Abstract
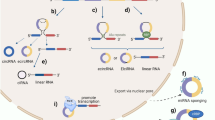
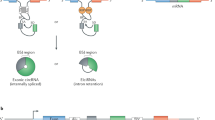
Circular RNAs (circRNAs) are emerging as a new class of non-coding RNA molecules. This unusual class of RNA species is generated by a back-splicing event of one or two exons, resulting in a covalently closed circRNA molecule. Owing to their circular form, circRNAs are protected from degradation by exonucleases and have greater stability than linear RNA. Advances in computational analysis of RNA sequencing have revealed that thousands of different circRNAs are expressed in a wide range of mammalian tissues, including the cardiovascular system. Moreover, numerous circRNAs are expressed in a disease-specific manner. A great deal of progress has been made in understanding the biogenesis and function of these circRNAs. In this Review, we discuss the current understanding of circRNA biogenesis and function, with a particular emphasis on the cardiovascular system.
Key points
-
Circular RNAs (circRNAs) are a large class of non-coding RNA molecules that form a covalently closed loop (unlike linear RNAs).
-
circRNAs are produced from precursor mRNA back-splicing, a process catalysed by the spliceosome machinery.
-
High-throughput sequencing has identified thousands of circRNAs in the human body, a great number of which are expressed in a tissue-specific or disease-specific manner.
-
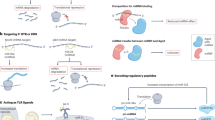
Although the biological functions of most circRNAs remain unknown, specific circRNAs have been shown to act as microRNA sponges, to interact with RNA-binding proteins, to regulate transcription or to be translated into proteins.
-
Preliminary studies have provided evidence that individual circRNAs have critical regulatory functions in the cardiovascular system.
-
Owing to their high stability and abundance in bodily fluids, circRNAs have potential as useful biomarkers for various diseases, including cardiovascular disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hsu, M.-T. & Coca-Prados, M. Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature 280, 339–340 (1979).
Sanger, H. L., Klotz, G., Riesner, D., Gross, H. J. & Kleinschmidt, A. K. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc. Natl Acad. Sci. USA 73, 3852–3856 (1976).
Cocquerelle, C., Mascrez, B., Hétuin, D. & Bailleul, B. Mis-splicing yields circular RNA molecules. FASEB J. 7, 155–160 (1993).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013).
Salzman, J., Chen, R. E., Olsen, M. N., Wang, P. L. & Brown, P. O. Cell-type specific features of circular RNA expression. PLOS Genet. 9, e1003777 (2013).
Wang, P. L. et al. Circular RNA is expressed across the eukaryotic tree of life. PLOS ONE 9, e90859 (2014).
Jeck, W. R. & Sharpless, N. E. Detecting and characterizing circular RNAs. Nat. Biotechnol. 32, 453–461 (2014).
Chen, X. et al. circRNADb: a comprehensive database for human circular RNAs with protein-coding annotations. Sci. Rep. 6, 34985 (2016).
Enuka, Y. et al. Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res. 44, 1370–1383 (2016).
Piwecka, M. et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science 357, eaam8526 (2017).
Aufiero, S. et al. Cardiac circRNAs arise mainly from constitutive exons rather than alternatively spliced exons. RNA 24, 815–827 (2018).
Rybak-Wolf, A. et al. Circular RNAs in the mammalian brain are highly abundant, conserved, and dynamically expressed. Mol. Cell 58, 870–885 (2015).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157 (2013).
Zheng, Q. et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple mi-RNAs. Nat. Commun. 7, 11215 (2016).
Salzman, J., Gawad, C., Wang, P. L., Lacayo, N. & Brown, P. O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLOS ONE 7, e30733 (2012).
Zhang, Y. et al. The biogenesis of nascent circular RNAs. Cell Rep. 15, 611–624 (2016).
Bachmayr-Heyda, A. et al. Correlation of circular RNA abundance with proliferation—exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci. Rep. 5, 8057 (2015).
Maass, P. G. et al. A map of human circular RNAs in clinically relevant tissues. J. Mol. Med. 95, 1179–1189 (2017).
Guo, J. U., Agarwal, V., Guo, H. & Bartel, D. P. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 15, 409 (2014).
Haque, S. & Harries, L. W. Circular RNAs (circRNAs) in health and disease. Genes 8, E353 (2017).
Szabo, L. et al. Statistically based splicing detection reveals neural enrichment and tissue-specific induction of circular RNA during human fetal development. Genome Biol. 16, 126 (2015).
