Abstract
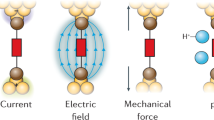
Chemical reactivity underlies our fundamental understanding of many physical and biological phenomena. Chemical reactions are typically initiated by heat, electric current or light. Albeit far less studied, mechanical force is yet another way to orthogonally catalyse chemical reactions. An applied force can substantially reduce the reaction energy barrier, thus enabling reaction pathways that are too slow (or even forbidden) according to the laws of thermodynamics. Single-molecule nanomechanical techniques, including optical and magnetic tweezers and atomic force microscopy, offer the possibility to apply a directional force on an individual chemical bond. In non-covalent (or soft) mechanochemistry, low, sub-nN forces trigger bond rotation or hydrogen-bond rupture. By contrast, in covalent mechanochemistry, higher forces typically result in the breaking and re-forming of individual bonds. This Review focuses on the advances in our mechanistic understanding of single-bond mechanochemistry resulting from single-molecule measurements, as well as on the exciting new perspectives that we envision for this burgeoning field in the near future.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Ribas-Arino, J. & Marx, D. Covalent mechanochemistry: theoretical concepts and computational tools with applications to molecular nanomechanics. Chem. Rev. 112, 5412–5487 (2012).
Turro, N. J., Ramamurthy, V. & Scaiano, J. C. Principles of Molecular Photochemistry (Univ. Sci. Books, 2009).
Bard, A. J. & Faulkner, L. R. Electrochemical Methods: Fundamentals and Applications 2nd edn (Wiley, 2001).
Beyer, M. K. & Clausen-Schaumann, H. Mechanochemistry: the mechanical activation of covalent bonds. Chem. Rev. 105, 2921–2948 (2005).
Hernandez, J. G. & Bolm, C. Altering product selectivity by mechanochemistry. J. Org. Chem. 82, 4007–4019 (2017).
May, P. A. & Moore, J. S. Polymer mechanochemistry: techniques to generate molecular force via elongational flows. Chem. Soc. Rev. 42, 7497–7506 (2013).
Ribas-Arino, J., Shiga, M. & Marx, D. Mechanochemical transduction of externally applied forces to mechanophores. J. Am. Chem. Soc. 132, 10609–10614 (2010).
Ribas-Arino, J., Shiga, M. & Marx D. Understanding covalent mechanochemistry. Angew. Chem. Int. Ed. 48, 4190–4193 (2009).
Ong, M. T., Leiding, J., Tao, H., Virshup, A. M. & Martinez, T. J. First principles dynamics and minimum energy pathways for mechanochemical ring opening of cyclobutene. J. Am. Chem. Soc. 131, 6377–6379 (2009).
Hickenboth, C. R. et al. Biasing reaction pathways with mechanical force. Nature 446, 423–427 (2007).
Lenhardt, J. M., Black, A. L. & Craig, S. L. gem-Dichlorocyclopropanes as abundant and efficient mechanophores in polybutadiene copolymers under mechanical stress. J. Am. Chem. Soc. 131, 10818–10819 (2009).
Lenhardt, J. M. et al. Trapping a diradical transition state by mechanochemical polymer extension. Science 329, 1057–1060 (2010).
Klukovich, H. M. et al. Tension trapping of carbonyl ylides facilitated by a change in polymer backbone. J. Am. Chem. Soc. 134, 9577–9580 (2012).
Huang, Z. & Boulatov, R. Chemomechanics: chemical kinetics for multiscale phenomena. Chem. Soc. Rev. 40, 2359–2384 (2011).
Ladenthin, J. N. et al. Force-induced tautomerization in a single molecule. Nat. Chem. 8, 935–940 (2016).
Schuler, B. et al. Reversible Bergman cyclization by atomic manipulation. Nat. Chem. 8, 220–224 (2016).
Pavlicˇek, N. et al. Synthesis and characterization of triangulene. Nat. Nanotechnol 12, 308–311 (2017).
Pavlicˇek, N. & Gross, L. Generation, manipulation and characterization of molecules by atomic force microscopy. Nat. Rev. Chem. 1, 11 (2017).
Neuman, K. C. & Nagy, A. Single-molecule force spectroscopy: optical tweezers, magnetic tweezers and atomic force microscopy. Nat. Methods 5, 491–505 (2008).
Clausen-Schaumann, H., Seitz, M., Krautbauer, R. & Gaub, H. E. Force spectroscopy with single bio-molecules. Curr. Opin. Chem. Biol. 4, 524–530 (2000).
