Abstract
Advances in our understanding of the metabolism and molecular functions of polyamines and their alterations in cancer have led to resurgence in the interest of targeting polyamine metabolism as an anticancer strategy. Increasing knowledge of the interplay between polyamine metabolism and other cancer-driving pathways, including the PTEN–PI3K–mTOR complex 1 (mTORC1), WNT signalling and RAS pathways, suggests potential combination therapies that will have considerable clinical promise. Additionally, an expanding number of promising clinical trials with agents targeting polyamines for both therapy and prevention are ongoing. New insights into molecular mechanisms linking dysregulated polyamine catabolism and carcinogenesis suggest additional strategies that can be used for cancer prevention in at-risk individuals. In addition, polyamine blocking therapy, a strategy that combines the inhibition of polyamine biosynthesis with the simultaneous blockade of polyamine transport, can be more effective than therapies based on polyamine depletion alone and may involve an antitumour immune response. These findings open up new avenues of research into exploiting aberrant polyamine metabolism for anticancer therapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



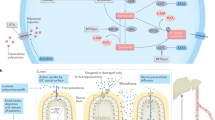
Figure adapted with permission from ref.17, Portland Press.

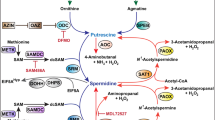
Figure adapted from ref.46, Springer Nature Limited.

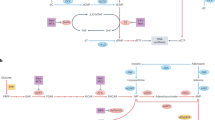
Figure adapted with permission from ref.151, Elsevier.
Similar content being viewed by others
References
Pegg, A. E. & Casero, R. A. Jr. Current status of the polyamine research field. Methods Mol. Biol. 720, 3–35 (2011).
Terui, Y. et al. Polyamines protect nucleic acids against depurination. Int. J. Biochem. Cell Biol. 99, 147–153 (2018).
Kurata, H. T., Akrouh, A., Li, J. B., Marton, L. J. & Nichols, C. G. Scanning the topography of polyamine blocker binding in an inwardly rectifying potassium channel. J. Biol. Chem. 288, 6591–6601 (2013).
Rao, J. N. et al. Polyamines regulate intestinal epithelial restitution through TRPC1-mediated Ca(2)+ signaling by differentially modulating STIM1 and STIM2. Am. J. Physiol. Cell Physiol. 303, C308–C317 (2012).
Murray-Stewart, T. R., Woster, P. M. & Casero, R. A. Jr. Targeting polyamine metabolism for cancer therapy and prevention. Biochem. J. 473, 2937–2953 (2016).
Mandal, A., Mandal, S. & Park, M. H. Genome-wide analyses and functional classification of proline repeat-rich proteins: potential role of eIF5A in eukaryotic evolution. PLOS ONE 9, e111800 (2014).
Park, M. H., Nishimura, K., Zanelli, C. F. & Valentini, S. R. Functional significance of eIF5A and its hypusine modification in eukaryotes. Amino Acids 38, 491–500 (2010).
Shin, B. S. et al. Amino acid substrates impose polyamine, eIF5A, or hypusine requirement for peptide synthesis. Nucleic Acids Res. 45, 8392–8402 (2017).
Nakanishi, S. & Cleveland, J. L. Targeting the polyamine-hypusine circuit for the prevention and treatment of cancer. Amino Acids 48, 2353–2362 (2016).
Huter, P. et al. Structural basis for polyproline-mediated ribosome stalling and rescue by the translation elongation factor EF-P. Mol. Cell 68, 515–527 (2017).
Pelechano, V. & Alepuz, P. eIF5A facilitates translation termination globally and promotes the elongation of many non polyproline-specific tripeptide sequences. Nucleic Acids Res. 45, 7326–7338 (2017).
Hoque, M. et al. Regulation of gene expression by translation factor eIF5A: hypusine-modified eIF5A enhances nonsense-mediated mRNA decay in human cells. Translation 5, e1366294 (2017).
Mathews, M. B. & Hershey, J. W. The translation factor eIF5A and human cancer. Biochim. Biophys. Acta 1849, 836–844 (2015).
Ikeguchi, Y., Bewley, M. C. & Pegg, A. E. Aminopropyltransferases: function, structure and genetics. J. Biochem. 139, 1–9 (2006).
Pegg, A. E. Regulation of ornithine decarboxylase. J. Biol. Chem. 281, 14529–14532 (2006).
Pegg, A. E. S-adenosylmethionine decarboxylase. Essays Biochem. 46, 25–45 (2009).
Kahana, C. Protein degradation, the main hub in the regulation of cellular polyamines. Biochem. J. 473, 4551–4558 (2016).
Wu, H. Y. et al. Structural basis of antizyme-mediated regulation of polyamine homeostasis. Proc. Natl Acad. Sci. USA 112, 11229–11234 (2015).
Kahana, C. Antizyme and antizyme inhibitor, a regulatory tango. Cell. Mol. Life Sci. 66, 2479–2488 (2009).
