towards standardization of protocol. J. Thromb. Haemost. 10, 437–446 (2012).
Gardiner, C. et al. Techniques used for the isolation and characterization of extracellular vesicles: results of a worldwide survey. J. Extracell. Vesicles 5, 32945 (2016).
Lin, J. et al. Exosomes: novel biomarkers for clinical diagnosis. ScientificWorldJournal 2015, 657086 (2015).
Tauro, B. J. et al. Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods 56, 293–304 (2012).
Rekker, K. et al. Comparison of serum exosome isolation methods for microRNA profiling. Clin. Biochem. 47, 135–138 (2014).
Alvarez, M. L., Khosroheidari, M., Kanchi Ravi, R. & DiStefano, J. K. Comparison of protein, microRNA, and mRNA yields using different methods of urinary exosome isolation for the discovery of kidney disease biomarkers. Kidney Int. 82, 1024–1032 (2012).
Melo, S. A. et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 523, 177–182 (2015).
Bednarz-Knoll, N., Alix-Panabieres, C. & Pantel, K. Plasticity of disseminating cancer cells in patients with epithelial malignancies. Cancer Metastasis Rev. 31, 673–687 (2012).
Rupp, A. K. et al. Loss of EpCAM expression in breast cancer derived serum exosomes: role of proteolytic cleavage. Gynecol. Oncol. 122, 437–446 (2011).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer Statistics, 2017. CA Cancer J. Clin. 67, 7–30 (2017).
Han, J. G. et al. A novel panel of serum miR-21/miR-155/miR-365 as a potential diagnostic biomarker for breast cancer. Ann. Surg. Treat. Res. 92, 55–66 (2017).
Usmani, A., Shoro, A. A., Memon, Z., Hussain, M. & Rehman, R. Diagnostic, prognostic and predictive value of MicroRNA-21 in breast cancer patients, their daughters and healthy individuals. Am. J. Cancer Res. 5, 2484–2490 (2015).
Gao, Y. et al. MicroRNA-21 as a potential diagnostic biomarker for breast cancer patients: a pooled analysis of individual studies. Oncotarget 7, 34498–34506 (2016).
Wang, H. et al. Circulating MiR-125b as a marker predicting chemoresistance in breast cancer. PLoS ONE 7, e34210 (2012).
Liu, B. et al. Serum miR-21 and miR-125b as markers predicting neoadjuvant chemotherapy response and prognosis in stage II/III breast cancer. Hum. Pathol. 64, 44–52 (2017).
Li, Q. et al. Circulating miR-19a and miR-205 in serum may predict the sensitivity of luminal A subtype of breast cancer patients to neoadjuvant chemotherapy with epirubicin plus paclitaxel. PLoS ONE 9, e104870 (2014).
Sochor, M. et al. Oncogenic microRNAs: miR-155, miR-19a, miR-181b, and miR-24 enable monitoring of early breast cancer in serum. BMC Cancer 14, 448 (2014).
Shi, G. L., Chen, Y., Sun, Y., Yin, Y. J. & Song, C. X. Significance of serum microRNAs in the auxiliary diagnosis of non-small cell lung cancer. Clin. Lab 63, 133–140 (2017).
Cui, E. H. et al. Serum microRNA 125b as a diagnostic or prognostic biomarker for advanced NSCLC patients receiving cisplatin-based chemotherapy. Acta Pharmacol. Sin. 34, 309–313 (2013).
Yuxia, M., Zhennan, T. & Wei, Z. Circulating miR-125b is a novel biomarker for screening non-small-cell lung cancer and predicts poor prognosis. J. Cancer Res. Clin. Oncol. 138, 2045–2050 (2012).
Wei, J. et al. Identification of plasma microRNA-21 as a biomarker for early detection and chemosensitivity of non-small cell lung cancer. Chin. J. Cancer 30, 407–414 (2011).
Liu, X. G. et al. High expression of serum miR-21 and tumor miR-200c associated with poor prognosis in patients with lung cancer. Med. Oncol. 29, 618–626 (2012).
