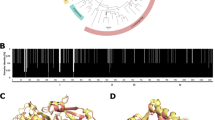
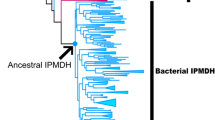
Abstract
Earth has several environments that are potentially hostile to life. The survival of organisms has required the expression of proteins that are adapted to function under extreme temperature, pH, pressure or ionic strength. However, the origin of such adaptations remains, in most cases, an open question. This Review presents a detailed analysis of the specialized enzymes that are able to maintain high catalytic rates at low temperatures and highlights the important role that computational studies have in uncovering the evolutionary principles behind the cold adaptation of enzymes. Although often highly homologous to their mesophilic counterparts, these cold-adapted enzymes have characteristic and universal properties that reflect their evolutionary optimization. In addition to exhibiting maximum reaction rates at lower temperatures, cold-adapted enzymes are more heat-labile and their catalytic mechanisms have distinct signatures in terms of the thermodynamic activation parameters. The structural origins of these properties have been elusive but are hypothesized to be related to protein flexibility.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
van den Burg, B. Extremophiles as a source for novel enzymes. Curr. Opin. Microbiol. 6, 213–218 (2003).
Gomes, J. & Steiner, W. The biocatalytic potential of extremophiles and extremozymes. Food Technol. Biotechnol. 42, 223–235 (2004).
Hough, D. W. & Danson, M. J. Extremozymes. Curr. Opin. Chem. Biol. 3, 39–46 (1999).
Danson, M. J. & Hough, D. W. The structural basis of protein halophilicity. Comp. Biochem. Physiol. A 117, 307–312 (1997).
Madern, D., Ebel, C. & Zaccai, G. Halophilic adaptation of enzymes. Extremophiles 4, 91–98 (2000).
Gros, M. & Jaenicke, R. Proteins under pressure. Eur. J. Biochem. 221, 617–630 (1994).
Daniel, I., Oger, P. & Winter, R. Origins of life and biochemistry under high-pressure conditions. Chem. Soc. Rev. 35, 858–875 (2006).
Lauro, F. M., Chastain, R. A., Blankenship, L. E., Yayanos, A. A. & Bartlett, D. H. The unique 16S rRNA genes of piezophiles reflect both phylogeny and adaptation. Appl. Environ. Microbiol. 73, 838–845 (2007).
Stetter, K. O. Extremophiles and their adaptation to hot environments. FEBS Lett. 452, 22–25 (1999).
Hurst, L. D. & Merchant, A. R. High guanine–cytosine content is not an adaptation to high temperature: a comparative analysis amongst prokaryotes. Proc. R. Soc. Lond. B 268, 493–497 (2000).
Nakashima, H., Fukuchi, S. & Nishikawa, K. Compositional changes in RNA, DNA and proteins for bacterial adaptation to higher and lower temperatures. J. Biochem. 133, 507–513 (2003).
Khachane, A. N., Timmis, K. N. & Martins dos Santos, V. A. P. Uracil content of 16S rRNA of thermophilic and psychrophilic prokaryotes correlates inversely with their optimal growth temperatures. Nucl. Acids Res. 33, 4016–4022 (2005).
Feller, G. & Gerday, C. Psychrophilic enzymes: hot topics in cold adadptation. Nat. Rev. Microbiol. 1, 200–208 (2003).
D’Amico, S., Marx, J. C., Gerday, C. & Feller, G. Activity–stability relationships in extremophilic enzymes. J. Biol. Chem. 278, 7891–7896 (2003).
Dias, C. L. et al. The hydrophobic effect and its role in cold denaturation. Cryobiology 60, 91–99 (2010).
Sicheri, F. & Yang, D. S. C. Ice-binding structure and mechanism of an antifreeze protein from winter flounder. Nature 375, 427–431 (1995).
Yeh, Y. & Feeney, R. E. Antifreeze proteins: structures and mechanisms of function. Chem. Rev. 96, 601–617 (1996).
Jaenicke, R. & Böhm, G. The stability of proteins in extreme environments. Curr. Opin. Struct. Biol. 8, 738–748 (1998).
