Abstract
Frequent outbreaks of coronaviruses underscore the need for antivirals and vaccines that can counter a broad range of coronavirus types. We isolated a human antibody named 76E1 from a COVID-19 convalescent patient, and report that it has broad-range neutralizing activity against multiple α- and β-coronaviruses, including the SARS-CoV-2 variants. 76E1 also binds its epitope in peptides from γ- and δ-coronaviruses. 76E1 cross-protects against SARS-CoV-2 and HCoV-OC43 infection in both prophylactic and therapeutic murine animal models. Structural and functional studies revealed that 76E1 targets a unique epitope within the spike protein that comprises the highly conserved S2’ site and the fusion peptide. The epitope that 76E1 binds is partially buried in the structure of the SARS-CoV-2 spike trimer in the prefusion state, but is exposed when the spike protein binds to ACE2. This observation suggests that 76E1 binds to the epitope at an intermediate state of the spike trimer during the transition from the prefusion to the postfusion state, thereby blocking membrane fusion and viral entry. We hope that the identification of this crucial epitope, which can be recognized by 76E1, will guide epitope-based design of next-generation pan-coronavirus vaccines and antivirals.
Similar content being viewed by others
Main
On the basis of antigenic and genetic criteria, coronaviruses are organized into four genera: α-, β-, γ- and δ- coronaviruses1. There are three major public health threats caused by highly pathogenic coronaviruses: severe acute respiratory syndrome coronavirus (SARS-CoV) in 2002–20032,3, Middle East respiratory syndrome coronavirus (MERS-CoV) in 20124 and SARS-CoV-2, which was first identified in 20195,6. The COVID-19 pandemic caused by SARS-CoV-2 has had a profound impact on the world economy and global health. In addition, several common epidemic human coronaviruses including the β-coronaviruses HCoV-OC43, HCoV-HKU1 and the distantly related α-coronaviruses HCoV-229E and HCoV-NL63 circulate annually and cause mild or moderate upper respiratory diseases7,8. The recurrent spillover events of coronaviruses into humans highlight the need to develop broad therapeutics and preventions.
One current research focus is the mutation of the SARS-CoV-2 during transmission9. A series of mutants have been identified within the spike (S) protein, which may weaken the protective activity of vaccines or antibodies that were designed on the basis of the original SARS-CoV-210,11. The identification of resistant variants suggests potential risks in the application of current vaccines and antibodies12,13,14.
Most neutralizing antibodies against SARS-CoV-2, SARS-CoV and MERS-CoV target the receptor-binding domain (RBD) of S1, thus blocking viral attachment to target cells. However, RBD antibodies are usually subgenus or strain specific, and pose a selective pressure causing the rapid emergence of new variants15,16,17,18. Nevertheless, only a few RBD antibodies have shown a narrow cross-reactivity, which is limited within the β-coronavirus genus19,20,21. Broad neutralizing antibodies (bnAbs) targeting conserved functional domains of viral surface proteins have been discovered to protect against global circulating viruses, such as influenza A virus (IAV), hepatitis C virus (HCV) and human immunodeficiency virus (HIV)22,23,24,25,26,27,28. The S2 subunit is more conserved than the S1 subunit among diverse coronaviruses. S2 antibodies (B6 and S2P6) have shown cross-reactivity to β-coronaviruses, but not to α-coronaviruses as reported by targeting the conserved stem helix region, prevent S2 subunit refolding from the pre- to the postfusion state and thus block virus entry by inhibiting membrane fusion21,29,30,31,32. Here we report a human antibody 76E1 with cross-neutralization to the representative strains from α- and β-coronavirus and even has cross-binding activity to the peptides containing the antibody epitope from γ- and δ-coronavirus genera. This antibody showed cross-protection in mice against SARS-CoV-2 and phylogenetically distant HCoV-OC43. Structural and functional analyses indicate that the epitope recognized by 76E1, including the fusion peptide and the S2’ site, was highly conserved among α-, β-, γ- and δ-coronaviruses. The recognition pattern is also interesting as the epitope is partially buried in the structure of the prefusion S trimer but could be exposed during the receptor-binding process. 76E1 inhibits membrane fusion possibly by blocking S2’ cleavage. The identification of the conserved epitopes will guide the design of pan-coronavirus vaccines and antivirals against both current and future emerging SARS-CoV-2 variants and other coronavirus genera.
Results
Isolation of S2-targeting human antibodies
We initially measured the antigen binding potencies of the plasma from five COVID-19 convalescent patients as compared with that of healthy people. The S2 regions are conserved among multiple human coronaviruses (Extended Data Fig. 1a). Antibodies that were cross-reactive to the S2 regions of SARS-CoV-2, SARS-CoV and MERS-CoV were detected, especially in sample 76 (Extended Data Fig. 1b–d). The tested plasmas also reacted well to the S proteins of HCoV-NL63, HCoV-OC43, HCoV-229E and HCoV-HKU-1, with higher antibody endpoint titres than the healthy donors’ plasmas (Extended Data Fig. 1e), indicating the potential existence of bnAbs, mostly targeting the S2 region in COVID-19 convalescent patients.
We then conducted single-cell PCR experiments to isolate human monoclonal antibodies (mAbs) from memory B cells of the COVID-19 convalescent patients by using the SARS-CoV-2 extracellular domain of S protein (S-ECD), without introduction of any mutations and modifications, as the bait. CD19+IgG+SARS-CoV-2 S-ECD+-specific single memory B cells were isolated by flow cytometry. The mAbs generated directly from single memory B cells were screened by the binding activities to RBD, S1, S2 and full S-ECD proteins, followed by assessing the pseudovirus neutralizing activities against SARS-CoV-2, SARS-CoV and MERS-CoV (Supplementary Tables 1 and 2). We identified 25 antibodies, 9 of which reacted to the SARS-CoV-2 S2 (Supplementary Tables 1 and 2). Three antibodies (555D6, 125C1 and 76E1) showed cross-reactivity to both SARS-CoV and MERS-CoV S2/S-ECD. Most of the S2 antibodies could not neutralize SARS-CoV-2 pseudovirus, except for 76E1 and 555E6 which had IC50 values (the concentration of antibody required for 50% viral inhibition) of 0.418 µg ml−1 and 9.79 µg ml−1, respectively (Supplementary Table 1). 76E1 was derived from donor 76 and engages IGHV3-43*02 and IGLV2-8*01 as the germline genes (Supplementary Table 3).
76E1 neutralizes SARS-CoV-2 and its variants
76E1 was selected for further in-depth characterization and was found to react well to the S-ECD proteins of SARS-CoV-2 variants of concern including B.1.1.7 (Alpha), B.1.351 (Beta), P.1 (Gamma), B.1.617.1 (Kappa), B.1.617.2 (Delta) and B.1.1.529 (Omicron, BA.1) (Fig. 1a and Extended Data Fig. 2). We subsequently observed that 76E1 could neutralize all the pseudotyped SARS-CoV-2 variants tested (Fig. 1b). 76E1 also showed comparable potency in Vero-E6, HeLa-hACE2 (human ACE2) and Calu-3 cells against authentic SARS-CoV-2, with IC50 values ranging from 0.373 to 0.727 μg ml−1 (Fig. 1c).
