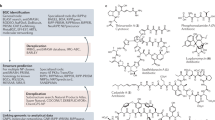
Abstract
Bacterial natural products display astounding structural diversity, which, in turn, endows them with a remarkable range of biological activities that are of significant value to modern society. Such structural features are generated by biosynthetic enzymes that construct core scaffolds or perform peripheral modifications, and can thus define natural product families, introduce pharmacophores and permit metabolic diversification. Modern genomics approaches have greatly enhanced our ability to access and characterize natural product pathways via sequence-similarity-based bioinformatics discovery strategies. However, many biosynthetic enzymes catalyse exceptional, unprecedented transformations that continue to defy functional prediction and remain hidden from us in bacterial (meta)genomic sequence data. In this Review, we highlight exciting examples of unusual enzymology that have been uncovered recently in the context of natural product biosynthesis. These suggest that much of the natural product diversity, including entire substance classes, awaits discovery. New approaches to lift the veil on the cryptic chemistries of the natural product universe are also discussed.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs from 1981 to 2014. J. Nat. Prod. 79, 629–661 (2016).
Demain, A. L. Importance of microbial natural products and the need to revitalize their discovery. J. Ind. Microbiol. Biotechnol. 41, 185–201 (2014).
Cantrell, C. L., Dayan, F. E. & Duke, S. O. Natural products as sources for new pesticides. J. Nat. Prod. 75, 1231–1242 (2012).
Davies, J. & Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 74, 417–433 (2010).
Oerke, E.-C. & Dehne, H.-W. Safeguarding production- losses in major crops and the role of crop protection. Crop Prot. 23, 275–285 (2004).
Katz, L. & Baltz, R. H. Natural product discovery: past, present, and future. J. Ind. Microbiol. Biotechnol. 43, 155–176 (2016).
Bornscheuer, U. T. et al. Engineering the third wave of biocatalysis. Nature 485, 185–194 (2012).
Sheldon, R. A. & Woodley, J. M. Role of biocatalysis in sustainable chemistry. Chem. Rev. 118, 801–838 (2018).
Harvey, A. L., Edrada-Ebel, R. & Quinn, R. J. The re-emergence of natural products for drug discovery in the genomics era. Nat. Rev. Drug Discov. 14, 111–129 (2015).
Medema, M. H. et al. Minimum information about a biosynthetic gene cluster. Nat. Chem. Biol. 11, 625–631 (2015).
Medema, M. H. & Fischbach, M. A. Computational approaches to natural product discovery. Nat. Chem. Biol. 11, 639–648 (2015).
Skinnider, M. A. et al. Genomic charting of ribosomally synthesized natural product chemical space facilitates targeted mining. Proc. Natl. Acad. Sci. USA 113, E6343–E6351 (2016).
Blin, K. et al. antiSMASH 4.0-improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res. 45, W36–W41 (2017).
Skinnider, M. A., Merwin, N. J., Johnston, C. W. & Magarvey, N. A. PRISM 3: expanded prediction of natural product chemical structures from microbial genomes. Nucleic Acids Res. 45, W49–W54 (2017).
Tietz, J. I. et al. A new genome-mining tool redefines the lasso peptide biosynthetic landscape. Nat. Chem. Biol. 13, 470–478 (2017).
Gross, H. et al. The genomisotopic approach: a systematic method to isolate products of orphan biosynthetic gene clusters. Chem. Biol. 14, 53–63 (2007).
Bernhardt, P. & O’Connor, S. E. Opportunities for enzyme engineering in natural product biosynthesis. Curr. Opin. Chem. Biol. 13, 35–42 (2009).
Fischbach, M. A. & Walsh, C. T. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: logic, machinery, and mechanisms. Chem. Rev. 106, 3468–3496 (2006).
Süssmuth, R. D. & Mainz, A. Nonribosomal peptide synthesis-principles and prospects. Angew. Chem. Int. Ed. 56, 3770–3821 (2017).
Donadio, S., Staver, M. J., McAlpine, J. B., Swanson, S. J. & Katz, L. Modular organization of genes required for complex polyketide biosynthesis. Science 252, 675–679 (1991).
Walsh, C. T., O’Brien, R. V. & Khosla, C. Nonproteinogenic amino acid building blocks for nonribosomal peptide and hybrid polyketide scaffolds. Angew. Chem. Int. Ed. 52, 7098–7124 (2013).
Scott, T. A., Heine, D., Qin, Z. & Wilkinson, B. An L-threonine transaldolase is required for L-threo-β-hydroxy-α-amino acid assembly during obafluorin biosynthesis. Nat. Commun. 8, 15935 (2017).
Schaffer, J. E., Reck, M. R., Prasad, N. K. & Wencewicz, T. A. β-Lactone formation during product release from a nonribosomal peptide synthetase. Nat. Chem. Biol. 13, 737–744 (2017).
Herbert, R. B. & Knaggs, A. R. Biosynthesis of the antibiotic obafluorin from r-aminophenylalanine and glycine (glyoxylate). J. Chem. Soc. Perkin Trans. 1 https://doi.org/10.1039/P19920000109 (1992).
Deng, H., Cross, S. M., McGlinchey, R. P., Hamilton, J. T. & O’Hagan, D. In vitro reconstituted biotransformation of 4-fluorothreonine from fluoride ion: application of the fluorinase. Chem. Biol. 15, 1268–1276 (2008).
Barnard-Britson, S. et al. Amalgamation of nucleosides and amino acids in antibiotic biosynthesis: discovery of an L-threonine:uridine-5ʹ-aldehyde transaldolase. J. Am. Chem. Soc. 134, 18514–18517 (2012).
Zelyas, N. J., Cai, H., Kwong, T. & Jensen, S. E. Alanylclavam biosynthetic genes are clustered together with one group of clavulanic acid biosynthetic genes in Streptomyces clavuligerus. J. Bacteriol. 190, 7957–7965 (2008).
Jensen, S. E. Biosynthesis of clavam metabolites. J. Ind. Microbiol. Biotechnol. 39, 1407–1419 (2012).