Kumar, S. & Nussinov, R. How do thermophilic proteins deal with heat? Cell. Mol. Life Sci. 58, 1216–1233 (2001).
Vieille, C. & Zeikus, G. J. Hyperthermophilic enzymes: sources, uses, and molecular mechanisms for thermostability. Microbiol. Mol. Biol. Rev. 65, 1–43 (2001).
Berezovsky, I. N. & Shakhnovich, E. I. Physics and evolution of thermophilic adaptation. Proc. Natl Acad. Sci. USA 102, 12742–12747 (2005).
Lazaridis, T., Lee, I. & Karplus, M. Dynamics and unfolding pathways of a hyperthermophilic and a mesophilic rubredoxin. Protein Sci. 6, 2589–2605 (1997).
Kovacic, F., Mandrysch, A., Poojari, C., Strodel, B. & Jaeger, K. E. Structural features determining thermal adaptation of esterases. Protein Eng. Des. Sel. 29, 65–76 (2016).
Dominy, B. N., Minoux, H. & Brooks, C. L. III . An electrostatic basis for the stability of thermophilic proteins. Proteins 57, 128–141 (2004).
Missimer, J. H. et al. Configurational entropy elucidates the role of salt-bridge networks in protein thermostability. Protein Sci. 16, 1349–1359 (2007).
Roca, M., Liu, H., Messer, B. & Warshel, A. On the relationship between thermal stability and catalytic power of enzymes. Biochemistry 46, 15067–15088 (2007).
Bjelic, S., Brandsdal, B. O. & Åqvist, J. Cold adaptation of enzyme reaction rates. Biochemistry 47, 10049–10057 (2008).
Isaksen, G. V., Åqvist, J. & Brandsdal, B. O. Protein surface softness is the origin of enzyme cold-adaptation of trypsin. PLoS Comput. Biol. 8, e1003813 (2014).
Isaksen, G. V., Åqvist, J. & Brandsdal, B. O. Enzyme surface rigidity tunes the temperature dependence of catalytic rates. Proc. Natl Acad. Sci. USA 113, 7822–7827 (2016).
Åqvist, J., Wennerström, P., Nervall, M., Bjelic, S. & Brandsdal, B. O. Molecular dynamics simulations of water and biomolecules with a Monte Carlo constant pressure algorithm. Chem. Phys. Lett. 384, 288–294 (2004).
Kitchen, D. B., Reed, L. H. & Levy, R. M. Molecular dynamics simulation of solvated protein at high pressure. Biochemistry 31, 10083–10093 (1992).
Paci, E. High pressure simulations of biomolecules. Biochim. Biophys. Acta 1595, 185–200 (2002).
Low, P. S., Bada, J. L. & Somero, G. N. Temperature adaptation of enzymes: roles of the free energy, the enthalpy and the entropy of activation. Proc. Natl Acad. Sci. USA 70, 430–432 (1973).
Lonhienne, T., Gerday, C. & Feller, G. Psychrophilic enzymes: revisiting the thermodynamic parameters of activation may explain local flexibility. Biochim. Biophys. Acta 1543, 1–10 (2000).
Siddiqui, K. S. & Cavicchioli, R. Cold-adapted enzymes. Annu. Rev. Biochem. 75, 403–433 (2006).
Fields, P. A. & Somero, G. N. Hot spots in cold adaptation: localized increases in conformational flexibility in lactate dehydrogenase A4 orthologs of Antarctic notothenoid fishes. Proc. Natl Acad. Sci. USA 95, 11476–11481 (1998).
Altermark, B., Niiranen, L., Willasen, N. P., Smalås, A. O. & Moe, E. Comparative studies of endonuclease I from cold-adapted Vibrio salmonicida and mesophilic Vibrio cholerae. FEBS J. 274, 252–263 (2007).
Liang, Z. X., Tsigos, I., Bouriotis, V. & Klinman, J. P. Impact of protein flexibility on hydride-transfer parameters in thermophilic and psychrophilic alcohol dehydrogenases. J. Am. Chem. Soc. 126, 9500–9501 (2004).
Daniel, R. M. & Danson, M. J. A new understanding of how temperature affects the catalytic activity of enzymes. Trends Biochem. Sci. 35, 584–591 (2010).
