Abstract
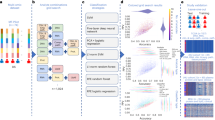
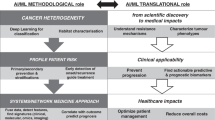
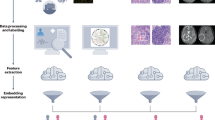
Advances in quantitative biomarker development have accelerated new forms of data-driven insights for patients with cancer. However, most approaches are limited to a single mode of data, leaving integrated approaches across modalities relatively underdeveloped. Multimodal integration of advanced molecular diagnostics, radiological and histological imaging, and codified clinical data presents opportunities to advance precision oncology beyond genomics and standard molecular techniques. However, most medical datasets are still too sparse to be useful for the training of modern machine learning techniques, and significant challenges remain before this is remedied. Combined efforts of data engineering, computational methods for analysis of heterogeneous data and instantiation of synergistic data models in biomedical research are required for success. In this Perspective, we offer our opinions on synthesizing complementary modalities of data with emerging multimodal artificial intelligence methods. Advancing along this direction will result in a reimagined class of multimodal biomarkers to propel the field of precision oncology in the coming decade.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
AACR Project GENIE Consortium. AACR Project GENIE: powering precision medicine through an international consortium. Cancer Discov. 7, 818–831 (2017).
Vasan, N. et al. Double PIK3CA mutations in cis increase oncogenicity and sensitivity to PI3Kα inhibitors. Science 366, 714–723 (2019).
Razavi, P. et al. The genomic landscape of endocrine-resistant advanced breast cancers. Cancer Cell 34, 427–438.e6 (2018).
Jonsson, P. et al. Genomic correlates of disease progression and treatment response in prospectively characterized gliomas. Clin. Cancer Res. 25, 5537–5547 (2019).
Soumerai, T. E. et al. Clinical utility of prospective molecular characterization in advanced endometrial cancer. Clin. Cancer Res. 24, 5939–5947 (2018).
Cui, M. & Zhang, D. Y. Artificial intelligence and computational pathology. Lab. Invest. 101, 412–422 (2021).
Shen, S. Y. et al. Sensitive tumour detection and classification using plasma cell-free DNA methylomes. Nature 563, 579–583 (2018).
Cristiano, S. et al. Genome-wide cell-free DNA fragmentation in patients with cancer. Nature 570, 385–389 (2019).
Klupczynska, A. et al. Study of early stage non-small-cell lung cancer using Orbitrap-based global serum metabolomics. J. Cancer Res. Clin. Oncol. 143, 649–659 (2017).
Helland, T. et al. Serum concentrations of active tamoxifen metabolites predict long-term survival in adjuvantly treated breast cancer patients. Breast Cancer Res. 19, 125 (2017).
Luo, P. et al. A large-scale, multicenter serum metabolite biomarker identification study for the early detection of hepatocellular carcinoma. Hepatology 67, 662–675 (2018).
Medina-Martínez, J. S. et al. Isabl Platform, a digital biobank for processing multimodal patient data. BMC Bioinforma. 21, 549 (2020).
Bhinder, B., Gilvary, C., Madhukar, N. S. & Elemento, O. Artificial intelligence in cancer research and precision medicine. Cancer Discov. 11, 900–915 (2021).
Hosny, A., Parmar, C., Quackenbush, J., Schwartz, L. H. & Aerts, H. J. W. L. Artificial intelligence in radiology. Nat. Rev. Cancer 18, 500–510 (2018).
Bera, K., Schalper, K. A., Rimm, D. L., Velcheti, V. & Madabhushi, A. Artificial intelligence in digital pathology — new tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 16, 703–715 (2019).
Gutman, D. A. et al. MR imaging predictors of molecular profile and survival: multi-institutional study of the TCGA glioblastoma data set. Radiology 267, 560–569 (2013).
Zwanenburg, A. et al. The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping. Radiology 295, 328–338 (2020).
Sun, R. et al. A radiomics approach to assess tumour-infiltrating CD8 cells and response to anti-PD-1 or anti-PD-L1 immunotherapy: an imaging biomarker, retrospective multicohort study. Lancet Oncol. 19, 1180–1191 (2018).
Rizzo, S. et al. Radiomics of high-grade serous ovarian cancer: association between quantitative CT features, residual tumour and disease progression within 12 months. Eur. Radiol. 28, 4849–4859 (2018).
Pisapia, J. M. et al. Predicting pediatric optic pathway glioma progression using advanced magnetic resonance image analysis and machine learning. Neurooncol. Adv. 2, vdaa090 (2020).
Chang, K. et al. Residual convolutional neural network for the determination of IDH status in low- and high-grade gliomas from MR imaging. Clin. Cancer Res. 24, 1073–1081 (2018).