Stauch, T. & Dreuw, A. Advances in quantum mechanochemistry: electronic structure methods and force analysis. Chem. Rev. 116, 14137–14180 (2016).
Grandbois, M., Beyer, M., Rief, M., Clausen-Schaumann, H. & Gaub, H. E. How strong is a covalent bond? Science 283, 1727–1730 (1999).
Marszalek, P. E. & Dufrene, Y. F. Stretching single polysaccharides and proteins using atomic force microscopy. Chem. Soc. Rev. 41, 3523–3534 (2012).
Rief, M., Oesterhelt, F., Heymann, B. & Gaub, H. E. Single molecule force spectroscopy on polysaccharides by atomic force microscopy. Science 275, 1295–1297 (1997).
Li, H. B. et al. Single-molecule force spectroscopy on polysaccharides by AFM — nanomechanical fingerprint of α-(1,4)-linked polysaccharides. Chem. Phys. Lett. 305, 197–201 (1999).
Marszalek, P. E., Oberhauser, A. F., Pang, Y. P. & Fernandez, J. M. Polysaccharide elasticity governed by chair–boat transitions of the glucopyranose ring. Nature 396, 661–664 (1998).
Marszalek, P. E., Li, H. & Fernandez, J. M. Fingerprinting polysaccharides with single-molecule atomic force microscopy. Nat. Biotechnol. 19, 258–262 (2001).
Marszalek, P. E., Li, H., Oberhauser, A. F. & Fernandez, J. M. Chair–boat transitions in single polysaccharide molecules observed with force-ramp AFM. Proc. Natl Acad. Sci. USA 99, 4278–4283 (2002).
Valiaev, A., Lim, D. W., Oas, T. G., Chilkoti, A. & Zauscher, S. Force-induced prolyl cis–trans isomerization in elastin-like polypeptides. J. Am. Chem. Soc. 129, 6491–6497 (2007).
Rognoni, L., Most, T., Zoldak, G. & Rief, M. Force-dependent isomerization kinetics of a highly conserved proline switch modulates the mechanosensing region of filamin. Proc. Natl Acad. Sci. USA 111, 5568–5573 (2014).
Hugel, T. et al. Single-molecule optomechanical cycle. Science 296, 1103–1106 (2002).
Smith, S. B., Cui, Y. & Bustamante, C. Overstretching B-DNA: the elastic response of individual double-stranded and single-stranded DNA molecules. Science 271, 795–799 (1996).
Bustamante, C., Smith, S. B., Liphardt, J. & Smith, D. Single-molecule studies of DNA mechanics. Curr. Opin. Struct. Biol. 10, 279–285 (2000).
Bianco, P., Bongini, L., Melli, L., Dolfi, M. & Lombardi, V. PicoNewton-millisecond force steps reveal the transition kinetics and mechanism of the double-stranded DNA elongation. Biophys. J. 101, 866–874 (2011).
Rief, M., Clausen-Schaumann, H. & Gaub, H. E. Sequence-dependent mechanics of single DNA molecules. Nat. Struct. Biol. 6, 346–349 (1999).
Woodside, M. T. et al. Direct measurement of the full, sequence-dependent folding landscape of a nucleic acid. Science 314, 1001–1004 (2006).
Woodside, M. T. et al. Nanomechanical measurements of the sequence-dependent folding landscapes of single nucleic acid hairpins. Proc. Natl Acad. Sci. USA 103, 6190–6195 (2006).
Marszalek, P. E. et al. Mechanical unfolding intermediates in titin modules. Nature 402, 100–103 (1999).
Lu, H. & Schulten, K. The key event in force-induced unfolding of Titin's immunoglobulin domains. Biophys. J. 79, 51–65 (2000).
Carrion-Vazquez, M. et al. Mechanical and chemical unfolding of a single protein: a comparison. Proc. Natl Acad. Sci. USA 96, 3694–3699 (1999).
Hughes, M. L. & Dougan, L. The physics of pulling polyproteins: a review of single molecule force spectroscopy using the AFM to study protein unfolding. Rep. Prog. Phys. 79, 076601 (2016).
Valbuena, A. et al. On the remarkable mechanostability of scaffoldins and the mechanical clamp motif. Proc. Natl Acad. Sci. USA 106, 13791–13796 (2009).