a, The EC50 values of 76E1 binding to the soluble S-ECD proteins of SARS-CoV-2 and its variants of concern, including B.1.1.7, P.1, B.1.351, B.1.617.1, B.1.617.2 and B.1.1.529 (Omicron BA.1). An IAV antibody 28-1253 was included as isotype control (Isoctrl). b, 76E1-mediated neutralization of pseudotyped SARS-CoV-2 and its variants in 293T-hACE2 cells. Unrelated VSV-G pseudovirus was included as a negative control. c, Authentic SARS-CoV-2 neutralization of 76E1 in Vero-E6, HeLa-hACE2 and Calu-3 target cells. n = 3. Representative data are shown from 2 independent experiments (a–c). d–f, Protective efficacy of 76E1 against authentic SARS-CoV-2 infection in hACE2 transgenic mice. d, The mouse study outline. Prophylaxis: the mice were infused with 15 mg kg−1 76E1 (n = 7) or PBS (n = 4) intraperitoneally 1 d before inoculating with 1 × 104 p.f.u. SARS-CoV-2 via intranasal infection. Therapy: the mice were treated with 25 mg kg−1 76E1 8 h after SARS-CoV-2 infection (n = 4). The prophylactic and therapeutic experiments were concurrently conducted and the PBS-administered mice were included as control for both experiments. The mice body weight from prophylactic (e) and therapeutic (f) groups were monitored for 3 d after infection. Lung virus titres of each mouse at 3 d.p.i. were determined by qRT-PCR with 3 replicates. Results are depicted as mean ± s.d. (c,e,f). Two-sided unpaired t-test; ***P < 0.001, ****P < 0.0001.
To evaluate the prophylactic efficacy of 76E1 in vivo, human ACE2 transgenic mice were intraperitoneally (i.p.) administered 15 mg kg−1 mAbs or PBS 24 h before challenge with a dose of 104 p.f.u. SARS-CoV-2 via intranasal inoculation. All mice were monitored for body weight daily and euthanized at 3 d post-infection (d.p.i) to allow for measurement of lung virus titres (Fig. 1d). Mice pretreated with 15 mg kg−1 76E1 showed minor weight loss and apparently reduced (104 times) lung viral titres compared with control mice (Fig. 1e). 76E1 at 50 and 150 mg kg−1 also exhibited similar protective efficacy in weight loss and lung virus titres (Extended Data Fig. 3). For the therapeutic experiment, mice were treated with 25 mg kg−1 mAbs 8 h after intranasal challenge with 104 p.f.u. of SARS-CoV-2 (Fig. 1d) and found to display minor weight loss and ~103 times lower lung virus titres than control mice (Fig. 1f). We conclude that 76E1 treatment can reduce virus burden in mice infected by SARS-CoV-2.
Broad neutralization of multiple human coronaviruses by 76E1
Seven coronaviruses have historically spilled over into humans and caused severe or mild diseases. We further assessed the broad reactivity of 76E1 to S-ECD proteins of seven human coronaviruses consisting of two α-coronaviruses (HCoV-229E and HCoV-NL63) and five β-coronaviruses (SARS-CoV-2, SARS-CoV, MERS-CoV, HCoV-OC43 and HCoV-HKU1) (Fig. 2a). 76E1 bound to all the S-ECD proteins tested with high activities (Fig. 2b and Extended Data Fig. 2). Moreover, 76E1 neutralized the pseudotyped or authentic coronaviruses, with IC50 values ranging from 0.373 to 4.821 µg ml−1 (Fig. 2c–h). To determine the cross-protective efficacy of 76E1 on an HCoV-OC43 infection mouse model, new-born mice were prophylactically treated with 15 or 50 mg kg−1 of 76E1, 50 mg kg−1 of isotype control antibody or with PBS, before challenge with a lethal dose of HCoV-OC43. The weight loss and survival rate were monitored daily for 14 d. Parallel groups of mice were euthanized at day 5 to determine viral titre in brains. We found that 50 mg kg−1 76E1 was sufficient to protect mice from weight loss and death, while the body weight of mice pretreated with 15 mg kg−1 slightly decreased since day 7 and recovered soon after at day 9, causing 66.7% protection (Fig. 2i,j). For the therapeutic experiment, 50 mg kg−1 76E1 resulted in 50% survival protection in mice (Fig. 2l,m). Moreover, 76E1 treatment also resulted in a significant reduction in brain viral loads on the 5th day, compared with the isotype control and PBS groups (Fig. 2k,n).
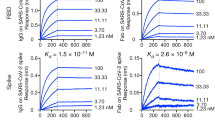
a, Phylogenetic tree of 28 coronaviruses constructed via MEGA (Molecular Evolutionary Genetics Analysis) and maximum likelihood analysis of full S amino acid sequences downloaded from the NCBI database. Seven human coronaviruses are denoted in filled red diamonds. b, Binding affinity of 76E1 to soluble S-ECD proteins of the seven human coronaviruses measured by biolayer interferometry (BLI) and enzyme linked immunosorbent assay (ELISA). n.d., not determined. c–e, 76E1-mediated neutralization of pseudotyped SARS-CoV in 293T-hACE2 cells, MERS-CoV and HCoV-229E in HuH-7 cells. n = 3. f–h, 76E1-mediated neutralization of authentic HCoV-OC43 in RD cells, HCoV-NL63 in LLC-MK2 cells and HCoV-229E in HuH-7 cells. n = 3. i–k, Prophylactic efficacy of 76E1 in mice against HCoV-OC43. New-born mice were treated with 76E1, isotype control antibody (15 or 50 mg kg−1) or PBS 24 h before challenge with a lethal dose of HCoV-OC43. Body weight (i) and survival rate (j) were monitored for 14 d after infection. n = 6. k, Virus titre in brain of each mouse was determined by calculating CPE on RD cells and expressed as TCID50 g−1. l–n, Protective efficacy of 76E1 treatment after HCoV-OC43 infection in mice. New-born mice were treated with 76E1, isotype control antibody (50 mg kg−1) or PBS 2 h after challenge with HCoV-OC43. Body weight (l) and survival rate (m) were monitored for 14 d after infection. n = 6. n, Virus titre in brain of each mouse. n = 5. Representative data from 2 independent experiments are shown (b–n). Results are depicted as mean ± s.d. Two-sided unpaired t-tests; **P < 0.01, ****P < 0.0001; NS, not significant.
Structural and functional basis for the broad reactivity of 76E1
We initially found that 76E1 reacts well to the denatured S-ECD protein (Extended Data Fig. 2e), indicating that this antibody might recognize a continuous amino acid sequence. We aligned amino acid sequences of the S proteins from 2 α-coronaviruses and 5 β-coronaviruses, and identified 7 conserved sequences in the S2 subunit. Among these conserved sequence motifs, only peptide809–833 (SARS-CoV-2 numbering) shows reactivity to 76E1, indicating that the S peptide809–833 comprises the epitope recognized by 76E1 (Fig. 3a, Extended Data Fig. 4 and Supplementary Table 4). To investigate the molecular basis for the broad specificity of 76E1 and the interaction between the 76E1 fragment of antigen binding (Fab) and the epitope, we carried out crystal structural study of the 76E1 Fab in complex with the S peptide809–833. The crystal structure of the 76E1 Fab-peptide complex was solved at a resolution of 2.6 Å (Supplementary Table 5). There are two Fab-peptide complexes in an asymmetry unit, related by a 2-fold non-crystallographic symmetry axis. The overall structure of the 76E1 Fab shows a canonical β sandwich immunoglobulin fold, with the heavy chain folding into the VH and CH domains and the light chain into the VL and CL domains, and an elbow angle of ~210° (Fig. 3b). Residues 812–826 of the S peptide are well defined in the electron density map (Fig. 3c). Disordering of the residues at both ends of the peptide indicates that these residues have relatively higher flexibility and are not required for binding. This is consistent with the isothermal titration calorimetry (ITC) analysis results showing that the 76E1 Fab has a higher binding affinity (about 9-fold) for peptide809–828 and peptide809–833 than peptide809–823 (Extended Data Fig. 5).
