Abstract
Achieving autonomous motion is a central objective in designing artificial cells that mimic biological cells in form and function. Cellular motion often involves complex multiprotein machineries, which are challenging to reconstitute in vitro. Here we achieve persistent motion of cell-sized liposomes. These small artificial vesicles are driven by a direct mechanochemical feedback loop between the MinD and MinE protein systems of Escherichia coli and the liposome membrane. Membrane-binding Min proteins self-organize asymmetrically around the liposomes, which results in shape deformation and generates a mechanical force gradient leading to motion. The protein distribution responds to the deformed liposome shape through the inherent geometry sensitivity of the reaction–diffusion dynamics of the Min proteins. We show that such a mechanochemical feedback loop between liposome and Min proteins is sufficient to drive continuous motion. Our combined experimental and theoretical study provides a starting point for the future design of motility features in artificial cells.
Similar content being viewed by others
Main
Autonomous movement is considered one of the hallmarks of life and one of the critical features in constructing artificial cells1, as well as other man-made microrobotic objects. A plethora of propulsion strategies have already been exploited for inorganic colloidal microparticles2, and there is a growing interest in implementing self-propulsion in soft and organic synthetic compartments, too, using strategies such as bubble propulsion3, diffusiophoresis4,5, magnetic force6 and the Marangoni effect7. However, implementing cell-like motion by the functional in vitro reconstitution of active biological elements is challenging because it usually involves multiple mechanochemical feedback loops that span a wide range of timescales and length scales8,9,10.
The fundamental requirement for a system to exhibit cell-like motion is the conversion of molecular chemical energy into macroscopic mechanical work (mechanochemical coupling)11. Cell-sized liposomes, also referred to as giant unilamellar vesicles, are so far the best representations of biomimetic compartments12 to transfer biochemical energy into a mechanical response on the cellular scale. One possible strategy to couple a biological system to energy flow is through self-organizing reaction–diffusion systems, which are driven far from equilibrium due to the action of NTPases13. Here we use the MinDE protein system from Escherichia coli, which has received great attention as a paradigmatic biological reaction–diffusion system14,15,16,17, as a mediator between chemical and mechanical energy. In vitro, the formation of protein gradients and patterns on lipid membranes requires MinD, MinE and ATP14. On ATP binding, the ATPase MinD dimerizes and binds to the lipid membrane by two individually weak membrane-targeting sequences18,19. Above a critical local concentration, membrane-bound MinD recruits MinE to the membrane. MinE then catalyses the hydrolysis of ATP in MinD, resulting in the dissociation of both MinD and MinE from the membrane20,21.
On flat supported membranes, Min protein self-organization yields travelling22 or standing23,24 protein patterns without mechanically affecting the membrane. In contrast, on free-standing membrane structures, such as liposomes, tubes and sheets, large-scale membrane deformations have been observed as a result of periodic protein binding and dissociation25,26. From a theoretical perspective, it is known that pattern formation on dynamic surfaces can induce large-scale deformations27,28. Existing models lack realistic protein dynamics—particularly bulk–boundary interactions that were found essential for Min protein patterns17—because of the technical complexity of simultaneously simulating dynamic volumes and surfaces. Here we study how cell-sized liposomes susceptible to membrane deformations on flat supported membranes interact with dynamic Min protein patterns on the supported membranes. Surprisingly, we observe that the liposomes engage in persistent cell-like motion in the presence of the pattern-forming Min protein system (Fig. 1a and Supplementary Video 1), indicative of a mechanochemical feedback loop (Fig. 1c). Experiments and simulations consistently demonstrate that the feedback mechanism between the liposome and self-organizing MinE proteins is based on geometric cues: spatial gradients of membrane-binding Min proteins induce a deformation of initially spherical liposomes into asymmetric shapes through spatially varying membrane adhesion linked to the distribution of Min proteins (Fig. 1b). This asymmetry yields mechanical force gradients resulting in directional liposome movement, which, in turn, causes a dynamic reorganization of the protein pattern due to the inherent geometry sensitivity of Min reaction–diffusion dynamics29, closing the feedback loop (Fig. 1c).
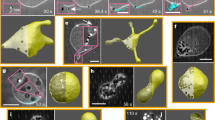
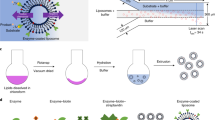
a, A 3D image showing liposome motion in the presence of Min protein patterns. Two 3D images were superimposed to show the motion sequence indicated by the arrow (time interval, 1 h 22 min; dimensions of the 3D image, 21.68 × 21.84 × 5.40 µm3). b, Schematic of the liposome motion driven by Min protein gradients. The MinE proteins asymmetrically accumulate at the flattened side of the liposome, and the liposome moves against the regions with MinE accumulation. c, Mechanochemical feedback loop that drives the liposome motion.
MinD–ATP induces differential liposome adhesion to SLBs
To investigate how Min proteins mechanically affect the free-standing liposome membrane, we first examined the effect of MinD in the absence of MinE. When placing liposomes on supported lipid bilayers (SLBs), they maintain spherical shapes in the absence of Min proteins or with no ATP added (Fig. 2a,b). In the presence of MinD and ATP, allowing MinD to bind to the membrane, we observed that liposomes were symmetrically flattened into dome shapes (Fig. 2b), which can be characterized by contact angle θ (Fig. 2c).
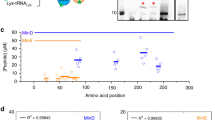
a, Schematic and experimental illustration of liposome flattening on SLBs (both grey) facilitated by MinD (magenta) membrane binding. MinD–ATP addition brings the liposomes close to the SLBs (top; the rows show snapshots with increasing time; the interaction is indicated by magenta curves), allowing the lipid membranes to adhere (centre; chain symbols). MinD proteins are excluded from the liposome–SLB contact areas (bottom). Fluorescence snapshots show MinD (magenta) and liposome membrane (grey) at the subsequent stages of liposome attachment. Scale bar, 5 μm; b, Liposome flattening requires both MinD and ATP (right), and no flattening is observed when only MinD (left) or only ATP (centre) is present. Scale bar, 5 μm. c, Characterizing liposome flattening by contact angle θ (defined in b). The actual contact angle of an ideal sphere should be 180°, but due to limited image resolution, the measured contact angle of a perfect sphere would be around 155° (Methods). Liposomes did not substantially deform in Min buffers with different MgCl2 concentrations, Min buffer with ATP addition (2.5 mM) and Min buffer with MinD addition (2.5 µM). However, θ substantially decreased in the simultaneous presence of MinD (2.5 µM) and ATP (2.5 mM). From left to right, the database for the measurements is n = 19, one experiment; n = 45, three experiments; n = 44, two experiments; n = 73, four experiments; n = 35, two experiments; n = 60, three experiments. For the box figure, whiskers are 1.5× the interquartile range, the median is shown as a black line and the mean is shown as a red line. Formation of Min quasi-stationary pattern in vitro and in silico. d, Schematic of MinE persistent binding model21,38,39,40,41. MinD–ATP (magenta) autocatalytically binds to the membrane, where it can bind MinE (green). MinE induces the dissociation of MinD from the membrane. MinE is present on the membrane not only as a part of a MinDE complex ① but also as a MinE dimer isolated from MinD ②. e, Quasi-stationary MinD (left) and MinE (right) protein patterns as observed in experiments (top) and simulations (bottom; Methods). The simulations show the local protein concentration, whereas the experiments show the protein fluorescence intensity. Scale bar, 10 μm.
As demonstrated earlier, liposome flattening on supported membranes is not only induced by membrane-binding proteins but also by other factors, such as electrostatic interactions30 (Extended Data Fig. 1). The exact adhesion mechanism is determined by a complex interplay between these different factors at the biochemical level, including the reduction in membrane rigidity31, actual formation of oligomers linking the opposing lipid bilayers32 or a change in the local electrostatic potential of the membranes33. MinD is apparently not directly required at the adhesive interface to maintain the flattened liposome shapes, as we observed that the protein was largely excluded from the liposome–SLB contacting areas (Fig. 2a and Supplementary Fig. 1). This suggests that the adhesion is maintained by attractive electrostatic forces rather than a direct linking of the two membranes by (oligomeric) MinD sandwiched between them. This view is supported by the attraction of like-charged membranes immersed in a solution of multivalent oppositely charged ions34,35. Indeed, in our system, the liposomes and SLBs were fabricated with a negative net charge, whereas the Min buffer contains a high cation concentration (Methods). Importantly, MinD is simultaneously needed to flatten the liposomes (Fig. 2c), meaning that both membrane charge and Min protein binding were required to flatten the liposomes in our system.
Quasi-stationary MinDE patterns support liposome motion
Although the liposomes readily adhered to the SLB in the presence of MinD–ATP alone, they showed no dynamics. Dynamic deformation and sustained movement were only observed after the addition of MinE and the subsequent formation of protein patterns that create the required heterogeneous environment on a liposome-sized scale. Therefore, we determined which features of the Min patterns were associated with liposome motion. In general, depending on various experimental conditions14, Min proteins can exhibit both travelling waves22 and quasi-stationary patterns on SLBs23. Intriguingly, for travelling Min waves, we found no evidence of a sustained mechanical effect on the liposomes that would support directional movement (Supplementary Video 2). We hypothesize that the Min proteins must locally interact with the liposome membrane over a sufficiently long period of time—thereby building up a mechanical strain through adhesion—for the liposomes to start moving. Therefore, we established conditions for quasi-stationary Min patterns to develop, in which protein concentrations at a given position remain approximately constant over time (Methods)23.