Gao, W. et al. MiRNA-21: a biomarker predictive for platinum-based adjuvant chemotherapy response in patients with non-small cell lung cancer. Cancer Biol. Ther. 13, 330–340 (2012).
Li, B. et al. MiR-21 overexpression is associated with acquired resistance of EGFR-TKI in non-small cell lung cancer. Lung Cancer 83, 146–153 (2014).
Shen, H. et al. Alteration in Mir-21/PTEN expression modulates gefitinib resistance in non-small cell lung cancer. PLoS ONE 9, e103305 (2014).
Wang, S. et al. Identification of plasma microRNA profiles for primary resistance to EGFR-TKIs in advanced non-small cell lung cancer (NSCLC) patients with EGFR activating mutation. J. Hematol. Oncol. 8, 127 (2015).
Sozzi, G. et al. Clinical utility of a plasma-based miRNA signature classifier within computed tomography lung cancer screening: a correlative MILD trial study. J. Clin. Oncol. 32, 768–773 (2014).
Stuopelyte, K. et al. The utility of urine-circulating miRNAs for detection of prostate cancer. Br. J. Cancer 115, 707–715 (2016).
Foj, L. et al. Exosomal and non-exosomal urinary miRNAs in prostate cancer detection and prognosis. Prostate 77, 573–583 (2017).
Bryant, R. J. et al. Changes in circulating microRNA levels associated with prostate cancer. Br. J. Cancer 106, 768–774 (2012).
Nguyen, H. C. et al. Expression differences of circulating microRNAs in metastatic castration resistant prostate cancer and low-risk, localized prostate cancer. Prostate 73, 346–354 (2013).
Huang, X. et al. Exosomal miR-1290 and miR-375 as prognostic markers in castration-resistant prostate cancer. Eur. Urol. 67, 33–41 (2015).
Al-Qatati, A. et al. Plasma microRNA signature is associated with risk stratification in prostate cancer patients. Int. J. Cancer 141, 1231–1239 (2017).
Bastaminejad, S. et al. Investigation of microRNA-21 expression levels in serum and stool as a potential non-invasive biomarker for diagnosis of colorectal cancer. Iran. Biomed. J. 21, 106–113 (2017).
Peng, Q. et al. The clinical role of microRNA-21 as a promising biomarker in the diagnosis and prognosis of colorectal cancer: a systematic review and meta-analysis. Oncotarget 8, 44893–44909 (2017).
Tsukamoto, M., Iinuma, H., Yagi, T., Matsuda, K. & Hashiguchi, Y. Circulating exosomal microRNA-21 as a biomarker in each tumor stage of colorectal cancer. Oncology 92, 360–370 (2017).
Kanemaru, H. et al. The circulating microRNA-221 level in patients with malignant melanoma as a new tumor marker. J. Dermatol. Sci. 61, 187–193 (2011).
Li, P., He, Q. Y., Luo, C. Q. & Qian, L. Y. Circulating miR-221 expression level and prognosis of cutaneous malignant melanoma. Med. Sci. Monit. 20, 2472–2477 (2014).
Greenberg, E. et al. A comparative analysis of total serum miRNA profiles identifies novel signature that is highly indicative of metastatic melanoma: a pilot study. Biomarkers 18, 502–508 (2013).
Nguyen, T. et al. Downregulation of microRNA-29c is associated with hypermethylation of tumor-related genes and disease outcome in cutaneous melanoma. Epigenetics 6, 388–394 (2011).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Gupta, R. A. et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071–1076 (2010).
Zhang, Y. et al. Circulating long non-coding HOX transcript antisense intergenic ribonucleic acid in plasma as a potential biomarker for diagnosis of breast cancer. Thorac. Cancer 7, 627–632 (2016).
Zhang, K. et al. Circulating lncRNA H19 in plasma as a novel biomarker for breast cancer. Cancer Biomark 17, 187–194 (2016).