a, ELISA-based binding activity of 76E1 to 7 conserved peptides within 7 human coronaviruses. Data are presented as mean values of duplicates. b, Overall structure of the 76E1 Fab-peptide complex in ribbon diagram. Heavy and light chains of the 76E1 Fab are shown in light blue and pink, respectively. The SARS-CoV-2 S peptide809–833 is shown in green. c, Representative simulated annealing composite omit map (contoured at 1.0 σ level) for the peptide. d, Structure of the Fab-peptide interaction interface showing that the peptide assumes an α-helical conformation and is embedded into a cleft formed by the Fab CDR loops H1, H2, H3 and L3. The interacting residues of the peptide are shown in stick model. e, Hydrogen-bonding interactions between the peptide and the Fab. f, Residues Leu822 and Phe823 of the peptide are located in a hydrophobic pocket of the Fab. g, ELISA-based alanine scanning on the S peptide809–833 of SARS-CoV-2, shown as EC50 values (ng ml−1). h, Sequence alignment of peptide809–833 (SARS-CoV-2 numbering) among 28 coronaviruses from α, β, γ and δ genera. The epitope residues are shown in red colour. i, ELISA-based binding activity of 76E1 to the S peptides809–833 of coronavirus from α, β, γ and δ genera, shown as EC50 values. j, Overlay of the crystal structure of the 76E1 Fab-peptide complex (cyan) onto the cryo-EM structure of the SARS-CoV-2 prefusion S trimer (grey) (PDB code 6XR8) based on superposition of the α-helical peptide. The zoomed-in section shows the superposition of the α-helical peptide in the crystal structure of the 76E1 Fab-peptide complex (green) onto the cryo-EM structure of the SARS-CoV-2 S trimer (orange). k, The binding affinity of 76E1 and 76F6 (an RBD antibody isolated in this project) to the prefusion S trimer as determined by BLI. Representative data from 2 independent experiments are shown (a,g,i,k).
In the structure of the 76E1 Fab-peptide complex, the peptide folds into an α-helix and is embedded in a binding pocket formed by the complementary determinant region (CDR) loops H1, H2, H3 and L3 of 76E1 Fab (Fig. 3d). The peptide residues interact with the 76E1 Fab residues via a network of salt bridges, hydrogen bonds and hydrophobic interactions, burying 676.7 Å2 of the solvent-accessible surface area of the Fab (Fig. 3d). The sidechain of Arg815 inserts into the cleft between the Fab heavy chain CDRs H1 and H3, forming hydrogen bonds with the mainchain carbonyls of Leu99H, Asp108H and Phe110H (the suffixes H and L refer to the Fab residues in heavy chain and light chain, respectively). A salt bridge triad is formed by the peptide residues Glu819 and Asp820 and the Fab residue Arg50H. Leu822 and Phe823 of the peptide are accommodated in a hydrophobic pocket formed by the Fab residues His35H, Arg50H, Ile101H, Phe34L, Tyr93L and Leu99L (Fig. 3e). In particular, the sidechain of Phe823 is stabilized by π-π interaction with Tyr93L and cation-π interaction with Arg50H. In addition, Lys825 makes several hydrogen bonds with the sidechains of Asn97L and Tyr32L (Fig. 3f). To verify the functional roles of the critical epitope residues involved in 76E1 binding, we performed ELISA-based alanine substitutions on the peptide809–833. The R815A, E819A and F823A peptide mutants showed complete loss of reactivity to 76E1, confirming the importance of the interaction between 76E1 and these residues. Additionally, the D820A, L822A and K825A peptide mutants partially disrupted the binding to 76E1 (Fig. 3g). These residues are highly conserved among the sequences aligned from four coronavirus genera, including α-, β-, γ- and δ-coronavirus (Fig. 3h). Of note, Arg815 is the S2’ cleavage site, which is critical for the cleavage event that initiates fusion activation33,34. As expected, 76E1 bound well to peptides809–833 from α-, β-, γ- and δ-coronaviruses (Fig. 3i), indicating a wide breadth of this antibody.
To further validate the conservation of the epitope recognized by 76E1, we analysed the natural viral mutations within the 76E1 epitope. Most of the mutations exhibit extremely lower frequency among circulating SARS-CoV-2 isolates, with less than 100 out of 5,506,473 sequences as of 19 November 2021 for each substitution, except for L822F (1,525 isolates) (Extended Data Fig. 6a,d). Most of the mutations completely or partially reduced the infectivity of SARS-CoV-2, indicating that these mutations were not able to induce widespread transmission (Extended Data Fig. 6b). Although L822F did not affect virus infectivity, 76E1 neutralized L822F pseudovirus comparably with the wild-type virus. For other pseudoviruses with natural mutations, 76E1 also exhibited comparable neutralizing IC50 values with wild-type virus (Extended Data Fig. 6c,d). These data suggest that 76E1 might have a high barrier for the pressure selection of SARS-CoV-2 escape mutants.
The epitope of 76E1 is partially buried in the prefusion S trimer
Structural comparison shows that the conformation of the S peptide in the Fab-peptide complex is slightly different from that of the corresponding region in the cryo-EM structure of the SARS-CoV-2 prefusion S trimer (Protein Data Bank (PDB) code 6XR8)34. In the structure of the prefusion S trimer, residues 817–823 fold into an α-helix, while residues 814–816 and residues 824–826 assume loop conformations (Fig. 3j). Intriguingly, the key epitope residues involved in the 76E1 binding are fully or partially buried in the interior core of the prefusion S trimer. In particular, Arg815 is partially shielded by the upstream loop (residues 806–804), and the sidechains of Ser816, Glu819, Leu822 and Phe823 are oriented towards the interior core of the S trimer and thus are solvent inaccessible (Extended Data Fig. 7a,b). Overlay of the Fab-peptide complex onto the structure of the prefusion S trimer based on superposition of the peptide shows clear steric clash between the Fab and the prefusion S trimer (Fig. 3j). Consistently, our in vitro binding assay shows that 76E1 has no measurable binding with the S trimer in the prefusion state; in contrast, 76F6 which targets the RBD of the S trimer exhibits tight binding (Fig. 3k). These results suggest that a conformational change should take place to expose the epitope during the transition from the prefusion state to the postfusion state, and 76E1 would bind the epitope at the intermediate state of the S protein during the transition process.
ACE2 promotes the exposure of the 76E1 epitope in SARS-CoV-2
We further set out to elucidate how 76E1 gains access to the cryptic epitope within the S2’ site and the fusion peptide as this region is partially buried in the prefusion S trimer. Other groups proposed that receptor binding would promote the exposure of the S2’ cleavage site and the fusion peptide33,35,36, which is also the epitope of 76E1. We constructed a cell line with stable expression of SARS-CoV-2 S protein on A549 cells (A549-S). A549-S cells were treated with recombinant hACE2 protein before trypsin cleavage. Interestingly, hACE2 increases S2’ cleavage by trypsin, resulting in more S2’ fragments than observed in the control (Fig. 4a). Notably, 76E1 competes with trypsin and inhibits hACE2-induced S2’ cleavage (Fig. 4b). We further performed flow cytometry to confirm whether hACE2 would enhance 76E1 binding to A549-S cells. A substantially higher frequency of 76E1 binding (1.5-fold) and mean fluorescence intensity (MFI, 4-fold) are observed for hACE2-treated cells than for untreated cells. The frequency of RBD antibody-binding cells decreases upon treatment with hACE2, as hACE2 competes with this antibody for the epitope (Fig. 4c,d). In conclusion, the receptor-binding process promotes the exposure of 76E1 epitope and 76E1 shows obvious inhibitory effect on S2’ cleavage. Since 76E1 inhibits S2’ cleavage by targeting the S2’ site (Arg815), it also strongly prevents membrane fusion induced by HeLa-S cells and HeLa-hACE2 cells in a dose-dependent manner, while the isotype control antibody shows no inhibitory effect (Fig. 4e).