Fujimori, D. G. et al. Cloning and characterization of the biosynthetic gene cluster for kutznerides. Proc. Natl. Acad. Sci. USA 104, 16498–16503 (2007).
Blair, L. M. & Sperry, J. Natural products containing a nitrogen-nitrogen bond. J. Nat. Prod. 76, 794–812 (2013).
Du, Y. L., He, H. Y., Higgins, M. A. & Ryan, K. S. A heme-dependent enzyme forms the nitrogen-nitrogen bond in piperazate. Nat. Chem. Biol. 13, 836–838 (2017).
Park, S. R. et al. Discovery of cahuitamycins as biofilm inhibitors derived from a convergent biosynthetic pathway. Nat. Commun. 7, 10710 (2016).
Sit, C. S. et al. Variable genetic architectures produce virtually identical molecules in bacterial symbionts of fungus-growing ants. Proc. Natl. Acad. Sci. USA 112, 13150–13154 (2015).
Leipoldt, F. et al. Warhead biosynthesis and the origin of structural diversity in hydroxamate metalloproteinase inhibitors. Nat. Commun. 8, 1965 (2017).
Du, Y. L., Dalisay, D. S., Andersen, R. J. & Ryan, K. S. N-Carbamoylation of 2,4-diaminobutyrate reroutes the outcome in padanamide biosynthesis. Chem. Biol. 20, 1002–1011 (2013).
Qu, X. et al. Cloning, sequencing and characterization of the biosynthetic gene cluster of sanglifehrin A, a potent cyclophilin inhibitor. Mol. Biosyst. 7, 852–861 (2011).
Ma, J. et al. Biosynthesis of himastatin: assembly line and characterization of three cytochrome P450 enzymes involved in the post-tailoring oxidative steps. Angew. Chem. Int. Ed. 50, 7797–7802 (2011).
Matsuda, K. et al. Discovery of unprecedented hydrazine-forming machinery in bacteria. J. Am. Chem. Soc. 140, 9083–9086 (2018).
Verdier-Pinard, P. et al. Structure-activity analysis of the interaction of curacin A, the potent colchicine site antimitotic agent, with tubulin and effects of analogs on the growth of MCF-7 breast cancer cells. Mol. Pharmacol. 53, 62–76 (1998).
Chang, Z. et al. Biosynthetic pathway and gene cluster analysis of curacin A, an antitubulin natural product from the tropical marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 67, 1356–1367 (2004).
Gu, L. et al. Polyketide decarboxylative chain termination preceded by O-sulfonation in curacin a biosynthesis. J. Am. Chem. Soc. 131, 16033–16035 (2009).
Gehret, J. J. et al. Terminal alkene formation by the thioesterase of curacin A biosynthesis: structure of a decarboxylating thioesterase. J. Biol. Chem. 286, 14445–14454 (2011).
Fiers, W. D., Dodge, G. J., Sherman, D. H., Smith, J. L. & Aldrich, C. C. Vinylogous dehydration by a polyketide dehydratase domain in curacin biosynthesis. J. Am. Chem. Soc. 138, 16024–16036 (2016).
Edwards, D. J. et al. Structure and biosynthesis of the jamaicamides, new mixed polyketide-peptide neurotoxins from the marine cyanobacterium Lyngbya majuscula. Chem. Biol. 11, 817–833 (2004).
Gu, L. et al. Metamorphic enzyme assembly in polyketide diversification. Nature 459, 731–735 (2009).
Khare, D. et al. Conformational switch triggered by α-ketoglutarate in a halogenase of curacin A biosynthesis. Proc. Natl. Acad. Sci. USA 107, 14099–14104 (2010).
Khare, D. et al. Structural basis for cyclopropanation by a unique enoyl-acyl carrier protein reductase. Structure 23, 2213–2223 (2015).
Gaudelli, N. M., Long, D. H. & Townsend, C. A. β-Lactam formation by a non-ribosomal peptide synthetase during antibiotic biosynthesis. Nature 520, 383–387 (2015).
Long, D. H. & Townsend, C. A. Mechanism of integrated β-lactam formation by a nonribosomal peptide synthetase during antibiotic synthesis. Biochemistry 57, 3353–3358 (2018).
Gaudelli, N. M. & Townsend, C. A. Epimerization and substrate gating by a TE domain in β-lactam antibiotic biosynthesis. Nat. Chem. Biol. 10, 251–258 (2014).
Keatinge-Clay, A. T. The uncommon enzymology of cis-acyltransferase assembly lines. Chem. Rev. 117, 5334–5366 (2017).
Minami, A. & Oikawa, H. Recent advances of Diels-Alderases involved in natural product biosynthesis. J. Antibiot. 69, 500–506 (2016).
Kim, H. J., Ruszczycky, M. W., Choi, S. H., Liu, Y. N. & Liu, H. W. Enzyme-catalysed [4 + 2] cycloaddition is a key step in the biosynthesis of spinosyn A. Nature 473, 109–112 (2011).
Fage, C. D. et al. The structure of SpnF, a standalone enzyme that catalyzes [4 + 2] cycloaddition. Nat. Chem. Biol. 11, 256–258 (2015).
Hashimoto, T. et al. Biosynthesis of versipelostatin: identification of an enzyme-catalyzed [4 + 2]-cycloaddition required for macrocyclization of spirotetronate-containing polyketides. J. Am. Chem. Soc. 137, 572–575 (2015).
Tian, Z. et al. An enzymatic [4 + 2] cyclization cascade creates the pentacyclic core of pyrroindomycins. Nat. Chem. Biol. 11, 259–265 (2015).
Byrne, M. J. et al. The catalytic mechanism of a natural Diels-Alderase revealed in molecular detail. J. Am. Chem. Soc. 138, 6095–6098 (2016).
Jeon, B. S., Wang, S. A., Ruszczycky, M. W. & Liu, H. W. Natural [4 + 2]-cyclases. Chem. Rev. 117, 5367–5388 (2017).
Schorn, M. et al. Genetic basis for the biosynthesis of the pharmaceutically important class of epoxyketone proteasome inhibitors. ACS Chem. Biol. 9, 301–309 (2014).