Li, Z., Wang, Y., Yu, J., Guo, Y. & Cao, W. Deep learning based radiomics (DLR) and its usage in noninvasive IDH1 prediction for low grade glioma. Sci. Reports 7, 5467 (2017).
Lu, C.-F. et al. Machine learning-based radiomics for molecular subtyping of gliomas. Clin. Cancer Res. 24, 4429–4436 (2018).
Wang, S. et al. Predicting EGFR mutation status in lung adenocarcinoma on computed tomography image using deep learning. Eur. Respir. J. 53, 1800986 (2019).
Khosravi, P., Lysandrou, M., Eljalby, M. & Li, Q. A deep learning approach to diagnostic classification of prostate cancer using pathology–radiology fusion. J. Magn. Reson. 54, 462–471 (2021).
Hosny, A. et al. Deep learning for lung cancer prognostication: a retrospective multi-cohort radiomics study. PLoS Med. 15, e1002711 (2018).
Rajpurkar, P. et al. AppendiXNet: deep learning for diagnosis of appendicitis from a small dataset of CT exams using video pretraining. Sci. Rep. 10, 3958 (2020).
Khosravi, P., Kazemi, E., Imielinski, M., Elemento, O. & Hajirasouliha, I. Deep convolutional neural networks enable discrimination of heterogeneous digital pathology images. EBioMedicine 27, 317–328 (2018).
Coudray, N. et al. Classification and mutation prediction from non–small cell lung cancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
Fu, Y. et al. Pan-cancer computational histopathology reveals mutations, tumor composition and prognosis. Nat. Cancer 1, 800–810 (2020).
Kather, J. N. et al. Pan-cancer image-based detection of clinically actionable genetic alterations. Nat. Cancer 1, 789–799 (2020).
Ding, K. et al. Feature-enhanced graph networks for genetic mutational prediction using histopathological images in colon cancer. in Medical Image Computing and Computer Assisted Intervention 294–304 (Springer, 2020).
Rutledge, W. C. et al. Tumor-infiltrating lymphocytes in glioblastoma are associated with specific genomic alterations and related to transcriptional class. Clin. Cancer Res. 19, 4951–4960 (2013).
Kather, J. N. et al. Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat. Med. 25, 1054–1056 (2019).
Echle, A. et al. Clinical-grade detection of microsatellite instability in colorectal tumors by deep learning. Gastroenterology 159, 1406–1416.e11 (2020).
Saltz, J. et al. Spatial organization and molecular correlation of tumor-infiltrating lymphocytes using deep learning on pathology images. Cell Rep. 23, 181–193.e7 (2018).
Diao, J. A. et al. Human-interpretable image features derived from densely mapped cancer pathology slides predict diverse molecular phenotypes. Nat. Commun. 12, 1613 (2021).
Corredor, G. et al. Spatial architecture and arrangement of tumor-infiltrating lymphocytes for predicting likelihood of recurrence in early-stage non-small cell lung cancer. Clin. Cancer Res. 25, 1526–1534 (2019).
AbdulJabbar, K. et al. Geospatial immune variability illuminates differential evolution of lung adenocarcinoma. Nat. Med. 26, 1054–1062 (2020).
Kong, J. et al. Machine-based morphologic analysis of glioblastoma using whole-slide pathology images uncovers clinically relevant molecular correlates. PLoS ONE 8, e81049 (2013).
Rao, A., Barkley, D., França, G. S. & Yanai, I. Exploring tissue architecture using spatial transcriptomics. Nature 596, 211–220 (2021).
Lewis, S. M. et al. Spatial omics and multiplexed imaging to explore cancer biology. Nat. Methods https://doi.org/10.1038/s41592-021-01203-6 (2021).
Flaherty, K. T. et al. Improved survival with MEK inhibition in BRAF-mutated melanoma. N. Engl. J. Med. 367, 107–114 (2012).
Maemondo, M. et al. Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N. Engl. J. Med. 362, 2380–2388 (2010).
Slamon, D. J. et al. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N. Engl. J. Med. 344, 783–792 (2001).
DiNardo, C. D. et al. Durable remissions with ivosidenib in IDH1-mutated relapsed or refractory AML. N. Engl. J. Med. 378, 2386–2398 (2018).
Mirza, M. R. et al. Niraparib maintenance therapy in platinum-sensitive, recurrent ovarian cancer. N. Engl. J. Med. 375, 2154–2164 (2016).
de Bono, J. et al. Olaparib for metastatic castration-resistant prostate cancer. N. Engl. J. Med. 382, 2091–2102 (2020).
Drilon, A. et al. Efficacy of larotrectinib in TRK fusion-positive cancers in adults and children. N. Engl. J. Med. 378, 731–739 (2018).