a, hACE2 promoted trypsin-induced S2’ cleavage. A549-S cells were incubated with hACE2 before treatment with 0.4, 0.2 or 0.1 µg ml−1 trypsin. b, 76E1 inhibited hACE2 and trypsin-induced S2’ cleavage. A549-S cells were incubated with hACE2 and 76E1 before trypsin treatment at 0.4 µg ml−1. S2, approximately 110 kDa; S2’, approximately 70 kDa. c,d, hACE2 facilitated 76E1 binding to the cell surface-expressed S proteins of SARS-CoV-2, as determined by flow cytometry. The original cell frequencies (c) and the MFI values (d) are shown. Data are presented as mean values of duplicates (d). e, 76E1 inhibits spike- and hACE2-mediated cell-cell membrane fusion. Scale bar, 100 µm. f, The pseudovirus neutralizing activity of 76E1 and an RBD antibody CB6 against SARS-CoV-2 at pre- and post-attachment steps. In a standard neutralization assay, virus was incubated with the mAbs before adding onto HeLa-hACE2 cells, and the infectivity was determined after a 2 d incubation at 37 °C. For pre-attachment assay, mAbs were incubated with virus before adding onto cells at 4 °C for 1 h. Unbound virus was washed and virus infectivity was determined as above. For post-attachment assay, cells were infected with virus at 4 °C for 1 h, and unbound virus was washed before antibodies were incubated with cells. CB6 was included as a control. g, The synergistic effect of 76E1 and hACE2 in neutralizing authentic SARS-CoV-2. Neutralizing curves (left) of 76E1 and hACE2 alone and in combination at a constant ratio with serial threefold-diluted concentration from 81 times IC50. A dose of 1 represents the concentration at IC50. The constant ratio indicates the ratio between 76E1 and hACE2 at IC50. CI values (right) at ED50, ED75 and ED90 (50%, 75% and 90% effective dose) were calculated using the CompuSyn programme. The red dotted line represents CI = 1; CI < 1 is defined as synergism; CI > 1 is defined as antagonism. h, The synergistic effect of 76E1 and CB6 in neutralizing authentic SARS-CoV-2 (left) and CI values (right) at ED50, ED75 and ED90. Results are depicted as mean ± s.d. from 3 replicate wells (f–h). For all data, representative data from 2 independent experiments are shown.
As 76E1 mainly binds S protein after the receptor-binding process, we thus proposed that the RBD antibodies might lose antiviral activity earlier than 76E1. Time-of-addition experiments were conducted, in which SARS-CoV-2 pseudovirus was incubated with 76E1 before (pre-attachment) or after (post-attachment) virus absorption to the surface of HeLa-hACE2 cells at 4 °C. The data showed that the RBD antibody CB6 lost the neutralizing activity at the post-attachment step, while 76E1 still retained the neutralizing ability (Fig. 4f).
The synergistic effect of 76E1 and hACE2 or RBD antibodies
We further determined whether 76E1 was synergistic with hACE2 in neutralizing authentic SARS-CoV-2 using the CompuSyn programme which has been described previously37,38. Dose-dependent neutralizing activity of 76E1, hACE2 or their combination was then evaluated by serial 3-fold dilutions in concentrations from 81 times the IC50 (Fig. 4g). The combination index (CI) was analysed to evaluate the synergy of the two drugs. The CI values of the mixture at ED50, ED75 and ED90 are presented as 0.623, 0.67 and 0.722, respectively, which are less than 1—a hallmark of synergism (Fig. 4g and Supplementary Table 6). The dose reduction index was calculated by comparing the dose required to reach 50%, 75% or 90% neutralization when 76E1 or hACE2 was treated alone and in combination. The data showed that 76E1 and hACE2 in combination caused 2.474~3.687-fold reduction at IC50, IC75 or IC90, further demonstrating the synergy of 76E1 and hACE2 (Supplementary Table 6). We also determined the synergistic effect between 76E1 and an RBD antibody CB6 in neutralizing authentic SARS-CoV-2. Similar to hACE2, CB6 is synergistic with 76E1, with CI values of the mixture being <1 at ED50, ED75 and ED90 (Fig. 4h and Supplementary Table 7). Interestingly, we also found that CB6 is an ACE2-mimicking antibody, which could promote the binding of 76E1 to A549-S cells (Extended Data Fig. 7a). P2C-1F11 is a previously reported antibody with ACE2 mimicry feature39. From pseudovirus synergistic neutralization experiments, we confirmed that both P2C-1F11 and CB6 were synergistic with 76E1 (Extended Data Fig. 8b–g). Another RBD antibody with lower ability in promoting 76E1 binding to A549-S cells also exhibits a certain degree of synergistic effect (Extended Data Fig. 8h–j). These data indicate a potential combination use for 76E1 and hACE2 or RBD antibodies to combat SARS-CoV-2.
Proposed model for 76E1 neutralization of SARS-CoV-2
On the basis of the mechanism study, we propose a model for 76E1 neutralization. The closed RBD in the prefusion S trimer cannot recognize the hACE2 receptor of the host cell while occasionally, one RBD may hold up and bind to hACE2 through conformational change. The process of receptor binding to ACE2 induces obvious conformational change of the prefusion S trimer, especially for the exposure of the fusion peptide and the S2’ site. Once the fusion peptide and S2’ site is exposed, 76E1 will immediately target the epitope and inhibit S2’ cleavage, thus avoiding a cascade of rearrangement of S protein and blocking virus–host cell membrane fusion and viral entry. Most of the RBD antibodies, such as CB616, compete with the S protein in binding the cell surface-expressed receptor ACE2. Another class of antibodies target the peptide 1147SFKEELDKYFKNKTS1161 within the stem helix of the S2’ fragment, such as S2P6 and B6, which are reported to disrupt the stem helix bundle and prevent S2’ fragment refolding from the pre- to the postfusion state30,32 (Fig. 5). Therefore, 76E1 represents a new class of neutralizing antibody against SARS-CoV-2.
The closed RBD in the prefusion S trimer could not recognize the hACE2 receptor of the host cell. The receptor binding process induces obvious conformational change of the S trimer, especially for the exposure of the fusion peptide (FP) and the S2’ site. Once the fusion peptide and S2’ sites are exposed, 76E1 will immediately target the sites and inhibit S2’ cleavage, thus avoiding S2’ fragment refolding and blocking the virus–host cell membrane fusion. Different from previously reported RBD mAb CB6 and S2 stem helix mAbs S2P6 and B6, 76E1 represents a new class of antibody against SARS-CoV-2.
Discussion
The S2 region of coronaviruses is more conserved than the S1 region and is thus a promising target for broad coronavirus antibodies and vaccines. Several reported S2 antibodies, such as S2P6, B6, 28D9 and CC40.8 have shown cross-reactivity to β-coronaviruses, but not other coronavirus genera21,29,30,31. In our work, 76E1 shows broad reactivity against pan-coronavirus genera, including α- and β-coronaviruses, and even several γ- and δ-coronaviruses. The highly conserved epitope recognized by 76E1 determines the broad reactivity of this antibody. Less than 0.032% of the isolated SARS-CoV-2 sequences have mutations within the epitope residues of 76E1, and all the SARS-CoV-2 variants of concern harbour no residue mutations within this epitope. Notably, the major natural mutations within the 76E1 epitope did not alter the neutralizing potency of 76E1. Some mutations even severely reduced the virus infectivity, which might restrict virus spread. The broad spectrum defines 76E1 as an attractive agent to combat the challenge of viral escape mutations and cope with a possible new coronavirus outbreak in the future. It must be emphasized that the highly conserved key 76E1 epitope residue Arg815 is the ‘fusion activation’ proteolytic site (S2’) of SARS-CoV-2, which indicates that the S2’ site is a promising target for broad coronavirus inhibitors.