Owen, J. G. et al. Multiplexed metagenome mining using short DNA sequence tags facilitates targeted discovery of epoxyketone proteasome inhibitors. Proc. Natl. Acad. Sci. USA 112, 4221–4226 (2015).
Keller, L. et al. Macyranones: structure, biosynthesis, and binding mode of an unprecedented epoxyketone that targets the 20S proteasome. J. Am. Chem. Soc. 137, 8121–8130 (2015).
Zabala, D. et al. A flavin-dependent decarboxylase-dehydrogenase-monooxygenase assembles the warhead of α,β-epoxyketone proteasome inhibitors. J. Am. Chem. Soc. 138, 4342–4345 (2016).
Jomon, K., Kuroda, Y., Ajisaka, M. & Sakai, H. A new antibiotic, ikarugamycin. J. Antibiot. 25, 271–280 (1972).
Hasumi, K., Shinohara, C., Naganuma, S. & Endo, A. Inhibition of the uptake of oxidized low-density lipoprotein in macrophage J774 by the antibiotic ikarugamycin. Eur. J. Biochem. 205, 841–846 (1992).
Luo, T., Fredericksen, B. L., Hasumi, K., Endo, A. & Garcia, J. V. Human immunodeficiency virus type 1 Nef-induced CD4 cell surface downregulation is inhibited by ikarugamycin. J. Virol. 75, 2488–2492 (2001).
Blodgett, J. A. et al. Common biosynthetic origins for polycyclic tetramate macrolactams from phylogenetically diverse bacteria. Proc. Natl. Acad. Sci. USA 107, 11692–11697 (2010).
Popescu, R. et al. Ikarugamycin induces DNA damage, intracellular calcium increase, p38 MAP kinase activation and apoptosis in HL-60 human promyelocytic leukemia cells. Mutat. Res. 709–710, 60–66 (2011).
Luo, Y. et al. Activation and characterization of a cryptic polycyclic tetramate macrolactam biosynthetic gene cluster. Nat. Commun. 4, 2894 (2013).
Ding, Y. et al. Alteramide B is a microtubule antagonist of inhibiting Candida albicans. Biochim. Biophys. Acta 1860, 2097–2106 (2016).
Ding, Y. et al. HSAF-induced antifungal effects in Candida albicans through ROS-mediated apoptosis. RSC Adv. 6, 30895–30904 (2016).
Malcomson, B. et al. Connectivity mapping (ssCMap) to predict A20-inducing drugs and their antiinflammatory action in cystic fibrosis. Proc. Natl. Acad. Sci. USA 113, E3725–E3734 (2016).
Antosch, J., Schaefers, F. & Gulder, T. A. Heterologous reconstitution of ikarugamycin biosynthesis in E. coli. Angew. Chem. Int. Ed. 53, 3011–3014 (2014).
Zhang, G. et al. Mechanistic insights into polycycle formation by reductive cyclization in ikarugamycin biosynthesis. Angew. Chem. Int. Ed. 53, 4840–4844 (2014).
Greunke, C., Glöckle, A., Antosch, J. & Gulder, T. A. Biocatalytic total synthesis of ikarugamycin. Angew. Chem. Int. Ed. 56, 4351–4355 (2017).
Scott, J. D. & Williams, R. M. Chemistry and biology of the tetrahydroisoquinoline antitumor antibiotics. Chem. Rev. 102, 1669–1730 (2002).
Koketsu, K., Watanabe, K., Suda, H., Oguri, H. & Oikawa, H. Reconstruction of the saframycin core scaffold defines dual Pictet-Spengler mechanisms. Nat. Chem. Biol. 6, 408–410 (2010).
Helfrich, E. J. & Piel, J. Biosynthesis of polyketides by trans-AT polyketide synthases. Nat. Prod. Rep. 33, 231–316 (2016).
Butcher, R. A. et al. The identification of bacillaene, the product of the PksX megacomplex in Bacillus subtilis. Proc. Natl. Acad. Sci. USA 104, 1506–1509 (2007).
Ueoka, R., Bortfeld-Miller, M., Morinaka, B. I., Vorholt, J. A. & Piel, J. Toblerols: cyclopropanol-containing polyketide modulators of antibiosis in Methylobacteria. Angew. Chem. Int. Ed. 57, 977–981 (2018).
Heine, D., Bretschneider, T., Sundaram, S. & Hertweck, C. Enzymatic polyketide chain branching to give substituted lactone, lactam, and glutarimide heterocycles. Angew. Chem. Int. Ed. 53, 11645–11649 (2014).
Sundaram, S., Heine, D. & Hertweck, C. Polyketide synthase chimeras reveal key role of ketosynthase domain in chain branching. Nat. Chem. Biol. 11, 949–951 (2015).
Pöplau, P., Frank, S., Morinaka, B. I. & Piel, J. An enzymatic domain for the formation of cyclic ethers in complex polyketides. Angew. Chem. Int. Ed. 52, 13215–13218 (2013).
Luhavaya, H. et al. Enzymology of pyran ring a formation in salinomycin biosynthesis. Angew. Chem. Int. Ed. 127, 13826–13829 (2015).
Sung, K. H. et al. Insights into the dual activity of a bifunctional dehydratase-cyclase domain. Angew. Chem. Int. Ed. 57, 343–347 (2018).
Ma, M., Lohman, J. R., Liu, T. & Shen, B. C-S bond cleavage by a polyketide synthase domain. Proc. Natl. Acad. Sci. USA 112, 10359–10364 (2015).
Wang, H., Fewer, D. P., Holm, L., Rouhiainen, L. & Sivonen, K. Atlas of nonribosomal peptide and polyketide biosynthetic pathways reveals common occurrence of nonmodular enzymes. Proc. Natl. Acad. Sci. USA 111, 9259–9264 (2014).
Iqbal, H. A., Low-Beinart, L., Obiajulu, J. U. & Brady, S. F. Natural product discovery through improved functional metagenomics in Streptomyces. J. Am. Chem. Soc. 138, 9341–9344 (2016).
Crits-Christoph, A., Diamond, S., Butterfield, C. N., Thomas, B. C. & Banfield, J. F. Novel soil bacteria possess diverse genes for secondary metabolite biosynthesis. Nature 558, 440–444 (2018).