Canon, J. et al. The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity. Nature 575, 217–223 (2019).
Hallin, J. et al. The KRASG12C inhibitor MRTX849 provides insight toward therapeutic susceptibility of KRAS-mutant cancers in mouse models and patients. Cancer Discov. 10, 54–71 (2020).
André, F. et al. Alpelisib for PIK3CA-mutated, hormone receptor-positive advanced breast cancer. N. Engl. J. Med. 380, 1929–1940 (2019).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Priestley, P. et al. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature 575, 210–216 (2019).
Vöhringer, H., Van Hoeck, A., Cuppen, E. & Gerstung, M. Learning mutational signatures and their multidimensional genomic properties with TensorSignatures. Nat. Commun. 12, 3628 (2021).
Macintyre, G. et al. Copy number signatures and mutational processes in ovarian carcinoma. Nat. Genet. 50, 1262–1270 (2018).
Funnell, T. et al. Integrated structural variation and point mutation signatures in cancer genomes using correlated topic models. PLoS Comput. Biol. https://doi.org/10.1371/journal.pcbi.1006799 (2019).
Liu, Y. et al. High-spatial-resolution multi-omics sequencing via deterministic barcoding in tissue. Cell 183, 1665–1681.e18 (2020).
Maniatis, S. et al. Spatiotemporal dynamics of molecular pathology in amyotrophic lateral sclerosis. Science 364, 89–93 (2019).
Payne, A. C. et al. In situ genome sequencing resolves DNA sequence and structure in intact biological samples. Science 371, eaay3446 (2021).
Wang, W., Tran, D. & Feiszli, M. What makes training multi-modal classification networks hard? in Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition 12695–12705 (IEEE, 2020).
Raghu, M., Zhang, C., Kleinberg, J. & Bengio, S. Transfusion: understanding transfer learning for medical imaging. Preprint at arXiv https://arxiv.org/abs/1902.07208 (2019).
Zhang, L. et al. Deep learning-based multi-omics data integration reveals two prognostic subtypes in high-risk neuroblastoma. Front. Genet. 9, 477 (2018).
Chaudhary, K., Poirion, O. B., Lu, L. & Garmire, L. X. Deep learning-based multi-omics integration robustly predicts survival in liver cancer. Clin. Cancer Res. 24, 1248–1259 (2018).
Ramazzotti, D., Lal, A., Wang, B., Batzoglou, S. & Sidow, A. Multi-omic tumor data reveal diversity of molecular mechanisms that correlate with survival. Nat. Commun. 9, 4453 (2018).
Poirion, O. B., Chaudhary, K. & Garmire, L. X. Deep Learning data integration for better risk stratification models of bladder cancer. AMIA Jt. Summits Transl. Sci. Proc. 2017, 197–206 (2018).
Žitnik, M. & Zupan, B. Survival regression by data fusion. Syst. Biomed. 2, 47–53 (2014).
The Cancer Genome Atlas Research Network. Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N. Engl. J. Med. 372, 2481–2498 (2015).
Cantini, L. et al. Benchmarking joint multi-omics dimensionality reduction approaches for the study of cancer. Nat. Commun. 12, 124 (2021).
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. 20, 257–272 (2019).
Hasin, Y., Seldin, M. & Lusis, A. Multi-omics approaches to disease. Genome Biol. 18, 83 (2017).
Sun, D., Wang, M. & Li, A. A multimodal deep neural network for human breast cancer prognosis prediction by integrating multi-dimensional data. IEEE/ACM Trans. Comput. Biol. Bioinform. 3, 841–850 (2018).
Huang, Z. et al. SALMON: survival analysis learning with multi-omics neural networks on breast cancer. Front. Genet. 10, 166 (2019).
Lee, B. et al. DeepBTS: prediction of recurrence-free survival of non-small cell lung cancer using a time-binned deep neural network. Sci. Rep. 10, 1952 (2020).
Choi, E., Bahadori, M. T., Schuetz, A., Stewart, W. F. & Sun, J. Doctor AI: predicting clinical events via recurrent neural networks. in Proceedings of the 1st Machine Learning for Healthcare Conference. Proceedings of Machine Learning Research 56, 301–318 (PLMR, 2016).
Tomašev, N. et al. A clinically applicable approach to continuous prediction of future acute kidney injury. Nature 572, 116–119 (2019).
Yang, J. et al. MIA-prognosis: a deep learning framework to predict therapy response. in Medical Image Computing and Computer Assisted Intervention 211–220 (Springer, 2020).
Srivastava, R. K., Greff, K. & Schmidhuber, J. Highway networks. in Advances in Neural Information Processing Systems 28 https://arxiv.org/abs/1505.00387 (NIPS, 2015).