Yordanova, M. M. et al. AMD1 mRNA employs ribosome stalling as a mechanism for molecular memory formation. Nature 553, 356–360 (2018).
Casero, R. A. & Pegg, A. E. Polyamine catabolism and disease. Biochem. J. 421, 323–338 (2009).
Pegg, A. E. Spermidine/spermine-N(1)-acetyltransferase: a key metabolic regulator. Am. J. Physiol. Endocrinol. Metab. 294, E995–E1010 (2008).
Pegg, A. E. Toxicity of polyamines and their metabolic products. Chem. Res. Toxicol. 26, 1782–1800 (2013).
Masuko, T. et al. N(1)-Nonyl-1,4-diaminobutane ameliorates brain infarction size in photochemically induced thrombosis model mice. Neurosci. Lett. 13, 118–122 (2018).
Albers, E. Metabolic characteristics and importance of the universal methionine salvage pathway recycling methionine from 5′-methylthioadenosine. IUBMB Life 61, 1132–1142 (2009).
Bertino, J. R., Waud, W. R., Parker, W. B. & Lubin, M. Targeting tumors that lack methylthioadenosine phosphorylase (MTAP) activity: current strategies. Cancer Biol. Ther. 11, 627–632 (2011).
Poulin, R., Casero, R. A. & Soulet, D. Recent advances in the molecular biology of metazoan polyamine transport. Amino Acids 42, 711–723 (2012).
Soulet, D., Gagnon, B., Rivest, S., Audette, M. & Poulin, R. A fluorescent probe of polyamine transport accumulates into intracellular acidic vesicles via a two-step mechanism. J. Biol. Chem. 279, 49355–49366 (2004).
Belting, M. et al. Glypican-1 is a vehicle for polyamine uptake in mammalian cells: a pivital role for nitrosothiol-derived nitric oxide. J. Biol. Chem. 278, 47181–47189 (2003).
Uemura, T., Stringer, D. E., Blohm-Mangone, K. A. & Gerner, E. W. Polyamine transport is mediated by both endocytic and solute carrier transport mechanisms in the gastrointestinal tract. Am. J. Physiol. Gastrointest. Liver Physiol. 299, G517–G522 (2010).
Kahana, C. Regulation of cellular polyamine levels and cellular proliferation by antizyme and antizyme inhibitor. Essays Biochem. 46, 47–61 (2009).
Bello-Fernandez, C., Packham, G. & Cleveland, J. L. The ornithine decarboxylase gene is a transcriptional target of c-Myc. Proc. Natl Acad. Sci. USA 90, 7804–7808 (1993).
Ozfiliz, P. et al. Bag-1 promotes cell survival through c-Myc-mediated ODC upregulation that is not preferred under apoptotic stimuli in MCF-7 cells. Cell Biochem. Funct. 33, 293–307 (2015).
Koomoa, D. L. et al. DFMO/eflornithine inhibits migration and invasion downstream of MYCN and involves p27Kip1 activity in neuroblastoma. Int. J. Oncol. 42, 1219–1228 (2013).
Funakoshi-Tago, M., Sumi, K., Kasahara, T. & Tago, K. Critical roles of Myc-ODC axis in the cellular transformation induced by myeloproliferative neoplasm-associated JAK2 V617F mutant. PLOS ONE 8, e52844 (2013).
Hogarty, M. D. et al. ODC1 is a critical determinant of MYCN oncogenesis and a therapeutic target in neuroblastoma. Cancer Res. 68, 9735–9745 (2008).
Rimpi, S. & Nilsson, J. A. Metabolic enzymes regulated by the Myc oncogene are possible targets for chemotherapy or chemoprevention. Biochem. Soc. Trans. 35, 305–310 (2007).
Roy, U. K., Rial, N. S., Kachel, K. L. & Gerner, E. W. Activated K-RAS increases polyamine uptake in human colon cancer cells through modulation of caveolar endocytosis. Mol. Carcinog. 47, 538–553 (2008).
Ignatenko, N. A., Babbar, N., Mehta, D., Casero, R. A. Jr & Gerner, E. W. Suppression of polyamine catabolism by activated Ki-ras in human colon cancer cells. Mol. Carcinog. 39, 91–102 (2004).
Davies, H. et al. Mutations of the BRAF gene in human cancer. Nature 417, 949–954 (2002).
Solit, D. B. & Rosen, N. Resistance to BRAF inhibition in melanomas. N. Engl. J. Med. 364, 772–774 (2011).
Peters, M. C., Minton, A., Phanstiel Iv, O. & Gilmour, S. K. A. Novel polyamine-targeted therapy for BRAF mutant melanoma tumors. Med. Sci. 6, https://doi.org/10.3390/medsci6010003 (2018).
Kucharzewska, P., Welch, J. E., Svensson, K. J. & Belting, M. The polyamines regulate endothelial cell survival during hypoxic stress through PI3K/AKT and MCL-1. Biochem. Biophys. Res. Commun. 380, 413–418 (2009).