Rief, M., Gautel, M., Oesterhelt, F., Fernandex, J. M. & Gaub, H. E. Reversible unfolding of individual titin immunoglobulin domains by AFM. Science 276, 1109–1112 (1997).
Kellermayer, M. S., Smith, S. B., Granzier, H. L. & Bustamante, C. Folding–unfolding transitions in single titin molecules characterized with laser tweezers. Science 276, 1112–1116 (1997).
Carrion-Vazquez, M. et al. The mechanical stability of ubiquitin is linkage dependent. Nat. Struct. Biol. 10, 738–743 (2003).
Brockwell, D. J. et al. Pulling geometry defines the mechanical resistance of a β-sheet protein. Nat. Struct. Biol. 10, 731–737 (2003).
Dietz, H., Berkemeier, F., Bertz, M. & Rief, M. Anisotropic deformation response of single protein molecules. Proc. Natl Acad. Sci. USA 103, 12724–12728 (2006).
Stirnemann, G., Kang, S. G., Zhou, R. H. & Berne, B. J. How force unfolding differs from chemical denaturation. Proc. Natl Acad. Sci. USA 111, 3413–3418 (2014).
Berkovich, R. et al. Rate limit of protein elastic response is tether dependent. Proc. Natl Acad. Sci. USA 109, 14416–14421 (2012).
Schoeler, C. et al. Mapping mechanical force propagation through biomolecular complexes. Nano Lett. 15, 7370–7376 (2015).
Stacklies, W., Vega, M. C., Wilmanns, M. & Grater, F. Mechanical network in titin immunoglobulin from force distribution analysis. PLoS Comput. Biol. 5, e1000306 (2009).
Schlierf, M., Li, H. & Fernandez, J. M. The unfolding kinetics of ubiquitin captured with single-molecule force-clamp techniques. Proc. Natl Acad. Sci. USA 101, 7299–7304 (2004).
Brujic, J., Hermans, R. I., Garcia-Manyes, S., Walther, K. A. & Fernandez, J. M. Dwell-time distribution analysis of polyprotein unfolding using force-clamp spectroscopy. Biophys. J. 92, 2896–2903 (2007).
Garcia-Manyes, S., Brujic, J., Badilla, C. L. & Fernandez, J. M. Force-clamp spectroscopy of single-protein monomers reveals the individual unfolding and folding pathways of I27 and ubiquitin. Biophys. J. 93, 2436–2446 (2007).
Lannon, H., Vanden-Eijnden, E. & Brujic, J. Force-clamp analysis techniques give highest rank to stretched exponential unfolding kinetics in ubiquitin. Biophys. J. 103, 2215–2222 (2012).
Kuo, T. L. et al. Probing static disorder in Arrhenius kinetics by single-molecule force spectroscopy. Proc. Natl Acad. Sci. USA 107, 11336–11340 (2010).
Garcia-Manyes, S., Kuo, T. L. & Fernandez, J. M. Contrasting the individual reactive pathways in protein unfolding and disulfide bond reduction observed within a single protein. J. Am. Chem. Soc. 133, 3104–3113 (2011).
Brujic, J., Hermans, R. I., Walther, K. A. & Fernandez, J. M. Single-molecule force spectroscopy reveals signatures of glassy dynamics in the energy landscape of ubiquitin. Nat. Phys. 2, 282–286 (2006).
Bell, G. I. Models for the specific adhesion of cells to cells. Science 200, 618–627 (1978).
Zheng, P. & Li, H. Highly covalent ferric–thiolate bonds exhibit surprisingly low mechanical stability. J. Am. Chem. Soc. 133, 6791–6798 (2011).
Cecconi, C., Shank, E. A., Bustamante, C. & Marqusee, S. Direct observation of the three-state folding of a single protein molecule. Science 309, 2057–2060 (2005).
Jagannathan, B., Elms, P. J., Bustamante, C. & Marqusee, S. Direct observation of a force-induced switch in the anisotropic mechanical unfolding pathway of a protein. Proc. Natl Acad. Sci. USA 109, 17820–17825 (2012).
Garcia-Manyes, S., Dougan, L., Badilla, C. L., Brujic, J. & Fernandez, J. M. Direct observation of an ensemble of stable collapsed states in the mechanical folding of ubiquitin. Proc. Natl Acad. Sci. USA 106, 10534–10539 (2009).
Garcia-Manyes, S., Dougan, L. & Fernandez, J. M. Osmolyte-induced separation of the mechanical folding phases of ubiquitin. Proc. Natl Acad. Sci. USA 106, 10540–10545 (2009).