Nguyen, T. et al. Exploiting the mosaic structure of trans-acyltransferase polyketide synthases for natural product discovery and pathway dissection. Nat. Biotechnol. 26, 225–233 (2008).
Meoded, R. A. et al. A polyketide synthase component for oxygen insertion into polyketide backbones. Angew. Chem. Int. Ed. 57, 11644–11648 (2018).
Cociancich, S. et al. The gyrase inhibitor albicidin consists of p-aminobenzoic acids and cyanoalanine. Nat. Chem. Biol. 11, 195–197 (2015).
Chun, S. W., Hinze, M. E., Skiba, M. A. & Narayan, A. R. H. Chemistry of a unique polyketide-like synthase. J. Am. Chem. Soc. 140, 2430–2433 (2018).
Cohen, D. R. & Townsend, C. A. A dual role for a polyketide synthase in dynemicin enediyne and anthraquinone biosynthesis. Nat. Chem. 10, 231–236 (2018).
Cohen, D. R. & Townsend, C. A. Characterization of an anthracene intermediate in dynemicin biosynthesis. Angew. Chem. Int. Ed. 57, 5650–5654 (2018).
Pidot, S., Ishida, K., Cyrulies, M. & Hertweck, C. Discovery of clostrubin, an exceptional polyphenolic polyketide antibiotic from a strictly anaerobic bacterium. Angew. Chem. Int. Ed. 53, 7856–7859 (2014).
Behnken, S. & Hertweck, C. Cryptic polyketide synthase genes in non-pathogenic Clostridium spp. PLoS One 7, e29609 (2012).
Vattekkatte, A., Garms, S., Brandt, W. & Boland, W. Enhanced structural diversity in terpenoid biosynthesis: enzymes, substrates and cofactors. Org. Biomol. Chem. 16, 348–362 (2018).
Yamada, Y. et al. Terpene synthases are widely distributed in bacteria. Proc. Natl. Acad. Sci. USA 112, 857–862 (2015).
Baunach, M., Franke, J. & Hertweck, C. Terpenoid biosynthesis off the beaten track: unconventional cyclases and their impact on biomimetic synthesis. Angew. Chem. Int. Ed. 54, 2604–2626 (2015).
Dickschat, J. S. Bacterial terpene cyclases. Nat. Prod. Rep. 33, 87–110 (2016).
Heide, L. Prenyl transfer to aromatic substrates: genetics and enzymology. Curr. Opin. Chem. Biol. 13, 171–179 (2009).
Li, S. et al. Hapalindole/ambiguine biogenesis is mediated by a Cope rearrangement, C-C bond-forming cascade. J. Am. Chem. Soc. 137, 15366–15369 (2015).
Li, S. et al. Decoding cyclase-dependent assembly of hapalindole and fischerindole alkaloids. Nat. Chem. Biol. 13, 467–469 (2017).
Zhu, Q. & Liu, X. Discovery of a calcium-dependent enzymatic cascade for the selective assembly of hapalindole-type alkaloids: on the biosynthetic origin of hapalindole U. Angew. Chem. Int. Ed. 56, 9062–9066 (2017).
Newmister, S. A. et al. Structural basis of the Cope rearrangement and cyclization in hapalindole biogenesis. Nat. Chem. Biol. 14, 345–351 (2018).
Komatsu, M., Tsuda, M., Omura, S., Oikawa, H. & Ikeda, H. Identification and functional analysis of genes controlling biosynthesis of 2-methylisoborneol. Proc. Natl. Acad. Sci. USA 105, 7422–7427 (2008).
Rinkel, J., Lauterbach, L., Rabe, P. & Dickschat, J. S. Two diterpene synthases for spiroalbatene and cembrene A from Allokutzneria albata. Angew. Chem. Int. Ed. 57, 3238–3241 (2018).
Yamada, Y. et al. Novel terpenes generated by heterologous expression of bacterial terpene synthase genes in an engineered Streptomyces host. J. Antibiot. 68, 385–394 (2015).
Domik, D. et al. A terpene synthase is involved in the synthesis of the volatile organic compound sodorifen of Serratia plymuthica 4Rx13. Front. Microbiol. 7, 737 (2016).
von Reuss, S. et al. Sodorifen biosynthesis in the rhizobacterium Serratia plymuthica involves methylation and cyclization of MEP-derived farnesyl pyrophosphate by a SAM-dependent C-methyltransferase. J. Am. Chem. Soc. 140, 11855–11862 (2018).
Weise, T. et al. VOC emission of various Serratia species and isolates and genome analysis of Serratia plymuthica 4Rx13. FEMS Microbiol. Lett. 352, 45–53 (2014).
Ozaki, T. et al. Enzymatic formation of a skipped methyl-substituted octaprenyl side chain of longestin (KS-505a): involvement of homo-IPP as a common extender unit. Angew. Chem. Int. Ed. 57, 6629–6632 (2018).
Awakawa, T. et al. A methyltransferase initiates terpene cyclization in teleocidin B biosynthesis. J. Am. Chem. Soc. 136, 9910–9913 (2014).
Nelson, D. R. Cytochrome P450 diversity in the tree of life. Biochim. Biophys. Acta Proteins Proteom. 1866, 141–154 (2018).
Zhang, X. & Li, S. Expansion of chemical space for natural products by uncommon P450 reactions. Nat. Prod. Rep. 34, 1061–1089 (2017).
Greule, A., Stok, J. E., De Voss, J. J. & Cryle, M. J. Unrivalled diversity: the many roles and reactions of bacterial cytochromes P450 in secondary metabolism. Nat. Prod. Rep. 35, 757–791 (2018).
Duan, L., Jogl, G. & Cane, D. E. The cytochrome P450-catalyzed oxidative rearrangement in the final step of pentalenolactone biosynthesis: substrate structure determines mechanism. J. Am. Chem. Soc. 138, 12678–12689 (2016).