Cheerla, A. & Gevaert, O. Deep learning with multimodal representation for pancancer prognosis prediction. Bioinformatics 35, i446–i454 (2019).
Gevaert, O. et al. Imaging-AMARETTO: An imaging genomics software tool to interrogate multiomics networks for relevance to radiography and histopathology imaging biomarkers of clinical outcomes. JCO Clin. Cancer Inf. 4, 421–435 (2020).
Zhu, X. et al. Imaging-genetic data mapping for clinical outcome prediction via supervised conditional Gaussian graphical model. in 2016 IEEE International Conference on Bioinformatics and Biomedicine https://doi.org/10.1109/BIBM.2016.7822559 (IEEE, 2016).
Popovici, V. et al. Joint analysis of histopathology image features and gene expression in breast cancer. BMC Bioinformatics 17, 209 (2016).
Mobadersany, P. et al. Predicting cancer outcomes from histology and genomics using convolutional networks. Proc. Natl Acad. Sci. USA 115, E2970–E2979 (2018).
Chen, R. J. et al. Pathomic fusion: an integrated framework for fusing histopathology and genomic features for cancer diagnosis and prognosis. IEEE Trans. Med. Imaging https://doi.org/10.1109/TMI.2020.3021387 (2020).
Zadeh, A., Chen, M., Poria, S., Cambria, E. & Morency, L.-P. Tensor fusion network for multimodal sentiment analysis. in Proceedings of the 2017 Conference on Empirical Methods in Natural Language Processing 1103–1114 (Biblio, 2017).
Yuan, Y., Giger, M. L., Li, H., Bhooshan, N. & Sennett, C. A. Multimodality computer-aided breast cancer diagnosis with FFDM and DCE-MRI. Acad. Radiol. 17, 1158–1167 (2010).
Chan, H.-W., Weng, Y.-T. & Huang, T.-Y. Automatic classification of brain tumor types with the MRI scans and histopathology images. in Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries 353–359 (Springer, 2017).
Rathore, S. et al. Radiomic MRI signature reveals three distinct subtypes of glioblastoma with different clinical and molecular characteristics, offering prognostic value beyond IDH1. Sci. Rep. 8, 5087 (2018).
Donini, M. et al. Combining heterogeneous data sources for neuroimaging based diagnosis: re-weighting and selecting what is important. Neuroimage 195, 215–231 (2019).
Gonen, M. & Alpaydin, E. Multiple kernel learning algorithms. J. Mach. Learn. Res. 12, 2211–2268 (2011).
Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: images are more than pictures, they are data. Radiology 278, 563–577 (2016).
Duanmu, H. et al. Prediction of pathological complete response to neoadjuvant chemotherapy in breast cancer using deep learning with integrative imaging, molecular and demographic data. in Medical Image Computing and Computer Assisted Intervention 242–252 (Springer, 2020).
Bhattacharya, I. et al. CorrSigNet: learning correlated prostate cancer SIGnatures from radiology and pathology images for improved computer aided diagnosis. in Medical Image Computing and Computer Assisted Intervention — MICCAI 2020 315–325 (Springer, 2020).
Deng, J. et al. ImageNet: a large-scale hierarchical image database. in IEEE Conference on Computer Vision and Pattern Recognition https://doi.org/10.1109/CVPR.2009.5206848 (IEEE, 2009).
Kay, W. et al. The kinetics human action video dataset. Preprint at arXiv http://arxiv.org/abs/1705.06950 (2017).
Ardila, D. et al. End-to-end lung cancer screening with three-dimensional deep learning on low-dose chest computed tomography. Nat. Med. 25, 954–961 (2019).
Zhou, Z.-H. A brief introduction to weakly supervised learning. Natl Sci. Rev. 5, 44–53 (2018).
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25, 1301–1309 (2019).
Suphavilai, C., Bertrand, D. & Nagarajan, N. Predicting cancer drug response using a recommender system. Bioinformatics 34, 3907–3914 (2018).
Joachims, T., Swaminathan, A. & de Rijke, M. Deep learning with logged bandit feedback. in Proceedings of the International Conference on Learning Representations (ICLR) (ICLR, 2018).
Lee, J. S. et al. Synthetic lethality-mediated precision oncology via the tumor transcriptome. Cell 184, 2487–2502.e13 (2021).
Gundersen, G., Dumitrascu, B., Ash, J. T. & Engelhardt, B. E. End-to-end training of deep probabilistic CCA for joint modeling of paired biomedical observations. in Proceedings of the 35th Uncertainty in Artificial Intelligence Conference 945–955 (PMLR, 2020).
Li, Y. et al. Inferring multimodal latent topics from electronic health records. Nat. Commun. 11, 2536 (2020).
