Abstract
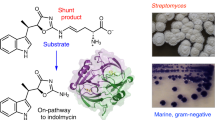
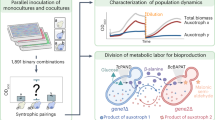
The biosynthesis of secondary metabolites by ‘group effort’ — in which two or more species cooperate to generate a hybrid small molecule — has been vastly underappreciated. In the laboratory, biosynthetic studies typically focus on a single species that is responsible for the expression of the necessary enzymes and assembly of the small-molecule product. However, in natural environments, microorganisms live in tight associations and are surrounded by a dynamic and intricate exchange of small molecules. The biosynthetic paradigm that is emerging for these conditions is one in which exogenous signals can act as inducers of silent pathways or as alternative substrates that give rise to hybrid small molecules. Examples of such secondary metabolites of dual origin are highlighted in this Perspective article. Aside from demonstrating the remarkable metabolic economy that microorganisms use to create complex molecules, these examples also highlight the benefits of studying biosynthesis using a multispecies approach. This integrative mindset reveals not only the chemistry that underlies symbiotic biosynthetic pathways but also the naturally evolved functions of secondary metabolites that mediate or modulate interspecies interactions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Curtis, T. P., Sloan, W. T. & Scannell, J. W. Estimating prokaryotic diversity and its limits. Proc. Natl Acad. Sci. USA 9, 10494–10499 (2002).
Young, I. M. & Crawford, J. W. Interactions and self-organization in the soil–microbe complex. Science 304, 1634–1637 (2004).
Traxler, M. F. & Kolter, R. Natural products in soil microbe interactions and evolution. Nat. Prod. Rep. 32, 956–970 (2015).
Fuqua, C. & Greenberg, E. P. Listening in on bacteria: acyl-homoserine lactone signaling. Nat. Rev. Mol. Cell Biol. 3, 685–695 (2002).
Waters, C. M. & Bassler, B. L. Quorum sensing: cell-to-cell communication in bacteria. Annu. Rev. Cell Dev. Biol. 21, 319–346 (2005).
Pearson, J. P., Passador, L., Iglewski, B. H. & Greenberg, E. P. A second N-acylhomoserine lactone signal produced by Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 92, 1490–1494 (1995).
Fuqua, W. C. & Winans, S. C. A. LuxR–LuxI type regulatory system activates Agrobacterium Ti plasmid conjugal transfer in the presence of a plant tumor metabolite. J. Bacteriol. 176, 2796–2806 (1994).
Nett, M., Ikeda, H. & Moore, B. S. Genomic basis for natural product biosynthetic diversity in the actinomycetes. Nat. Prod. Rep. 26, 1362–1384 (2009).
Schaefer, A. L. et al. A new class of homoserine lactone quorum-sensing signals. Nature 454, 595–599 (2008).
Ahlgren, N. A., Harwood, C. S., Schaefer, A. L., Giraud, E. & Greenberg, E. P. Aryl-homoserine lactone quorum sensing in stem-nodulating photosynthetic bradyrhizobia. Proc. Natl Acad. Sci. USA 108, 7183–7188 (2011).
Schmidt, E. W. Trading molecules and tracking targets in symbiotic interactions. Nat. Chem. Biol. 4, 466–473 (2008).
Schink, B. Synergistic interactions in the microbial world. Antonie Van Leeuwenhoek 81, 257–261 (2002).
Dubilier, N., Bergin, C. & Lott, C. Symbiotic diversity in marine animals: the art of harnessing chemosynthesis. Nat. Rev. Microbiol. 6, 725–740 (2008).
Stewart, F. J., Newton, I. L. & Cavanaugh, C. M. Chemosynthetic endosymbioses: adaptations to oxic–anoxic interfaces. Trends Microbiol. 13, 439–448 (2005).
Croft, M. T., Lawrence, A. D., Raux-Deery, E., Warren, M. J. & Smith, A. G. Algae acquire vitamin B12 through a symbiotic relationship with bacteria. Nature 438, 90–93 (2005).
Stupp, G. S. et al. Chemical detoxification of small molecules by Caenorhabditis elegans. ACS Chem. Biol. 8, 309–313 (2013).
Wright, G. D. The antibiotic resistome: the nexus of chemical and genetic diversity. Nat. Rev. Microbiol. 5, 175–186 (2007).
Pan, C. et al. Characterization of anaerobic catabolism of p-coumarate in Rhodopseudomonas palustris by integrating transcriptomics and quantitative proteomics. Mol. Cell Proteomics 7, 938–948 (2008).
Larimer, F. W. et al. Complete genome sequence of the metabolically versatile photosynthetic bacterium Rhodopseudomonas palustris. Nat. Biotechnol. 22, 55–61 (2004).
Palmer, A. G. & Blackwell, H. E. Deciphering a protolanguage for bacteria–host communication. Nat. Chem. Biol. 4, 452–454 (2008).