Wang, C. et al. Spermidine/spermine N1-acetyltransferase regulates cell growth and metastasis via AKT/beta-catenin signaling pathways in hepatocellular and colorectal carcinoma cells. Oncotarget 8, 1092–1109 (2017).
Zabala-Letona, A. et al. Corrigendum: mTORC1-dependent AMD1 regulation sustains polyamine metabolism in prostate cancer. Nature 554, 554 (2018).
Zabala-Letona, A. et al. mTORC1-dependent AMD1 regulation sustains polyamine metabolism in prostate cancer. Nature 547, 109–113 (2017). This is an important new study describing the dysregulation of AdoMetDC in prostate cancer and suggesting new targets for therapeutic intervention.
Gomes, A. P., Schild, T. & Blenis, J. Adding polyamine metabolism to the mTORC1 toolkit in cell growth and cancer. Dev. Cell 42, 112–114 (2017).
Tomasi, M. L. et al. Polyamine and methionine adenosyltransferase 2A crosstalk in human colon and liver cancer. Exp. Cell Res. 319, 1902–1911 (2013).
D’Amico, D. et al. Non-canonical hedgehog/AMPK-mediated control of polyamine metabolism supports neuronal and medulloblastoma cell growth. Dev. Cell 35, 21–35 (2015).
Sandsmark, E. et al. A novel non-canonical Wnt signature for prostate cancer aggressiveness. Oncotarget 8, 9572–9586 (2017).
Ou, Y., Wang, S. J., Li, D., Chu, B. & Gu, W. Activation of SAT1 engages polyamine metabolism with p53-mediated ferroptotic responses. Proc. Natl Acad. Sci. USA 113, E6806–E6812 (2016).
Cao, J. Y. & Dixon, S. J. Mechanisms of ferroptosis. Cell. Mol. Life Sci. 73, 2195–2209 (2016).
Tormey, D. C. et al. Biological markers in breast carcinoma. I. Incidence of abnormalities of CEA, HCG, three polyamines, and three minor nucleosides. Cancer 35, 1095–1100 (1975).
Miolo, G. et al. Pharmacometabolomics study identifies circulating spermidine and tryptophan as potential biomarkers associated with the complete pathological response to trastuzumab-paclitaxel neoadjuvant therapy in HER-2 positive breast cancer. Oncotarget 7, 39809–39822 (2016).
Liu, R. et al. Plasma N-acetylputrescine, cadaverine and 1,3-diaminopropane: potential biomarkers of lung cancer used to evaluate the efficacy of anticancer drugs. Oncotarget 8, 88575–88585 (2017).
Xu, H. et al. Polyamine metabolites profiling for characterization of lung and liver cancer using an LC-tandem MS method with multiple statistical data mining strategies: discovering potential cancer biomarkers in human plasma and urine. Molecules 21, E1040 (2016).
Kato, M. et al. Prognostic significance of urine N1, N12-diacetylspermine in patients with non-small cell lung cancer. Anticancer Res. 34, 3053–3059 (2014).
Takahashi, Y. et al. Urinary N1, N12-diacetylspermine is a non-invasive marker for the diagnosis and prognosis of non-small-cell lung cancer. Br. J. Cancer 113, 1493–1501 (2015).
Niemi, R. J. et al. Urinary polyamines as biomarkers for ovarian cancer. Int. J. Gynecol. Cancer 27, 1360–1366 (2017).
Dejea, C. M. & Sears, C. L. Do biofilms confer a pro-carcinogenic state? Gut Microbes 7, 54–57 (2016).
Johnson, C. H. et al. Metabolism links bacterial biofilms and colon carcinogenesis. Cell Metab. 21, 891–897 (2015).
Nakajima, T. et al. Urinary polyamine biomarker panels with machine-learning differentiated colorectal cancers, benign disease, and healthy controls. Int. J. Mol. Sci. 19, E756 (2018). This paper describes a novel approach to improve the use of polyamine biomarker panels in determining disease states in detection of colon cancers.
Liu, R., Lin, X., Li, Z., Li, Q. & Bi, K. Quantitative metabolomics for investigating the value of polyamines in the early diagnosis and therapy of colorectal cancer. Oncotarget 9, 4583–4592 (2018).
Asai, Y. et al. Elevated polyamines in saliva of pancreatic cancer. Cancers 10, E43 (2018).
Fitzgerald, B. L. et al. Elucidating the structure of N(1)-acetylisoputreanine: a novel polyamine catabolite in human urine. ACS Omega 2, 3921–3930 (2017).
Giskeodegard, G. F. et al. Spermine and citrate as metabolic biomarkers for assessing prostate cancer aggressiveness. PLOS ONE 8, e62375 (2013).
Millward, M. J. et al. Multi-centre phase II trial of the polyamine synthesis inhibitor SAM486A (CGP48664) in patients with metastatic melanoma. Invest. New Drugs 23, 253–256 (2005).