We performed numerical simulations of molecular reaction–diffusion models in realistic geometry to rationalize how the Min proteins interact with the liposomes, focusing on obtaining quasi-stationary Min patterns. One widely used model, the so-called Min skeleton model, can qualitatively reproduce the Min oscillations in vivo36,37 and in vitro16, but is insufficient to achieve the quasi-stationary Min patterns. From our experimental results, the concentration of MinE is high in regions where the concentration of MinD is low (Fig. 2e), which suggests that MinE can remain on the membrane after the MinD–MinE heterodimer has disassociated. This is not accounted for in the Min skeleton model16,36,37, where membrane-bound MinE only occurs as a part of the MinDE complex, and the MinD concentration is always larger than that of MinE. Previous experimental and theoretical studies have proposed MinE persistent binding models, where MinE can remain on the membrane after the MinD dimer is disassociated, and membrane-bound MinE can capture and dissociate multiple membrane-bound MinD proteins (Fig. 2d)21,38,39,40,41. Inspired by this previous work and our experimental results, we extended the Min skeleton model to include MinE persistent binding and found that it then indeed exhibits quasi-stationary patterns (Fig. 2e and Methods). In addition, fluorescence recovery after photobleaching measurements indicated that the diffusion of MinE is more than one order of magnitude slower than that of MinD on SLBs (Extended Data Fig. 2). To account for this observation, we reduced the diffusion constant of MinE (De) and found that only when De was at most about half as large as the diffusion coefficient of MinD, the resulting pattern reproduced the experimental results (Extended Data Fig. 3). The pattern becomes more pronounced on lowering De, with transient mesh patterns emerging when MinE diffusion is slower than MinD diffusion by more than one order of magnitude (Supplementary Fig. 2).
A geometry effect induces asymmetric accumulation of MinE
Next, we wanted to quantify the effect of liposome membrane on the Min protein pattern. We first prepared quasi-stationary Min patterns on SLBs and then added the liposomes. On settling down on the SLBs, the liposomes got quickly covered with MinD, but without MinE-induced pattern formation on the liposome, at least on the timescales of observation (Fig. 3a). In this way, Min pattern formation on SLBs could be separated from protein binding to the liposomes. Once the liposomes were in contact with the SLBs, SLB-associated MinE accumulated at the edges of the liposome–SLB contact areas; because of the pattern-induced spatial variation of protein concentration on the surface, it adopted an asymmetric distribution on the liposome membrane in the contact area (Fig. 3a). In particular, this asymmetric MinE distribution correlated with an asymmetric deformation of the liposomes, flattening them on the sides of high MinE concentration (Fig. 3a).
a, Schematic (top) and experimental (bottom; MinD fluorescence is shown in magenta, MinE fluorescence is shown in green and SLB is indicated in grey) illustration of asymmetric MinE accumulation around the liposome–SLB contact edges. Liposomes were added to the preformed Min patterns on the SLBs (left), and then nearby SLB-associated MinE accumulated to the liposome–SLB contact edges (centre; pattern dynamics are indicated by the green arrow). Because of the preformed patterns, MinE adopted an asymmetric distribution on the liposome membrane, and the liposomes were asymmetrically deformed (right). Shortly before the liposome settles (t = 0 s), the shape appears distorted due to the high liposome velocity (compared with the image acquisition speed). The Min protein pattern was transiently affected because of the addition of liposome solution containing sucrose (Methods). The side views were obtained by slicing 3D images from the positions shown with the white dashed lines in the top-down images. The buffer contained 25 mM Tris (pH 7.5) and 100 mM KCl. Scale bar, 5 μm. b, Schematic of a narrow bulk volume close to the contact edges confined by the SLB and liposome lipid bilayers. The net protein concentration fluxes and protein (un-)binding are indicated by the magenta and black arrows, respectively. The high membrane-to-bulk ratio alters the reactive equilibria42 and reduces the MinD attachment rates per membrane area29,43. Slowly diffusing MinE can accumulate in this region on the membrane, whereas the MinD concentration is reduced. c, Symmetric MinE accumulation around a symmetrically deformed liposome (marked with a square) in a 3D simulation (top-down view) (top). A 3D image of the area marked in the top panel (bottom). The liposome shape is indicated as a grey hatched surface (Methods). d, Asymmetric MinE accumulation at the more flattened side of asymmetrically deformed liposomes.
Without having to make assumptions about the interaction of MinE with the liposome membrane in the contact areas, our theoretical considerations suggest that a geometry effect may be responsible for a self-enhanced MinE accumulation at the liposome–SLB contact edges. Here the term ‘geometry effect’ relates to the fact that in this region, the membrane-to-bulk ratio—which depends on the geometry—is strongly increased, which alters the reactive equilibria42 and effectively reduces the MinD attachment rates per membrane area29,43. As a result, a concentration gradient of MinD proteins emerges on the membrane, leading to a net flux of MinD, particularly MinDE complexes towards the contact edge (Fig. 3b). Due to persistent MinE binding, individual MinE proteins are expected to remain on the membrane after triggering MinD dissociation (Fig. 2d), and MinE can accumulate at the liposome–SLB contact edges as the MinD concentration reduces (Fig. 3b). To test these heuristic arguments, we examined the Min protein dynamics in finite element simulations in a geometry emulating a liposome resting on an SLB. These simulations confirmed that MinE accumulation is indeed biased towards regions of high membrane-to-bulk ratios, whereas the protein pattern remains unperturbed elsewhere (Fig. 3c,d). In addition, when the simulated liposome was asymmetrically deformed, MinE preferentially accumulated at the flattened side (Fig. 3e). MinE binding, in turn, enhances further liposome flattening (Supplementary Fig. 3), which may arise from a change in membrane mechanical properties by several plausible mechanisms44,45. Once MinE is recruited by the membrane-bound MinD, the membrane-targeting sequence of MinE is exposed44,45. The insertion of the membrane-targeting sequence into the lipid bilayer may induce a spontaneous membrane curvature, which has been suggested to account for pulsating shape deformations in osmotically deflated vesicles25. Moreover, it has been shown that helix insertion reduces membrane bending rigidity and enhances the adhesion of liposomes to substrates31. In addition, enhanced protein binding increases membrane occupancy, decreasing protein diffusion on the membrane46. The reduced diffusivity of MinE enhances the pattern contrast (Extended Data Fig. 3), stabilizing the observed geometry sensitivity. Therefore, we conclude that MinE accumulation and liposome deformation promote each other, suggesting a positive mechanochemical feedback mechanism.
Asymmetric MinE distribution induces liposome motion
Having established that the deformation of liposomes leads to the differential accumulation of MinE at the liposome–SLB contact edges, we asked how the patterns affect the movement of liposomes. To this end, we adopted a different experimental procedure where we first prepared the liposome–SLB membrane system and then added the Min proteins (Fig. 4a). We observed that MinE accumulation is correlated with liposome movement in a direction pointing away from the MinE accumulation sides (Fig. 4b and Supplementary Video 3). To quantify this effect, we inferred the motion direction by correlating the liposome and MinE centroids (Methods), which fits well with the observed direction even for curved liposome trajectories (Fig. 4c and Extended Data Fig. 4). From this, we conclude that the asymmetric MinE distribution drives the liposome motion, which was also confirmed through a fluorescence characterization (Extended Data Fig. 5). The Min protein concentrations used in our system are substantially higher than those used in physiological assays14 (Methods) to generate stable stationary patterns23. When sequentially decreasing MinE concentration, the MinE patterns turned into spots or lines instead of meshes or labyrinths. Under these conditions, motile liposomes were rarely observed (Supplementary Videos 4 and 5 and Supplementary Fig. 4). We assume that sufficient MinE binding or the continuity of zones of high MinE concentration is critical for sustaining liposome motion. When reducing MinD concentrations, the protein patterns are no longer stable (Supplementary Video 6), and no motile liposomes were observed (Supplementary Fig. 4).
a, Schematic and experimental illustration of liposome motion driven by Min quasi-stationary patterns. The Min proteins (MinD, magenta; MinE, green) are added into the liposome–SLB membrane (grey) system; subsequently, the liposomes deform asymmetrically and move. The asymmetric shapes of the liposomes promote asymmetric MinE distribution due to a generic diffusional limitation effect. Thus, liposome motion (grey arrow) and Min pattern reorganization (green arrow) set up a mechanochemical feedback loop. b, Snapshots from Supplementary Video 3 illustrating liposome motion. The side views were obtained by slicing 3D images from the positions, as shown by the white dashed lines in the top-down view. The buffer contained 25 mM Tris (pH 7.5) and 100 mM KCl. Scale bar, 5 μm. c, We infer the direction of liposome motion by correlating the centroids of liposome and MinE distribution. The inferred direction fits well with the observed one during the whole motion process (Methods and Extended Data Fig. 4). d, Conceptual self-regulating feedback loop between protein pattern reorganization and liposome motion (shown in a) is sufficient to fuel the persistent motion in a 2D phase-field simulation. The liposome is modelled as an excluded volume (white area). During continuous motion, MinE gradually accumulated towards the rear part of the liposome. The corresponding kymograph of the redistribution of MinE (green shadow) during liposome motion (white shadow) (right), where the times and positions of the side-view snapshots (left) are indicated by the white dashed lines. Scale bars, 5 μm; 5,000 s.