Agrawal, P., Khater, S., Gupta, M., Sain, N. & Mohanty, D. RiPPMiner: a bioinformatics resource for deciphering chemical structures of RiPPs based on prediction of cleavage and cross-links. Nucleic Acids Res. 45, W80–W88 (2017).
van Heel, A. J. et al. BAGEL4: a user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res. 46, W278–W281 (2018).
Arnison, P. G. et al. Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 30, 108–160 (2013).
Banerjee, S. & Hansen, J. N. Structure and expression of a gene encoding the precursor of subtilin, a small protein antibiotic. J. Biol. Chem. 263, 9508–9514 (1988).
Kelly, W. L., Pan, L. & Li, C. Thiostrepton biosynthesis: prototype for a new family of bacteriocins. J. Am. Chem. Soc. 131, 4327–4334 (2009).
Freeman, M. F. et al. Metagenome mining reveals polytheonamides as posttranslationally modified ribosomal peptides. Science 338, 387–390 (2012).
Walsh, C. T. Blurring the lines between ribosomal and nonribosomal peptide scaffolds. ACS Chem. Biol. 9, 1653–1661 (2014).
Müller, M. M. Post-translational modifications of protein backbones: unique functions, mechanisms, and challenges. Biochemistry 57, 177–185 (2018).
Burkhart, B. J., Schwalen, C. J., Mann, G., Naismith, J. H. & Mitchell, D. A. YcaO-dependent posttranslational amide activation: biosynthesis, structure, and function. Chem. Rev. 117, 5389–5456 (2017).
Crone, W. J. et al. Dissecting bottromycin biosynthesis using comparative untargeted metabolomics. Angew. Chem. Int. Ed. 55, 9639–9643 (2016).
Franz, L., Adam, S., Santos-Aberturas, J., Truman, A. W. & Koehnke, J. Macroamidine formation in bottromycins is catalyzed by a divergent YcaO enzyme. J. Am. Chem. Soc. 139, 18158–18161 (2017).
Schwalen, C. J. et al. In vitro biosynthetic studies of bottromycin expand the enzymatic capabilities of the YcaO superfamily. J. Am. Chem. Soc. 139, 18154–18157 (2017).
Metelev, M. et al. Klebsazolicin inhibits 70S ribosome by obstructing the peptide exit tunnel. Nat. Chem. Biol. 13, 1129–1136 (2017).
Travin, D. Y. et al. Biosynthesis of translation inhibitor klebsazolicin proceeds through heterocyclization and N-terminal amidine formation catalyzed by a single YcaO enzyme. J. Am. Chem. Soc. 140, 5625–5633 (2018).
Schwalen, C. J., Hudson, G. A., Kille, B. & Mitchell, D. A. Bioinformatic expansion and discovery of thiopeptide antibiotics. J. Am. Chem. Soc. 140, 9494–9501 (2018).
Frattaruolo, L., Lacret, R., Cappello, A. R. & Truman, A. W. A genomics-based approach identifies a thioviridamide-like compound with selective anticancer activity. ACS Chem. Biol. 12, 2815–2822 (2017).
Izawa, M., Kawasaki, T. & Hayakawa, Y. Cloning and heterologous expression of the thioviridamide biosynthesis gene cluster from Streptomyces olivoviridis. Appl. Environ. Microbiol. 79, 7110–7113 (2013).
Izumikawa, M. et al. Novel thioviridamide derivative - JBIR-140: heterologous expression of the gene cluster for thioviridamide biosynthesis. J. Antibiot. 68, 533–536 (2015).
Kawahara, T. et al. Neothioviridamide, a polythioamide compound produced by heterologous expression of a Streptomyces sp. cryptic RiPP biosynthetic gene cluster. J. Nat. Prod. 81, 264–269 (2018).
Kjaerulff, L. et al. Thioholgamides: thioamide-containing cytotoxic RiPP natural products. ACS Chem. Biol. 12, 2837–2841 (2017).
Lincke, T., Behnken, S., Ishida, K., Roth, M. & Hertweck, C. Closthioamide: an unprecedented polythioamide antibiotic from the strictly anaerobic bacterium Clostridium cellulolyticum. Angew. Chem. Int. Ed. 49, 2011–2013 (2010).
Dunbar, K. L. et al. Genome editing reveals novel thiotemplated assembly of polythioamide antibiotics in anaerobic bacteria. Angew. Chem. Int. Ed. 57, 14080–14084 (2018).
Coyne, S. et al. Biosynthesis of the antimetabolite 6-thioguanine in Erwinia amylovora plays a key role in fire blight pathogenesis. Angew. Chem. Int. Ed. 52, 10564–10568 (2013).
Litomska, A. et al. Enzymatic thioamide formation in a bacterial antimetabolite pathway. Angew. Chem. Int. Ed. 57, 11574–11578 (2018).
Broderick, J. B., Duffus, B. R., Duschene, K. S. & Shepard, E. M. Radical S-adenosylmethionine enzymes. Chem. Rev. 114, 4229–4317 (2014).
Gerlt, J. A. Genomic enzymology: web tools for leveraging protein family sequence-function space and genome context to discover novel functions. Biochemistry 56, 4293–4308 (2017).
Byer, A. S. et al. Paradigm shift for radical S-adenosyl-L-methionine reactions: the organometallic intermediate Ω is central to catalysis. J. Am. Chem. Soc. 140, 8634–8638 (2018).
Mehta, A. P. et al. Radical S-adenosylmethionine (SAM) enzymes in cofactor biosynthesis: a treasure trove of complex organic radical rearrangement reactions. J. Biol. Chem. 290, 3980–3986 (2015).
Mahanta, N., Hudson, G. A. & Mitchell, D. A. Radical S-adenosylmethionine enzymes involved in RiPP biosynthesis. Biochemistry 56, 5229–5244 (2017).
Zhang, Q. et al. Radical-mediated enzymatic carbon chain fragmentation-recombination. Nat. Chem. Biol. 7, 154–160 (2011).
Nicolet, Y., Zeppieri, L., Amara, P. & Fontecilla-Camps, J. C. Crystal structure of tryptophan lyase (NosL): evidence for radical formation at the amino group of tryptophan. Angew. Chem. Int. Ed. 53, 11840–11844 (2014).