Kadariya, Y. et al. Mice heterozygous for germ-line mutations in methylthioadenosine phosphorylase (MTAP) die prematurely of T cell lymphoma. Cancer Res. 69, 5961–5969 (2009).
Evageliou, N. F. et al. Polyamine antagonist therapies inhibit neuroblastoma initiation and progression. Clin. Cancer Res. 22, 4391–4404 (2016). This paper describes the underlying rationale for and the current status of clinical trials of inhibitors of polyamine biosynthesis in the treatment of neuroblastomas.
Cheng, X. Y. et al. Deletion and downregulation of MTAP contribute to the motility of esophageal squamous carcinoma cells. Onco Targets Ther. 10, 5855–5862 (2017).
Chaturvedi, S., Hoffman, R. M. & Bertino, J. R. Exploiting methionine restriction for cancer treatment. Biochem. Pharmacol. 154, 170–173 (2018).
Tang, B., Lee, H. O., An, S. S., Cai, K. Q. & Kruger, W. D. Specific targeting of MTAP-deleted tumors with a combination of 2-fluoroadenine and 5′-methylthioadenosine. Cancer Res. https://doi.org/10.1158/0008-5472.CAN-18-0814 (2018).
Marjon, K. et al. MTAP deletions in cancer create vulnerability to targeting of the MAT2A/PRMT5/RIOK1 axis. Cell Rep. 15, 574–587 (2016). This paper provides a description of the novel targeting of methylthioadenosine metabolism.
Mavrakis, K. J. et al. Disordered methionine metabolism in MTAP/CDKN2A-deleted cancers leads to dependence on PRMT5. Science 351, 1208–1213 (2016).
Bistulfi, G. et al. The essential role of methylthioadenosine phosphorylase in prostate cancer. Oncotarget 7, 14380–14393 (2016).
Evans, G. B. et al. Tight binding enantiomers of pre-clinical drug candidates. Bioorg. Med. Chem. 23, 5326–5333 (2015).
Basu, I. et al. Growth and metastases of human lung cancer are inhibited in mouse xenografts by a transition state analogue of 5′-methylthioadenosine phosphorylase. J. Biol. Chem. 286, 4902–4911 (2011).
Qu, N. et al. Inhibition of human ornithine decarboxylase activity by enantiomers of difluoromethylornithine. Biochem. J. 375, 465–470 (2003).
Danzin, C., Casara, P., Claverie, N., Metcalf, B. W. & Jung, M. J. (2R,5R)-6-heptyne-2,5-diamine, an extremely potent inhibitor of mammalian ornithine decarboxylase. Biochem. Biophys. Res. Commun. 116, 237–243 (1983).
Mamont, P. S., Duchesne, M. C., Grove, J. & Bey, P. Anti-proliferative properties of DL-alpha-difluoromethyl ornithine in cultured cells. A consequence of the irreversible inhibition of ornithine decarboxylase. Biochem. Biophys. Res. Commun. 81, 58–66 (1978).
Levin, V. A., Ictech, S. E. & Hess, K. R. Clinical importance of eflornithine (alpha-difluoromethylornithine) for the treatment of malignant gliomas. CNS Oncol. 7, CNS16 (2018). This paper provides an excellent description of the historical and current attempts to treat glioblastomas and other brain tumours with antagonists of polyamine biosynthesis.
Bassiri, H. et al. Translational development of difluoromethylornithine (DFMO) for the treatment of neuroblastoma. Transl Pediatr. 4, 226–238 (2015).
Saulnier Sholler, G. L. et al. A phase I trial of DFMO targeting polyamine addiction in patients with relapsed/refractory neuroblastoma. PLOS ONE 10, e0127246 (2015).
Laukaitis, C. M., Erdman, S. H. & Gerner, E. W. Chemoprevention in patients with genetic risk of colorectal cancers. Colorectal Cancer 1, 225–240 (2012).
Raj, K. P. et al. Role of dietary polyamines in a phase III clinical trial of difluoromethylornithine (DFMO) and sulindac for prevention of sporadic colorectal adenomas. Br. J. Cancer 108, 512–518 (2013).
Madeo, F., Eisenberg, T., Pietrocola, F. & Kroemer, G. Spermidine in health and disease. Science 359, eaan2788 (2018).
Kennedy, P. G. Clinical features, diagnosis, and treatment of human African trypanosomiasis (sleeping sickness). Lancet Neurol. 12, 186–194 (2013).
Vissing, A. C., Taudorf, E. H., Haak, C. S., Philipsen, P. A. & Haedersdal, M. Adjuvant eflornithine to maintain IPL-induced hair reduction in women with facial hirsutism: a randomized controlled trial. J. Eur. Acad. Dermatol. Venereol. 30, 314–319 (2016).
Rounbehler, R. J. et al. Targeting ornithine decarboxylase impairs development of MYCN-amplified neuroblastoma. Cancer Res. 69, 547–553 (2009).