Choplin, R. H., Boehme, J. M. 2nd & Maynard, C. D. Picture archiving and communication systems: an overview. Radiographics 12, 127–129 (1992).
Puchalski, R. B. et al. An anatomic transcriptional atlas of human glioblastoma. Science 360, 660–663 (2018).
Weigelt, B. et al. Radiogenomics analysis of intratumor heterogeneity in a patient with high-grade serous ovarian cancer. JCO Precis. Oncol. 3, 1–9 (2019).
Jiménez-Sánchez, A. et al. Unraveling tumor-immune heterogeneity in advanced ovarian cancer uncovers immunogenic effect of chemotherapy. Nat. Genet. 52, 582–593 (2020).
Hersh, W. R. et al. Caveats for the use of operational electronic health record data in comparative effectiveness research. Med. Care 51, S30–S37 (2013).
Allison, J. J. et al. The art and science of chart review. Jt. Comm. J. Qual. Improv. 26, 115–136 (2000).
Vassar, M. & Holzmann, M. The retrospective chart review: important methodological considerations. J. Educ. Eval. Health Prof. 10, 12 (2013).
Hripcsak, G. et al. Observational health data sciences and informatics (OHDSI): opportunities for observational researchers. Stud. Health Technol. Inform. 216, 574–578 (2015).
Stein, B. & Morrison, A. The enterprise data lake: better integration and deeper analytics. PwC Technol. Forecast Rethinking Integr. 1, 18 (2014).
Armbrust, M. et al. Delta lake: high-performance ACID table storage over cloud object stores. in Proceedings of the VLDB Endowment Vol. 13 3411–3424 (ACM, 2020).
Zagan, E. & Danubianu, M. Cloud DATA LAKE: the new trend of data storage. in 2021 3rd International Congress on Human-Computer Interaction, Optimization and Robotic Applications (HORA) 1–4 (IEEE, 2021).
Rieke, N. et al. The future of digital health with federated learning. NPJ Digital Med. 3, 119 (2020).
Andreux, M., Manoel, A., Menuet, R., Saillard, C. & Simpson, C. Federated survival analysis with discrete-time Cox models. in FL-ICML 2020: International Workshop on Federated Learning for User Privacy and Data Confidentiality in Conjunction with ICML 2020 https://arxiv.org/abs/2006.08997 (ICML 2020).
Lin, J.-H. & Haug, P. J. Exploiting missing clinical data in Bayesian network modeling for predicting medical problems. J. Biomed. Inform. 41, 1–14 (2008).
Khan, A., Atzori, M., Otálora, S., Andrearczyk, V. & Müller, H. Generalizing convolution neural networks on stain color heterogeneous data for computational pathology. in Medical Imaging 2020: Digital Pathology https://doi.org/10.1117/12.2549718 (International Society for Optics and Photonics, 2020).
Glatz-Krieger, K., Spornitz, U., Spatz, A., Mihatsch, M. J. & Glatz, D. Factors to keep in mind when introducing virtual microscopy. Virchows Arch. 448, 248–255 (2006).
Janowczyk, A., Basavanhally, A. & Madabhushi, A. Stain Normalization using Sparse AutoEncoders (StaNoSA): application to digital pathology. Comput. Med. Imaging Graph. 57, 50–61 (2017).
Lacroix, M. et al. Correction for magnetic field inhomogeneities and normalization of voxel values are needed to better reveal the potential of MR radiomic features in lung cancer. Front. Oncol. 10, 43 (2020).
Macenko, M. et al. A method for normalizing histology slides for quantitative analysis. in 2009 IEEE International Symposium on Biomedical Imaging: From Nano to Macro https://doi.org/10.1109/ISBI.2009.5193250 (IEEE, 2009).
Srinidhi, C. L., Ciga, O. & Martel, A. L. Deep neural network models for computational histopathology: a survey. Med. Image Anal. 67, 101813 (2021).
Hu, Z., Tang, A., Singh, J., Bhattacharya, S. & Butte, A. J. A robust and interpretable end-to-end deep learning model for cytometry data. Proc. Natl Acad. Sci. USA 117, 21373–21380 (2020).
Kleppe, A. et al. Designing deep learning studies in cancer diagnostics. Nat. Rev. Cancer 21, 199–211 (2021).
Lopez, K., Fodeh, S. J., Allam, A., Brandt, C. A. & Krauthammer, M. Reducing annotation burden through multimodal learning. Front. Big Data 3, 19 (2020).
Gundersen, O. E. & Kjensmo, S. State of the art: reproducibility in artificial intelligence. in Thirty-Second AAAI Conference on Artificial Intelligence (AAAI, 2018).
Courtiol, P. et al. Deep learning-based classification of mesothelioma improves prediction of patient outcome. Nat. Med. 25, 1519–1525 (2019).