You, X. et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat. Neurosci. 18, 603–610 (2015).
Westholm, J. O. et al. Genomewide analysis of Drosophila circular RNAs reveals their structural and sequence properties and age-dependent neural accumulation. Cell Rep. 9, 1966–1980 (2014).
Siede, D. et al. Identification of circular RNAs with host gene-independent expression in human model systems for cardiac differentiation and disease. J. Mol. Cell. Cardiol. 109, 48–56 (2017).
Du, W. W. et al. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur. Heart J. 38, 1402–1412 (2017).
Starke, S. et al. Exon circularization requires canonical splice signals. Cell Rep. 10, 103–111 (2015).
Dubin, R. A., Kazmi, M. A. & Ostrer, H. Inverted repeats are necessary for circularization of the mouse testis Sry transcript. Gene 167, 245–248 (1995).
Ivanov, A. et al. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep. 10, 170–177 (2015).
Zhang, X.-O. et al. Complementary sequence-mediated exon circularization. Cell 159, 134–147 (2014).
Liang, D. & Wilusz, J. E. Short intronic repeat sequences facilitate circular RNA production. Genes Dev. 28, 2233–2247 (2014).
Ashwal-Fluss, R. et al. circRNA biogenesis competes with pre-mRNA splicing. Mol. Cell 56, 55–66 (2014).
Conn, S. J. et al. The RNA binding protein quaking regulates formation of circRNAs. Cell 160, 1125–1134 (2015).
Kramer, M. C. et al. Combinatorial control of Drosophila circular RNA expression by intronic repeats, hnRNPs, and SR proteins. Genes Dev. 29, 2168–2182 (2015).
Li, X. et al. Coordinated circRNA biogenesis and function with NF90/NF110 in viral infection. Mol. Cell 67, 214–227 (2017).
Zhang, Y. et al. Circular intronic long noncoding RNAs. Mol. Cell 51, 792–806 (2013).
Zaphiropoulos, P. G. Circular RNAs from transcripts of the rat cytochrome P450 2C24 gene: correlation with exon skipping. Proc. Natl Acad. Sci. USA 93, 6536–6541 (1996).
Khan, M. A. F. et al. RBM20 regulates circular RNA production from the titin gene. Circ. Res. 119, 996–1003 (2016).
Zeng, X., Lin, W., Guo, M. & Zou, Q. A comprehensive overview and evaluation of circular RNA detection tools. PLOS Comput. Biol. 13, e1005420 (2017).
Li, X., Chu, C., Pei, J., Mandoiu, I. & Wu, Y. CircMarker: a fast and accurate algorithm for circular RNA detection. BMC Genomics 19, 572 (2018).
Gao, Y. & Zhao, F. Computational strategies for exploring circular RNAs. Trends Genet. 34, 389–400 (2018).
Hansen, T. B., Venø, M. T., Damgaard, C. K. & Kjems, J. Comparison of circular RNA prediction tools. Nucleic Acids Res. 44, e58 (2016).
Szabo, L. & Salzman, J. Detecting circular RNAs: bioinformatic and experimental challenges. Nat. Rev. Genet. 17, 679–692 (2016).
Suzuki, H. et al. Characterization of RNase R-digested cellular RNA source that consists of lariat and circular RNAs from pre-mRNA splicing. Nucleic Acids Res. 34, e63 (2006).
Yang, L., Duff, M. O., Graveley, B. R., Carmichael, G. G. & Chen, L.-L. Genomewide characterization of non-polyadenylated RNAs. Genome Biol. 12, R16 (2011).
Legnini, I. et al. Circ-ZNF609 is a circular RNA that can be translated and functions in myogenesis. Mol. Cell 66, 22–37 (2017).
Boeckel, J.-N. et al. Identification and characterization of hypoxia-regulated endothelial circular RNA. Circ. Res. 117, 884–890 (2015).
Wang, K. et al. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur. Heart J. 37, 2602–2611 (2016).
Holdt, L. M. et al. Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat. Commun. 7, 12429 (2016).
Kamens, J. The Addgene repository: an international nonprofit plasmid and data resource. Nucleic Acids Res. 43, D1152–D1157 (2015).
Hansen, T. B. et al. miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J. 30, 4414–4422 (2011).
Hansen, T. B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388 (2013).
Bosson, A. D., Zamudio, J. R. & Sharp, P. A. Endogenous miRNA and target concentrations determine susceptibility to potential ceRNA competition. Mol. Cell 56, 347–359 (2014).