Gamble, L. D. et al. Polyamine pathway inhibition as a novel therapeutic approach to treating neuroblastoma. Front. Oncol. 2, 162 (2012).
Geerts, D. et al. The polyamine metabolism genes ornithine decarboxylase and antizyme 2 predict aggressive behavior in neuroblastomas with and without MYCN amplification. Int. J. Cancer 126, 2012–2024 (2010).
Depuydt, P. et al. Genomic amplifications and distal 6q loss: novel markers for poor survival in high-risk neuroblastoma patients. J. Natl Cancer Inst. https://doi.org/10.1093/jnci/djy022 (2018).
Schultz, C. R. et al. Synergistic drug combination GC7/DFMO suppresses hypusine/spermidine-dependent eIF5A activation and induces apoptotic cell death in neuroblastoma. Biochem. J. 475, 531–545 (2018). This paper describes preclinical studies combining an inhibitor of hypusine synthesis with an inhibitor of polyamine synthesis and demonstrates synergy with the combination.
Fujimura, K. et al. Eukaryotic translation initiation factor 5A (EIF5A) regulates pancreatic cancer metastasis by modulating RhoA and Rho-associated kinase (ROCK) protein expression levels. J. Biol. Chem. 290, 29907–29919 (2015).
Muramatsu, T. et al. The hypusine cascade promotes cancer progression and metastasis through the regulation of RhoA in squamous cell carcinoma. Oncogene 35, 5304–5316 (2016).
Sarhan, S., Knodgen, B. & Seiler, N. The gastrointestinal tract as polyamine source for tumor growth. Anticancer Res. 9, 215–223 (1989).
Burns, M. R., Graminski, G. F., Weeks, R. S., Chen, Y. & O’Brien, T. G. Lipophilic lysine-spermine conjugates are potent polyamine transport inhibitors for use in combination with a polyamine biosynthesis inhibitor. J. Med. Chem. 52, 1983–1993 (2009).
Weeks, R. S. et al. Novel lysine-spermine conjugate inhibits polyamine transport and inhibits cell growth when given with DFMO. Exp. Cell Res. 261, 293–302 (2000).
Samal, K. et al. AMXT-1501, a novel polyamine transport inhibitor, synergizes with DFMO in inhibiting neuroblastoma cell proliferation by targeting both ornithine decarboxylase and polyamine transport. Int. J. Cancer 133, 1323–1333 (2013).
Gitto, S. B. et al. Difluoromethylornithine combined with a polyamine transport inhibitor is effective against gemcitabine resistant pancreatic cancer. Mol. Pharm. 15, 369–376 (2018). This paper provides a description of the successful use of the polyamine blocking therapy strategy in the treatment of a pancreatic ductal carcinoma model.
Hayes, C. S. et al. Polyamine-blocking therapy reverses immunosuppression in the tumor microenvironment. Cancer Immunol. Res. 2, 274–285 (2014).
Alexander, E. T., Minton, A., Peters, M. C., Phanstiel, O. t. & Gilmour, S. K. A novel polyamine blockade therapy activates an anti-tumor immune response. Oncotarget 8, 84140–84152 (2017).
Singh, K. et al. Ornithine decarboxylase in macrophages exacerbates colitis and promotes colitis-associated colon carcinogenesis by impairing M1 immune responses. Cancer Res. https://doi.org/10.1158/0008-5472.CAN-18-0116 (2018).
Hardbower, D. M. et al. Ornithine decarboxylase regulates M1 macrophage activation and mucosal inflammation via histone modifications. Proc. Natl Acad. Sci. USA 114, E751–E760 (2017).
Chaturvedi, R. et al. Polyamines impair immunity to Helicobacter pylori by inhibiting L-arginine uptake required for nitric oxide production. Gastroenterology 139, 1686–1698 (2010).
Chaturvedi, R., de Sablet, T., Coburn, L. A., Gobert, A. P. & Wilson, K. T. Arginine and polyamines in Helicobacter pylori-induced immune dysregulation and gastric carcinogenesis. Amino Acids 42, 627–640 (2012).
Hesterberg, R. S., Cleveland, J. L. & Epling-Burnette, P. K. Role of polyamines in immune cell functions. Med. Sci. 6, E22 (2018).
Feith, D. J., Pegg, A. E. & Fong, L. Y. Targeted expression of ornithine decarboxylase antizyme prevents upper aerodigestive tract carcinogenesis in p53-deficient mice. Carcinogenesis 34, 570–576 (2013).
Feith, D. J. et al. Mouse skin chemical carcinogenesis is inhibited by antizyme in promotion-sensitive and promotion-resistant genetic backgrounds. Mol. Carcinog. 46, 453–465 (2007).
Nowotarski, S. L., Feith, D. J. & Shantz, L. M. Skin carcinogenesis studies using mouse models with altered polyamines. Cancer Growth Metastasis 8, 17–27 (2015).