McKinney, S. M. et al. Addendum: international evaluation of an AI system for breast cancer screening. Nature 586, E19 (2020).
Haibe-Kains, B. et al. Transparency and reproducibility in artificial intelligence. Nature 586, E14–E16 (2020).
Hosny, A. et al. ModelHub.AI: dissemination platform for deep learning models. Preprint at arXiv http://arxiv.org/abs/1911.13218 (2019).
Beede, E. et al. A human-centered evaluation of a deep learning system deployed in clinics for the detection of diabetic retinopathy. in Proceedings of the 2020 CHI Conference on Human Factors in Computing Systems 1–12 (ACM, 2020).
Cruz Rivera, S. et al. Guidelines for clinical trial protocols for interventions involving artificial intelligence: the SPIRIT-AI extension. Nat. Med. 26, 1351–1363 (2020).
Moher, D. et al. CONSORT 2010 explanation and elaboration: updated guidelines for reporting parallel group randomised trials. BMJ 340, c869 (2010).
Liu, X. et al. Reporting guidelines for clinical trial reports for interventions involving artificial intelligence: the CONSORT-AI extension. Nat. Med. 26, 1364–1374 (2020).
Lauritsen, S. M. et al. Explainable artificial intelligence model to predict acute critical illness from electronic health records. Nat. Commun. 11, 3852 (2020).
Rudin, C. Stop explaining black box machine learning models for high stakes decisions and use interpretable models instead. Nat. Mach. Intell. 1, 206–215 (2019).
Pavel, M. A., Petersen, E. N., Wang, H., Lerner, R. A. & Hansen, S. B. Studies on the mechanism of general anesthesia. Proc. Natl Acad. Sci. USA 117, 13757–13766 (2020).
Wang, F., Kaushal, R. & Khullar, D. Should health care demand interpretable artificial intelligence or accept “black box” medicine? Ann. Intern. Med. 172, 59–60 (2020).
Castro, D. C., Walker, I. & Glocker, B. Causality matters in medical imaging. Nat. Commun. 11, 3673 (2020).
Zhou, B., Khosla, A., Lapedriza, A., Oliva, A. & Torralba, A. Learning deep features for discriminative localization. in Proceedings of the IEEE conference on computer vision and pattern recognition 2921–2929 (IEEE, 2016).
Olah, C. et al. The building blocks of interpretability. Distill https://distill.pub/2018/building-blocks/ (2018).
Graziani, M., Andrearczyk, V. & Müller, H. Visualizing and interpreting feature reuse of pretrained CNNs for histopathology. in Irish Machine Vision and Image Processing Conference (IMVIP, 2019).
Burns, C., Thomason, J. & Tansey, W. Interpreting black box models via hypothesis testing. in FODS ’20: Proceedings of the 2020 ACM-IMS on Foundations of Data Science Conference 47–57 (ACM, 2019).
Donoghue, M. T. A., Schram, A. M., Hyman, D. M. & Taylor, B. S. Discovery through clinical sequencing in oncology. Nat. Cancer 1, 774–783 (2020).
Kehl, K. L. et al. Assessment of deep natural language processing in ascertaining oncologic outcomes from radiology reports. JAMA Oncol. 5, 1421–1429 (2019).
Cosgriff, C. V., Stone, D. J., Weissman, G., Pirracchio, R. & Celi, L. A. The clinical artificial intelligence department: a prerequisite for success. BMJ Health Care Inf. 27, e100183 (2020).
Zadeh, A. et al. Memory fusion network for multi-view sequential learning. in Proceedings of the Thirty-Second AAAI Conference on Artificial Intelligence (AAAI, 2018).
Zadeh, A., Liang, P. P., Poria, S., Cambria, E. & Morency, L.-P. Multimodal language analysis in the wild: CMU-MOSEI dataset and interpretable dynamic fusion graph. in Proceedings of the 56th Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers) 2236–2246 (ACL Anthology, 2018).
Zadeh, A. et al. Multi-attention recurrent network for human communication comprehension. in Thirty-Second AAAI Conference on Artificial Intelligence Vol. 2018 5642–5649 (AAAI, 2018).
Kumar, A., Srinivasan, K., Cheng, W.-H. & Zomaya, A. Y. Hybrid context enriched deep learning model for fine-grained sentiment analysis in textual and visual semiotic modality social data. Inf. Process. Manag. 57, 102141 (2020).
Liang, P. P., Zadeh, A. & Morency, L.-P. Multimodal local-global ranking fusion for emotion recognition. in Proceedings of the 20th ACM International Conference on Multimodal Interaction 472–476 (ACM, 2018).
Liu, Z. et al. Efficient low-rank multimodal fusion with modality-specific factors. in Proceedings of the 56th Annual Meeting of the Association for Computational Linguistics (Long Papers) 2247–2256 (ACL Anthology, 2018).