Abdelmohsen, K. et al. Identification of HuR target circular RNAs uncovers suppression of PABPN1 translation by CircPABPN1. RNA Biol. 14, 361–369 (2017).
Congrains, A., Kamide, K., Ohishi, M. & Rakugi, H. ANRIL: molecular mechanisms and implications in human health. Int. J. Mol. Sci. 14, 1278–1292 (2013).
Li, Z. et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 22, 256–264 (2015).
Liang, D. et al. The output of protein-coding genes shifts to circular RNAs when the pre-mRNA processing machinery is limiting. Mol. Cell 68, 940–954 (2017).
Chen, C. Y. & Sarnow, P. Initiation of protein synthesis by the eukaryotic translational apparatus on circular RNAs. Science 268, 415–417 (1995).
Perriman, R. & Ares, M. Circular mRNA can direct translation of extremely long repeating-sequence proteins in vivo. RNA 4, 1047–1054 (1998).
Wang, Y. & Wang, Z. Efficient backsplicing produces translatable circular mRNAs. RNA 21, 172–179 (2015).
Pamudurti, N. R. et al. Translation of circRNAs. Mol. Cell 66, 9–21 (2017).
Yang, Y. et al. Extensive translation of circular RNAs driven by N6-methyladenosine. Cell Res. 27, 626–641 (2017).
Jakobi, T., Czaja-Hasse, L. F., Reinhardt, R. & Dieterich, C. Profiling and validation of the circular RNA repertoire in adult murine hearts. Genomics Proteomics Bioinformatics 14, 216–223 (2016).
Werfel, S. et al. Characterization of circular RNAs in human, mouse and rat hearts. J. Mol. Cell. Cardiol. 98, 103–107 (2016).
Tan, W. L. W. et al. A landscape of circular RNA expression in the human heart. Cardiovasc. Res. 113, 298–309 (2017).
Dong, R., Ma, X.-K., Chen, L.-L. & Yang, L. Increased complexity of circRNA expression during species evolution. RNA Biol. 14, 1064–1074 (2016).
Li, X. F. & Lytton, J. A circularized sodium-calcium exchanger exon 2 transcript. J. Biol. Chem. 274, 8153–8160 (1999).
Anderson, B. R. & Granzier, H. L. Titin-based tension in the cardiac sarcomere: molecular origin and physiological adaptations. Prog. Biophys. Mol. Biol. 110, 204–217 (2012).
Haas, J. et al. Atlas of the clinical genetics of human dilated cardiomyopathy. Eur. Heart J. 36, 1123–1135a (2015).
Guo, W. et al. RBM20, a gene for hereditary cardiomyopathy, regulates titin splicing. Nat. Med. 18, 766–773 (2012).
Maatz, H. et al. RNA-binding protein RBM20 represses splicing to orchestrate cardiac pre-mRNA processing. J. Clin. Invest. 124, 3419–3430 (2014).
Gupta, S. K. et al. Quaking inhibits doxorubicin-mediated cardiotoxicity through regulation of cardiac circular RNA expression. Circ. Res. 122, 246–254 (2018).
Tang, C.-M. et al. CircRNA_000203 enhances the expression of fibrosis-associated genes by derepressing targets of miR-26b-5p, Col1a2 and CTGF, in cardiac fibroblasts. Sci. Rep. 7, 40342 (2017).
Zeng, Y. et al. A circular RNA binds to and activates AKT phosphorylation and nuclear localization reducing apoptosis and enhancing cardiac repair. Theranostics 7, 3842–3855 (2017).
Memczak, S., Papavasileiou, P., Peters, O. & Rajewsky, N. Identification and characterization of circular RNAs as a new class of putative biomarkers in human blood. PLOS ONE 10, e0141214 (2015).
Li, Y. et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res. 25, 981–984 (2015).
Koh, W. et al. Noninvasive in vivo monitoring of tissue-specific global gene expression in humans. Proc. Natl Acad. Sci. USA 111, 7361–7366 (2014).
Bahn, J. H. et al. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin. Chem. 61, 221–230 (2015).
Vausort, M. et al. Myocardial infarction-associated circular RNA predicting left ventricular dysfunction. J. Am. Coll. Cardiol. 68, 1247–1248 (2016).
Zhang, J. et al. Plasma circular RNAs, Hsa_circRNA_025016, predict postoperative atrial fibrillation after isolated off-pump coronary artery bypass grafting. J. Am. Heart Assoc. 7, e006642 (2018).
Melo, S. A. et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 523, 177–182 (2015).