Ji, X., Li, Y., Ding, W. & Zhang, Q. Substrate-tuned catalysis of the radical S-adenosyl-L-methionine enzyme NosL involved in nosiheptide biosynthesis. Angew. Chem. Int. Ed. 54, 9021–9024 (2015).
Sicoli, G. et al. Fine-tuning of a radical-based reaction by radical S-adenosyl-L-methionine tryptophan lyase. Science 351, 1320–1323 (2016).
Bhandari, D. M., Fedoseyenko, D. & Begley, T. P. Mechanistic studies on the radical SAM enzyme tryptophan lyase (NosL). Meth. Enzymol. 606, 155–178 (2018).
Badding, E. D. et al. Rerouting the pathway for the biosynthesis of the side ring system of nosiheptide: the roles of NosI, NosJ, and NosK. J. Am. Chem. Soc. 139, 5896–5905 (2017).
Ding, W. et al. Biosynthesis of the nosiheptide indole side ring centers on a cryptic carrier protein NosJ. Nat. Commun. 8, 437 (2017).
Qiu, Y. et al. Thiolation protein-based transfer of indolyl to a ribosomally synthesized polythiazolyl peptide intermediate during the biosynthesis of the side-ring system of nosiheptide. J. Am. Chem. Soc. 139, 18186–18189 (2017).
LaMattina, J. W. et al. NosN, a radical S-adenosylmethionine methylase, catalyzes both C1 transfer and formation of the ester linkage of the side-ring system during the biosynthesis of nosiheptide. J. Am. Chem. Soc. 139, 17438–17445 (2017).
Yokoyama, K. & Lilla, E. A. C-C bond forming radical SAM enzymes involved in the construction of carbon skeletons of cofactors and natural products. Nat. Prod. Rep. 35, 660–694 (2018).
Schramma, K. R., Bushin, L. B. & Seyedsayamdost, M. R. Structure and biosynthesis of a macrocyclic peptide containing an unprecedented lysine-to-tryptophan crosslink. Nat. Chem. 7, 431–437 (2015).
Davis, K. M. et al. Structures of the peptide-modifying radical SAM enzyme SuiB elucidate the basis of substrate recognition. Proc. Natl. Acad. Sci. USA 114, 10420–10425 (2017).
Schramma, K. R. & Seyedsayamdost, M. R. Lysine-tryptophan-crosslinked peptides produced by radical SAM enzymes in pathogenic streptococci. ACS Chem. Biol. 12, 922–927 (2017).
Bushin, L. B., Clark, K. A., Pelczer, I. & Seyedsayamdost, M. R. Charting an unexplored streptococcal biosynthetic landscape reveals a unique peptide cyclization motif. J. Am. Chem. Soc. 140, 17674–17684 (2018).
Caruso, A., Bushin, L. B., Clark, K. A., Martinie, R. J. & Seyedsayamdost, M. R. Radical approach to enzymatic b-thioether bond formation. J. Am. Chem. Soc. 141, 990–997 (2018).
Schramma, K. R., Forneris, C. C., Caruso, A. & Seyedsayamdost, M. R. Mechanistic investigations of lysine-tryptophan cross-link formation catalyzed by streptococcal radical S-adenosylmethionine enzymes. Biochemistry 57, 461–468 (2018).
Bruender, N. A. & Bandarian, V. The radical S-adenosyl-L-methionine enzyme MftC catalyzes an oxidative decarboxylation of the C-terminus of the MftA peptide. Biochemistry 55, 2813–2816 (2016).
Khaliullin, B., Ayikpoe, R., Tuttle, M. & Latham, J. A. Mechanistic elucidation of the mycofactocin-biosynthetic radical S-adenosylmethionine protein, MftC. J. Biol. Chem. 292, 13022–13033 (2017).
Flühe, L. et al. The radical SAM enzyme AlbA catalyzes thioether bond formation in subtilosin A. Nat. Chem. Biol. 8, 350–357 (2012).
Flühe, L. et al. Two [4Fe-4S] clusters containing radical SAM enzyme SkfB catalyze thioether bond formation during the maturation of the sporulation killing factor. J. Am. Chem. Soc. 135, 959–962 (2013).
Wieckowski, B. M. et al. The PqqD homologous domain of the radical SAM enzyme ThnB is required for thioether bond formation during thurincin H maturation. FEBS Lett. 589, 1802–1806 (2015).
Wilson, M. C. et al. An environmental bacterial taxon with a large and distinct metabolic repertoire. Nature 506, 58–62 (2014).
Mori, T. et al. Single-bacterial genomics validates rich and varied specialized metabolism of uncultivated Entotheonella sponge symbionts. Proc. Natl. Acad. Sci. USA 115, 1718–1723 (2018).
Hamada, T., Matsunaga, S., Yano, G. & Fusetani, N. Polytheonamides A and B, highly cytotoxic, linear polypeptides with unprecedented structural features, from the marine sponge. Theonella swinhoei. J. Am. Chem. Soc. 127, 110–118 (2005).
Freeman, M. F., Helf, M. J., Bhushan, A., Morinaka, B. I. & Piel, J. Seven enzymes create extraordinary molecular complexity in an uncultivated bacterium. Nat. Chem. 9, 387–395 (2017).
Morinaka, B. I. et al. Radical S-adenosyl methionine epimerases: regioselective introduction of diverse D-amino acid patterns into peptide natural products. Angew. Chem. Int. Ed. 53, 8503–8507 (2014).
Morinaka, B. I., Verest, M., Freeman, M. F., Gugger, M. & Piel, J. An orthogonal D2O-based induction system that provides insights into D-amino acid pattern formation by radical S-adenosylmethionine peptide epimerases. Angew. Chem. Int. Ed. 56, 762–766 (2017).
Renevey, A. & Riniker, S. The importance of N-methylations for the stability of the b6.3-helical conformation of polytheonamide B. Eur. Biophys. J. 46, 363–374 (2017).
Akiva, E. et al. The structure-function linkage database. Nucleic Acids Res. 42, D521–D530 (2014).
Haft, D. H. & Basu, M. K. Biological systems discovery in silico: radical S-adenosylmethionine protein families and their target peptides for posttranslational modification. J. Bacteriol. 193, 2745–2755 (2011).
