Abstract
The ongoing monkeypox virus (MPXV) outbreak is the largest ever recorded outside of Africa. We isolated and sequenced a virus from the first clinical MPXV case diagnosed in France (May 2022). We report that tecovirimat (ST-246), a US Food and Drug Administration approved drug, is efficacious against this isolate in vitro at nanomolar concentrations, whereas cidofovir is only effective at micromolar concentrations. Our results support the use of tecovirimat in ongoing human clinical trials.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
All data supporting the findings of this study are available in the manuscript. The MPXV genomic data analysed here are deposited in GenBank (https://www.ncbi.nlm.nih.gov/genbank/, ON755039). The raw sequencing data of isolate MPXV/France/IRBA2211i/2022 have been deposited in the sequence read archive (https://www.ncbi.nlm.nih.gov/sra) under Accession PRJNA864836. The isolate MPXV/France/IRBA2211i/2022 is available upon request, as it may be useful for evaluating diagnostic tools, antivirals or vaccines (contact: irba-cnropv.accueil.fct@def.gouv.fr). Source data are provided with this paper.
References
McCollum, A. M. & Damon, I. K. Human monkeypox. Clin. Infect. Dis. 58, 260–267 (2014).
Monkeypox Outbreak (WHO, 2022).
Antinori, A. et al. Epidemiological, clinical and virological characteristics of four cases of monkeypox support transmission through sexual contact, Italy, May 2022. Euro Surveill. 27, 220042 (2022).
Basgoz, N. et al. Case 24-2022: a 31-year-old man with perianal and penile ulcers, rectal pain, and rash. N. Engl. J. Med. 387, 547–556 (2022).
Girometti, N. et al. Demographic and clinical characteristics of confirmed human monkeypox virus cases in individuals attending a sexual health centre in London, UK: an observational analysis. Lancet Infect. Dis. 22, 1321–1328 (2022).
Scaramozzino, N. et al. Real-time PCR to identify variola virus or other human pathogenic orthopox viruses. Clin. Chem. 53, 606–613 (2007).
Li, Y., Zhao, H., Wilkins, K., Hughes, C. & Damon, I. K. Real-time PCR assays for the specific detection of monkeypox virus West African and Congo Basin strain DNA. J. Virol. Methods 169, 223–227 (2010).
Happi, C. et al. Urgent need for a non-discriminatory and non-stigmatizing nomenclature for monkeypox virus. PLoS Biol. 20, e3001769 (2022).
Likos, A. M. et al. A tale of two clades: monkeypox viruses. J. Gen. Virol. 86, 2661–2672 (2005).
De Baetselier, I. et al. Asymptomatic monkeypox virus infections among male sexual health clinic attendees in Belgium. Preprint at medRxiv https://doi.org/10.1101/2022.07.04.22277226 (2022).
Miura, F. et al. Estimated incubation period for monkeypox cases confirmed in the Netherlands, May 2022. Euro Surveill. 27, 2200448 (2022).
Huggins, J. et al. Nonhuman primates are protected from smallpox virus or monkeypox virus challenges by the antiviral drug ST-246. Antimicrob. Agents Chemother. 53, 2620–2625 (2009).
Jordan, R. et al. ST-246 antiviral efficacy in a nonhuman primate monkeypox model: determination of the minimal effective dose and human dose justification. Antimicrob. Agents Chemother. 53, 1817–1822 (2009).
Grosenbach, D. W. et al. Oral tecovirimat for the treatment of smallpox. N. Engl. J. Med. 379, 44–53 (2018).
Delaune, D. & Iseni, F. Drug development against smallpox: present and future. Antimicrob. Agents Chemother. 64, e01683-19 (2020).
Adler, H. et al. Clinical features and management of human monkeypox: a retrospective observational study in the UK. Lancet Infect. Dis. 22, 1153–1162 (2022).
Tecovirimat SIGA (EMA, 2022).
Duraffour, S. et al. ST-246 is a key antiviral to inhibit the viral F13L phospholipase, one of the essential proteins for orthopoxvirus wrapping. J. Antimicrob. Chemother. 70, 1367–1380 (2015).
Yang, G. et al. An orally bioavailable antipoxvirus compound (ST-246) inhibits extracellular virus formation and protects mice from lethal orthopoxvirus challenge. J. Virol. 79, 13139–13149 (2005).
Lederman, E. R. et al. Progressive vaccinia: case description and laboratory-guided therapy with vaccinia immune globulin, ST-246, and CMX001. J. Infect. Dis. 206, 1372–1385 (2012).
Reed, L. J. & Muench, H. A simple method of estimating fifty per cent endpoints. Am. J. Epidemiol. 27, 493–497 (1938).
Isidro, J. et al. Phylogenomic characterization and signs of microevolution in the 2022 multi-country outbreak of monkeypox virus. Nat. Med. 28, 1569–1572 (2022).