Dudko, O. K., Hummer, G. & Szabo, A. Intrinsic rates and activation free energies from single-molecule pulling experiments. Phys. Rev. Lett. 96, 108101 (2006).
Dudko, O. K., Hummer, G. & Szabo, A. Theory, analysis, and interpretation of single-molecule force spectroscopy experiments. Proc. Natl Acad. Sci. USA 105, 15755–15760 (2008).
Popa, I., Fernandez, J. M. & Garcia-Manyes, S. Direct quantification of the attempt frequency determining the mechanical unfolding of ubiquitin protein. J. Biol. Chem. 286, 31072–31079 (2011).
Chung, J., Kushner, A. M., Weisman, A. C. & Guan, Z. Direct correlation of single-molecule properties with bulk mechanical performance for the biomimetic design of polymers. Nat. Mater. 13, 1055–1062 (2014).
Caruso, M. M. et al. Mechanically-induced chemical changes in polymeric materials. Chem. Rev. 109, 5755–5798 (2009).
Lee, B., Niu, Z., Wang, J., Slebodnick, C. & Craig, S. L. Relative mechanical strengths of weak bonds in sonochemical polymer mechanochemistry. J. Am. Chem. Soc. 137, 10826–10832 (2015).
Li, J., Nagamani, C. & Moore, J. S. Polymer mechanochemistry: from destructive to productive. Acc. Chem. Res. 48, 2181–2190 (2015).
Wu, D., Lenhardt, J. M., Black, A. L., Akhremitchev, B. B. & Craig, S. L. Molecular stress relief through a force-induced irreversible extension in polymer contour length. J. Am. Chem. Soc. 132, 15936–15938 (2010).
Wang, J. P. et al. Inducing and quantifying forbidden reactivity with single-molecule polymer mechanochemistry. Nat. Chem. 7, 323–327 (2015).
Wang, J. P., Kouznetsova, T. B. & Craig, S. L. Reactivity and mechanism of a mechanically activated anti-Woodward–Hoffmann–DePuy reaction. J. Am. Chem. Soc. 137, 11554–11557 (2015).
Wang, J. et al. Catch and release: orbital symmetry guided reaction dynamics from a freed “tension trapped transition state”. J. Org. Chem. 80, 11773–11778 (2015).
Wang, J., Kouznetsova, T. B. & Craig, S. L. Single-molecule observation of a mechanically activated cis-to-trans cyclopropane isomerization. J. Am. Chem. Soc. 138, 10410–10412 (2016).
Gossweiler, G. R., Kouznetsova, T. B. & Craig, S. L. Force-rate characterization of two spiropyran-based molecular force probes. J. Am. Chem. Soc. 137, 6148–6151 (2015).
Davis, D. A. et al. Force-induced activation of covalent bonds in mechanoresponsive polymeric materials. Nature 459, 68–72 (2009).
Huang, W. et al. Single molecule study of force-induced rotation of carbon–carbon double bonds in polymers. ACS Nano 11, 194–203 (2017).
Hanson, D. E. & Martin, R. L. How far can a rubber molecule stretch before breaking? Ab initio study of tensile elasticity and failure in single-molecule polyisoprene and polybutadiene. J. Chem. Phys. 130, 064903 (2009).
Li, H. & Walker, G. C. Twist and shout: single-molecule mechanochemistry. ACS Nano 11, 28–30 (2017).
Chen, Z. et al. Mechanochemical unzipping of insulating polyladderene to semiconducting polyacetylene. Science 357, 475–479 (2017).
Ainavarapu, S. R., Wiita, A. P., Huang, H. H. & Fernandez, J. M. A single-molecule assay to directly identify solvent-accessible disulfide bonds and probe their effect on protein folding. J. Am. Chem. Soc. 130, 436–437 (2008).
Lantz, M. A. et al. Quantitative measurement of short-range chemical bonding forces. Science 291, 2580–2583 (2001).
Glaser, T., Hedman, B., Hodgson, K. O. & Solomon, E. I. Ligand K-edge X-ray absorption spectroscopy: a direct probe of ligand–metal covalency. Acc. Chem. Res. 33, 859–868 (2000).
Zheng, P. & Li, H. Direct measurements of the mechanical stability of zinc-thiolate bonds in rubredoxin by single-molecule atomic force microscopy. Biophys. J. 101, 1467–1473 (2011).