Gilmour, S. K. Polyamines and nonmelanoma skin cancer. Toxicol. Appl. Pharmacol. 224, 249–256 (2007).
Guo, Y., Cleveland, J. L. & O’Brien, T. G. Haploinsufficiency for ODC modifies mouse skin tumor susceptibility. Cancer Res. 65, 1146–1149 (2005).
Meyskens, F. L. Jr et al. Difluoromethylornithine plus sulindac for the prevention of sporadic colorectal adenomas: a randomized placebo-controlled, double-blind trial. Cancer Prev. Res. 1, 32–38 (2008). This study shows the effectiveness of the combination of an NSAID with DFMO in the prevention of sporadic colorectal adenomas.
Zell, J. A. et al. Role of obesity in a randomized placebo-controlled trial of difluoromethylornithine (DFMO) + sulindac for the prevention of sporadic colorectal adenomas. Cancer Causes Control 23, 1739–1744 (2012).
Chandra, S., Nymeyer, A. C., Rice, P. F., Gerner, E. W. & Barton, J. K. Intermittent dosing with sulindac provides effective colorectal cancer chemoprevention in the azoxymethane-treated mouse model. Cancer Prev. Res. 10, 459–466 (2017).
Sinicrope, F. A. et al. Evaluation of difluoromethylornithine for the chemoprevention of Barrett’s esophagus and mucosal dysplasia. Cancer Prev. Res. 4, 829–839 (2011).
Witherspoon, M., Chen, Q., Kopelovich, L., Gross, S. S. & Lipkin, S. M. Unbiased metabolite profiling indicates that a diminished thymidine pool is the underlying mechanism of colon cancer chemoprevention by alpha-difluoromethylornithine. Cancer Discov. 3, 1072–1081 (2013).
Meyskens, F. L. Jr, Simoneau, A. R. & Gerner, E. W. Chemoprevention of prostate cancer with the polyamine synthesis inhibitor difluoromethylornithine. Recent Results Cancer Res. 202, 115–120 (2014).
Xu, H. et al. Spermine oxidation induced by Helicobacter pylori results in apoptosis and DNA damage: implications for gastric carcinogenesis. Cancer Res. 64, 8521–8525 (2004).
Babbar, N. & Casero, R. A. Jr. Tumor necrosis factor-alpha increases reactive oxygen species by inducing spermine oxidase in human lung epithelial cells: a potential mechanism for inflammation-induced carcinogenesis. Cancer Res. 66, 11125–11130 (2006).
Goodwin, A. C. et al. Increased spermine oxidase expression in human prostate cancer and prostatic intraepithelial neoplasia tissues. Prostate 68, 766–772 (2008).
Chaturvedi, R. et al. Spermine oxidase mediates the gastric cancer risk associated with Helicobacter pylori CagA. Gastroenterology 141, 1696–1708 (2011).
Goodwin, A. C. et al. Polyamine catabolism contributes to enterotoxigenic Bacteroides fragilis-induced colon tumorigenesis. Proc. Natl Acad. Sci. USA 108, 15354–15359 (2011).
Hong, S. K. et al. Increased expression and cellular localization of spermine oxidase in ulcerative colitis and relationship to disease activity. Inflamm. Bowel Dis. 16, 1557–1566 (2010).
Smirnova, O. A. et al. Hepatitis C virus alters metabolism of biogenic polyamines by affecting expression of key enzymes of their metabolism. Biochem. Biophys. Res. Commun. 483, 904–909 (2017).
Pledgie, A. et al. Spermine oxidase SMO(PAOh1), not N1-acetylpolyamine oxidase PAO, is the primary source of cytotoxic H2O2 in polyamine analogue-treated human breast cancer cell lines. J. Biol. Chem. 280, 39843–39851 (2005).
Chaturvedi, R. et al. Increased Helicobacter pylori-associated gastric cancer risk in the Andean region of Colombia is mediated by spermine oxidase. Oncogene 34, 3429–3440 (2015).
Murray-Stewart, T. et al. Epigenetic silencing of miR-124 prevents spermine oxidase regulation: implications for Helicobacter pylori-induced gastric cancer. Oncogene 35, 5480–5488 (2016). This study defines an epigenetic mechanism by which SMOX expression is dysregulated in individuals with Helicobacter pylori infection who are at high risk of developing gastric cancer.
Zahedi, K. et al. Activation of endoplasmic reticulum stress response by enhanced polyamine catabolism is important in the mediation of cisplatin-induced acute kidney injury. PLOS ONE 12, e0184570 (2017).
Seiler, N. How important is the oxidative degradation of spermine?: minireview article. Amino Acids 26, 317–319 (2004).
Porter, C. W. & Bergeron, R. J. Regulation of polyamine biosynthetic activity by spermidine and spermine analogs — a novel antiproliferative strategy. Adv. Exp. Med. Biol. 250, 677–690 (1988).