The neutralizing potency of 76E1 against SARS-CoV-2 is lower than that of some of the potent RBD antibodies. Indeed, the reported S2-targeting antibodies also showed moderate neutralization activity29,30,31. This phenomenon is consistent with influenza virus, in which the neutralizing potency of antibodies targeting the conserved haemagglutinin stem region is lower than the ones targeting the head region22. Our in vivo data showed that 76E1 displayed both prophylactic and therapeutic effects against SARS-CoV-2 and HCoV-OC43 infection, indicating its potential for clinical application.
Our structural data show that several key residues involved in the antibody–antigen interactions are fully or partially buried in the prefusion S trimer. The moderate neutralization potency of 76E1 might be the result of limited epitope exposure. This suggests that a conformational change has to take place during the transition from the prefusion state to the postfusion state, leading to the exposure of the epitope to permit the binding of 76E1. We found that binding to ACE2 promoted the exposure of the S2’ site and the fusion peptide, perhaps by structural rearrangement of the S protein, thus removing the obstacles shielding this epitope and providing accessibility for 76E133,35. 76E1 is synergistic with hACE2 in preventing SARS-CoV-2 infection, which might indicate two possible mechanisms: a synergy where 76E1 binds to the S2’ site and the fusion peptide in an S conformation that needs to be primed by receptor binding; another synergy between the receptor blocking process and the membrane fusion process. We also found that some RBD antibodies might mimic ACE2 to promote 76E1 binding to the A549-S cells, with different synergistic potencies. Our data provide a potential combination of 76E1 and hACE2 or RBD antibodies to prevent or treat SARS-CoV-2 infection.
Epitope-based vaccine design has shed light for the control of respiratory syncytial virus, influenza virus and other viruses40,41. We found that 11 out of 36 plasmas from COVID-19 convalescent patients showed obvious reactivity to peptide809–833 (Extended Data Fig. 9). Although 3 (76E1, 555D6 and 125C1) out of 9 S2-targeting mAbs were isolated in this study and showed binding activity to peptide809–833, 555D6 and 125C1 could not neutralize SARS-CoV-2 pseudovirus. The non-neutralizing property of 555D6 and 125C1 might be because they were not able to inhibit S2’ cleavage (Extended Data Fig. 10). Thus, although it is possible to isolate 76E1-like antibodies using peptide809–833 as a probe, not all the peptide809–833 binding mAbs possess neutralizing activities. These findings suggest that eliciting high titres of 76E1-like mAbs by vaccination is a big challenge for developing pan-coronavirus vaccines. Epitope-based de novo antigen design and immunization strategy design might be promising routes to expose the epitope in a proper state and induce broad coronavirus antibodies.
Methods
Ethics statement
This study was approved by the Shanghai Public Health Clinical Center Ethics Committee of Fudan University. Blood was collected from patients who had recovered from SARS-CoV-2 infection after they had signed the informed consent form. The animal experiments were approved by the Institutional Laboratory Animal Care and Use Committee at Fudan University. All manipulations were strictly conducted in compliance with ethics guidelines and approved protocols.
Cells and viruses
HeLa, RD, LLC-MK2, HuH-7 and HEK 293T cells were all obtained from the American Type Culture collection and cultured in Dulbecco’s modified Eagle’s medium (DMEM, Gibco) with 10% fetal bovine serum (FBS). CHO cells (Thermo Fisher) were cultured in Expi CHO expression medium (Thermo Fisher, A2910001). HEK 293T cells stably expressing human ACE2 (293T-hACE2) were generated and cultured in DMEM with 10% FBS. A549 cells stably expressing SARS-CoV-2 spike were generated and cultured in DMEM with 10% FBS. The SARS-CoV-2 clinical isolate nCoV-SH01 (GenBank: MT121215.1) was amplified in Vero-E6 cells.
Recombinant proteins and antibodies
To generate recombinant human ACE2 protein, a DNA fragment encoding the extracellular domain of human ACE2 (residues S19 to S740) was cloned into a refined pGT128 vector containing an antibody signal peptide in the N terminus and human IgG1 crystallizable fragment (Fc) in the C terminus. A flexible ‘GSGGGG’ linker was inserted between ACE2 and human IgG1 Fc. The recombinant hACE2 protein was expressed in CHO cells and purified with Protein A (MabSelect Prism A, GE Healthcare) and size exclusion chromatography using a Superdex 200 10/300 column (GE Healthcare).
To generate the prefusion S trimer, the extracellular domain of the S protein (1–1,208 amino acids) was cloned into the pCAG vector with a ‘GSAS’ substitution at residues 682–685, two proline substitutions at residues 986 and 987, and a T4 fibritin trimerization motif in the C terminus.
The S-ECD is the extracellular domain of the S protein lacking the transmembrane region and without any mutations and modifications in the sequence. The following S-ECD proteins were expressed either from baculovirus or mammalian expression systems and utilized for binding assays: SARS-CoV-2 S-ECD-His (Sino Biological, 40589-V08B1), SARS-CoV S-ECD-His (Sino Biological, 40634-V08B), MERS-CoV-2 S-ECD-His (Sino Biological, 40069-V08B), HCoV-NL63 S-ECD-His (Sino Biological, 40604-V08B), HCoV-229E S-ECD-His (Sino Biological, 40605-V08B), HCoV-OC43 S-ECD-His (Sino Biological, 40607-V08B), HCoV-HKU1 S-ECD-His (Sino Biological, 40606-V08B), SARS-CoV-2 S1-His (Sino Biological, 40150-V08B1), SARS-CoV-2 S2-His (Sino Biological, 40590-V08B), SARS-CoV-2 RBD-SD1–His (Novoprotein, DRA42), SARS-CoV S1-His (Sino Biological, 40150-V08B1), SARS-CoV S2-His (Sino Biological, 40150-V08B3), SARS-CoV RBD-His (Sino Biological, 40150-V08B2), MERS-CoV S1-His (Sino Biological, 40069-V08H), MERS-CoV S2-His (Sino Biological, 40070-V08B) and MERS-CoV RBD-His (Sino Biological, 40071-V08B1); B.1.1.7 S-ECD-His (Sino Biological, 40589-V08B6), P.1 S-ECD-His (Sino Biological, 40589-V08B8), B.1.351 S-ECD-His (Sino Biological, 40589-V08B7), B.1.617.1 S-ECD-His (Sino Biological, 40589-V08B15), B.1.617.2 S-ECD-His (Sino Biological, 40589-V08B16) and B.1.1.529 S-ECD-His (Sino Biological, 40589-V08B33).