During directional motion, the liposomes assume asymmetric shapes (Fig. 4b) that, in turn, promote asymmetric MinE distribution due to the geometry effect described above (Fig. 3b). Therefore, Min pattern reorganization and liposome motion set up a mechanochemical feedback loop (Fig. 4a). To test whether this feedback loop is sufficient to sustain persistent motion, we incorporated this feedback loop in a simplified two-dimensional (2D) simulation, where the liposome is represented by a 2D cross section on a one-dimensional membrane (Methods). Since the 2D system does not have rotational symmetry around the axis normal to the membrane, we neglect the asymmetric liposome shape in this simulation. In this reduced feedback loop, pattern reorganization induced by liposome motion emerges from the inherent geometry sensitivity of the Min reaction–diffusion dynamics in the persistent binding model (Fig. 3b). Since the biochemical details of the protein–membrane interactions that lead to liposome motion are presently not fully understood on a molecular level, we generically linked liposome movement to pattern reorganization in the sense that the simulated liposome was designed to move away from high MinE concentrations (Fig. 4b). Only by combining these two components, persistent liposome motion accompanied by a continuous redistribution of MinE against the moving direction can be achieved (Fig. 4d). In contrast, when the mechanochemical feedback loop is interrupted, the liposome motion decays quickly (Extended Data Fig. 6). This shows that a conceptual self-regulating feedback loop between protein pattern reorganization and liposome motion is necessary and sufficient to drive persistent motion.
Note that the timescales of pattern formation and liposome movement need to be comparable for the feedback loop to be effective. We numerically studied the cases where the liposome moves much faster or slower than the pattern adapts (Extended Data Fig. 7). This is achieved either by varying some of the reaction rates or by changing the manual coupling of the liposome movement to the pattern (Methods). No chemical asymmetry can be maintained when the pattern adapts too fast, and no liposome movement is observed in simulations. In contrast, when the pattern cannot adapt to the moving liposome in time, no persistent continuous motion is established, but instead, the liposome is observed to rapidly move between patches of high MinE accumulation (Extended Data Fig. 8). To some extent, this is reminiscent of the experimental observation that liposome motion seems to be strongly affected by the mesh pattern, where liposomes tend to hop between mesh cells rather than moving smoothly across the surface (Supplementary Video 1).
Membrane mechanical properties determine liposome motility
In general, a sustained imbalance of forces on the front and rear sides of the liposomes is required for directional motion. As evident from our experiments, this imbalance is induced by the asymmetric distribution of MinE around the liposomes. How exactly this chemical protein gradient translates into a force is a fascinating yet unsolved question. An in-depth theoretical investigation of this question would require detailed structural insights into the dynamics of Min protein binding and a new theoretical framework that should incorporate pattern formation, dynamic and deformable geometries and adaptive mechanical properties47.
To still gain insights into the possible mechanisms of motility induction, we recorded the percentage of motile vesicles (Supplementary Fig. 4a) dependent on various plausible factors. During liposome motion, membrane trails were often left behind (Fig. 4b), consuming the membrane available for Min protein binding and thus increasing membrane tension. Therefore, we first investigated the general importance of membrane deformability. We fabricated lipid membranes onto silica beads (liposome beads), not allowing any membrane deformation. We found that these liposome-sized beads did not move (Supplementary Video 7), suggesting that membrane deformability is a crucial determinant of motion.
In the absence of sophisticated probe technology, such as optical or magnetic tweezers, a rough estimate of membrane deformation can be given by the liposome contact angle that reflects membrane mechanical properties, such as membrane tension48 and membrane bending rigidity31. Therefore, we tracked the contact angle at the rear part of moving liposomes to map changes in the membrane mechanical properties. Interestingly, liposome velocities decreased with increasing contact angles (Extended Data Fig. 9). With increasing membrane adhesion (decreasing contact angle), lateral membrane tension is supposed to substantially increase48. Membrane tension consequently elevates the Laplace pressure and generates a positive outward pressure, which can be balanced by a traction force generated by the SLBs48. Asymmetric MinE binding decreased the contact angle at the liposome rear (Extended Data Fig. 9), and the traction pressure difference is expected to drive liposome motion and affect the velocity. Therefore, the membrane mechanical properties characterized by the contact angles in our system affect liposome motility. In addition, we quantified the speed in relation to liposome size, and larger liposomes tend to move faster (Supplementary Fig. 5). Since the Laplace pressure counteracting deviations from an ideal sphere are inversely proportional to the liposome radius, large liposomes are generally more easily deformable than small ones. On the basis of these observations, we propose that larger liposomes may be more susceptible to structural asymmetries, thereby facilitating liposome motion.
Liposome sliding is probably mediated by membrane adhesion
Previous studies have shown that liposomes can either roll49 or slide50 across substrates, which may at least hint at potential mechanisms of motion. To distinguish these two types of liposome movement, we fabricated phase-separated liposomes (Fig. 5a). We find that the relative position of non-labelled lipid-ordered (Lo) domains on the liposomes remained unchanged during directional motion, indicating liposome sliding (Fig. 5a and Supplementary Videos 8–11). The rolling of liposomes was observed only when the directional motion was restricted by obstacles, such as random membrane aggregates (Extended Data Fig. 10 and Supplementary Video 12). Liposome motion by sliding was suggested to arise from the kinetic trapping of cations at the liposome–SLB contact area50. This effect may also be responsible for liposome movement on Min protein patterns, which is supported by our observation that liposome flattening is mediated by membrane adhesion through charge interactions (Extended Data Fig. 1). In addition, at high MgCl2 concentrations, liposomes were constricted during motion (Supplementary Video 13), which may arise from higher membrane adhesion. When there is no MgCl2 in the Min buffer, the liposomes could move without noticeable constriction (Supplementary Videos 1 and 3), and reducing cation concentration facilitated the directional motion (Supplementary Fig. 6).
a, Schematic and experimental illustration of phase-separated liposomes sliding on the SLBs with the non-labelled Lo domain remaining at the rear of the liposomes. Here Lo refers to the lipid-ordered domain (grey), whereas the lipid-disordered domain (Ld) is labelled in red. The rows show two subsequent snapshots of a liposome moving (arrows) in the presence of MinE (green). Scale bar, 5 µm. The buffer contained 25 mM Tris (pH 7.5) and 150 mM KCl. b, Schematic of the proposed force generation mechanisms. MinD binding flattens the liposomes, stabilizing the shape with an adhesive force (chain symbol). Asymmetrically distributed MinE proteins lead to an adhesion gradient, which leads to a horizontal force that displaces the liposome (grey arrow), and the asymmetry is maintained by pattern reorganization (green arrow). c, In another scenario, asymmetrically distributed MinE proteins lead to the deformation of the liposome. The deformation is counteracted by membrane surface tension (small arrows inside the liposome), resulting in a retraction of the rear part of the liposome and a displacement of the liposome–SLB contact area. Then, the MinE pattern adapts to the displaced liposome and maintains the interfacial energy gradient.
Liposome motion due to an adhesive gradient has been previously reported in the context of self-induced haptotaxis50. A similar effect may occur in our system, where we expect the negatively charged liposomes and SLBs to be linked through attractive electrostatic forces34,35. Since Min proteins are charged33, they could interfere with the electrostatic forces. In addition, it has recently been shown that MinDE complexes interact more strongly with the lipid membrane than MinD alone32. Consequently, MinE protein accumulation could locally change the mechanical membrane properties44,45 and interfere with the adhesive forces. Therefore, MinE proteins may be able to decrease the adhesive interactions at the rear and induce liposome drifting towards regions of higher adhesive energy50 (Fig. 5b). Another plausible scenario is that the asymmetric MinE distribution on the liposome could locally change the membrane tension, which has been referred to as interfacial energy gradient51. The liposome membrane tension could then counteract the local deformation, leading to a retraction of the liposome-exposed tails and a lateral displacement of the liposome–SLB contact area away from regions of high MinE concentration (Fig. 5c).
Having established that membrane properties generally determine liposome motility, we further investigated how lipid composition affects liposome motion. The typical lipid composition in our system contains 30 mol% 1,2-dioleoyl-sn-glycero-3-phospho-rac-(1-glycerol) (DOPG), as the negative charge of DOPG is critical to generate Min protein patterns14 (Methods). Changing the lipid composition simultaneously changes membrane adhesion and protein pattern formation, affecting liposome motion. At 10 mol% DOPG, both membrane adhesion and protein binding to the membrane33 are reduced. Under these conditions, the Min stationary patterns are no longer stable and transform into travelling waves over time, terminating liposome motion (Supplementary Video 14; two independent experiments). At 50 mol% DOPG, both membrane adhesion and protein binding to the membrane33 increase. Under these conditions, the yield of motile liposomes was reduced, but the velocity did not considerably change compared with 30 mol% DOPG (Supplementary Fig. 4).
Given the striking effects of MinDE protein patterns on large membrane structures such as vesicles, it is plausible to ask whether the reported transport phenomenon may serve a so far undetected physiological purpose. A cell’s healthy growth and homoeostasis rely on the precise spatial organization of molecules. Reaction–diffusion mechanisms can facilitate this organization not only on a molecular scale but also on a larger scale, as evidenced by ParA-directed plasmid movement and actin- or microtubule-based cargo transportation. These systems have previously been proposed to function through a filament-based mechanism52,53. However, non-filament-based mechanisms for the transport and positioning of large cellular cargo have gradually been revealed. A previous study recapitulated the directed movement by the ParA system, which is well described by a diffusion ratchet mechanism, where the cargo dynamically establishes and interacts with a concentration gradient of the ATPase54. Another work demonstrated that the diffusive fluxes of MinDE can transport and sort functionally unrelated cargo on membranes via a diffusiophoretic mechanism55. These observations hint at an underappreciated cargo transport mechanism that does not rely on stable molecular assemblies but depends on weak and transient interactions between a sufficiently large number of ATPases and ATPase effectors. The system observed here accomplished directional liposome motion driven by dynamic interactions between ATPase MinD and its stimulator MinE, which further strengthens the possible relevance of non-filament-based mechanisms.