Charlson, R. J., Lovelock, J. E., Andreae, M. O. & Warren, S. G. Oceanic phytoplankton, atmospheric sulphur, cloud albedo and climate. Nature 326, 655–661 (1987).
Holligan, P. M., Viollier, M., Harbour, D. S., Camus, P. & Champagne-Philippe, M. Satellite and ship studies of coccolithophore production along a continental shelf edge. Nature 304, 339–342 (1983).
Wagner-Döbler, I. & Biebl, H. Environmental biology of the marine Roseobacter lineage. Annu. Rev. Microbiol. 60, 255–280 (2006).
Geng, H. & Belas, R. Molecular mechanisms underlying Roseobacter–phytoplankton symbioses. Curr. Opin. Biotechnol. 21, 332–338 (2010).
Brinkhoff, T., Giebel, H. A. & Simon, M. Diversity, ecology, and genomics of the Rosebacter clade: a short overview. Arch. Microbiol. 189, 531–539 (2008).
Kiene, R. P., Linn, L. J., González, J., Moran, M. A. & Bruton, J. A. Dimethylsulfoniopropionate and methanethiol are important precursors of methionine and protein–sulfur in marine bacterioplankton. Appl. Environ. Microbiol. 65, 4549–4558 (1999).
Reisch, C. R. et al. Novel pathway for assimilation of dimehtylsulphoniopropionate widespread in marine bacteria. Nature 473, 208–211 (2011).
Thiel, V. et al. Identification and biosynthesis of tropone derivatives and sulfur volatiles produced by bacteria of the marine Roseobacter clade. Org. Biomol. Chem. 8, 234–246 (2010).
Brinkhoff, T. et al. Antibiotic production by a Roseobacter clade-affiliated species from the German Wadden Sea and its antagonistic effects on indigenous isolates. Appl. Environ. Microbiol. 70, 2560–2565 (2004).
Seyedsayamdost, M. R., Case, R. J., Kolter, R. & Clardy, J. The Jekyll-and-Hyde chemistry of Phaeobacter gallaeciensis. Nat. Chem. 3, 331–335 (2011).
Seyedsayamdost, M. R., Carr, G., Kolter, R. & Clardy, J. Roseobacticides: small molecule modulators of an algal–bacterial symbiosis. J. Am. Chem. Soc. 133, 18343–18349 (2011).
González, J. M. et al. Bacterial community structure associated with a dimethylsulfoniopropionate-producing north Atlantic algal bloom. Appl. Environ. Microbiol. 66, 4237–4246 (2000).
Lamy, D. et al. Temporal changes of major bacterial groups and bacterial heterotrophic activity during a Phaeocystic globosa bloom in the eastern English Channel. Aquat. Microb. Ecol. 58, 95–107 (2009).
Seyedsayamdost, M. R., Wang, R., Kolter, R. & Clardy, J. Hybrid biosynthesis of roseobacticides from algal and bacterial precursor molecules. J. Am. Chem. Soc. 136, 15150–15153 (2014).
Wang, R., Gallant, É. & Seyedsayamdost, M. R. Investigation of the genetics and biochemistry of roseobacticide production in the Roseobacter clade bacterium Phaeobacter inhibens. mBio 7, e02118-15 (2016).
Trost, B. M. The atom economy — a search for synthetic efficiency. Science 254, 1471–1477 (1991).
Alborn, H. T. et al. An elicitor of plant volatiles from beet armyworm oral secretion. Science 276, 945–949 (1997).
Paré, P. W., Alborn, H. T. & Tumlinson, J. H. Concerted biosynthesis of an insect elicitor of plant volatiles. Proc. Natl Acad. Sci. USA 95, 13971–13975 (1998).
Lait, C. G., Alborn, H. T., Teal, P. E. & Tumlinson, J. H. Rapid biosynthesis of N-linolenoyl-L-glutamine, an elicitor of plant volatiles, by membrane-associated enzyme(s) in Manduca sexta. Proc. Natl Acad. Sci. USA 100, 7027–7032 (2003).
Yoshinaga, N., Morigaki, N., Matsuda, F., Nishida, R. & Mori, N. In vitro biosynthesis of volicitin in Spodoptera litura. Insect Biochem. Mol. Biol. 35, 175–184 (2005).
Paré, P. W. & Tumlinson, J. H. Induced synthesis of plant volatiles. Nature 385, 30–31 (1997).
De Moraes, C. M., Lewis, W. J., Paré, P. W., Alborn, H. T. & Tumlinson, J. H. Herbivore-infested plants selectively attract parasitoids. Nature 393, 570–573 (1998).
Turlings, T. C. J., Tumlinson, J. H. & Lewis, W. J. Exploitation of herbivore-induced plant odors by host-seeking parasitic wasps. Science 250, 1251–1253 (1990).
Fazeau-Braesch, S., Genin, E., Jullien, R., Knowles, E. & Papin, C. Composition and role of volatile substances in atmosphere surrounding two gregarious locusts, Locusta migratoria and Schistocerca gregaria. J. Chem. Ecol. 14, 1023–1033 (1988).