Tantai, J., Hu, D., Yang, Y. & Geng, J. Combined identification of long non-coding RNA XIST and HIF1A-AS1 in serum as an effective screening for non-small cell lung cancer. Int. J. Clin. Exp. Pathol. 8, 7887–7895 (2015).
Liang, W. et al. Circulating long noncoding RNA GAS5 is a novel biomarker for the diagnosis of nonsmall cell lung cancer. Medicine 95, e4608 (2016).
Tan, Q. et al. Identification of circulating long non-coding RNA GAS5 as a potential biomarker for non-small cell lung cancer diagnosisnon-small cell lung cancer, long non-coding RNA, plasma, GAS5, biomarker. Int. J. Oncol. 50, 1729–1738 (2017).
Zhu, Q. et al. Long non-coding RNA 00312 regulated by HOXA5 inhibits tumour proliferation and promotes apoptosis in Non-small cell lung cancer. J. Cell. Mol. Med. 21, 2184–2198 (2017).
Hessels, D. et al. DD3(PCA3)-based molecular urine analysis for the diagnosis of prostate cancer. Eur. Urol. 44, 8–16 (2003).
Day, J. R., Jost, M., Reynolds, M. A., Groskopf, J. & Rittenhouse, H. PCA3: from basic molecular science to the clinical lab. Cancer Lett. 301, 1–6 (2011).
Lee, G. L., Dobi, A. & Srivastava, S. Prostate cancer: diagnostic performance of the PCA3 urine test. Nat. Rev. Urol. 8, 123–124 (2011).
Sanda, M. G. et al. Association between combined TMPRSS2:ERG and PCA3 RNA urinary testing and detection of aggressive prostate cancer. JAMA Oncol. 3, 1085–1093 (2017).
Rigau, M. et al. A three-gene panel on urine increases PSA specificity in the detection of prostate cancer. Prostate 71, 1736–1745 (2011).
Wang, F. et al. Development and prospective multicenter evaluation of the long noncoding RNA MALAT-1 as a diagnostic urinary biomarker for prostate cancer. Oncotarget 5, 11091–11102 (2014).
Ren, S. et al. Long non-coding RNA metastasis associated in lung adenocarcinoma transcript 1 derived miniRNA as a novel plasma-based biomarker for diagnosing prostate cancer. Eur. J. Cancer 49, 2949–2959 (2013).
Crea, F. et al. Identification of a long non-coding RNA as a novel biomarker and potential therapeutic target for metastatic prostate cancer. Oncotarget 5, 764–774 (2014).
Ren, S. et al. RNA-seq analysis of prostate cancer in the Chinese population identifies recurrent gene fusions, cancer-associated long noncoding RNAs and aberrant alternative splicings. Cell Res. 22, 806–821 (2012).
Zhang, W. et al. A novel urinary long non-coding RNA transcript improves diagnostic accuracy in patients undergoing prostate biopsy. Prostate 75, 653–661 (2015).
Graham, L. D. et al. Colorectal Neoplasia Differentially Expressed (CRNDE), a novel gene with elevated expression in colorectal adenomas and adenocarcinomas. Genes Cancer 2, 829–840 (2011).
Kogo, R. et al. Long noncoding RNA HOTAIR regulates polycomb-dependent chromatin modification and is associated with poor prognosis in colorectal cancers. Cancer Res. 71, 6320–6326 (2011).
Ozawa, T. et al. CCAT1 and CCAT2 long noncoding RNAs, located within the 8q.24.21 ‘gene desert’, serve as important prognostic biomarkers in colorectal cancer. Ann. Oncol. 28, 1882–1888 (2017).
Zhao, W., Song, M., Zhang, J., Kuerban, M. & Wang, H. Combined identification of long non-coding RNA CCAT1 and HOTAIR in serum as an effective screening for colorectal carcinoma. Int. J. Clin. Exp. Pathol. 8, 14131–14140 (2015).
Wang, C. et al. Long non-coding RNAs LOC285194, RP11-462C24.1 and Nbla12061 in serum provide a new approach for distinguishing patients with colorectal cancer from healthy controls. Oncotarget 7, 70769–70778 (2016).