Marinelli, R. J. et al. The Stanford Tissue Microarray Database. Nucleic Acids Res. 36, D871–D877 (2008).
Clark, K. et al. The Cancer Imaging Archive (TCIA): maintaining and operating a public information repository. J. Digit. Imaging 26, 1045–1057 (2013).
Newitt, D. & Hylton, N. Single site breast DCE-MRI data and segmentations from patients undergoing neoadjuvant chemotherapy. The Cancer Imaging Archive https://doi.org/10.7937/K9/TCIA.2016.QHsyhJKy (2016).
Wolpert, D. H. The lack of a priori distinctions between learning algorithms. Neural Comput. 8, 1341–1390 (1996).
Wolpert, D. H. & Macready, W. G. No free lunch theorems for optimization. IEEE Trans. Evol. Comput. 1, 67–82 (1997).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. in 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) https://doi.org/10.1109/CVPR.2016.90 (IEEE, 2016).
Iandola, F. N. et al. SqueezeNet: AlexNet-level accuracy with 50x fewer parameters and <0.5MB model size. Preprint at arXiv http://arxiv.org/abs/1602.07360 (2016).
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J. & Wojna, Z. Rethinking the inception architecture for computer vision. in 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (IEEE, 2016).
Huang, G., Liu, Z., van der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. in 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) https://arxiv.org/abs/1608.06993 (IEEE, 2016).
Cho, K. et al. Learning phrase representations using RNN encoder–decoder for statistical machine translation. in Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing (EMNLP) (Association for Computational Linguistics, 2014).
Hochreiter, S. & Schmidhuber, J. Long short-term memory. Neural Comput. 9, 1735–1780 (1997).
Vaswani, A. et al. Attention is all you need. in 31st Conference on Neural Information Processing Systems (NIPS, 2017).
Lee, G., Kang, B., Nho, K., Sohn, K.-A. & Kim, D. MildInt: deep learning-based multimodal longitudinal data integration framework. Front. Genet. 10, 617 (2019).
Acknowledgements
The authors thank N. Rusk and W. Tansey for helpful comments on the manuscript. S.P.S. is supported by the Nicholls-Biondi Endowed Chair in Computational Oncology and the Susan G. Komen Scholars programme. K.M.B. is supported by the National Cancer Institute (NCI) of the US National Institutes of Health (NIH) under award number F30CA257414, the Jonathan Grayer Fellowship of Gerstner Sloan Kettering Graduate School of Biomedical Sciences and a Medical Scientist Training Program Grant from the National Institute of General Medical Sciences of the NIH under award number T32GM007739 to the Weill Cornell/Rockefeller/Sloan Kettering Tri-Institutional MD-PhD Program. MSK MIND is generously supported by Cycle for Survival. All authors are supported by NIH NCI Cancer Center Support Grant P30 CA008748.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
S.P.S is a shareholder in and consultant for Canexia Health Inc. K.M.B., P.K., R.V. and J.J.G. declare no competing interests.
Additional information
Peer review information
Nature Reviews Cancer thanks Anant Madabhushi, who co-reviewed with Nathaniel Braman; Benjamin Haibe-Kains; and Aristotelis Tsirigos for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Area under the receiver operating characteristic curve
-
(AUROC). A measurement of the ability of a binary classifier to separate the populations of interest. It describes the increase in the true positive rate relative to the increase in the false positive rate over the range of score thresholds chosen to separate the two classes. The highest value obtainable is 1, and random performance is associated with a value of 0.5.
- Artificial intelligence
-
(AI). A broad field of computer science concerned with developing computational tools to perform tasks historically requiring human-level intelligence.
- Autoencoders
-
Unsupervised neural network architectures trained to represent data in a lower-dimensional space. They are a form of lossy compression (reducing the size of data representations, but with some loss of information) that can be used to uncover latent structure in the data or reduce computational needs before further analysis.
- Bayesian inference
-
A statistical method that refers to the application of Bayes’s theorem in determining the updated probability of a hypothesis given new information. Bayesian inference allows the posterior probability to be calculated given the prior probability of a hypothesis and a likelihood function.
- Biomarkers
-
Measurements that indicate a biological state. Cancer biomarkers can be categorized into diagnostic (disease progression), predictive (treatment response) and prognostic (survival).
- Concordance index
-
(c-index). An index that generalizes the area under the receiver operating characteristic curve (AUROC) to measure the ability of a model to separate censored data. As with the AUROC, the baseline value for a model with arbitrary predictions is 0.5, and the ceiling value for a perfect prediction model is 1.0.