Torto, B., Obeng-Ofori, D., Njagi, P. G., Hassanali, A. & Amiani, H. Aggregation pheromone system of adult gregarious desert locust Schistocerca gregaria (forskal). J. Chem. Ecol. 20, 1749–1762 (1994).
Obeng-Ofori, D., Torto, B., Njagi, P. G., Hassanali, A. & Amiani, H. Fecal volatiles as part of the aggregation pheromone complex of the desert locust, Schistocerca gregaria (forskal) (Orthoptera: Acrididae). J. Chem. Ecol. 20, 2077–2087 (1994).
Charnley, A. K., Hunt, J. & Dillon, R. J. The germ-free culture of desert locusts, Schistocerca gregaria. J. Insect. Physiol. 31, 477–485 (1985).
Dillon, R. J., Vennard, C. T. & Charnley, A. K. Pheromones: exploitation of gut bacteria in the locust. Nature 403, 851 (2000).
Dillon, R. J., Vennard, C. T. & Charnley, A. K. A note: gut bacteria produce components of a locust cohesion pheromone. J. Appl. Microbiol. 92, 759–763 (2002).
Schmidt, E. W. & Donia, M. S. Life in cellulose houses: symbiotic bacterial biosynthesis of ascidian drugs and drug leads. Curr. Opin. Biotechnol. 21, 827–833 (2010).
Sings, H. L. & Rinehart, K. L. Compounds produced from potential tunicate-blue-green algal symbiosis: a review. J. Ind. Microbiol. 17, 385–396 (1996).
Sings, H. L., Bible, K. C. & Rinehart, K. L. Acyl tunichlorins: a new class of nickel chlorins isolated from the Caribbean tunicate Trididemnum solidum. Proc. Natl Acad. Sci. USA 93, 10560–10565 (1996).
Kustin, K. & McLeod, G. C. Interactions between metal ions and living organisms in sea water. Top. Curr. Chem. 69, 1–37 (1977).
Rayner-Canham, G. W., Van Roode, M. & Burke, J. Nickel and cobalt concentrations in the tunicate Halocynthia pyriformis: evidence for essentiality of the two metals. Inorg. Chim. Acta 106, L37–L38 (1985).
Pettit, G. R. et al. The isolation and structure of a remarkable marine animal antineoplastic constituent: dolastatin 10. J. Am. Chem. Soc. 109, 6883–6885 (1987).
Pukatzki, S. & Provenzano, D. Vibrio cholerae as a predator: lessons from evolutionary principles. Front. Microbiol. 4, 384 (2013).
Bassler, B. L. & Losick, R. Bacterially speaking. Cell 125, 237–246 (2006).
Chen, X. et al. Structural identification of a bacterial quorum-sensing signal containing boron. Nature 415, 545–549 (2002).
Higgins, D. A. et al. The major Vibrio cholerae autoinducer and its role in virulence factor production. Nature 450, 883–886 (2007).
Papenfort, K., Förstner, K. U., Cong, J. P., Sharma, C. M. & Bassler, B. L. Differential RNA-seq of Vibrio cholerae identifies the VqmR small RNA as a regulator of biofilm formation. Proc. Natl Acad. Sci. USA 112, E766–E775 (2015).
Papenfort, K. et al. Vibrio cholerae autoinducer—receptor pair that controls biofilm formation. Nat. Chem. Biol. (in the press).
McGuckin, M. A., Lindén, S. K., Sutton, P. & Florin, T. H. Mucin dynamics and enteric pathogens. Nat. Rev. Microbiol. 9, 265–278 (2011).
Millet, Y. A. et al. Insights into Vibrio cholerae intestinal colonization from monitoring fluorescently labeled bacteria. PLoS Pathog. 10, e1004405 (2014).
Crawford, J. M., Kontnik, R. & Clardy, J. Regulating alternative lifestyles in entomopathogenic bacteria. Curr. Biol. 20, 69–74 (2010).
Fang, A., Keables, P. & Demain, A. L. Unexpected enhancement of β-lactam antibiotic formation in Streptomyces clavuligerus by very high concentrations of exogenous lysine. Appl. Microbiol. Biotechnol. 44, 705–709 (1996).
Acknowledgements
Research in the author's laboratory on microbial symbiotic interactions was generously supported by the US National Institutes of Health (grant GM098299) and the Pew Biomedical Scholars Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Wang, R., Seyedsayamdost, M. Opinion: Hijacking exogenous signals to generate new secondary metabolites during symbiotic interactions. Nat Rev Chem 1, 0021 (2017). https://doi.org/10.1038/s41570-017-0021
Published:
DOI: https://doi.org/10.1038/s41570-017-0021
This article is cited by
-
Toward a global picture of bacterial secondary metabolism
Journal of Industrial Microbiology and Biotechnology (2019)
-
Best practices for analysing microbiomes
Nature Reviews Microbiology (2018)