Wang, R. et al. Identification of long noncoding RNAs as potential novel diagnosis and prognosis biomarkers in colorectal cancer. J. Cancer Res. Clin. Oncol. 142, 2291–2301 (2016).
Khaitan, D. et al. The melanoma-upregulated long noncoding RNA SPRY4-IT1 modulates apoptosis and invasion. Cancer Res. 71, 3852–3862 (2011).
Liu, T. et al. Clinical significance of long noncoding RNA SPRY4-IT1 in melanoma patients. FEBS Open Bio 6, 147–154 (2016).
Cantile, M. et al. HOTAIR role in melanoma progression and its identification in the blood of patients with advanced disease. J. Cell Physiol. 232, 3422–3432 (2017).
Chen, X. et al. Long noncoding RNA PVT1 as a novel diagnostic biomarker and therapeutic target for melanoma. Biomed. Res. Int. 2017, 7038579 (2017).
Alix-Panabieres, C. & Pantel, K. Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov. 6, 479–491 (2016).
Bardelli, A. & Pantel, K. Liquid biopsies, what we do not know (yet). Cancer Cell 31, 172–179 (2017).
Sawyers, C. L. The cancer biomarker problem. Nature 452, 548–552 (2008).
de Bono, J. S. & Ashworth, A. Translating cancer research into targeted therapeutics. Nature 467, 543–549 (2010).
Gorges, T. M. et al. Heterogeneous PSMA expression on circulating tumor cells: a potential basis for stratification and monitoring of PSMA-directed therapies in prostate cancer. Oncotarget 7, 34930–34941 (2016).
Gauwerky, C. E., Huebner, K., Isobe, M., Nowell, P. C. & Croce, C. M. Activation of MYC in a masked t(8;17) translocation results in an aggressive B cell leukemia. Proc. Natl Acad. Sci. USA 86, 8867–8871 (1989).
Bullrich, F. et al. Characterization of the 13q14 tumor suppressor locus in CLL: identification of ALT1, an alternative splice variant of the LEU2 gene. Cancer Res. 61, 6640–6648 (2001).
EL Andaloussi, S., Mager, I., Breakefield, X. O. & Wood, M. J. Extracellular vesicles: biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 12, 347–357 (2013).
Goldie, B. J. et al. Activity-associated miRNA are packaged in Map1b-enriched exosomes released from depolarized neurons. Nucleic Acids Res. 42, 9195–9208 (2014).
Waldenstrom, A. & Ronquist, G. Role of exosomes in myocardial remodeling. Circ. Res. 114, 315–324 (2014).
Ghidoni, R. et al. Cystatin C is released in association with exosomes: a new tool of neuronal communication which is unbalanced in Alzheimer’s disease. Neurobiol. Aging 32, 1435–1442 (2011).
Vella, L. J., Hill, A. F. & Cheng, L. Focus on extracellular vesicles: exosomes and their role in protein trafficking and biomarker potential in Alzheimer’s and Parkinson’s disease. Int. J. Mol. Sci. 17, 173 (2016).
Gutierrez-Vazquez, C., Villarroya-Beltri, C., Mittelbrunn, M. & Sanchez-Madrid, F. Transfer of extracellular vesicles during immune cell-cell interactions. Immunol. Rev. 251, 125–142 (2013).
Ratajczak, J. et al. Embryonic stem cell-derived microvesicles reprogram hematopoietic progenitors: evidence for horizontal transfer of mRNA and protein delivery. Leukemia 20, 847–856 (2006).
Aliotta, J. M. et al. Alteration of marrow cell gene expression, protein production, and engraftment into lung by lung-derived microvesicles: a novel mechanism for phenotype modulation. Stem Cells 25, 2245–2256 (2007).
Morhayim, J., Rudjito, R., van Leeuwen, J. P. & van Driel, M. Paracrine signaling by extracellular vesicles via osteoblasts. Curr. Mol. Biol. Rep. 2, 48–55 (2016).