- Convolutional neural networks
-
(CNNs). A form of deep neural network typically used to analyse images. CNNs are named for their use of convolutions, a mathematical operation involving the input data and a smaller matrix known as a kernel. This parameter sharing reduces the number of parameters to be learned and encourages the learning of features which are invariant to image shifts.
- Counterfactual ML
-
A set of techniques for machine learning (ML) based on the paradigm of modelling situations that did not factually occur. These techniques are often deployed for interpretable models or to learn from biased logged data. For example, a counterfactual analysis could involve using a model developed to predict a disease outcome using a set of measurements to predict scenarios where the input measurements are perturbed to study their causal relationship. This paradigm has also been harnessed to learn unbiased recommenders from logged data, such as user purchases on online marketplaces, despite changes in how products are recommended over time and the lack of a controlled experimental setup.
- Cox proportional hazards (CPH) model
-
A regression model used to associate censored temporal outcomes, such as time to survival, and potential predictor variables, such as age or cancer stage. It is the most common method to evaluate prognostic variables in survival analyses of patients with cancer.
- Data lakes
-
Places to store relational and non-relational data from a vast pool of raw data. The structure of the data or schema is not defined when data are captured. Different types of analytics on data such as structured query language (SQL) queries, big data analytics, full text search, real-time analytics and machine learning can be used to uncover insights.
- Data parallelism
-
The approach of performing a computing task in parallel using multiple processors. It focuses on distributing data across various cores and enabling simultaneous subcomputations.
- Deep learning
-
(DL). Comprises a class of machine learning methods based on artificial neural networks, which use multiple non-linear layers to derive progressively higher-order features from data.
- Deep neural network
-
(DNN). A form of deep learning, namely artificial neural networks with more than one hidden layer between the input and output layers.
- Federated learning
-
A training strategy wherein the model to be trained is passed around among institutions instead of data being centrally amalgamated. Each institution then updates the model parameters on the basis of the local dataset. This strategy enables multi-institutional model training without data sharing among institutions.
- Kernel
-
A similarity function often used to transform input data implicitly into a form more suitable for machine learning tasks. For example, a two-dimensional pattern-based kernel could be used to identify the presence of specific shapes in an image, and a one-dimensional Gaussian kernel could be used to impute a smoothed trendline on the basis of noisy data points.
- Layer-wise relevance propagation
-
(LRP). One of the most prominent techniques in explainable machine learning. LRP decomposes the network’s output score into the individual contributions of the input neurons using model parameters (that is, weights) and neuron activations.
- Machine learning
-
(ML). A type of artificial intelligence that aims to discover patterns in data that are not explicitly programmed. ML models typically use a dataset for pattern discovery, known as ‘training’, to make predictions on unseen data, known as ‘inference’.
- Recommender systems
-
Systems that aim to predict items relevant to users by building a model from past behaviour. In precision medicine, recommender systems can be used to predict the preferred treatment for a disease on the basis of multiple patient measurements.
- Recurrent neural networks
-
(RNNs). A form of deep neural network optimized for time series data. An RNN analyses each element of the input sequence in succession and updates its representation of the data on the basis of previous elements.
- Sentiment analysis
-
A field seeking to characterize human emotional states from text, images and sounds by the use of machine learning models.
- Supervised learning
-
A machine learning paradigm that aims to elucidate the relationship between input data variables and predefined classes (‘classification’) or continuous labels (‘regression’) of interest. By contrast, unsupervised learning aims to identify patterns in a dataset without the use of such labels or classes.
- Voxel
-
The volume element defined by the x, y and z coordinates in three-dimensional space used in medical imaging modalities. Its dimensions are given by the pixel, together with the thickness of the slice.
- Weight decay
-
A regularization strategy to improve the generalizability of models whereby high estimated values of model parameters are penalized despite marginal increases in accuracy on the training set.
Rights and permissions
About this article
Cite this article
Boehm, K.M., Khosravi, P., Vanguri, R. et al. Harnessing multimodal data integration to advance precision oncology. Nat Rev Cancer 22, 114–126 (2022). https://doi.org/10.1038/s41568-021-00408-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41568-021-00408-3
This article is cited by
-
Pathogenomics for accurate diagnosis, treatment, prognosis of oncology: a cutting edge overview
Journal of Translational Medicine (2024)
-
Transfer learning–based PET/CT three-dimensional convolutional neural network fusion of image and clinical information for prediction of EGFR mutation in lung adenocarcinoma
BMC Medical Imaging (2024)
-
Standardizing digital biobanks: integrating imaging, genomic, and clinical data for precision medicine
Journal of Translational Medicine (2024)
-
Guardrails for the use of generalist AI in cancer care
Nature Reviews Cancer (2024)
-
Artificial intelligence in neuro-oncology: advances and challenges in brain tumor diagnosis, prognosis, and precision treatment
npj Precision Oncology (2024)