Compared with the above in vitro systems using beads54 or DNA origami55, our system further incorporated the mechanical factor of membrane deformation and demonstrates that a minimal mechanochemical feedback mechanism consisting of a deformable liposome and a non-equilibrium protein system is sufficient to realize persistent and directional motion of cell-like objects. The pattern-forming reaction–diffusion system is a generic mechanism for continuously supplying a non-equilibrium system with energy56, which can then be transformed into directional motion or deformations. The reaction–diffusion Belousov–Zhabotinsky system has been harnessed to directionally transport a CO2 bubble, which intermittently moves in the direction of chemical wave propagation57. In our system, liposomes move away from high MinE concentration, generating a MinE concentration gradient by using the setup of a flowcell41 may guide the liposome towards a specific direction. Mechanochemical coupling has recently been appreciated as an advanced strategy to achieve life-like properties of artificial cells in vitro58. Although uncovering the mechanical details that comprise this feedback loop on a molecular level remains an open research topic, we are confident that our system will serve as a modelling platform for designing artificial cells with advanced motion functions. Moreover, further studies of such mechanochemical feedback between reaction–diffusion systems and their effects on the mechanical properties of soft materials will greatly facilitate our understanding of complex life-like behaviours and the functional evolution of living organisms.
Methods
Protein purification
The plasmids pET28a–His–MinD_MinE (ref. 22), pET28a–mCherry–MinD, pET28a–MinE–KCK–His (ref. 41) and pET28a–MinE–His (ref. 41) were used for the purification of His–MinD, mCherry–MinD, MinE–KCK–His and MinE–His, respectively59. In brief, proteins were expressed in E. coli BL21 (DE3) and then purified via Ni-NTA affinity and size-exclusion chromatography in a storage buffer (50.0 mM HEPES/KOH at pH 7.2, 150.0 mM KCl, 10% glycerol, 0.1 mM EDTA and 0.4 mM tris(2-carboxyethyl)phosphine). Proteins were snap frozen in liquid nitrogen and stored in small aliquots at –80 °C.
Protein concentration measurements
All the protein concentrations were determined via a linearized Bradford assay60 using bovine serum albumin as a reference. Measurements were done in 96-well plates, measuring the absorbance at 595 nm using a TECAN plate reader (Tecan Group), and triplicates of each measured concentration were taken. A minimum of two different dilutions of the same protein stock were measured.
Protein labelling
The labelling of MinE–KCK–His with Atto488 maleimide (ATTO-TEC; order no. AD 488-41) was performed according to the dye manufacturer’s instructions. In brief, the protein stock solution was mixed with 1.3-fold molar excess of reactive dye per sulfhydryl group in anhydrous, amine-free dimethylformamide under gentle shaking. The reaction mixture was incubated for 2 h at room temperature protected from light. A dialysis cassette (Thermo Scientific; prod. no. 66380) was used to remove most of the unbound dye, followed by an overnight dialysis against excess storage buffer.
SUV preparation
Here 20.0 mol% DOPE (Avanti, >99%; (Δ9-cis) PE; CAS, 4004-05-1), 50.0 mol% DOPC (Avanti, 18:1 (Δ9-cis) PC; SKU, 850375P), 30.0 mol% DOPG (Avanti, 18:1 (Δ9-cis) PG, SKU: 840475 P) and 0.1 mol% Atto655–PE were dissolved in chloroform. Chloroform was then evaporated under a nitrogen flow, followed by vacuuming for 2 h. The dry lipid film was rehydrated in phosphate-buffered saline (PBS) (1×, pH 7.4) (final lipid concentration, 1 mM). Lipid dispersions were sonicated to clarity (10 min). Small unilamellar vesicle (SUV) solutions are stored in −20 °C and sonicated for 10 min before usage.
SLB preparation
The sample well was plasma cleaned for 10 min and then 20 μl SUV solution and 60 μl Min buffer (25 mM Tris at pH 7.5, 150 mM KCl and 5 mM MgCl2) were added. After incubation for 10 min at 37 °C, excess non-fused SUVs were washed away with PBS (1× PBS, pH 7.4) 10 times and then the PBS was replaced twice with the respective buffer.
Cell-sized liposome preparation
Cell-sized liposomes were produced by electro formation in PTFE chambers with Pt electrodes61. Then, 6 µl of lipid mixture (20.0 mol% DOPE, 50.0 mol% DOPC, 30.0 mol% DOPG and 0.1 mol% Atto655–PE) was spread onto two Pt wires and dried in a desiccator for 30 min. The chamber was filled with 350 μl sucrose (final lipid concentration, 1 mg ml−1), and the sucrose solution was mixed with an equal osmolality of the respective imaging buffer. An a.c. electric field of 2 V (r.m.s.) was applied at a frequency of 10 Hz for 1.0 h, followed by 2 Hz for 0.5 h. The liposomes were kept at 4 °C for achieving equilibrium of the negatively charged DOPG on both lipid leaflets61. Then, 5 µl of the liposome suspension (prediluted 1:50 in sucrose) was added into the imaging chambers. For phase-separated liposomes, a lipid composition of 40.0 mol% DOPC, 20.0 mol% DOPG, 20.0 mol% DSPC, 20.0 mol% cholesterol and 0.1 mol% Atto655–PE was used (final lipid concentration, 1 mg ml−1). Atto655–PE was used to label the Ld domain. Further, the PTFE chambers were incubated within a temperature controller at 60 °C during electro formation. After electro formation, the phase-separated liposomes were slowly cooled down to room temperature overnight.
Reconstitution of the Min proteins on the SLB–liposome membrane system
Min proteins and ATP were premixed and added into the sample well containing SLB–liposomes. To test how the liposome membrane affects the Min protein patterns, we adopted another experimental procedure. We preformed Min protein patterns on SLBs and then added liposomes to the preformed Min pattern. For labelling, MinD was doped with 20 mol% mCherry–MinD, and MinE was doped with 20 mol% Atto488–KCK–MinE–His, if not stated separately. Mg–ATP solution (100.0 mM ATP and 100.0 mM MgCl2) was mixed with a final concentration of 2.5 mM Min. Quasi-stationary patterns were achieved by moving the protein modification His tag of MinE from the N terminus (His–MinE) to the C terminus (MinE–His; this is referred to as MinE in the main text and Methods)23. For all the experiments, the final volumes were 100 μl. The concentrations of MinD and MinE were 2.5 and 5.0 µM, and the Min buffers contained 25 mM Tris at pH 7.5, 150 mM KCl and 5 mM MgCl2, if not stated separately.
Fabrication of silica-bead-supported liposomes
SUVs were prepared as above, but the dry lipid film was rehydrated in water instead of PBS. Then, 10 µl of the bead solution (Corpuscular; C-SIO-1.0; C-SIO-2.0; C-SIO-5.0 (ref. 62); 5 × 108 beads per ml) was mixed with 20 µl SUVs (1 mM), 20 µl NaCl (5 M) and Milli-Q water to a total volume of 100 µl. We have used silica beads of 1, 2 and 5 µm in size. Then, the beads were incubated at room temperature for 30 min with intermittent mixing (Eppendorf, ThermoMixer C, 400 r.p.m.). Then, 1 ml Milli-Q water was added to wash off the excess unbound liposomes. Thereafter, the beads were spun at 260×g for 2 min at room temperature. The washing procedure was repeated three times. Subsequently, 1 µl silica-bead-supported liposomes were added to SLB containing the imaging chambers. Min proteins were reconstituted as described above. For each size, two independent experiments were conducted.
Microscopy
All the images were taken on a Zeiss LSM800 confocal laser scanning microscope using a Zeiss C-Apochromat 40×/1.20 water-immersion objective (Carl Zeiss). The longer time series were acquired using the built-in definite focus system. Atto488–KCK–MinE–His was excited using a 488 nm argon laser; mCherry–MinD, using a 561 nm diode-pumped solid-state laser; and Atto655, using a 633 nm He–Ne laser. To reduce the interference of the laser on the Min proteins, an oxygen scavenger was applied (final concentrations of 3.7 U ml–1 pyranose oxidase, 90.0 U ml–1 catalase and 0.8% glucose; all the reagents were purchased from Sigma).
Calibration of fluorescence intensity across SLBs and liposomes
In Fiji 70 (v1.51q) software, we marked circles with the same area on both liposome–SLB contact areas and SLBs. Then, we compared their fluorescence intensities, that is, the MinD intensity on the SLBs was 1 and the calibrated MinD intensity of the contact area indicated the protein exclusion degree.
Fluorescence recovery after photobleaching measurement
We applied an exponential fit to determine the diffusion coefficient, which would only accurately describe the recovery in diffusion-dominated systems opposed to a reaction-dominated system63. Therefore, the fluorescence recovery after photobleaching results obtained here were just used to roughly compare the diffusion dynamics of Min proteins. For this measurement, MinE was doped with 10 mol% Atto488–KCK–MinE–His. The bleaching sizes were 31.95 µm × 31.95 µm. Two circles with the same area (diameter, 2 μm) were selected. One of the circles was bleached after recording 5 frames and then 45 frames were recorded (0.16 min per frame). To reduce any photoinduced protein aggregation, we used an oxygen scavenger for all the measurements and we reduced the photobleaching laser power for the MinE channel (25%).
Measurement of liposome motion speeds and contact angles
Liposome motion was detected by eye. The liposome–SLB contact areas were obtained from the ‘analyze particles’ function of ImageJ (Fiji 70, v1.51q). The images from the membrane channel were subjected to ‘subtract background’, ‘median filter’ and ‘adjust threshold’ to obtain the liposome outlines. The TrackMate plugin in ImageJ was used to track the moving liposomes64,65. Using a Laplacian of Gaussian filter, the liposomes were identified and their trajectories were formed. The tracks with liposome collisions were eliminated. The LBADSA plugin was used to measure the contact angle66. The liposome radius was either manually measured or calculated by using chord length, c = 2Rsin(θ/2); the chord length was manually measured. Because a limited z-stack resolution was used for determining the contact angle, we did not measure liposomes smaller than 4 µm. Data collection and analysis were not performed blind to the conditions of the experiments. No statistical methods were used to predetermine the sample sizes, but our sample sizes are similar to those reported in previous publications22,55. Data distribution was assumed to be normal, but this was not formally tested.