Jost, I. et al. Functional sequestration of microRNA-122 from hepatitis C virus by circular RNA sponges. RNA Biol. 15, 1032–1039 (2018).
Wang, K. et al. Circular RNA mediates cardiomyocyte death via miRNA-dependent upregulation of MTP18 expression. Cell Death Differ. 24, 1111–1120 (2017).
Geng, H.-H. et al. The circular RNA Cdr1as promotes myocardial infarction by mediating the regulation of miR-7a on its target genes expression. PLOS ONE 11, e0151753 (2016).
Wang, Y. et al. A Zfp609 circular RNA regulates myoblast differentiation by sponging miR-194-5p. Int. J. Biol. Macromol. 121, 1308–1313 (2018).
Zhou, B. & Yu, J.-W. A novel identified circular RNA, circRNA_010567, promotes myocardial fibrosis via suppressing miR-141 by targeting TGF-β1. Biochem. Biophys. Res. Commun. 487, 769–775 (2017).
Shan, K. et al. Circular noncoding RNA HIPK3 mediates retinal vascular dysfunction in diabetes mellitus. Circulation 136, 1629–1642 (2017).
Liu, C. et al. Silencing of circular RNA-ZNF609 ameliorates vascular endothelial dysfunction. Theranostics 7, 2863–2877 (2017).
He, Q. et al. circ-SHKBP1 regulates the angiogenesis of U87 glioma-exposed endothelial cells through miR-544a/FOXP1 and miR-379/FOXP2 pathways. Mol. Ther. Nucleic Acids 10, 331–348 (2018).
Sun, Y. et al. A novel regulatory mechanism of smooth muscle α-actin expression by NRG-1/circACTA2/miR-548f-5p axis. Circ. Res. 121, 628–635 (2017).
Dang, R.-Y., Liu, F.-L. & Li, Y. Circular RNA hsa_circ_0010729 regulates vascular endothelial cell proliferation and apoptosis by targeting the miR-186/HIF-1α axis. Biochem. Biophys. Res. Commun. 490, 104–110 (2017).
Li, C.-Y., Ma, L. & Yu, B. Circular RNA hsa_circ_0003575 regulates oxLDL induced vascular endothelial cells proliferation and angiogenesis. Biomed. Pharmacother. 95, 1514–1519 (2017).
Chen, J., Cui, L., Yuan, J., Zhang, Y. & Sang, H. Circular RNA WDR77 target FGF-2 to regulate vascular smooth muscle cells proliferation and migration by sponging miR-124. Biochem. Biophys. Res. Commun. 494, 126–132 (2017).
Zhang, S.-J. et al. Identification and characterization of circular RNAs as a new class of putative biomarkers in diabetes retinopathy. Invest. Ophthalmol. Vis. Sci. 58, 6500–6509 (2017).
Pan, L. et al. Human circular RNA-0054633 regulates high glucose-induced vascular endothelial cell dysfunction through the microRNA-218/roundabout 1 and microRNA-218/heme oxygenase-1 axes. Int. J. Mol. Med. 42, 597–606 (2018).
Acknowledgements
Reviewer information
Nature Reviews Cardiology thanks G. Condorelli, C. Dieterich and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.A. and E.E.C. contributed to researching data for the article, discussion of content, writing, reviewing and editing the manuscript before submission. Y.J.R. contributed to researching data for the article, writing, reviewing and editing the manuscript before submission. Y.M.P. contributed to reviewing and editing the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
circBase: http://www.circbase.org
Rights and permissions
About this article
Cite this article
Aufiero, S., Reckman, Y.J., Pinto, Y.M. et al. Circular RNAs open a new chapter in cardiovascular biology. Nat Rev Cardiol 16, 503–514 (2019). https://doi.org/10.1038/s41569-019-0185-2
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41569-019-0185-2
This article is cited by
-
Non-coding RNAs in disease: from mechanisms to therapeutics
Nature Reviews Genetics (2024)
-
Circular RNA EPHA3 suppresses progression and metastasis in prostate cancer through the miR-513a-3p/BMP2 axis
Journal of Translational Medicine (2023)
-
CircPrime: a web-based platform for design of specific circular RNA primers
BMC Bioinformatics (2023)
-
CircRNA DICAR as a novel endogenous regulator for diabetic cardiomyopathy and diabetic pyroptosis of cardiomyocytes
Signal Transduction and Targeted Therapy (2023)
-
CircTrim37 Ameliorates Intracerebral Hemorrhage Outcomes by Modulating Microglial Polarization via the miR-30c-5p/SOCS3 Axis
Molecular Neurobiology (2023)