Solomon, E. I., Gorelsky, S. I. & Dey, A. Metal–thiolate bonds in bioinorganic chemistry. J. Comput. Chem. 27, 1415–1428 (2006).
Perales-Calvo, J., Lezamiz, A. & Garcia-Manyes, S. The mechanochemistry of a structural zinc finger. J. Phys. Chem. Lett. 6, 3335–3340 (2015).
Beedle, A. E. M., Lezamiz, A., Stirnemann, G. & Garcia-Manyes, S. The mechanochemistry of copper reports on the directionality of unfolding in model cupredoxin proteins. Nat. Commun. 6, 7894 (2015).
Solomon, E. I., Hare, J. W. & Gray, H. B. Spectroscopic studies and a structural model for blue copper centers in proteins. Proc. Natl Acad. Sci. USA 73, 1389–1393 (1976).
Liu, J. et al. Metalloproteins containing cytochrome, iron–sulfur, or copper redox centers. Chem. Rev. 114, 4366–4469 (2014).
Wei, W. et al. Structural insights and the surprisingly low mechanical stability of the Au–S bond in the gold-specific protein GolB. J. Am. Chem. Soc. 137, 15358–15361 (2015).
Xue, Y. R., Li, X., Li, H. B. & Zhang, W. K. Quantifying thiol–gold interactions towards the efficient strength control. Nat. Commun. 5, 4348 (2014).
Zheng, P., Takayama, S. J., Mauk, A. G. & Li, H. Hydrogen bond strength modulates the mechanical strength of ferric–thiolate bonds in rubredoxin. J. Am. Chem. Soc. 134, 4124–4131 (2012).
Zheng, P., Chou, C. C., Guo, Y., Wang, Y. & Li, H. Single molecule force spectroscopy reveals the molecular mechanical anisotropy of the FeS4 metal center in rubredoxin. J. Am. Chem. Soc. 135, 17783–17792 (2013).
Lei, H. et al. Reversible unfolding and folding of the metalloprotein ferredoxin revealed by single-molecule atomic force microscopy. J. Am. Chem. Soc. 139, 1538–1544 (2017).
Zheng, P., Wang, Y. & Li, H. Reversible unfolding–refolding of rubredoxin: a single-molecule force spectroscopy study. Angew. Chem. Int. Ed. 53, 14060–14063 (2014).
Arantes, G. M., Bhattacharjee, A. & Field, M. J. Homolytic cleavage of Fe–S bonds in rubredoxin under mechanical stress. Angew. Chem. Int. Ed. 52, 8144–8146 (2013).
Zheng, P., Arantes, G. M., Field, M. J. & Li, H. Force-induced chemical reactions on the metal centre in a single metalloprotein molecule. Nat. Commun. 6, 7569 (2015).
Aktah, D. & Frank, I. Breaking bonds by mechanical stress: when do electrons decide for the other side? J. Am. Chem. Soc. 124, 3402–3406 (2002).
Wiita, A. P., Ainavarapu, S. R., Huang, H. H. & Fernandez, J. M. Force-dependent chemical kinetics of disulfide bond reduction observed with single-molecule techniques. Proc. Natl Acad. Sci. USA 103, 7222–7227 (2006).
Anfinsen, C. B. & Haber, E. Studies on the reduction and re-formation of protein disulfide bonds. J. Biol. Chem. 236, 1361–1363 (1961).
Ainavarapu, S. R. et al. Contour length and refolding rate of a small protein controlled by engineered disulfide bonds. Biophys. J. 92, 225–233 (2007).
Carl, P., Kwok, C. H., Manderson, G., Speicher, D. W. & Discher, D. E. Forced unfolding modulated by disulfide bonds in the Ig domains of a cell adhesion molecule. Proc. Natl Acad. Sci. USA 98, 1565–1570 (2001).
Hogg, P. J. Disulfide bonds as switches for protein function. Trends Biochem. Sci. 28, 210–214 (2003).
Bach, R. D., Dmitrenko, O. & Thorpe, C. Mechanism of thiolate–disulfide interchange reactions in biochemistry. J. Org. Chem. 73, 12–21 (2008).
Whitesides, G. M., Houk, J. & Patterson, M. A. K. Activation parameters for thiolate disulfide interchange reactions in aqueous-solution. J. Org. Chem. 48, 112–115 (1983).
Nagy, P. Kinetics and mechanisms of thiol-disulfide exchange covering direct substitution and thiol oxidation-mediated pathways. Antioxid. Redox Signal. 18, 1623–1641 (2013).