Hadfield, J. et al. Nextstrain: real-time tracking of pathogen evolution. Bioinformatics 34, 4121–4123 (2018).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35, 518–522 (2018).
Nalca, A. et al. Evaluation of orally delivered ST-246 as postexposure prophylactic and antiviral therapeutic in an aerosolized rabbitpox rabbit model. Antiviral Res. 79, 121–127 (2008).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Acknowledgements
We thank all the healthcare workers, public health employees and scientists involved in the MPX outbreak response; the technical skills of C. Castellarin for preparing the samples for transmission electron microscopy; and the authors, and originating and submitting laboratories of the Orthopoxvirus sequences deposited in GenBank. This work used the computational and storage services provided by the IT department at Institut Pasteur, Paris.
The E.S.-L. laboratory acknowledges funding from the INCEPTION programme (Investissements d’Avenir grant ANR-16-CONV-0005), the NIH PICREID programme (Award Number U01AI151758) and the Labex IBEID (ANR-10-LABX-62-IBEID). O.F., A.F. and J.-N.T. acknowledge the support from Santé Publique France for the CNR-LE Orthopoxvirus. F.I. acknowledges funding from the Direction Générale de l’Armement (DGA) (Biomedef PDH-2-NBC-5-B-4120). F.G. acknowledges funding from the Agence Innovation Défense (AID) (RAPID DENALPOVIR 192906106).
Author information
Authors and Affiliations
Contributions
F.I., J.-N.T. and E.S.-L. conceived and designed the experiments; J.L. and R.K. provided patient care; O.F., A.F. and G.F.-V. conducted virus isolation, production and titration; A.-L.F., C.E. and X.H. conducted electron microscopy; O.G. and F.N. performed sequencing; F.I. and G.F.-V. performed MPXV antiviral assay; F.G. and E.M. conducted VACV antiviral assay; D.W.G. and D.E.H. provided antiviral material; A.B. and E.S.-L. performed phylogenetic analysis; J.-N.T. and E.S.-L. wrote the original draft, and reviewed and edited the manuscript with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
F.G. and E.M. are employed by NeoVirTech SAS. F.G. is a shareholder of NeoVirTech SAS. Tecovirimat is manufactured by SIGA Technologies. D.E.H. and D.W.G. are employees of SIGA Technologies and hold shares of SIGA stock. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
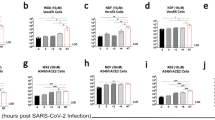
Extended Data Fig. 1 ST-246 inhibits VACV ANCHORTM-GFP propagation at nM concentration.
a, Reconstruction of 12 imaging fields captured on a CX7 CellInsight HCS microscope 48 h post infection with indicated concentration of ST-246. Blue: nuclei, green VACV ANCHORTM. (b,c) Quantification of the impact of ST-246 on the number of infected cells at 48 h (b) and 72 h (c) post-infection. IC50 and R2 are indicated. Data are presented as mean values +/- SD. Experiments were performed once in triplicate.
Extended Data Fig. 2 VP37 (F13L homologs) amino acid analysis.
VP37 (F13L homologs) amino acid alignment to MPVX clade IIb shows multiple differences, marked with black vertical bars, with other orthopoxviruses susceptible to tecovirimat. The E353K mutation in VP37 is a signature substitution of MPXV lineage B.1 (2022). Experimentally generated and verified mutations associated with resistance to ST-246 are marked in red below the alignment as well as in the corresponding OPV species in which they have been experimentally tested (vertical red bars). Importantly, these mutations (in red) are annotated in the alignment in the corresponding positions, while they do not exist in the original sequences.
Extended Data Fig. 3 Cytopathic effect on Vero cell line of MPXV/France/IRBA2211i/2022 strain (optical microscopy 20x).
Experiments were performed twice.
Supplementary information
Source data
Source Data Fig. 1
Source data.
Source Data Extended Data Fig. 1
Source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Frenois-Veyrat, G., Gallardo, F., Gorgé, O. et al. Tecovirimat is effective against human monkeypox virus in vitro at nanomolar concentrations. Nat Microbiol 7, 1951–1955 (2022). https://doi.org/10.1038/s41564-022-01269-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01269-8
This article is cited by
-
Identification of the myxobacterial secondary metabolites Aurachin A and Soraphinol A as promising inhibitors of thymidylate kinase of the Monkeypox virus
Molecular Diversity (2024)
-
Fluorescence-enhanced dual signal lateral flow immunoassay for flexible and ultrasensitive detection of monkeypox virus
Journal of Nanobiotechnology (2023)
-
Mpox virus infection and drug treatment modelled in human skin organoids
Nature Microbiology (2023)
-
Mpox (formerly monkeypox): pathogenesis, prevention, and treatment
Signal Transduction and Targeted Therapy (2023)