Gatti, S. et al. Microvesicles derived from human adult mesenchymal stem cells protect against ischaemia-reperfusion-induced acute and chronic kidney injury. Nephrol. Dial. Transplant. 26, 1474–1483 (2011).
Todorova, D., Simoncini, S., Lacroix, R., Sabatier, F. & Dignat-George, F. Extracellular vesicles in angiogenesis. Circ. Res. 120, 1658–1673 (2017).
Leidinger, P., Backes, C., Rheinheimer, S., Keller, A. & Meese, E. Towards clinical applications of blood-borne miRNA signatures: the influence of the anticoagulant EDTA on miRNA abundance. PLoS ONE 10, e0143321 (2015).
Meyer, A. et al. Evaluation of existing methods for human blood mRNA isolation and analysis for large studies. PLoS ONE 11, e0161778 (2016).
Pizzamiglio, S. et al. A methodological procedure for evaluating the impact of hemolysis on circulating microRNAs. Oncol. Lett. 13, 315–320 (2017).
Ohtsuka, M. et al. H19 noncoding RNA, an independent prognostic factor, regulates essential Rb-E2F and CDK8-beta-catenin signaling in colorectal cancer. EBioMedicine 13, 113–124 (2016).
Yang, F. et al. Up-regulated long non-coding RNA H19 contributes to proliferation of gastric cancer cells. FEBS J. 279, 3159–3165 (2012).
Zhou, W. et al. The lncRNA H19 mediates breast cancer cell plasticity during EMT and MET plasticity by differentially sponging miR-200b/c and let-7b. Sci. Signal. 10, eaak9557 (2017).
Zhou, Y., Sheng, B., Xia, Q., Guan, X. & Zhang, Y. Association of long non-coding RNA H19 and microRNA-21 expression with the biological features and prognosis of non-small cell lung cancer. Cancer Gene Ther. 24, 317–324 (2017).
Lv, M. et al. lncRNA H19 regulates epithelial-mesenchymal transition and metastasis of bladder cancer by miR-29b-3p as competing endogenous RNA. Biochim. Biophys. Acta 1864, 1887–1899 (2017).
Hernandez, J. M. et al. miR-675 mediates downregulation of Twist1 and Rb in AFP-secreting hepatocellular carcinoma. Ann. Surg. Oncol. 20 (Suppl. 3), S625–S635 (2013).
Imig, J. et al. miR-CLIP capture of a miRNA targetome uncovers a lincRNA H19-miR-106a interaction. Nat. Chem. Biol. 11, 107–114 (2015).
Gutschner, T. et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 73, 1180–1189 (2013).
Han, Y., Liu, Y., Nie, L., Gui, Y. & Cai, Z. Inducing cell proliferation inhibition, apoptosis, and motility reduction by silencing long noncoding ribonucleic acid metastasis-associated lung adenocarcinoma transcript 1 in urothelial carcinoma of the bladder. Urology 81, 209.e1–209.e7 (2013).
Meseure, D. et al. Prognostic value of a newly identified MALAT1 alternatively spliced transcript in breast cancer. Br. J. Cancer 114, 1395–1404 (2016).
Liu, S. P., Yang, J. X., Cao, D. Y. & Shen, K. Identification of differentially expressed long non-coding RNAs in human ovarian cancer cells with different metastatic potentials. Cancer Biol. Med. 10, 138–141 (2013).
Wang, X. et al. Silencing of long noncoding RNA MALAT1 by miR-101 and miR-217 inhibits proliferation, migration, and invasion of esophageal squamous cell carcinoma cells. J. Biol. Chem. 290, 3925–3935 (2015).
Hirata, H. et al. Long noncoding RNA MALAT1 promotes aggressive renal cell carcinoma through Ezh2 and interacts with miR-205. Cancer Res. 75, 1322–1331 (2015).
Ji, Q. et al. Long non-coding RNA MALAT1 promotes tumour growth and metastasis in colorectal cancer through binding to SFPQ and releasing oncogene PTBP2 from SFPQ/PTBP2 complex. Br. J. Cancer 111, 736–748 (2014).