Liang, J. & Fernandez, J. M. Mechanochemistry: one bond at a time. ACS Nano 3, 1628–1645 (2009).
Kucharski, T. J. et al. Kinetics of thiol/disulfide exchange correlate weakly with the restoring force in the disulfide moiety. Angew. Chem. Int. Ed. 48, 7040–7043 (2009).
Fernandes, P. A. & Ramos, M. J. Theoretical insights into the mechanism for thiol/disulfide exchange. Chemistry 10, 257–266 (2004).
Koti Ainavarapu, S. R., Wiita, A. P., Dougan, L., Uggerud, E. & Fernandez, J. M. Single-molecule force spectroscopy measurements of bond elongation during a bimolecular reaction. J. Am. Chem. Soc. 130, 6479–6487 (2008).
Li, W. & Grater, F. Atomistic evidence of how force dynamically regulates thiol/disulfide exchange. J. Am. Chem. Soc. 132, 16790–16795 (2010).
Liang, J. & Fernandez, J. M. Kinetic measurements on single-molecule disulfide bond cleavage. J. Am. Chem. Soc. 133, 3528–3534 (2011).
Eyring, H. The activated complex in chemical reactions. J. Chem. Phys. 3, 107–115 (1935).
Wiita, A. P. et al. Probing the chemistry of thioredoxin catalysis with force. Nature 450, 124–127 (2007).
Perez-Jimenez, R. et al. Diversity of chemical mechanisms in thioredoxin catalysis revealed by single-molecule force spectroscopy. Nat. Struct. Mol. Biol. 16, 890–896 (2009).
Garcia-Manyes, S., Liang, J., Szoszkiewicz, R., Kuo, T. L. & Fernandez, J. M. Force-activated reactivity switch in a bimolecular chemical reaction. Nat. Chem. 1, 236–242 (2009).
Dopieralski, P. et al. The Janus-faced role of external forces in mechanochemical disulfide bond cleavage. Nat. Chem. 5, 685–691 (2013).
Beyer, M. K. The mechanical strength of a covalent bond calculated by density functional theory. J. Chem. Phys. 112, 7307–7312 (2000).
Iozzi, M. F., Helgaker, T. & Uggerud, E. Influence of external force on properties and reactivity of disulfide bonds. J. Phys. Chem. A 115, 2308–2315 (2011).
Baldus, I. B. & Grater, F. Mechanical force can fine-tune redox potentials of disulfide bonds. Biophys. J. 102, 622–629 (2012).
Hofbauer, F. & Frank, I. Disulfide bond cleavage: a redox reaction without electron transfer. Chemistry 16, 5097–5101 (2010).
Iannuzzi, M., Laio, A. & Parrinello, M. Efficient exploration of reactive potential energy surfaces using Car–Parrinello molecular dynamics. Phys. Rev. Lett. 90, 238302 (2003).
Dopieralski, P., Ribas-Arino, J., Anjukandi, P., Krupicka, M. & Marx, D. Unexpected mechanochemical complexity in the mechanistic scenarios of disulfide bond reduction in alkaline solution. Nat. Chem. 9, 164–170 (2017).
Dopieralski, P., Ribas-Arino, J., Anjukandi, P., Krupicka, M. & Marx, D. Force-induced reversal of β-eliminations: stressed disulfide bonds in alkaline solution. Angew. Chem. Int. Ed. 55, 1304–1308 (2016).
Alegre-Cebollada, J., Kosuri, P., Rivas-Pardo, J. A. & Fernandez, J. M. Direct observation of disulfide isomerization in a single protein. Nat. Chem. 3, 882–887 (2011).
Kosuri, P. et al. Protein folding drives disulfide formation. Cell 151, 794–806 (2012).
Sevier, C. S. & Kaiser, C. A. Formation and transfer of disulphide bonds in living cells. Nat. Rev. Mol. Cell Biol. 3, 836–847 (2002).
Wilkinson, B. & Gilbert, H. F. Protein disulfide isomerase. Biochim. Biophys. Acta 1699, 35–44 (2004).
Kadokura, H., Tian, H., Zander, T., Bardwell, J. C. & Beckwith, J. Snapshots of DsbA in action: detection of proteins in the process of oxidative folding. Science 303, 534–537 (2004).
Kahn, T. B., Fernandez, J. M., Perez-Jimenez, R. Monitoring oxidative folding of a single protein catalyzed by the disulfide oxidoreductase DsbA. J. Biol. Chem. 290, 14518–14527 (2015).