Morinaka, B. I. et al. Natural noncanonical protein splicing yields products with diverse β-amino acid residues. Science 359, 779–782 (2018).
Wiebach, V. et al. The anti-staphylococcal lipolanthines are ribosomally synthesized lipopeptides. Nat. Chem. Biol. 14, 652–654 (2018).
Antunes, E. M., Copp, B. R., Davies-Coleman, M. T. & Samaai, T. Pyrroloiminoquinone and related metabolites from marine sponges. Nat. Prod. Rep. 22, 62–72 (2005).
Jordan, P. A. & Moore, B. S. Biosynthetic pathway connects cryptic ribosomally synthesized posttranslationally modified peptide genes with pyrroloquinoline alkaloids. Cell Chem. Biol. 23, 1504–1514 (2016).
Viehrig, K. et al. Structure and biosynthesis of crocagins: polycyclic posttranslationally modified ribosomal peptides from Chondromyces crocatus. Angew. Chem. Int. Ed. 56, 7407–7410 (2017).
Ogasawara, Y. & Dairi, T. Biosynthesis of oligopeptides using ATP-grasp enzymes. Chemistry 23, 10714–10724 (2017).
Ulrich, E. C. & van der Donk, W. A. Cameo appearances of aminoacyl-tRNA in natural product biosynthesis. Curr. Opin. Chem. Biol. 35, 29–36 (2016).
Fortin, P. D., Walsh, C. T. & Magarvey, N. A. A transglutaminase homologue as a condensation catalyst in antibiotic assembly lines. Nature 448, 824–827 (2007).
Wolf, F. et al. Biosynthesis of the β-lactone proteasome inhibitors belactosin and cystargolide. Angew. Chem. Int. Ed. 56, 6665–6668 (2017).
Meng, S. et al. A six-oxidase cascade for tandem C-H bond activation revealed by reconstitution of bicyclomycin biosynthesis. Angew. Chem. Int. Ed. 57, 719–723 (2018).
Patteson, J. B., Cai, W., Johnson, R. A., Santa Maria, K. C. & Li, B. Identification of the biosynthetic pathway for the antibiotic bicyclomycin. Biochemistry 57, 61–65 (2018).
Vior, N. M. et al. Discovery and biosynthesis of the antibiotic bicyclomycin in distantly related bacterial classes. Appl. Environ. Microbiol. 84, e02828–17 (2018).
Du, Y. L., Alkhalaf, L. M. & Ryan, K. S. In vitro reconstitution of indolmycin biosynthesis reveals the molecular basis of oxazolinone assembly. Proc. Natl. Acad. Sci. USA 112, 2717–2722 (2015).
Du, Y. L. et al. A pyridoxal phosphate-dependent enzyme that oxidizes an unactivated carbon-carbon bond. Nat. Chem. Biol. 12, 194–199 (2016).
Thaker, M. N. et al. Identifying producers of antibacterial compounds by screening for antibiotic resistance. Nat. Biotechnol. 31, 922–927 (2013).
Alanjary, M. et al. The Antibiotic Resistant Target Seeker (ARTS), an exploration engine for antibiotic cluster prioritization and novel drug target discovery. Nucleic Acids Res. 45, W42–W48 (2017).
Panter, F., Krug, D., Baumann, S. & Müller, R. Self-resistance guided genome mining uncovers new topoisomerase inhibitors from myxobacteria. Chem. Sci. 9, 4898–4908 (2018).
Jankowitsch, F. et al. A novel N,N-8-amino-8-demethyl-D-riboflavin dimethyltransferase (RosA) catalyzing the two terminal steps of roseoflavin biosynthesis in Streptomyces davawensis. J. Biol. Chem. 286, 38275–38285 (2011).
Jhulki, I., Chanani, P. K., Abdelwahed, S. H. & Begley, T. P. A remarkable oxidative cascade that replaces the riboflavin C8 methyl with an amino group during roseoflavin biosynthesis. J. Am. Chem. Soc. 138, 8324–8327 (2016).
Schwarz, J., Konjik, V., Jankowitsch, F., Sandhoff, R. & Mack, M. Identification of the key enzyme of roseoflavin biosynthesis. Angew. Chem. Int. Ed. 55, 6103–6106 (2016).
Doroghazi, J. R. et al. A roadmap for natural product discovery based on large-scale genomics and metabolomics. Nat. Chem. Biol. 10, 963–968 (2014).
Pye, C. R., Bertin, M. J., Lokey, R. S., Gerwick, W. H. & Linington, R. G. Retrospective analysis of natural products provides insights for future discovery trends. Proc. Natl. Acad. Sci. USA 114, 5601–5606 (2017).
Rutledge, P. J. & Challis, G. L. Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat. Rev. Microbiol. 13, 509–523 (2015).
Wang, B., Guo, F., Dong, S. H. & Zhao, H. Activation of silent biosynthetic gene clusters using transcription factor decoys. Nat. Chem. Biol. 15, 111–114 (2018).
Xu, F. et al. A genetics-free method for high-throughput discovery of cryptic microbial metabolites. Nat. Chem. Biol. 15, 161–168 (2019).
Bouslimani, A., Sanchez, L. M., Garg, N. & Dorrestein, P. C. Mass spectrometry of natural products: current, emerging and future technologies. Nat. Prod. Rep. 31, 718–729 (2014).
Krug, D. & Müller, R. Secondary metabolomics: the impact of mass spectrometry-based approaches on the discovery and characterization of microbial natural products. Nat. Prod. Rep. 31, 768–783 (2014).
Gruene, T. et al. Rapid structure determination of microcrystalline molecular compounds using electron diffraction. Angew. Chem. Int. Ed. 57, 16313–16317 (2018).
Jones, C. G. et al. The cryoEM method microED as a powerful tool for small molecule structure determination. ACS Cent. Sci. 4, 1587–1592 (2018).
Balskus, E. P. Colibactin: understanding an elusive gut bacterial genotoxin. Nat. Prod. Rep. 32, 1534–1540 (2015).