Okugawa, Y. et al. Metastasis-associated long non-coding RNA drives gastric cancer development and promotes peritoneal metastasis. Carcinogenesis 35, 2731–2739 (2014).
Ren, S. et al. Long noncoding RNA MALAT-1 is a new potential therapeutic target for castration resistant prostate cancer. J. Urol. 190, 2278–2287 (2013).
Kim, K. et al. HOTAIR is a negative prognostic factor and exhibits pro-oncogenic activity in pancreatic cancer. Oncogene 32, 1616–1625 (2013).
Loewen, G., Jayawickramarajah, J., Zhuo, Y. & Shan, B. Functions of lncRNA HOTAIR in lung cancer. J. Hematol. Oncol. 7, 90 (2014).
Li, P. et al. lncRNA HOTAIR contributes to 5FU resistance through suppressing miR-218 and activating NF-kappaB/TS signaling in colorectal cancer. Mol. Ther. Nucleic Acids 8, 356–369 (2017).
Pickard, M. R. & Williams, G. T. Molecular and cellular mechanisms of action of tumour suppressor GAS5 lncRNA. Genes 6, 484–499 (2015).
Zhang, Z. et al. Negative regulation of lncRNA GAS5 by miR-21. Cell Death Differ. 20, 1558–1568 (2013).
Liu, Z. et al. Downregulation of GAS5 promotes bladder cancer cell proliferation, partly by regulating CDK6. PLoS ONE 8, e73991 (2013).
Shi, X. et al. A critical role for the long non-coding RNA GAS5 in proliferation and apoptosis in non-small-cell lung cancer. Mol. Carcinog. 54 (Suppl. 1), E1–E12 (2015).
Yin, D. et al. Long noncoding RNA GAS5 affects cell proliferation and predicts a poor prognosis in patients with colorectal cancer. Med. Oncol. 31, 253 (2014).
Pickard, M. R., Mourtada-Maarabouni, M. & Williams, G. T. Long non-coding RNA GAS5 regulates apoptosis in prostate cancer cell lines. Biochim. Biophys. Acta 1832, 1613–1623 (2013).
Tu, Z. Q., Li, R. J., Mei, J. Z. & Li, X. H. Down-regulation of long non-coding RNA GAS5 is associated with the prognosis of hepatocellular carcinoma. Int. J. Clin. Exp. Pathol. 7, 4303–4309 (2014).
Lu, X. et al. Downregulation of gas5 increases pancreatic cancer cell proliferation by regulating CDK6. Cell Tissue Res. 354, 891–896 (2013).
Cao, S., Liu, W., Li, F., Zhao, W. & Qin, C. Decreased expression of lncRNA GAS5 predicts a poor prognosis in cervical cancer. Int. J. Clin. Exp. Pathol. 7, 6776–6783 (2014).
Chen, D. L. et al. Long noncoding RNA XIST expedites metastasis and modulates epithelial-mesenchymal transition in colorectal cancer. Cell Death Dis. 8, e3011 (2017).
Chang, S., Chen, B., Wang, X., Wu, K. & Sun, Y. Long non-coding RNA XIST regulates PTEN expression by sponging miR-181a and promotes hepatocellular carcinoma progression. BMC Cancer 17, 248 (2017).
Yildirim, E. et al. Xist RNA is a potent suppressor of hematologic cancer in mice. Cell 152, 727–742 (2013).
Tang, Y. et al. lncRNA XIST interacts with miR-140 to modulate lung cancer growth by targeting iASPP. Oncol. Rep. 38, 941–948 (2017).
Ma, L. et al. Long non-coding RNA XIST promotes cell growth and invasion through regulating miR-497/MACC1 axis in gastric cancer. Oncotarget 8, 4125–4135 (2017).
Yao, Y. et al. Knockdown of long non-coding RNA XIST exerts tumor-suppressive functions in human glioblastoma stem cells by up-regulating miR-152. Cancer Lett. 359, 75–86 (2015).