Gupta, V. & Carroll, K. S. Sulfenic acid chemistry, detection and cellular lifetime. Biochim. Biophys. Acta 1840, 847–875 (2014).
Beedle, A. E., Lynham, S. & Garcia-Manyes, S. Protein S-sulfenylation is a fleeting molecular switch that regulates non-enzymatic oxidative folding. Nat. Commun. 7, 12490 (2016).
Beedle, A. E. M., Mora, M., Lynham, S., Stirnemann, G. & Garcia-Manyes, S. Tailoring protein nanomechanics with chemical reactivity. Nat. Commun. 8, 15658 (2017).
Alegre-Cebollada, J. et al. S-Glutathionylation of cryptic cysteines enhances titin elasticity by blocking protein folding. Cell 156, 1235–1246 (2014).
Vogel, V. Mechanotransduction involving multimodular proteins: converting force into biochemical signals. Annu. Rev. Biophys. Biomol. Struct. 35, 459–488 (2006).
Eisenstein, M. Mechanobiology: a measure of molecular muscle. Nature 544, 255–257 (2017).
Puchner, E. M. & Gaub, H. E. Single-molecule mechanoenzymatics. Annu. Rev. Biophys. 41, 497–518 (2012).
Valle-Orero, J. et al. Mechanical deformation accelerates protein ageing. Angew. Chem. Int. Ed. 56, 9741–9746 (2017).
Mayans, O. et al. Structural basis for activation of the titin kinase domain during myofibrillogenesis. Nature 395, 863–869 (1998).
Grater, F., Shen, J., Jiang, H., Gautel, M. & Grubmuller, H. Mechanically induced titin kinase activation studied by force-probe molecular dynamics simulations. Biophys. J. 88, 790–804 (2005).
Puchner, E. M. et al. Mechanoenzymatics of titin kinase. Proc. Natl Acad. Sci. USA 105, 13385–13390 (2008).
Zhang, X., Halvorsen, K., Zhang, C. Z., Wong, W. P. & Springer, T. A. Mechanoenzymatic cleavage of the ultralarge vascular protein von Willebrand factor. Science 324, 1330–1334 (2009).
del Rio, A. et al. Stretching single talin rod molecules activates vinculin binding. Science 323, 638–641 (2009).
Yao, M. et al. Mechanical activation of vinculin binding to talin locks talin in an unfolded conformation. Sci. Rep. 4, 4610 (2014).
Yao, M. et al. Force-dependent conformational switch of α-catenin controls vinculin binding. Nat. Commun. 5, 4525 (2014).
Sawada, Y. et al. Force sensing by mechanical extension of the Src family kinase substrate p130Cas. Cell 127, 1015–1026 (2006).
Guilluy, C. et al. Isolated nuclei adapt to force and reveal a mechanotransduction pathway in the nucleus. Nat. Cell Biol. 16, 376–381 (2014).
Swift, J. et al. Nuclear lamin-A scales with tissue stiffness and enhances matrix-directed differentiation. Science 341, 1240104 (2013).
Popa, I. et al. Nanomechanics of HaloTag tethers. J. Am. Chem. Soc. 135, 12762–12771 (2013).
Popa, I. et al. HaloTag anchored ruler for week-long studies of protein dynamics. J. Am. Chem. Soc. 138, 10546–10553 (2016).
Pernigo, S. et al. Structural insight into M-band assembly and mechanics from the titin-obscurin-like-1 complex. Proc. Natl Acad. Sci. USA 107, 2908–2913 (2010).
Kim, J., Zhang, C. Z., Zhang, X. & Springer, T. A. A mechanically stabilized receptor-ligand flex-bond important in the vasculature. Nature 466, 992–995 (2010).
Pernigo, S. et al. Binding of myomesin to obscurin-like-1 at the muscle M-band provides a strategy for isoform-specific mechanical protection. Structure 25, 107–120 (2017).
Echelman, D. J., Lee, A. Q. & Fernandez, J. M. Mechanical forces regulate the reactivity of a thioester bond in a bacterial adhesin. J. Biol. Chem. 292, 8988–8997 (2017).
Allen, M. P. & Tildesley, D. J. (eds) Computer Simulation in Chemical Physics (Springer, 1993).
Frenkel, D. & Smit, B. Understanding Molecular Simulation: from Algorithms to Applications 2nd edn (Academic Press, 2002).