Isolation, cloning, expression and purification of 76E1
Fresh peripheral blood mononuclear cells were isolated from the collected blood using the Ficoll-Paque gradient (GE Healthcare, 17144002). Percp-CY5.5-CD4−, Percp-CY5.5-CD14−, Percp-CY5.5-CD8−, APC-Cy7-780-dead−, FITC-CD19+, APC-IgG+ and biotinylated SARS-CoV-2 S-ECD protein+-streptavidin-SA BV421+ specific single memory B cells were isolated by a Sony MA900 flow separator and sorted into a 96-well plate containing 10 μl RNase-free water (Sangon Biotech) and 20 U RNase inhibitor (Promega) in each well. Complementary DNA was produced using reverse transcriptase PCR kit (Yeasen). Antibody VH and VL genes were amplified by nested PCR using DNA polymerase (TransGen Biotech). The PCR products were sequenced by BioSune Biotech. Antibody sequences were then analysed using the National Center for Biotechnology Information (NCBI)-IgBLAST (https://www.ncbi.nlm.nih.gov/igblast/) and IMGT/V-QUEST (http://www.imgt.org/IMGT_vquest). The genes encoding VH and VL genes of the antibodies were cloned into vectors Abvec-HIgG1-VH for the heavy chain, and Abvec-HIgG1-VK and Abvec-HIgG1-Vλ for the light chain. The VH and VL plasmids were cotransfected into Expi CHO cells, which were cultured and collected according to the manufacturer’s instructions. The antibodies were purified using Protein A, followed by size exclusion chromatography.
ELISA
To determine the binding properties of the antibodies and sera, the RBD, S1, S2 or S-ECD proteins or peptides (0.5 µg ml−1 in 100 µl per well) were captured on 96-well plates overnight and blocked with 2% bovine serum albumin in PBS-Tween 20 (PBST) for 2 h. Antibodies or sera were serially diluted and incubated in the wells for 2 h. The samples were washed three times, and an anti-human Fc-HRP antibody (Sigma-Aldrich) was used to determine binding affinity, followed by incubation with tetramethylbenzidine (Beyotime), which was stopped by adding 1 M HCl. The absorbance at 450 nm was recorded by a plate reader (Bio-Tek Epoch 2). For the receptor competition assay, hACE2 (5 µg ml−1) was coated on the microplates. The isolated antibodies were serially diluted and incubated with biotinylated S protein (0.3 µg ml−1) for 1 h and then added to the wells after washing and blocking. Streptavidin-conjugated HRP was used as the detection antibody. To evaluate the reactivity of 76E1 to the denatured or untreated S-ECD protein, ELISA was performed as mentioned above. S-ECD protein was denatured with 0.1% SDS and 50 mM dithiothreitol (DTT) in a metal block at 100 °C for 5 min.
Biolayer interferometry (BLI) analysis
BLI was performed using OCTET DATA acquisition instrument (ForteBio). To test the binding affinity between 76E1 and different S-ECD proteins, 76E1 (15 µg ml−1) was immobilized on an anti-human IgG-Fc-coated biosensor surface for 300 s. The baseline interference phase was then read for 180 s in kinetics buffer (KB: 1× PBS), followed by subsequent association phase immersion of the sensors into wells containing 2-fold serial dilutions of S-ECD protein (from 200 nM to 6.25 nM) in KB for 600 s. Then, the sensors were immersed in KB for dissociation for up to 600 s. The mean association rate constant (Kon), dissociation rate constant (Koff) and dissociation constant (KD) values were calculated using the global fit to a 1:1 Langmuir binding model with an R2 ≥ 0.95 by Fortebio Data Analysis software. To determine the binding affinity of 76E1 to the prefusion S trimer, biotinylated S trimer (15 µg ml−1) was immobilized on a streptavidin biosensor for 300 s. The following phases were as described above except for the association phase in which the sensors were immersed into wells containing serially diluted 76E1 (500-250-125-62.5 nM) or control antibody 76F6 (100-50-25-12.5-6.25 nM).
Pseudovirus neutralization assay
The neutralization activity of 76E1 was determined against pseudotyped SARS-CoV (S gene GenBank: AAP13567.1), SARS-CoV-2 (S gene GenBank: MT121215.1), MERS-CoV (S gene GenBank: AFS88936.1) and HCoV-229E (S gene GenBank: APT69883.1) viruses as previously described42. Briefly, plasmids encoding the full-length S gene and pNL4-3.luc.RE were cotransfected into HEK 293T cells in 10 cm dishes. The medium was replaced after 6 h, and virus supernatants were collected 48 h after transfection. HEK 293T-hACE2 cells were plated in 96-well plates and grown overnight. The virus was then diluted in DMEM with 10% FBS mixed with an equal volume of serially diluted antibodies and incubated at 37 °C. One hour later, the mixtures were transferred to HEK 293T-hACE2 cells in 96-well plates. The cells were incubated at 37 °C for 48 h, followed by cell lysis and luciferase activity assays (Promega). Data were collected using BioTek Synergy NEO. The percent neutralization was calculated by comparing the luciferase value of the antibody-treated cells to that of the untreated control. IC50 values were analysed by GraphPad Prism 7.0 software. The amino acid sequences of S genes of the SARS-CoV-2 variants of concern used in the preparation of pseudoviruses are identical with Wuhan-Hu-1 isolates (NCBI reference sequence: YP_009724390.1) with mutations as summarized in Extended Data Fig. 2a. The neutralizing potency of 76E1 against these variants was determined as described above.
Authentic virus neutralization assay
The authentic SARS-CoV-2 experiments were performed in the BSL-3 laboratory of Fudan University under the guidelines of Fudan University. Vero-E6, HeLa-hACE2 and Calu-3 cells were plated into 96-well plates and grown overnight. One hundred 50% cell culture infectious dose (TCID50, 50 µl) of live SARS-CoV-2 virus (nCoV-SH01) was mixed with 50 µl of 3-fold serially diluted 76E1 and incubated at 37 °C for 1 h. The supernatants were discarded, and the antibody–virus mixture was mixed with the target cells. After incubation at 37 °C for 1 h, the DMEM containing 1% methylcellulose and 2% FBS was added. The cells were incubated at 37 °C for 24 h, fixed with 1% formaldehyde for 1 h, washed three times with PBS and permeabilized in 0.2% Triton-X in PBS with gentle shaking for 10 min at r.t. The wells were then blocked with 2% BSA for 2 h at r.t. The cells were first incubated with a primary antibody recognizing SARS-CoV-2 nuclear protein, followed by a secondary antibody incubation with AlexaFluor 488-conjugated goat anti-rabbit antibody. The data were collected by a high throughput imaging instrument Cytation/BioSpa8/Multiflo FX. IC50 was calculated by GraphPad Prism 7.
The inhibitory activity of antibodies against HCoV-OC43 (ATCC, VR-1558) infection in RD cells was assessed. Briefly, 100 TCID50 of HCoV-OC43 was mixed with a test antibody at indicated concentration and incubated at 37 °C for 30 min. The mixture was added to RD cells. After culture for an additional 2–3 d, Cell Counting Kit-8 (CCK8, Dojindo) assay was used to determine cytopathic effect (CPE). The inhibitory activity of the tested antibodies against HCoV-229E (ATCC, VR-740) infection in Huh-7 cells and HCoV-NL63 (Amsterdam strain) infection in LLC-MK2 cells was evaluated as described above. IC50 was calculated using GraphPad Prism 7.0.
The synergistic effect of 76E1 with hACE2 or RBD antibodies
The synergistic effect of 76E1 with hACE2 or RBD antibody CB6 in neutralizing authentic SARS-CoV-2 in Vero-E6 cells was determined as described previously37,38. 76E1, hACE2 or CB6 and their combination were 3-fold serially diluted from 81 times the IC50 and incubated with SARS-CoV-2 virus (100 TCID50) at 37 °C for 1 h. 76E1 and hACE2 or CB6 in combination were mixed at a constant ratio, which represented the dose at IC50. Neutralization was performed as described above. The CI and dose reduction index at ED50, ED75 and ED90 (effective dose at IC50, IC75 and IC90) were analysed to evaluate the degree of cooperation among the two drugs using the CompuSyn programme. The synergistic effects of 76E1 with RBD antibodies CB6, P2C-1F11 and 28–8L in neutralizing SARS-CoV-2 pseudovirus in 293T-hACE2 cells were also determined as described above.