Inference of liposome movement direction from MinE distribution
We performed image processing in Mathematica 12.1. From a z-stack time series, the liposome outlines and centres were generated from the giant unilamellar vesicle equators of the MinD channel. All the images were preprocessed by applying Gaussian filtering and binarization to reduce noise and artifacts. The liposomes were identified as the largest ring-like objects (non-simply connected regions), which were subsequently converted to a simply connected region by setting the values of all the enclosed pixels to 1. The liposome centre was defined as the centroid of this region, where p is the pixel coordinates and I(p) is the binary intensity of the corresponding pixels. The MinE distribution in the vicinity of the vesicle was obtained by multiplying the MinE image data at the SLB with the binary liposome mask obtained from the MinD data in a pixel-wise manner. The resulting data were preprocessed as described above, allowing to extract the MinE centroid. The liposome motion directions were extracted from the liposome centroid between subsequent snapshots. The liposome motion was predicted from the offset between the liposome centroid and the MinE centroid, with the direction vector pointed away from the MinE centroid.
Partial differential equations of the Min persistent binding model
We used a reaction–diffusion approach to reproduce the Min patterns observed in vitro38. In brief, three cytosolic protein species (MinE (uE), MinD–ADP (uDD) and MinD–ATP (uDT)), as well as three membrane-bound species (MinE (ue), MinD (ud) and MinDE complexes (ude)) were included. In the cytosol, MinD–ADP is converted into MinD–ATP at rate λ, corresponding to the nucleotide exchange. On the membrane, MinE forms complexes with membrane-bound MinD at rate ked. In addition, the protein cytosolic dynamics are coupled to the membrane dynamics via various attachment and detachment processes. Cytosolic MinD–ATP is recruited to the membrane by membrane-bound MinD at rate kdD, and it can also bind to the membrane in an unassisted manner at rate kD. Similarly, cytosolic MinE is recruited to the membrane by membrane-bound MinD at rate kdE, forming a MinDE complex. The MinDE complexes split up at rate kde, releasing MinD–ADP into the cytosol, whereas MinE remains on the membrane (persistent binding). These isolated MinE proteins dissociate from the membrane at rate ke. The general corresponding partial differential equations for the persistent binding model read as follows:
The three former equations represent the protein dynamics in the bulk volume, and the three latter equations represent the protein dynamics on the reactive boundary (SLB and parts of the liposome membrane). On the reactive membrane, the Min proteins interact with each other and form complexes according to the reaction terms:
Here \(\tilde u_{\mathrm{DT}}\) and \(\tilde u_\mathrm{E}\) correspond to the respective MinD and MinE bulk concentrations right on the membrane, that is, \(\tilde u_i = u_i\left. \right|_\mathrm{m}\).
Finally, the bulk protein dynamics were coupled to the membrane dynamics through reactive boundary conditions, which are expressed by a net diffusive flux ∇nui|m, where ∇n represents a spatial derivative normal to the reactive boundary:
Geometry of static liposomes in simulation
Equations (1)–(3) were numerically solved in a three-dimensional (3D) box geometry using COMSOL Multiphysics 5.6. For simplicity and to ensure numerical stability, we used no-flux boundary conditions at the sides and top of the box. The bottom boundary (z = 0) represented the reactive membrane, obeying the reactive boundary conditions (equation (2)). We deformed the bottom boundary into a liposome-shaped geometry. In line with the observation that MinE only interacted with the SLB and the lower part of the liposome (Fig. 3a), we rendered the upper part of the deformed boundary inert. To simulate the symmetric vesicles, we chose a deformation that qualitatively resembled the experimental observation (Fig. 2a). In the simulation of the asymmetric vesicles, we chose a deformation that qualitatively resembled the experimentally observed liposome shape (Fig. 3a).
Coupling liposome motion to MinE reorganization
In contrast to the simulations of a static liposome, where membrane deformation represented the liposome, we used a phase-field approach to generate moving liposomes. The reactive boundary represents the SLB, whereas the volume occupied by the liposome is modelled as a localized repulsive potential V(x, y; t). Although this approach allows to dynamically change the liposome position, the Min proteins now cannot bind to the liposome membrane (excluded volume). However, as the excluded volume changes the local membrane-to-bulk ratio close to the liposome, the geometry effect leading to MinE accumulation around the vesicle is still valid. Thus, the phase-field approach is sufficient to study the Min pattern–liposome interaction.
To couple the cytosolic protein dynamics to the liposomes, we included a drift term in the partial differential equations:
Here ξ is a scaling factor, which determines the strength of the drift term. We chose ξ to be as large as possible without causing numerical instabilities, representing an impermeable liposome.
We simulated the pattern–liposome interactions in a simplified 2D box geometry. We manually linked the MinE protein concentration close to the liposome (ue) to the liposome position along the reactive membrane (xV(t)), whose dynamics are defined by
where the integral kernel \({{{\mathcal{S}}}}\left( {x,t} \right)\) limits the integral to the vicinity of the liposome (radius R):
The negative sign in equation (5a) stems from the experimental observation that the liposome (excluded volume) moves away from high MinE concentrations. The factor v is a phenomenological proportionality constant representing the coupling strength between the MinE concentration gradient and the liposome speed. We manually broke the symmetry by imposing an exponentially decaying movement to the excluded volume at the beginning of the simulation (first ~2,000 s). This manual symmetry breaking was designed to move the excluded volume by exactly 1 μm, so that any additional displacement of the excluded volume is a result of the coupling of the protein dynamics.
Simulation parameter dependence of liposome motion
The observed mechanochemical coupling depends—to a limited degree—on the reaction rates and, in a straightforward way, on the speed proportionality factor v, which we will discuss in the following. In general, distinct liposome motion characteristics are observed when the timescale of pattern formation does not match the timescale of liposome motion.
First, the reaction rates used in the default simulations were chosen such that stationary patterns form in 3D simulations of the Min system. Previous studies showed that the observed pattern type crucially depends on the Min concentrations in experiments14 and on the reaction rates in simulations36,38. Therefore, most parameters are not eligible for sweep focusing on the mechanochemical coupling, as this would entirely destroy the stationary pattern. However, the MinE detachment rate ke, which essentially determines the strength of the persistent binding, can be tuned over various orders of magnitude without destroying the stationary pattern. On an abstract level, ke affects the timescale at which the pattern can adapt to a change in geometry. For lower ke, the pattern adapts more quickly to the liposome, so that the manually broken symmetry is not maintained and the liposome comes to a halt (Extended Data Fig. 8c). In contrast, for faster MinE detachment, the liposome moves faster than the pattern can adapt, meaning that the liposome ‘hops’ on the MinE pattern, changing the direction of movement whenever a high MinE concentration is encountered (Extended Data Fig. 8a).
Second, the characteristic timescale of the liposome motion is exclusively determined by the speed proportionality factor v. Intuitively, the liposome speed should be directly proportional to v, which we numerically confirm in the early phase of liposome motion (Extended Data Fig. 7). However, when the timescale of liposome motion strongly deviates from the timescale of pattern formation, similar effects as for changing the MinE detachment rate are observed. In the approximate range of 5 × 10–4 s–1 < v < 10–2 s–1, persistent motion is observed. For v ≲ 5 × 10–4 s–1, the liposome movement is too slow, and chemical symmetry is restored before the liposome can pick up speed. For v ≳ 10–2 s–1, the liposome moves faster than the pattern can adapt, resulting in similar ‘hopping’ characteristics as observed for fast MinE detachment. It should be noted that the speed proportionality factor is a phenomenological parameter, and the actual timescale of liposome motion needs to be deduced from a microscopic model of mechanical interaction between the liposome, SLB and Min proteins, which we do not specify here due to a lack of experimental evidence.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data that support the plots within this paper and other findings of this study are available via the Harvard Dataverse67 at https://doi.org/10.7910/dvn/ahyyye and from the corresponding authors upon request.
Code availability
The code that supports the plots within this paper are described in the Methods and Supplementary Information and is available from the corresponding authors upon request.
References
Zhang, S. et al. Engineering motile aqueous phase-separated droplets via liposome stabilisation. Nat. Commun. 12, 1673 (2021).
Yang, Q. et al. Recent advances in motion control of micro/nanomotors. Adv. Intell. Syst. 2, 2000049 (2020).
Sun, J., Mathesh, M., Li, W. & Wilson, D. A. Enzyme-powered nanomotors with controlled size for biomedical applications. ACS Nano 13, 10191–10200 (2019).
Zhang, Y. & Hess, H. Chemically-powered swimming and diffusion in the microscopic world. Nat. Rev. Chem. 5, 500–510 (2021).
Gentile, K., Somasundar, A., Bhide, A. & Sen, A. Chemically powered synthetic ‘living’ systems. Chem 6, 2174–2185 (2020).
Yu, S. et al. Self-propelled Janus microdimer swimmers under a rotating magnetic field. Nanomaterials 9, 1672 (2019).
Santiago, I. & Simmel, F. C. Self-propulsion strategies for artificial cell-like compartments. Nanomaterials 9, 1680 (2019).
Keren, K. et al. Mechanism of shape determination in motile cells. Nature 453, 475–480 (2008).
Cao, Y., Ghabache, E. & Rappel, W. J. Plasticity of cell migration resulting from mechanochemical coupling. eLife 8, e48478 (2019).
Siton-Mendelson, O. & Bernheim-Groswasser, A. Toward the reconstitution of synthetic cell motility. Cell Adh. Migr. 10, 461–474 (2016).
Cao, Y. et al. Cell motility dependence on adhesive wetting. Soft Matter 15, 2043–2050 (2019).
Litschel, T. & Schwille, P. Protein reconstitution inside giant unilamellar vesicles. Annu. Rev. Biophys. 50, 525–548 (2021).