Austen, K. et al. Extracellular rigidity sensing by talin isoform-specific mechanical linkages. Nat. Cell Biol. 17, 1597–1606 (2015).
Chowdhury, F. et al. Defining single molecular forces required for notch activation using nano yoyo. Nano Lett. 16, 3892–3897 (2016).
Xu, B. Q., Xiao, X. Y. & Tao, N. J. Measurements of single-molecule electromechanical properties. J. Am. Chem. Soc. 125, 16164–16165 (2003).
Xu, B. Q., Li, X. L., Xiao, X. Y., Sakaguchi, H. & Tao, N. J. Electromechanical and conductance switching properties of single oligothiophene molecules. Nano Lett. 5, 1491–1495 (2005).
Frei, M., Aradhya, S. V., Koentopp, M., Hybertsen, M. S. & Venkataraman, L. Mechanics and chemistry: single molecule bond rupture forces correlate with molecular backbone structure. Nano Lett. 11, 1518–1523 (2011).
Marszalek, P. E., Greenleaf, W. J., Li, H., Oberhauser, A. F. & Fernandez, J. M. Atomic force microscopy captures quantized plastic deformation in gold nanowires. Proc. Natl Acad. Sci. USA 97, 6282–6286 (2000).
Sotomayor, M. & Schulten, K. Single-molecule experiments in vitro and in silico. Science 316, 1144–1148 (2007).
Acknowledgements
The authors apologize to the many colleagues whose work could not be cited owing to space constraints. A.E.M.B. thanks the Engineering and Physical Sciences Research Council (EPSRC) for funding through an EPSRC DTP fellowship. This work was supported by a Marie Curie Career Integration Grant (No. 293462), a Biotechnology and Biological Sciences Research Council grant (No. J00992X/1), a Royal Society Research grant (No. RG120038), a British Heart Foundation grant (No. PG/13/50/30426), an EPSRC Fellowship (No. K00641X/1) and a Leverhulme Trust Research Leadership Award, all to S.G.-M.
Author information
Authors and Affiliations
Contributions
Both authors researched data for the article and contributed to discussion of content. A.E.M.B. wrote part of the covalent mechanochemistry section, and S.G.-M. wrote the rest of the article. Both authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Glossary
- Amylose
-
A homopolymer of α-D-glucopyranose rings in which C1 is bonded to C4 on the consecutive ring through a glycosidic bond.
- Dextran
-
A homopolymer formed by glycosidic bonds linking C1 and C6 of consecutive α-D-glucopyranose rings.
- Freely jointed chain (FJC) model
-
A model that describes the behaviour of a semi-flexible polymer composed of rigid and inextensible segments, which are free to rotate at any angle with no correlation between the directions of the neighbouring segments. Under purely thermal control, the polymer will reside in a maximum entropy random configuration. As mechanical force is applied, the polymer stretches against an entropic force resulting from the reduced number of available conformations under force.
- Worm-like chain (WLC) model
-
An extension of the freely jointed chain model that takes into account the energetic costs for bending the polymer chain. Single-molecule nanomechanical techniques are best suited for characterizing the force-dependent stretching behaviour of various polymers.
- Woodward–Hoffmann–DePuy (WHD) rule
-
A rule describing the conservation of orbital symmetry for reactions that have a transition state with cyclic geometry.
- Ligand K-edge X-ray absorption spectroscopy
-
An experimental technique that can be used to investigate the degree of covalency of a metal–ligand bond. The K-edge of the absorption spectra is generated by the excitation of the ligand 1s electron to an empty p orbital, providing information on the atomic arrangement at the active site of a metalloprotein.
Rights and permissions
About this article
Cite this article
Garcia-Manyes, S., Beedle, A. Steering chemical reactions with force. Nat Rev Chem 1, 0083 (2017). https://doi.org/10.1038/s41570-017-0083
Published:
DOI: https://doi.org/10.1038/s41570-017-0083
This article is cited by
-
Mechanocatalysis of CO to CO2 on TiO2 surface controlled at atomic scale
Nano Research (2024)
-
How polymers dance to the pulses of ultrasound
Nature Chemistry (2023)
-
Mechanochemical Synthesis of PZT Powders and the Effects of Mechanical Activation on Solid-State Sintering Kinetics
Transactions of the Indian Institute of Metals (2023)
-
Reactions in single-molecule junctions
Nature Reviews Materials (2022)
-
The role of single-protein elasticity in mechanobiology
Nature Reviews Materials (2022)