Tummala, R. et al. Combination effects of platinum drugs and N1, N11 diethylnorspermine on spermidine/spermine N1-acetyltransferase, polyamines and growth inhibition in A2780 human ovarian carcinoma cells and their oxaliplatin and cisplatin-resistant variants. Cancer Chemother. Pharmacol. 67, 401–414 (2011).
Hector, S. et al. Polyamine catabolism in colorectal cancer cells following treatment with oxaliplatin, 5-fluorouracil and N1, N11 diethylnorspermine. Cancer Chemother. Pharmacol. 62, 517–527 (2008).
Casero, R. A. Jr & Woster, P. M. Recent advances in the development of polyamine analogues as antitumor agents. J. Med. Chem. 52, 4551–4573 (2009).
Allen, W. L. et al. The role of spermidine/spermine N1-acetyltransferase in determining response to chemotherapeutic agents in colorectal cancer cells. Mol. Cancer Ther. 6, 128–137 (2007).
Hacker, A., Marton, L. J., Sobolewski, M. & Casero, R. A. Jr. In vitro and in vivo effects of the conformationally restricted polyamine analogue CGC-11047 on small cell and non-small cell lung cancer cells. Cancer Chemother. Pharmacol. 63, 45–53 (2008).
Murray-Stewart, T., Hanigan, C. L., Woster, P. M., Marton, L. J. & Casero, R. A. Jr. Histone deacetylase inhibition overcomes drug resistance through a miRNA-dependent mechanism. Mol. Cancer Ther. 12, 2088–2099 (2013).
Smith, M. A. et al. Initial testing (stage 1) of the polyamine analog PG11047 by the pediatric preclinical testing program. Pediatr. Blood Cancer 57, 268–274 (2011).
Pledgie-Tracy, A. et al. The role of the polyamine catabolic enzymes SSAT and SMO in the synergistic effects of standard chemotherapeutic agents with a polyamine analogue in human breast cancer cell lines. Cancer Chemother. Pharmacol. 65, 1067–1081 (2010).
Goyal, L. et al. Phase 1 study of N(1),N(11)diethylnorspermine (DENSPM) in patients with advanced hepatocellular carcinoma. Cancer Chemother. Pharmacol. 72, 1305–1314 (2013).
Wolff, A. C. et al. A phase II study of the polyamine analog N1,N11-diethylnorspermine (DENSpm) daily for five days every 21 days in patients with previously treated metastatic breast cancer. Clin. Cancer Res. 9, 5922–5928 (2003).
Hahm, H. A. et al. Phase I study of N(1),N(11)-diethylnorspermine in patients with non-small cell lung cancer. Clin. Cancer Res. 8, 684–690 (2002).
Streiff, R. R. & Bender, J. F. Phase 1 study of N1-N11-diethylnorspermine (DENSPM) administered TID for 6 days in patients with advanced malignancies. Invest. New Drugs 19, 29–39 (2001).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Hosseini, A. & Minucci, S. A comprehensive review of lysine-specific demethylase 1 and its roles in cancer. Epigenomics 9, 1123–1142 (2017).
Huang, Y., Marton, L. J., Woster, P. M. & Casero, R. A. Polyamine analogues targeting epigenetic gene regulation. Essays Biochem. 46, 95–110 (2009).
Nowotarski, S. L. et al. Structure-activity study for (bis)ureidopropyl- and (bis)thioureidopropyldiamine LSD1 inhibitors with 3-5-3 and 3-6-3 carbon backbone architectures. Bioorg. Med. Chem. 23, 1601–1612 (2015).
Schenk, T. et al. Inhibition of the LSD1 (KDM1A) demethylase reactivates the all-trans-retinoic acid differentiation pathway in acute myeloid leukemia. Nat. Med. 18, 605–611 (2012).
Sharma, S. K. et al. Bis)urea and (Bis)thiourea inhibitors of lysine-specific demethylase 1 as epigenetic modulators. J. Med. Chem. 53, 5197–5212 (2010).
Murray-Stewart, T. et al. Biochemical evaluation of the anticancer potential of the polyamine-based nanocarrier Nano11047. PLOS ONE 12, e0175917 (2017).
Xie, Y. et al. Self-immolative nanoparticles for simultaneous delivery of microRNA and targeting of polyamine metabolism in combination cancer therapy. J. Control. Release 246, 110–119 (2017). This is the first description of the use of a polyamine analogue as a backbone for nanoparticle construction, which results in a prodrug as a delivery system for therapeutic nucleic acids.
Leeuwenhoek, A. Natis e semine genitali animalculis. Phi. Trans. R. Soc. 12, 1040–1043 (1678).
Flynn, A. T. & Hogarty, M. D. Myc, oncogenic protein translation, and the role of polyamines. Med. Sci. 6, E41 (2018).
Tong, Y. et al. Crystal structure of human eIF5A1: insight into functional similarity of human eIF5A1 and eIF5A2. Proteins 75, 1040–1045 (2009).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03536728?term=NCT03536728&rank=1 (2018).