Halatek, J., Brauns, F. & Frey, E. Self-organization principles of intracellular pattern formation. Philos. Trans. R. Soc. B 373, 20170107 (2018).
Ramm, B., Heermann, T. & Schwille, P. The E. coli MinCDE system in the regulation of protein patterns and gradients. Cell. Mol. Life Sci. 76, 4245–4273 (2019).
Wettmann, L. & Kruse, K. The Min-protein oscillations in Escherichia coli: an example of self-organized cellular protein waves. Philos. Trans. R. Soc. B 373, 20170111 (2018).
Brauns, F. et al. Bulk-surface coupling identifies the mechanistic connection between Min-protein patterns in vivo and in vitro. Nat. Commun. 12, 3312 (2021).
Halatek, J. & Frey, E. Rethinking pattern formation in reaction–diffusion systems. Nat. Phys. 14, 507–514 (2018).
Hu, Z. & Lutkenhaus, J. Topological regulation of cell division in E. coli: spatiotemporal oscillation of MinD requires stimulation of its ATPase by MinE and phospholipid. Mol. Cell 7, 1337–1343 (2001).
Zhou, H. & Lutkenhaus, J. Membrane binding by MinD involves insertion of hydrophobic residues within the C-terminal amphipathic helix into the bilayer. J. Bacteriol. 185, 4326–4635 (2003).
Park, K.-T., Villar, M. T., Artigues, A. & Lutkenhaus, J. MinE conformational dynamics regulate membrane binding, MinD interaction, and Min oscillation. Proc. Natl Acad. Sci. USA 114, 7497–7504 (2017).
Ayed, S. H. et al. Dissecting the role of conformational change and membrane binding by the bacterial cell division regulator MinE in the stimulation of MinD ATPase activity. J. Biol. Chem. 292, 20732–20743 (2017).
Loose, M., Fischer-Friedrich, E., Ries, J., Kruse, K. & Schwille, P. Spatial regulators for bacterial cell division self-organize into surface waves in vitro. Science 320, 789–792 (2008).
Glock, P. et al. Stationary patterns in a two-protein reaction-diffusion system. ACS Synth. Biol. 8, 148–157 (2019).
Kretschmer, S., Heermann, T., Tassinari, A., Glock, P. & Schwille, P. Increasing MinD’s membrane affinity yields standing wave oscillations and functional gradients on flat membranes. ACS Synth. Biol. 10, 939–949 (2021).
Litschel, T., Ramm, B., Maas, R., Heymann, M. & Schwille, P. Beating vesicles: encapsulated protein oscillations cause dynamic membrane deformations. Angew. Chem. Int. Ed. 57, 16286–16290 (2018).
Fu, M., Franquelim, H. G., Kretschmer, S. & Schwille, P. Non-equilibrium large-scale membrane transformations driven by MinDE biochemical reaction cycles. Angew. Chem. Int. Ed. 60, 6496–6502 (2021).
Mietke, A., Julicher, F. & Sbalzarini, I. F. Self-organized shape dynamics of active surfaces. Proc. Natl Acad. Sci. USA 116, 29–34 (2019).
Tamemoto, N. & Noguchi, H. Pattern formation in reaction–diffusion system on membrane with mechanochemical feedback. Sci. Rep. 10, 19582 (2020).
Thalmeier, D., Halatek, J. & Frey, E. Geometry-induced protein pattern formation. Proc. Natl Acad. Sci. USA 113, 548–553 (2016).
Hardy, G. J., Nayak, R. & Zauscher, S. Model cell membranes: techniques to form complex biomimetic supported lipid bilayers via vesicle fusion. Curr. Opin. Colloid Interface Sci. 18, 448–458 (2013).
Gleisner, M. et al. Epsin N-terminal homology domain (ENTH) activity as a function of membrane tension. J. Biol. Chem. 291, 19953–19961 (2016).
Heermann, T., Steiert, F., Ramm, B., Hundt, N. & Schwille, P. Mass-sensitive particle tracking to elucidate the membrane-associated MinDE reaction cycle. Nat. Methods 18, 1239–1246 (2021).
Vecchiarelli, A. G., Li, M., Mizuuchi, M. & Mizuuchi, K. Differential affinities of MinD and MinE to anionic phospholipid influence Min patterning dynamics in vitro. Mol. Microbiol. 93, 453–463 (2014).
Netz, R. R. Electrostatistics of counter-ions at and between planar charged walls: from Poisson-Boltzmann to the strong-coupling theory. Eur. Phys. J. E 5, 557–574 (2001).
Komorowski, K. et al. Vesicle adhesion in the electrostatic strong-coupling regime studied by time-resolved small-angle X-ray scattering. Soft Matter 16, 4142–4154 (2020).
Halatek, J. & Frey, E. Highly canalized MinD transfer and MinE sequestration explain the origin of robust MinCDE-protein dynamics. Cell Rep. 1, 741–752 (2012).
Huang, K. C., Meir, Y. & Wingreen, N. S. Dynamic structures in Escherichia coli: spontaneous formation of MinE rings and MinD polar zones. Proc. Natl Acad. Sci. USA 100, 12724–12728 (2003).
Denk, J. et al. MinE conformational switching confers robustness on self-organized Min protein patterns. Proc. Natl Acad. Sci. USA 115, 4553–4558 (2018).
Loose, M., Fischer-Friedrich, E., Herold, C., Kruse, K. & Schwille, P. Min protein patterns emerge from rapid rebinding and membrane interaction of MinE. Nat. Struct. Mol. Biol. 18, 577–584 (2011).
Park, K. T. et al. The Min oscillator uses MinD-dependent conformational changes in MinE to spatially regulate cytokinesis. Cell 146, 396–407 (2011).
Vecchiarelli, A. G. et al. Membrane-bound MinDE complex acts as a toggle switch that drives Min oscillation coupled to cytoplasmic depletion of MinD. Proc. Natl Acad. Sci. USA 113, E1479–E1488 (2016).
Brauns, F., Halatek, J. & Frey, E. Phase-space geometry of mass-conserving reaction-diffusion dynamics. Phys. Rev. X 10, 041036 (2020).
Würthner, L. et al. Bridging scales in a multiscale pattern-forming system. Proc. Natl Acad. Sci. USA 119, e2206888119 (2022).
Shih, Y. L. et al. The N-terminal amphipathic helix of the topological specificity factor MinE is associated with shaping membrane curvature. PLoS ONE 6, e21425 (2011).
Zheng, M. et al. Self-assembly of MinE on the membrane underlies formation of the MinE ring to sustain function of the Escherichia coli Min system. J. Biol. Chem. 289, 21252–21266 (2014).
Horton, M. R., Höfling, F., Rädler, J. O. & Franosch, T. Development of anomalous diffusion among crowding proteins. Soft Matter 6, 2648–2656 (2010).
Burkart, T., Wigbers, M. C., Würthner, L. & Frey, E. Control of protein-based pattern formation via guiding cues. Nat. Rev. Phys. 4, 511–527 (2022).
Murrell, M. P. et al. Liposome adhesion generates traction stress. Nat. Phys. 10, 163–169 (2014).
Bartelt, S. M., Steinkuhler, J., Dimova, R. & Wegner, S. V. Light-guided motility of a minimal synthetic cell. Nano Lett. 18, 7268–7274 (2018).
Solon, J., Streicher, P., Richter, R., Brochard-Wyart, F. & Bassereau, P. Vesicles surfing on a lipid bilayer: self-induced haptotactic motion. Proc. Natl Acad. Sci. USA 103, 12382–12387 (2006).
Sommers, A. D., Brest, T. J. & Eid, K. F. Topography-based surface tension gradients to facilitate water droplet movement on laser-etched copper substrates. Langmuir 29, 12043–12050 (2013).
Ringgaard, S., van Zon, J., Howard, M. & Gerdes, K. Movement and equipositioning of plasmids by ParA filament disassembly. Proc. Natl Acad. Sci. USA 106, 19369–19374 (2009).
Ptacin, J. L. et al. A spindle-like apparatus guides bacterial chromosome segregation. Nat. Cell Biol. 12, 791–798 (2010).
Vecchiarelli, A. G., Neuman, K. C. & Mizuuchi, K. A propagating ATPase gradient drives transport of surface-confined cellular cargo. Proc. Natl Acad. Sci. USA 111, 4880–4885 (2014).
Ramm, B. et al. A diffusiophoretic mechanism for ATP-driven transport without motor proteins. Nat. Phys. 17, 850–858 (2021).
Cera, L. & Schalley, C. A. Under diffusion control: from structuring matter to directional motion. Adv. Mater. 30, e1707029 (2018).
Shiraki, Y. & Yoshida, R. Autonomous intestine-like motion of tubular self-oscillating gel. Angew. Chem. 124, 6216–6220 (2012).
Senoussi, A., Galas, J.-C. & Estevez-Torres, A. Programmed mechano-chemical coupling in reaction-diffusion active matter. Sci. Adv. 7, eabi9865 (2021).
Ramm, B., Glock, P. & Schwille, P. In vitro reconstitution of self-organizing protein patterns on supported lipid bilayers. J. Vis. Exp. 137, 58139 (2018).
Ernst, O. & Zor, T. Linearization of the Bradford protein assay. J. Vis. Exp. 38, 1918 (2010).
Steinkuhler, J., De Tillieux, P., Knorr, R. L., Lipowsky, R. & Dimova, R. Charged giant unilamellar vesicles prepared by electroformation exhibit nanotubes and transbilayer lipid asymmetry. Sci. Rep. 8, 11838 (2018).
Neumann, S., Pucadyil, T. J. & Schmid, S. L. Analyzing membrane remodeling and fission using supported bilayers with excess membrane reservoir. Nat. Protoc. 8, 213–222 (2013).