Castro-Falcón, G., Hahn, D., Reimer, D. & Hughes, C. C. Thiol probes to detect electrophilic natural products based on their mechanism of action. ACS Chem. Biol. 11, 2328–2336 (2016).
Maxson, T. et al. Targeting reactive carbonyls for identifying natural products and their biosynthetic origins. J. Am. Chem. Soc. 138, 15157–15166 (2016).
Machado, H., Tuttle, R. N. & Jensen, P. R. Omics-based natural product discovery and the lexicon of genome mining. Curr. Opin. Microbiol. 39, 136–142 (2017).
Berdy, B., Spoering, A. L., Ling, L. L. & Epstein, S. S. In situ cultivation of previously uncultivable microorganisms using the ichip. Nat. Protoc. 12, 2232–2242 (2017).
Wilson, M. C. & Piel, J. Metagenomic approaches for exploiting uncultivated bacteria as a resource for novel biosynthetic enzymology. Chem. Biol. 20, 636–647 (2013).
Charlop-Powers, Z., Milshteyn, A. & Brady, S. F. Metagenomic small molecule discovery methods. Curr. Opin. Microbiol. 19, 70–75 (2014).
Copp, J. N., Akiva, E., Babbitt, P. C. & Tokuriki, N. Revealing unexplored sequence-function space using sequence similarity networks. Biochemistry 57, 4651–4662 (2018).
Zhao, S. et al. Prediction and characterization of enzymatic activities guided by sequence similarity and genome neighborhood networks. eLife 3, e03275 (2014).
Calhoun, S. et al. Prediction of enzymatic pathways by integrative pathway mapping. eLife 7, e31097 (2018).
Cimermancic, P. et al. Insights into secondary metabolism from a global analysis of prokaryotic biosynthetic gene clusters. Cell 158, 412–421 (2014).
Perry, C., de Los Santos, E. L. C., Alkhalaf, L. M. & Challis, G. L. Rieske non-heme iron-dependent oxygenases catalyse diverse reactions in natural product biosynthesis. Nat. Prod. Rep. 35, 622–632 (2018).
Gao, S. S., Naowarojna, N., Cheng, R., Liu, X. & Liu, P. Recent examples of a-ketoglutarate-dependent mononuclear non-haem iron enzymes in natural product biosyntheses. Nat. Prod. Rep. 35, 792–837 (2018).
Du, Y. L. & Ryan, K. S. Pyridoxal phosphate-dependent reactions in the biosynthesis of natural products. Nat. Prod. Rep. 36, 430–457 (2018).
Tolmie, C., Smit, M. S. & Opperman, D. J. Native roles of Baeyer-Villiger monooxygenases in the microbial metabolism of natural compounds. Nat. Prod. Rep. 36, 326–353 (2018).
Cruz-Morales, P. et al. Phylogenomic analysis of natural products biosynthetic gene clusters allows discovery of arseno-organic metabolites in model Streptomycetes. Genome Biol. Evol. 8, 1906–1916 (2016).
Gerlt, J. A. et al. The enzyme function initiative. Biochemistry 50, 9950–9962 (2011).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
Staunton, J. & Weissman, K. J. Polyketide biosynthesis: a millennium review. Nat. Prod. Rep. 18, 380–416 (2001).
Hertweck, C. The biosynthetic logic of polyketide diversity. Angew. Chem. Int. Ed. 48, 4688–4716 (2009).
Yu, F. et al. Structure and biosynthesis of heat-stable antifungal factor (HSAF), a broad-spectrum antimycotic with a novel mode of action. Antimicrob. Agents Chemother. 51, 64–72 (2007).
Lou, L. et al. Biosynthesis of HSAF, a tetramic acid-containing macrolactam from Lysobacter enzymogenes. J. Am. Chem. Soc. 133, 643–645 (2011).
Xu, L., Wu, P., Wright, S. J., Du, L. & Wei, X. Bioactive polycyclic tetramate macrolactams from Lysobacter enzymogenes and their absolute configurations by theoretical ECD calculations. J. Nat. Prod. 78, 1841–1847 (2015).
Wang, X., Shi, J. & Liu, Y. Oxidative rearrangement mechanism of pentalenolactone F catalyzed by cytochrome P450 CYP161C2 (PntM). Inorg. Chem. 57, 8933–8941 (2018).
Acknowledgements
T.A.S. is financially supported by a European Molecular Biology Organization (EMBO) Long-Term Fellowship (ALTF 344-2018). The authors are additionally grateful for funding from the European Research Council (ERC; ERC Advanced Project SynPlex), the Swiss National Science Foundation (SNF; 31003A_146992/1 and NRP 72 ‘Antimicrobial resistance’, 407240_167051), the Helmut Horten Foundation and the Promedica Foundation. The authors would also like to thank S. Leopold-Messer for comments on the figures.
Reviewer information
Nature Reviews Chemistry thanks T. Simpson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
T.A.S. researched the literature and wrote the article, in addition to producing all of the figures. J.P. contributed to discussion, writing and reviewing/editing of the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Orphan
-
Biosynthetic genes that can be detected bioinformatically in a natural product biosynthetic gene cluster context but are functionally unassigned or have been assigned an incorrect or nonspecific function.
- Cryptic
-
Functionally unassigned genes not previously recognized as belonging to natural product biosynthesis.
Rights and permissions
About this article
Cite this article
Scott, T.A., Piel, J. The hidden enzymology of bacterial natural product biosynthesis. Nat Rev Chem 3, 404–425 (2019). https://doi.org/10.1038/s41570-019-0107-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41570-019-0107-1
This article is cited by
-
Genombasierte Wege zur Identifikation bioaktiver bakterieller Naturstoffe
BIOspektrum (2024)
-
Precision enzyme discovery through targeted mining of metagenomic data
Natural Products and Bioprospecting (2024)
-
A pyridoxal 5′-phosphate-dependent Mannich cyclase
Nature Catalysis (2023)
-
Computationally-guided exchange of substrate selectivity motifs in a modular polyketide synthase acyltransferase
Nature Communications (2021)
-
Co-occurrence of enzyme domains guides the discovery of an oxazolone synthetase
Nature Chemical Biology (2021)