Hobbs, J. K. et al. Change in heat capacity for enzyme catalysis determines temperature dependence of enzyme catalyzed rates. ACS Chem. Biol. 8, 2388–2393 (2008).
Arcus, V. L. Prentice, E. J. et al. On the temperature dependence of enzyme-catalyzed rates. Biochemistry 55, 1681–1688 (2016).
Nguyen, V. et al. Evolutionary drivers of thermoadaptation in enzyme catalysis. Science 355, 289–294 (2017).
Elias, M., Wieczorek, G., Rosenne, S. & Tawfik, D. S. The universality of enzymatic rate–temperature dependency. Trends Biochem. Sci. 39, 1–7 (2014).
Petrescu, I. et al. Xylanase from the psychrophilic yeast Cryptococcus adeliae. Extremophiles 4, 137–144 (2000).
Smalås, A. O., Heimstad, E. S., Hordvik, A., Willasen, W. P. & Male, R. Cold adaptation of enzymes: structural comparison between salmon and bovine trypsins. Proteins 20, 149–166 (1994).
Olufsen, M., Smalås, A. O., Moe, E. & Brandsdal, B. Increased flexibility as a strategy for cold adpatation. J. Biol. Chem. 280, 18042–18048 (2005).
Papaleo, E., Riccardi, L., Villa, C., Fantucci, P. & De Gioia, L. Flexibility and enzymatic cold-adaptation: a comparative molecular dynamics investigation of the elastase family. Biochim. Biophys. Acta 1764, 1397–1406 (2006).
Papaleo, E. et al. Protein flexibility in psychrophilic and mesophilic trypsins. Evidence of evolutionary conservation of protein dynamics in trypsin-like serine-proteases. FEBS Lett. 582, 1008–1018 (2008).
Åqvist, J., Kazemi, M., Isaksen, G. V. & Brandsdal, B. O. Entropy and enzyme catalysis. Acc. Chem. Res. 50, 199–207 (2017).
Kazemi, M. & Åqvist, J. Chemical reaction mechanisms in solution from brute force computational Arrhenius plots. Nat. Commun. 6, 7293 (2015).
Kazemi, M., Himo, F. & Åqvist, J. Enzyme catalysis by entropy without Circe effect. Proc. Natl Acad. Sci. USA 113, 2406–2411 (2016).
Åqvist, J. & Kamerlin, S. C. L. Exceptionally large entropy contributions enable the high rates of GTP hydrolysis on the ribosome. Sci. Rep. 5, 15817 (2015).
Åqvist, J. & Kamerlin, S. C. L. Conserved motifs in different classes of GTPases dictate their specific modes of catalysis. ACS Catal. 6, 1737–1743 (2016).
Warshel, A. Computer Modeling of Chemical Reactions in Enzymes and Solutions (Wiley, 1991).
Åqvist, J. & Warshel, A. Simulation of enzyme reactions using valence bond force fields and other hybrid quantum/classical approaches. Chem. Rev. 93, 2523–2544 (1993).
Snider, M. J., Gaunitz, S., Ridgway, C., Short, S. A. & Wolfenden, R. Temperature effects on the catalytic efficiency, rate enhancement, and transition state affinity of cytidine deaminase, and the thermodynamic consequences for catalysis of removing a substrate “anchor”. Biochemistry 39, 9746–9753 (2000).
Outzen, H., Berglund, G. I., Smalås, A. O. & Willasen, N. P. Temperature and pH sensitivity of trypsins from Atlantic salmon (Salmo salar) in comparison with bovine and porcine trypsin. Comp. Biochem. Physiol. B 115, 33–45 (1996).
Ghanem, M., Li, L., Wing, C. & Schramm, V. L. Altered thermodynamics from remote mutations altering human toward bovine purine nucleoside phosphorylase. Biochemistry 47, 2559–2564 (2008).
Isaksen, G. V., Åqvist, J. & Brandsdal, B. O. Thermodynamics of the purine nucleoside phosphorylase reaction revealed by computer simulations. Biochemistry 56, 306–312 (2017).
Leiros, H. K. S., McSweeney, S. M. & Smalås, A. O. Atomic resolution structures of trypsin provide insight into structural radiation damage. Acta Crystallogr. D 57, 488–497 (2001).