Weiss, M. Challenges and artifacts in quantitative photobleaching experiments. Traffic 5, 662–671 (2004).
Tarantino, N. et al. TNF and IL-1 exhibit distinct ubiquitin requirements for inducing NEMO–IKK supramolecular structures. J. Cell Biol. 204, 231–245 (2014).
Tinevez, J.-Y. & Herbert, S. The NEMO dots assembly: single-particle tracking and analysis. Bioimage Data Analysis Workflows 67–96 (2020).
Stalder, A. F. et al. Low-bond axisymmetric drop shape analysis for surface tension and contact angle measurements of sessile drops. Colloids Surf. A Physicochem. Eng. Asp. 364, 72–81 (2010).
Fu, M. Replication data for ‘Mechanochemical feedback loop drives persistent motion of liposomes’. Harvard Dataverse V3 https://doi.org/10.7910/dvn/ahyyye (2023).
Acknowledgements
We thank the MPIB Core Facility for assistance in protein purification. We thank P. Glock for labelling and measuring the concentration of KCK–MinE–His. We thank L. Würthner for helpful discussions and for providing code for the simulations. We acknowledge financial support from the German Research Foundation (DFG) through TRR 174 (project ID 269423233), SFB863 (project ID 111166240) and SFB1032 (project ID 201269156), as well as support from Germany’s Excellence Strategy, Excellence Cluster ORIGINS, EXC-2094-390783311. M.F. acknowledges support from the Humboldt Research Fellowship for Postdoctoral Researchers (ref. 3.5-1207791-CHN-HFST-P) and the Strategic Priority Research Program of the Chinese Academy of Sciences (grant no. XDB0480000). T.B. acknowledges support from the Joachim Herz Foundation. I.M. has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (Marie Skłodowska-Curie grant agreement no. 754388). I.M. acknowledges support from LMUexcellent, funded by the Federal Ministry of Education and Research (BMBF) and the Free State of Bavaria under the Excellence Strategy of the German Federal and State Governments. H.G.F. acknowledges support from the STARK project (project ID 46SKD023X), funded by the Federal Ministry for Economic Affairs and Energy (BMWi) and Saxon State Ministry of Science and Art (SMWK). P.S. acknowledges support from the research network MaxSynBio via a joint funding initiative of the German Federal Ministry of Education and Research (BMBF) and the Max Planck Society.
Funding
Open access funding provided by Max Planck Society.
Author information
Authors and Affiliations
Contributions
M.F., T.B., E.F. and P.S. conceived the study. T.B., I.M. and E.F. designed the theoretical analysis. T.B. conducted the theoretical analysis. M.F., H.G.F. and P.B. designed the experiments. M.F. performed all the experiments. A.M.-S. and M.R.-L. assisted with the supplementary experiments. M.F., T.B., E.F. and P.S. wrote the manuscript. All authors discussed and interpreted the results and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Physics thanks Jodi Camberg and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 MinD protein binding did not flatten the vesicle in the absence of SLBs.
When the coverslip was passivated with PLL-PEG (Poly(L-lysine)-graft-poly(ethyleneglycol), left column) and no SLBs formed, MinD protein binding (magenta) to the membrane (red) did not flatten the vesicle due to the lack of membrane adhesion. In contrast, vesicles flattened when an SLB was present (right column).
Extended Data Fig. 2 Diffusion of MinE is slower than that of MinD.
a) The fluorescence intensity recovery of MinE is slow (around minutes), and the diffusion of MinE decreased further at higher MinD concentration (2 experiments; n = 13, 14, 19, 18, 14, 12 from left to right). We applied an exponential fit to determine the diffusion coefficient (right); b) Diffusion coefficients of MinD (2 experiments; n = 12, 18, 24, 12, 12, 12 from left to right). The diffusion coefficient of MinD also decreased at high MinD concentration. For all the box figures, whiskers are 1.5 x IQR (interquartile range), median is shown as a black line, mean is shown as a red line.
Extended Data Fig. 3 Quasi-stationary labyrinth patterns at different MinE diffusion constants in simulation.
Snapshots of the stationary MinE patterns (total MinE concentration ue + ude shown in green) for varying MinE diffusion constants. All four simulations were initialized with the same initial conditions. Upon decreasing the MinE diffusion constant De with respect to that of MinD on the membrane, the protein patterns become more pronounced, meaning that low MinE diffusivity is needed to establish MinE protein patterns. From left to right: De = 0.013 μm2 s−1; De = 0.010 μm2 s−1; De = 0.008 μm2 s−1; De = 0.005 μm2 s−1. The diffusion constants for MinD is 0.013 μm2 s−1. Scale bar: 5 μm.
Extended Data Fig. 4 The inferred liposome motion direction fits well with the observed one during the whole motion process.
The liposome motion directions were inferred by correlating the centroids of the liposome and the MinE distribution (Methods). The inferred motion direction is closely correlated to the actual direction, indicating that the asymmetric MinE distribution may cause the persistent liposome motion. The experimental conditions were: a) normal liposome at Min buffer without MgCl2, 100 mM KCl; b) phase-separated liposome at Min buffer without MgCl2, 150 mM KCl; c) phase-separated liposome at Min buffer without MgCl2, 150 mM KCl.
Extended Data Fig. 5 Fluorescence intensity change of the Min proteins during liposome motion.
a) Snapshots of the liposome motion driven by the Min quasi-stationary pattern in a top-down view (top rows; MinD fluorescence shown in magenta, MinE shown in green) and from the side (bottom rows); b) The intensity of MinD at the liposome equator did not change notably after the initial binding process, indicating that the liposome dynamics did not result from the protein dynamics at the liposome body; c, d) At liposome-SLB contact areas, the protein intensities would increase during protein membrane binding as well as during membrane constriction (when membrane aggregated). To differentiate the fluorescence change during the time sequence, we calibrated the fluorescence intensities of Min proteins on the liposomes with that on the SLBs, and with that of the SLB membrane, that is, the protein intensities on one unit of SLB would not change with time and remained at 1. The calibrated intensities of MinD were the same at left and right sides of the liposome, but MinE accumulated faster on the left and drove the liposome to the right.
Extended Data Fig. 6 Kymograph of MinE protein pattern and liposome position.
Kymograph indicating persistent liposome motion accompanied by a redistribution of MinE proteins (green) in a 2D phase-field simulation. White areas represent the excluded volume (liposome). a) Only when the liposome motion is coupled to the Min patterns and vice versa, persistent liposome motion is observed. b) When the liposome motion is uncoupled from the Min proteins (v = 0, ξ > 0), the liposome does not move, but Min proteins still accumulate at the liposome-SLB contact area; c) When the Min protein dynamics are uncoupled from the liposome (ξ = 0, v > 0), the liposome movement decays quickly and does not affect the Min pattern.
Extended Data Fig. 7 Liposome movement depends on the mechanochemical coupling strength.
Simulated trajectories with different values of the speed proportionality factor v, and the default value is \(v_0 = 0.005\) [1/s]. The data were obtained by tracing the liposome center \(x_V\left( t \right)\) over the course of one simulation each. The initial conditions were identical for all simulations. a) Liposome position vs. rescaled time. The liposome trajectories approximately collapse upon rescaling time by the speed proportionality factor, \(t \to t \cdot v/v_0\). The trajectories are cropped when the liposome leaves the simulated area [0 μm, 60 μm]; b) For the same trajectories as in (a), the liposome speed vs. simulated time is plotted. No motion is observed for low proportionality factors, whereas quick movement is observed for large \(v\). To improve the legibility, the trajectories are cropped before the speed becomes negative for the first time.
Extended Data Fig. 8 Liposome movement characteristics depends on patterning time scale.
The liposome movement type changes when the Min pattern adapts to the liposome dynamic at different time scales. a) When MinE detachment rate is larger than the default value (\(k_{{{\mathrm{e}}}} = 10^{ - 3}\) [1/s]), the pattern cannot promptly adapt to the moving liposome, and the liposome “hops” between patches of high MinE concentration. This simulation result is in consistence with the experimental observation where the liposomes tend to hop between mesh cells rather than moving smoothly across the surface (Supplementary Video 1); b) At the default MinE detachment rate \(k_{{{\mathrm{e}}}} = 10^{ - 4}\) [1/s], persistent motion is observed; c) When MinE detachment rate is smaller than the default value (\(k_{{{\mathrm{e}}}} = 10^{ - 5}\) [1/s]), the pattern adapts to the moving liposome faster than the liposome can move, so that the chemical asymmetry is lost and no motion is observed.
Extended Data Fig. 9 Liposome motion speed in relation to contact angle.
a) Schematic representation of a moving liposome. Asymmetric MinE binding decreased the contact angle of the liposome rear part and consequently different traction pressure would drive liposome motion. b) Contact angles at the rear and front of moving liposomes (5 moving liposomes were analyzed, in different experimental conditions. All angles are given in degrees. Liposome 1: contact angle pair rear:front = 109.456:140.621; normal liposome at Min buffer without MgCl2, 100 mM KCl, 2.5 µM MinD-5 µM MinE. Liposome 2: contact angle pair rear:front = 80.072:111.618; normal liposome at Min buffer without MgCl2, 150 mM KCl, 2.5 µM MinD-5 µM MinE. Liposome 3: contact angle pair rear:front = 110.925:122.579; normal liposome at Min buffer without MgCl2, 100 mM KCl, 2.5 µM MinD-5 µM MinE; Liposome 4: contact angle pair rear:front = 83.244:103.342; normal liposome at Min buffer, 2.5 µM MinD-5 µM MinE); Liposome 5: contact angle pair rear:front = 97.928:124.056; normal liposome at Min buffer without MgCl2, 100 mM KCl, 2.5 µM MinD-5 µM MinE); For the box figure, whiskers are 1.5 x IQR (interquartile range), median is shown as a black line, mean is shown as a red line; c) The liposome velocities decreased with increasing contact angles (rear parts of moving liposomes) during liposome motion. Experimental conditions were all: normal liposome at Min buffer without MgCl2, 100 mM KCl, 2.5 µM MinD-5 µM MinE.