Flow cytometry
To assess whether hACE2 promoted the binding of 76E1 to spike protein, A549 cells stably expressing the spike protein of SARS-CoV-2 were detached with TrypLE Express (Gibco) and incubated with 20 µg ml−1 hACE2 on ice for 30 min. After washing and staining with FVS780-APC-CY7, the cells were incubated with 15 µg ml−1 biotinylated 76E1, RBD antibody 76F6 or isotype control antibody in FACS buffer on ice for 1 h. Antibodies still bound to surface-expressed spike protein after washing were then stained with streptavidin-conjugated BV421 (BD Biosciences), followed by flow cytometry using BD Fortessa. The mean fluorescence intensity (MFI) was calculated. To determine whether RBD antibodies (CB6, P2C-1F11 and 28-8 L) and isotype control antibody promote the binding activity of 76E1 to the spike proteins on A549 cells, similar experiments as described above were conducted. Data were analysed by Flowjo V10.
Protective efficacy of 76E1 in mice
Transgenic mice expressing human ACE2-luciferase driven by the CAG promoter Tgtn (CAG-human ACE2-IRES-Luciferase-WPRE-polyA) were purchased from Shanghai Model Organisms Center and used to test the protective efficacy of 76E1 in a SARS-CoV-2 infection model. Briefly, 10-week-old female mice were raised in the BSL-3 laboratory of Fudan University. For prophylactic experiments, the mice were intranasally infected with SARS-CoV-2 (1 × 104 p.f.u. in Fig. 1e and 3.4 × 104 p.f.u. in Extended Data Fig. 3) 24 h after intraperitoneal administration of 76E1 (15 mg kg−1 in Fig. 1e and 150 or 50 mg kg−1 in Extended Data Fig. 3). For therapeutic experiments, the mice were treated with 76E1 after SARS-CoV-2 infection. Weight loss was monitored for 3 d post-infection. The mice were euthanized at day 3 post-infection and the lungs were collected for virus titre examination. PBS inoculation was included as controls. Viral RNA was extracted from lung tissue with TRIzol reagent (Invitrogen) and reverse transcribed into cDNA (Toyobo). Real-time qPCR was performed using a SYBR Green kit (Toyobo) and ABI QuantStudio 6 Flex instrument. The SARS-CoV-2 N gene-specific primers were as follows: forward primer, 5’-GGGGAACTTCTCCTGCTAGAAT-3’; reverse primer, 5’-CAGACATTTTGCTCTCAAGCTG-3’. The lung samples for histological examination were stored in 4% neutral-buffered formalin for 7 d, embedded in paraffin, sectioned and stained with haematoxylin and eosin, followed by light microscopy.
To determine the protective efficacy of 76E1 against HCoV-OC43, pregnant BALB/c mice (18 d) were separated into four groups after delivery of their offspring and received humane care in compliance with the guidelines of the Animal Research Ethics Board of Fudan University. As described elsewhere43,44,45, for prophylactic experiments, 4 d new-born BALB/c mice were pretreated with 15 or 50 mg kg−1 antibodies or PBS 24 h before intranasal challenge with OC43 with a lethal dose (50 TCID50 in 1 µl DMEM). For therapeutic experiments, 7-day-old mice were treated with 50 mg kg−1 antibodies or PBS 2 h after intranasal challenge with OC43 (50 TCID50). Body weight and survival rate were monitored for 14 d post-infection. Another group of mice were euthanized at 5 d.p.i., and the brains were collected for virus titre examination by calculating CPE on RD cells; results were expressed as TCID50 g−1.
Syncytium inhibition assay
The experimental procedure was as previously described42 with some modifications. In brief, HeLa cells were transfected (when ~60–70% confluent in 6-well plates) with plasmids encoding the codon-optimized full-length SARS-CoV-2 S protein by Lipofectamine 2000 (HeLa-S). Another group of HeLa cells was transfected with plasmids encoding the codon-optimized full-length hACE2 (HeLa-hACE2). Forty-eight hours after transfection, the HeLa-S cells were detached and incubated with different concentrations of 76E1 or control antibody at 37 °C for 1 h. Then, the HeLa-hACE2 cells were detached and mixed with HeLa-S cells at a 1:1 ratio in 48-well plates. After 12 h of co-culture, fused cells were observed. After washing and fixing with 4% paraformaldehyde, the cells were dyed with crystal violet. Syncytium formation was observed with the Olympus IX73 inverted microscope and images were collected by Olympus cellSens Entry software.
Isothermal titration calorimetry (ITC) analysis
The KD of the 76E1 Fab with the SARS-CoV-2 spike peptides was determined by ITC using a MicroCal PEAQ-ITC calorimeter (Malvern). Purified 76E1 Fab was adjusted to a concentration of 20–30 μM in PBS buffer. Synthetic peptides were solubilized to 300-400 μM in the same buffer. An initial injection of 0.8 μl protein sample was discarded for each dataset to remove the effect of titrant diffusion across the syringe tip during the equilibration process. Then, 2 μl of the peptide solution was injected to the calorimeter cell containing the 76E1 Fab solution every 2 min. The stirring speed and temperature were maintained at 750 r.p.m. and 25 °C in the calorimeter cell. A background titration was performed using identical titrant with the buffer solution. The heat for each injection was measured and the KD value was determined by fitting of the titration curve using the nonlinear least-squares method in MicroCal PEAQ-ITC analysing software with the single-site binding model.
Crystallization, data collection and structure determination
The Fab fragment of 76E1 was generated by papain digestion of the 76E1 antibody and purified by Protein A agarose (GE Healthcare) and gel filtration on a Superdex 200 10/300 column (Cytiva) in PBS buffer. The S peptide809–833 was synthesized by GenScript. The purified 76E1 Fab (9 mg ml−1) and the S peptide809–833 were mixed at a molar ratio of 1:2 and incubated at 4 °C for 2 h before crystallization screening. Crystallization was carried out at 16 °C using the hanging-drop vapour diffusion method by mixing equal volumes (1 μl) of the Fab-peptide mixture solution and the reservoir solution. Crystals of the Fab-peptide complex were grown in drops consisting of the reservoir solution of 0.1 M sodium citrate (pH 4.5) and 20% (w/v) polyethylene glycol 4000. Crystals were transferred into the cryoprotectant consisting of the reservoir solution and 20% (v/v) glycerol for cryoprotection, followed by flash-cooling into liquid nitrogen. X-ray diffraction data were collected at 100K at BL18U1 beamline of the National Facility for Protein Science Shanghai. Data indexing, integration and scaling were performed using HKL300046. The structure of the 76E1 Fab-peptide complex was solved by molecular replacement method as implemented in Phenix47 using the MGG4 Fab (PDB code 6BQB)48 as the search model. Model building was performed using Coot49 and structure refinement was performed using Phenix47. The peptide was built on the basis of the electron density after the Fab was fully refined. Stereochemistry and quality of the structure model were analysed using programmes in CCP4 suite50. The structure figures were generated using CCP4MG51 and Pymol52. The statistics of the diffraction data, the structure refinement and the final structure model are summarized in Supplementary Table 5.
hACE2 promotes the exposure of the S2’ cleavage site
To determine whether hACE2 promotes exposure of the S2’ cleavage site, A549-S cells were plated into 12-well plates and incubated with 20 µg ml−1 hACE2 at 37 °C for 30 min. After washing, the cells were digested with 0.4, 0.2 or 0.1 µg ml−1 TPCK-treated trypsin at 37 °C for 15 min. Then, the cells were washed twice again and cleavage was stopped by boiling the sample in a 100 °C water bath. Samples were run on SDS–polyacrylamide electrophoresis gel under reducing conditions and blotted using a rabbit anti-SARS-CoV-2 S2 antibody (Sino Biological, 40590-T62). Recombinant hACE2 was detected by a goat anti-human IgG1 Fc-HRP antibody, and actin was detected by a mouse anti-actin antibody. To determine whether 76E1 inhibits hACE2-induced S2’ cleavage, 76E1 was incubated with A549-S cells before treatment with trypsin and the experiment was conducted as described above. A GE healthcare ImageQuant LAS4000 mini biomolecular imager was used for the detection and quantitation of chemiluminescence.