Sfanos, K. S., Isaacs, W. B. & De Marzo, A. M. Infections and inflammation in prostate cancer. Am. J. Clin. Exp. Urol. 1, 3–11 (2013).
Hartnett, L. & Egan, L. J. Inflammation, DNA methylation and colitis-associated cancer. Carcinogenesis 33, 723–731 (2012).
Arthur, J. C. et al. Intestinal inflammation targets cancer-inducing activity of the microbiota. Science 338, 120–123 (2012).
Babbar, N., Murray-Stewart, T. & Casero, R. A. Jr. Inflammation and polyamine catabolism: the good, the bad and the ugly. Biochem. Soc. Trans. 35, 300–304 (2007).
Noto, J. M. & Peek, R. M. Jr. The gastric microbiome, its interaction with Helicobacter pylori, and its potential role in the progression to stomach cancer. PLOS Pathog. 13, e1006573 (2017).
Hardbower, D. M., Peek, R. M. Jr & Wilson, K. T. At the Bench: Helicobacter pylori, dysregulated host responses, DNA damage, and gastric cancer. J. Leukoc. Biol. 96, 201–212 (2014).
Wu, S. et al. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat. Med. 15, 1016–1022 (2009).
Toprak, N. U. et al. A possible role of Bacteroides fragilis enterotoxin in the aetiology of colorectal cancer. Clin. Microbiol. Infect. 12, 782–786 (2006).
Ha, H. C. et al. The natural polyamine spermine functions directly as a free radical scavenger. Proc. Natl Acad. Sci. USA 95, 11140–11145 (1998).
Murray-Stewart, T. et al. Nuclear localization of human spermine oxidase isoforms — possible implications in drug response and disease etiology. FEBS J. 275, 2795–2806 (2008).
O’Hagan, H. M. et al. Oxidative damage targets complexes containing DNA methyltransferases, SIRT1, and polycomb members to promoter CpG Islands. Cancer Cell 20, 606–619 (2011).
Acknowledgements
Work in the Casero laboratory is supported by grants from the US National Institutes of Health (CA204345), the Maryland Cigarette Restitution Fund and the Samuel Waxman Cancer Research Foundation.
Author information
Authors and Affiliations
Contributions
All authors researched data for the article, contributed to the discussion of content, wrote the article and reviewed and/or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors receive US National Institutes of Health grant support (R.A.C.) and are patent holders (R.A.C. and A.E.P.). T.M.S. declares no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary terms
- Cytostasis
-
The process by which cells stop dividing without dying; it is a common response by normal cells to polyamine depletion, which can then recover growth once the polyamine block is removed.
- Hypusine
-
A unique amino acid found in all eukaryotes and some archaea that is the product of a post-translational modification of a specific lysine (K50 in humans) residue of the transcription factor eukaryotic initiation factor 5A isoform 1 (eIF5A), which is produced by the transfer of the aminobutyl moiety of spermidine to the lysine residue.
- 5′-methylthioadenosine phosphorylase
-
(MTAP). The enzyme that catalyses the initial step in methionine and adenosine nucleotide salvage from the methylthioadenosine product of both spermidine and spermine synthesis; the gene coding for this protein is frequently lost or silenced in tumours, thus facilitating a unique biochemical difference that may be exploited in many cancers.
- Protonated
-
The state of having accepted a proton; in the case of polyamines at physiological pH, all imine and amine nitrogens are positively charged.
- Proenzyme
-
The inactive precursor of an active enzyme; S‑adenosylmethionine decarboxylase (AdoMetDC) is a self-processing proenzyme that undergoes an autocatalytic serinolysis to produce the pyruvoyl group necessary for the activity of the mature AdoMetDC enzyme.
- Ferroptosis
-
An iron-dependent programmed cell death process that is characterized by an accumulation of lipid peroxidases.
- Prodrug
-
A formulation of an inactive compound that once metabolized or altered by the cellular environment produces the active form of the drug.
Rights and permissions
About this article
Cite this article
Casero, R.A., Murray Stewart, T. & Pegg, A.E. Polyamine metabolism and cancer: treatments, challenges and opportunities. Nat Rev Cancer 18, 681–695 (2018). https://doi.org/10.1038/s41568-018-0050-3
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41568-018-0050-3
This article is cited by
-
Antizyme inhibitor family: biological and translational research implications
Cell Communication and Signaling (2024)
-
Heterogeneous changes in gut and tumor microbiota in patients with pancreatic cancer: insights from clinical evidence
BMC Cancer (2024)
-
AMD1 promotes breast cancer aggressiveness via a spermidine-eIF5A hypusination-TCF4 axis
Breast Cancer Research (2024)
-
Polyamine-mediated ferroptosis amplification acts as a targetable vulnerability in cancer
Nature Communications (2024)
-
Depletion of SAM leading to loss of heterochromatin drives muscle stem cell ageing
Nature Metabolism (2024)