Tong, G. et al. Knockdown of HOXA-AS2 suppresses proliferation and induces apoptosis in colorectal cancer. Am. J. Transl Res. 9, 4545–4552 (2017).
Fang, Y. et al. Long non-coding RNA HOXA-AS2 promotes proliferation and invasion of breast cancer by acting as a miR-520c-3p sponge. Oncotarget 8, 46090–46103 (2017).
Wang, F. et al. HOX antisense lincRNA HOXA-AS2 promotes tumorigenesis of hepatocellular carcinoma. Cell Physiol. Biochem. 40, 287–296 (2016).
Ding, J. et al. Long noncoding RNA HOXA-AS2 represses P21 and KLF2 expression transcription by binding with EZH2, LSD1 in colorectal cancer. Oncogenesis 6, e288 (2017).
Xie, M. et al. Long noncoding RNA HOXA-AS2 promotes gastric cancer proliferation by epigenetically silencing P21/PLK3/DDIT3 expression. Oncotarget 6, 33587–33601 (2015).
Salameh, A. et al. PRUNE2 is a human prostate cancer suppressor regulated by the intronic long noncoding RNA PCA3. Proc. Natl Acad. Sci. USA 112, 8403–8408 (2015).
Acknowledgements
Work in G.A.C.’s laboratory is supported by US National Institutes of Health, National Center for Advancing Translational Sciences (NIH–NCATS) grant UH3TR00943-01 through the NIH Common Fund, Office of Strategic Coordination (OSC); by NIH National Cancer Institute (NCI) grant 1 R01 CA182905-01; by a U54 grant (University of Puerto Rico and The University of Texas MD Anderson Cancer Center (UPR–MDACC) Partnership for Excellence in Cancer Research 2016 Pilot Project); by a Team (US Department of Defense) grant (CA160445P1); by a Ladies Leukemia League grant; by a Chronic Lymphoblastic Leukemia Moonshot Flagship project grant; by a Sister Institution Network Fund (SINF) 2017 grant; and by the Estate of C. G. Johnson Jr. Work in K.P.’s laboratory is supported by the European Community’s Seventh Framework Program (FP7) under grant agreement Innovative Medicines Initiative (IMI) contract no. 115749 CANCER-ID (European Union’s FP7 and EFPIA companies’ in kind contribution); by European Research Council Advanced Investigator grant 269081 DISSECT; by European Research Council Proof-of-Concept grant no. 754453 CTCapture_2.0; by German Research Foundation (DFG) grant PA 341/19-2; and by Deutsche Krebshilfe grant 70112507. We also thank M. Dragomir for his critical review of this manucript.
Reviewer information
Nature Reviews Clinical Oncology thanks O. Slaby, J. Ellinger, L. Castellano, and the other, anonymous reviewer for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
All authors made substantial contributions to each stage of the preparation of the manuscript for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
geNorm: https://genorm.cmgg.be/
NormFinder: https://moma.dk/normfinder-software
ClinicalTrials.gov database: https://clinicaltrials.gov/
CANCER-ID: www.cancer-id.eu
Rights and permissions
About this article
Cite this article
Anfossi, S., Babayan, A., Pantel, K. et al. Clinical utility of circulating non-coding RNAs — an update. Nat Rev Clin Oncol 15, 541–563 (2018). https://doi.org/10.1038/s41571-018-0035-x
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41571-018-0035-x
This article is cited by
-
Non-coding RNAs in disease: from mechanisms to therapeutics
Nature Reviews Genetics (2024)
-
miR-Blood – a small RNA atlas of human blood components
Scientific Data (2024)
-
Circulating tumor nucleic acids: biology, release mechanisms, and clinical relevance
Molecular Cancer (2023)
-
Tumor educated platelet: the novel BioSource for cancer detection
Cancer Cell International (2023)
-
The emerging role of the piRNA/PIWI complex in respiratory tract diseases
Respiratory Research (2023)