Liebschner, D., Dauter, M., Brzuszkiewicz, A. & Dauter, Z. On the reproducibility of protein crystal structures: five atomic resolution structures of trypsin. Acta Crystallogr. D 69, 1447–1462 (2013).
Bellissent-Funel, M. C. et al. Water determines the structure and dynamics of proteins. Chem. Rev. 116, 7673–7697 (2016).
Pucci, F. & Rooman, M. Physical and molecular bases of protein thermal stability and cold-adaptation. Curr. Opin. Struct. Biol. 42, 117–128 (2017).
Rodriguez-Correa, D. & Dahlberg, A. E. Kinetic and thermodynamic studies of of peptidyltransferase in ribosomes from the extreme thermophile Thermus thermophilus. RNA 14, 2314–2318 (2008).
Siddiqui, K. S., Cavicchioli, R. & Thomas, T. Thermodynamic activation parameters of elongation factor 2 (EF-2) proteins from psychrotolerant and thermophilic Archaea. Extremophiles 6, 143–150 (2002).
Merlino, A. et al. Structure and flexibility in cold-adapted iron superoxide dismutases: the case of the enzyme isolated from Pseudoalteromonas haloplanktis. J. Struct. Biol. 172, 343–352 (2010).
Struvay, C. & Feller, G. Optimization to low temperature activity in psychrophilic enzymes. Int. J. Mol. Sci. 13, 11643–11665 (2012).
Harms, M. J. & Thornton, J. W. Evolutionary biochemistry: revealing the historical and physical causes of protein properties. Nat. Rev. Genet. 14, 559–571 (2013).
Shoichet, B. K., Baase, W. A., Kuroki, R. & Matthews, B. W. A relationship between protein stability and protein function. Proc. Natl Acad. Sci. USA 92, 452–456 (1995).
Dang, L. X., Merz Jr, K. M. & Kollman, P. A. Free energy calculations on protein stability: Thr-157 → Val-157 mutation of T4 lysozyme. J. Am. Chem. Soc. 111, 8505–8508 (1989).
Pan, Y. P. & Daggett, V. Direct comparison of experimental and calculated folding free energies for hydrophobic deletion mutants of chymotrypsin inhibitor 2. Biochemistry 40, 2723–2731 (2001).
Seeliger, D. & de Groot, B. L. Protein thermostability calculations using alchemical free energy simulations. Biophys. J. 98, 2309–2316 (2011).
Folch, B., Dehouck, Y. & Rooman, M. Thermo- and mesostabilizing protein interactions identified by temperature-dependent statistical potentials. Biophys. J. 98, 667–677 (2010).
Pucci, F., Bourgeas, R. & Rooman, M. Predicting protein thermal stability changes upon point mutations using statistical potentials: introducing HoTMuSiC. Sci. Rep. 6, 23257 (2016).
Eyring, H. The activated complex in chemical reactions. J. Chem. Phys. 3, 107–115 (1935).
Evans, M. G. & Polanyi, M. Some applications of the transition state method to the calculation of reaction velocities, especially in solution. Trans. Faraday Soc. 31, 875–893 (1935).
Bjelic, S. & Åqvist, J. Catalysis and linear free energy relationships in aspartic proteases. Biochemistry 45, 7709–7723 (2006).
Russell, R. J., Ferguson, J. M., Hough, D. W., Danson, M. J. & Taylor, G. L. The crystal structure of citrate synthase from the hyperthermophilic archaeon Pyrococcus furiosus at 1.9 Å resolution. Biochemistry 36, 9983–9994 (1997).
Russell, R. J., Gerike, U., Danson, M. J., Hough, D. W. & Taylor, G. L. Structural adaptations of the cold-active citrate synthase from an Antarctic bacterium. Structure 6, 351—361 (1998).
Leiros, H. K. S., Willassen, N. P. & Smalås, A. O. Structural comparison of psychrophilic and mesophilic trypsins: elucidating the molecular basis of cold-adaptation. Eur. J. Biochem. 267, 1039–1049 (2000).