Extended Data Fig. 10 Rotation of phase-separated liposomes.
When an obstacle (located at the right edge of the imaged region) hindered the directional motion, the liposome rotated without moving forward. The labeled Ld domain was represented as red while MinE was represented as green. The arrows indicate rotation of the Lo/Ld interface. The Min buffer contained 25 mM Tris, pH 7.5, 150 mM KCl.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7, Tables 1–3 and discussion.
Supplementary Video 1
Three-dimensional illustration of liposome motion driven by Min quasi-stationary patterns. MinD was doped with 20 mol% mCherry–MinD (magenta), and MinE was doped with 20 mol% Atto488–KCK–MinE–His (green). The times are shown in hours. The buffer contained 25 mM Tris at pH 7.5, 100 mM KCl and in the absence of MgCl2. MinD and MinE were used at 2.5 and 5.0 µM protein concentrations, respectively.
Supplementary Video 2
Min travelling waves cannot drive liposome motion. MinD and His–MinE were used at 2 and 1 µM protein concentrations, respectively, to generate the travelling protein pattern. In this condition, the protein dynamics were not able to drive the motion of the liposomes. MinD was doped with 20 mol% eGFP–MinD (green) and the membrane was labelled with Atto655–PE (grey). The times are shown in minutes.
Supplementary Video 3
Two-dimensional illustration of liposome motion driven by Min quasi-stationary patterns. The experimental condition is the same as that in Supplementary Video 1. MinD was doped with 20 mol% mCherry–MinD (magenta), and MinE was doped with 20 mol% Atto488–KCK–MinE–His (green). The times are shown in hours. The buffer contained 25 mM Tris at pH 7.5, 100 mM KCl and in the absence of MgCl2. MinD and MinE were used at 2.5 and 5.0 µM protein concentrations, respectively.
Supplementary Video 4
Liposome motion at reduced MinE concentration (1.25 µM). MinD and MinE were used at 2.50 and 1.25 µM protein concentrations, respectively, and Min line and spot patterns were formed. In this condition, the motion percentage was reduced (Supplementary Fig. 5). MinD was doped with 20 mol% mCherry–MinD (magenta), and MinE was doped with 20 mol% Atto488–KCK–MinE–His (green). The times are shown in hours. The buffer contained 25 mM Tris (pH 7.5), 150 mM KCl and 5 mM MgCl2.
Supplementary Video 5
Liposome motion at reduced MinE concentration (2.5 µM). MinD and MinE were used at 2.5 and 2.5 µM protein concentrations, respectively, and Min line patterns were formed. In this condition, the motion percentage was reduced (Supplementary Fig. 5). MinD was doped with 20 mol% mCherry–MinD (magenta), and MinE was doped with 20 mol% Atto488–KCK–MinE–His (green). The times are shown in hours. The buffer contained 25 mM Tris at pH 7.5, 150 mM KCl and 5 mM MgCl2.
Supplementary Video 6
Liposome motion at reduced MinD concentration (1 µM). MinD and MinE were used at 1 and 5 µM protein concentrations, respectively, and the Min pattern was no longer stable. In this condition, liposomes did not move. MinD was doped with 20 mol% mCherry–MinD (magenta), MinE was doped with 20 mol% Atto488–KCK–MinE–His (green) and the membrane was labelled with Atto655–PE (red). The times are shown in hours. The buffer contained 25 mM Tris at pH 7.5, 150 mM KCl and 5 mM MgCl2.
Supplementary Video 7
Liposome beads (2 µM) did not move. Liposome beads exhibit restricted membrane deformation ability, and they did move during Min protein pattern formation. The bead sizes of 1 and 5 µM were also tested and the data are available on request. MinD and MinE were used at 2.5 and 5.0 µM protein concentrations, respectively. MinD was doped with 20 mol% mCherry–MinD (magenta), MinE was doped with 20 mol% Atto488–KCK–MinE–His (green) and the membrane was labelled with Atto655–PE (red). The times are shown in hours. The buffer contained 25 mM Tris at pH 7.5, 150 mM KCl and 5 mM MgCl2.
Supplementary Video 8
Surfing of a phase-separated liposome driven by Min quasi-stationary patterns. MinD was doped with 20 mol% mCherry–MinD (magenta), and MinE was doped with 20 mol% Atto488–KCK–MinE–His (green). Min proteins were bound to the Ld domain. The times are shown in hours. The lipid composition was 40.0 mol% DOPC, 20.0 mol% DOPG, 20.0 mol% DSPC, 20.0 mol% cholesterol and 0.1 mol% Atto655–PE (final lipid concentration, 1 mg ml−1). The buffer contained 25 mM Tris and 150 mM KCl and the pH is 7.5. MinD and MinE were used at 2.5 and 5.0 µM protein concentrations, respectively.
Supplementary Video 9
Motion of phase-separated liposomes driven by Min quasi-stationary patterns. MinD was doped with 20 mol% mCherry–MinD (magenta), MinE was doped with 20 mol% Atto488–KCK–MinE–His (green) and the membrane was labelled with Atto655–PE (red). Min proteins were bound to the Ld domain. The times are shown in hours. The lipid composition was 40.0 mol% DOPC, 20.0 mol% DOPG, 20.0 mol% DSPC, 20.0 mol% cholesterol and 0.1 mol% Atto655–PE (final lipid concentration, 1 mg ml−1). The buffer contained 25 mM Tris and 150 mM KCl and the pH is 7.5. MinD and MinE were used at 2.5 and 5.0 µM protein concentrations, respectively.
Supplementary Video 10
Motion of phase-separated liposomes driven by Min quasi-stationary patterns. MinD was doped with 20 mol% mCherry–MinD (magenta), MinE was doped with 20 mol% Atto488–KCK–MinE–His (green) and the membrane was labelled with Atto655–PE (red). Min proteins were bound to the Ld domain. The times are shown in hours. The lipid composition was 40.0 mol% DOPC, 20.0 mol% DOPG, 20.0 mol% DSPC, 20.0 mol% cholesterol and 0.1 mol% Atto655–PE (final lipid concentration, 1 mg ml−1). The buffer contained 25 mM Tris and 150 mM KCl and the pH is 7.5. MinD and MinE were used at 2.5 and 5.0 µM protein concentrations, respectively.
Supplementary Video 11
Motion of phase-separated liposomes driven by Min quasi-stationary patterns. MinD was doped with 20 mol% mCherry–MinD (magenta), MinE was doped with 20 mol% Atto488–KCK–MinE–His (green) and the membrane was labelled with Atto655–PE (red). Min proteins were bound to the Ld domain. The times are shown in hours. The lipid composition was 40.0 mol% DOPC, 20.0 mol% DOPG, 20.0 mol% DSPC, 20.0 mol% cholesterol and 0.1 mol% Atto655–PE (final lipid concentration, 1 mg ml−1). The buffer contained 25 mM Tris and 150 mM KCl and the pH is 7.5. MinD and MinE were used at 2.5 and 5.0 µM protein concentrations, respectively.
Supplementary Video 12
Rotation of a phase-separated liposome driven by Min quasi-stationary patterns. MinD and MinE were used at 2.5 and 5.0 µM protein concentrations, respectively. MinE was doped with 20 mol% Atto488–KCK–MinE–His (green). The times are shown in hours. The lipid composition was 40.0 mol% DOPC, 20.0 mol% DOPG, 20.0 mol% DSPC, 20.0 mol% cholesterol and 0.1 mol% Atto655–PE (final lipid concentration, 1 mg ml−1), and the Ld domain was labelled with Atto655–PE (red). The buffer contained 25 mM Tris and 150 mM KCl and the pH is 7.5.
Supplementary Video 13
Liposome was constricted during motion at high MgCl2 concentration. The buffer contained 25 mM Tris, 5 mM MgCl2 and 150 mM KCl and the pH is 7.5. MinD and MinE were used at 2.5 and 5.0 µM protein concentrations, respectively. MinD was doped with 20 mol% mCherry–MinD (magenta), MinE was doped with 20 mol% Atto488–KCK–MinE–His (green) and the membrane was labelled with Atto655–PE (red). The times are shown in hours.
Supplementary Video 14
Liposome motion at 10 mol% DOPG. The lipid composition was 20.0 mol% DOPE, 70.0 mol% DOPC, 10.0 mol% DOPG and 0.1 mol% Atto655–PE. In this condition, the Min pattern was no longer stable and turned into a wave with time, inducing the stoppage of liposome motion. MinD and MinE were used at 2.5 and 5.0 µM protein concentrations, respectively. MinD was doped with 20 mol% mCherry–MinD (magenta) and MinE was doped with 20 mol% Atto488–KCK–MinE–His (green). The times are shown in hours. The buffer contained 25 mM Tris, 5 mM MgCl2 and 150 mM KCl and the pH is 7.5.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fu, M., Burkart, T., Maryshev, I. et al. Mechanochemical feedback loop drives persistent motion of liposomes. Nat. Phys. 19, 1211–1218 (2023). https://doi.org/10.1038/s41567-023-02058-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41567-023-02058-8
This article is cited by
-
Present and future of synthetic cell development
Nature Reviews Molecular Cell Biology (2024)
-
The intersection of bottom-up synthetic cell engineering and nanobiotechnology
Nature Nanotechnology (2024)
-
Cell-free expression with a quartz crystal microbalance enables rapid, dynamic, and label-free characterization of membrane-interacting proteins
Communications Biology (2024)
-
Synthetic cells on the move
Nature Physics (2023)