Mereghetti, P. et al. Near native-state conformational landscape of psychrophilic and mesophilic enzymes: probing the folding funnel model. J. Phys. Chem. B. 114, 7609–7619 (2010).
Olufsen, M., Brandsdal, B. O. & Smalås, A. O. Comparative unfolding studies of psychrophilic and mesophilic uracil DNA glycosylase: MD simulations show reduced thermal stability of the cold-adapted enzyme. J. Mol. Graph. Model. 26, 124–134 (2007).
Papaleo, E., Olufsen, M., De Gioia, L. & Brandsdal, B. O. Optimization of electrostatics as a strategy for cold-adaptation: a case study of cold- and warm-active elastases. J. Mol. Graph. Model. 26, 93–103 (2007).
Michetti, D. et al. A comparative study of cold- and warm-adapted Endonucleases A using sequence analyses and molecular dynamics simulations. PLoS ONE 12, e0169586 (2017).
Papaleo, E., Pasi, M., Tiberti, M. & De Gioia, L. Molecular dynamics of mesophilic-like mutants of a cold-adapted enzyme: insights into distal effects induced by the mutations. PLoS ONE 6, e24214 (2011).
Zanphorlin, L. M. et al. Oligomerization as a strategy for cold adaptation: structure and dynamics of the GH1 β-glucosidase from Exiguobacterium antarcticum B7. Sci. Rep. 6, 23776 (2016).
Parvizpour, S., Razmara, J., Ramli, A. N. M., Md Illias, R. & Shamsir, M. S. Structural and functional analysis of a novel psychrophilic β-mannanase from Glaciozyma antarctica PI12. J. Comput. Aided Mol. Des. 28, 685–698 (2014).
Kim, M. K. et al. Structure-based investigation into the functional roles of the extended loop and substrate-recognition sites in an endo-β-1,4-d-mannanase from the Antarctic springtail. Cryptopygus antarcticus. Proteins 82, 3217–3223 (2014).
Gatti-Lafranconi, P. et al. Evolution of stability in a cold-active enzyme elicits specificity relaxation and highlights substrate-related effects on temperature adaptation. J. Mol. Biol. 395, 155–166 (2010).
Sigtryggsdóttir, Á. R., Papaleo, E., Thorbjarnardóttir, S. H. & Kristjánsson, M. M. Flexibility of cold- and heat-adapted subtilisin-like serine proteinases evaluated with fluorescence quenching and molecular dynamics. Biochim. Biophys. Acta 1844, 705–712 (2014).
Xie, B. B. et al. Cold adaptation of zinc metalloproteases in the thermolysin family from deep sea and arctic sea ice bacteria revealed by catalytic and structural properties and molecular dynamics: new insights into relationship between conformational flexibility and hydrogen bonding. J. Biol. Chem. 284, 9257–9269 (2009).
Adekoya, O. A., Helland, R., Willassen, N. P. & Sylte, I. Comparative sequence and structure analysis reveal features of cold adaptation of an enzyme in the thermolysin family. Proteins 62, 435–449 (2006).
Acknowledgements
The authors gratefully acknowledge support from the Swedish Research Council (VR), the Knut and Alice Wallenberg Foundation and the Research Council of Norway (through a Centre of Excellence grant, 179568/V30).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Åqvist, J., Isaksen, G. & Brandsdal, B. Computation of enzyme cold adaptation. Nat Rev Chem 1, 0051 (2017). https://doi.org/10.1038/s41570-017-0051
Published:
DOI: https://doi.org/10.1038/s41570-017-0051
This article is cited by
-
High temperature delays and low temperature accelerates evolution of a new protein phenotype
Nature Communications (2024)
-
Comparative molecular dynamics simulations provided insights into the mechanisms of cold-adaption of alginate lyases from the PL7 family
Extremophiles (2024)
-
Genomic analyses reveal a low-temperature adapted clade in Halorubrum, a widespread haloarchaeon across global hypersaline environments
BMC Genomics (2023)
-
Functional activity of E. coli RNase R in the Antarctic Pseudomonas syringae Lz4W
Journal of Genetic Engineering and Biotechnology (2023)
-
Extremophile enzyme optimization for low temperature and high salinity are fundamentally incompatible
Extremophiles (2022)