Pre- and post-attachment of mAbs
The experiments were conducted with ice-cold media and the cell plates were cold treated at 4 °C for 1 h before the experiments. In a standard neutralization assay, SARS-CoV-2 pseudoviruses were incubated with the mAbs for 1 h at 4 °C before adding onto HeLa-ACE2 cells, and the infectivity was determined after a 2 d incubation at 37 °C. For pre-attachment assay, mAbs were incubated with virus before adding onto cells at 4 °C for 1 h and the mixture was then transferred to cell plates and incubated at 4 °C for 1 h. Unbound viruses were washed with ice-cold media followed by 1 h incubation at 4 °C, and virus infectivity was determined as above. For post-attachment assay, cells were first infected with virus at 4 °C for 1 h, and unbound virus was washed with ice-cold media before serial-diluted antibodies were incubated with cells at 4 °C for 1 h. The infectivity was determined after a 2 d incubation at 37 °C as described above. CB6, an RBD antibody, was included as a control.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The crystal structure of the 76E1 Fab in complex with SARS-CoV-2 S peptide809–833 has been deposited in the Protein Data Bank with accession code 7X9E (https://www.rcsb.org/structure/7X9E). The prefusion SARS-CoV-2 S trimer structure used for analysis is 6XR8 (https://www.rcsb.org/structure/6XR8)34, which was downloaded from the Protein Data Bank. All other data supporting the findings of this study are available within the Article and its supplementary files. Source data are provided with this paper.
References
Malik, Y. A. Properties of coronavirus and SARS-CoV-2. Malays. J. Pathol. 42, 3–11 (2020).
Drosten, C. et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 348, 1967–1976 (2003).
Ksiazek, T. G. et al. A novel coronavirus associated with severe acute respiratory syndrome. N. Engl. J. Med. 348, 1953–1966 (2003).
Zaki, A. M., van Boheemen, S., Bestebroer, T. M., Osterhaus, A. D. & Fouchier, R. A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 367, 1814–1820 (2012).
Layne, S. P., Hyman, J. M., Morens, D. M. & Taubenberger, J. K. New coronavirus outbreak: framing questions for pandemic prevention. Sci. Transl. Med. 12, eabb1469 (2020).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
Cui, J., Li, F. & Shi, Z. L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 17, 181–192 (2019).
Walls, A. C. et al. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 183, 1735 (2020).
Layne, S. P. & Taubenberger, J. K. Increasing threats from SARS-CoV-2 variants: time to establish global surveillance. Sci. Transl. Med. 13, eabj6984 (2021).
Grubaugh, N. D., Hodcroft, E. B., Fauver, J. R., Phelan, A. L. & Cevik, M. Public health actions to control new SARS-CoV-2 variants. Cell 184, 1127–1132 (2021).
Fenwick, C. et al. A high-throughput cell- and virus-free assay shows reduced neutralization of SARS-CoV-2 variants by COVID-19 convalescent plasma. Sci. Transl. Med. 13, eabi8452 (2021).
Liu, Y. et al. Neutralizing activity of BNT162b2-elicited serum – preliminary report. N. Engl. J. Med. 384, 1466–1468 (2021).
Wu, K. et al. Serum neutralizing activity elicited by mRNA-1273 vaccine - preliminary report. N. Engl. J. Med. 384, 1468–1470 (2021).
Liu, L. et al. Striking antibody evasion manifested by the Omicron variant of SARS-CoV-2. Nature 602, 676–681 (2021).
Cao, Y. et al. Potent neutralizing antibodies against SARS-CoV-2 identified by high-throughput single-cell sequencing of convalescent patients’ B cells. Cell 182, 73–84.e16 (2020).
Shi, R. et al. A human neutralizing antibody targets the receptor-binding site of SARS-CoV-2. Nature 584, 120–124 (2020).
Wu, Y. et al. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science 368, 1274–1278 (2020).
Barnes, C. O. et al. Structures of human antibodies bound to SARS-CoV-2 spike reveal common epitopes and recurrent features of antibodies. Cell 182, 828–842.e16 (2020).
Rappazzo, C. G. et al. Broad and potent activity against SARS-like viruses by an engineered human monoclonal antibody. Science 371, 823–829 (2021).
Lv, Z. et al. Structural basis for neutralization of SARS-CoV-2 and SARS-CoV by a potent therapeutic antibody. Science 369, 1505–1509 (2020).
Shiakolas, A. R. et al. Cross-reactive coronavirus antibodies with diverse epitope specificities and Fc effector functions. Cell. Rep. Med. 2, 100313 (2021).
Kallewaard, N. L. et al. Structure and function analysis of an antibody recognizing all influenza A subtypes. Cell 166, 596–608 (2016).
Wang, W. et al. Human antibody 3E1 targets the HA stem region of H1N1 and H5N6 influenza A viruses. Nat. Commun. 7, 13577 (2016).
Ekiert, D. C. et al. A highly conserved neutralizing epitope on group 2 influenza A viruses. Science 333, 843–850 (2011).
Yi, C. et al. Junctional and somatic hypermutation-induced CX4C motif is critical for the recognition of a highly conserved epitope on HCV E2 by a human broadly neutralizing antibody. Cell. Mol. Immunol. 18, 675–685 (2021).
Law, M. et al. Broadly neutralizing antibodies protect against hepatitis C virus quasispecies challenge. Nat. Med. 14, 25–27 (2008).
Huang, J. et al. Broad and potent neutralization of HIV-1 by a gp41-specific human antibody. Nature 491, 406–412 (2012).
Walker, L. M. et al. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 477, 466–470 (2011).
Wang, C. et al. A conserved immunogenic and vulnerable site on the coronavirus spike protein delineated by cross-reactive monoclonal antibodies. Nat. Commun. 12, 1715 (2021).
Sauer, M. M. et al. Structural basis for broad coronavirus neutralization. Nat. Struct. Mol. Biol. 28, 478–486 (2021).
Zhou, P. et al. A human antibody reveals a conserved site onbeta-coronavirus spike proteins and confers protection against SARS-CoV-2 infection. Sci. Transl. Med. 14, eabi9215 (2022).
Pinto, D. et al. Broad betacoronavirus neutralization by a stem helix-specific human antibody. Science 373, 1109–1116 (2021).
Benton, D. J. et al. Receptor binding and priming of the spike protein of SARS-CoV-2 for membrane fusion. Nature 588, 327–330 (2020).
Cai, Y. et al. Distinct conformational states of SARS-CoV-2 spike protein. Science 369, 1586–1592 (2020).
Zhu, Y., Yu, D., Yan, H., Chong, H. & He, Y. Design of potent membrane fusion inhibitors against SARS-CoV-2, an emerging coronavirus with high fusogenic activity. J. Virol. 94, e00635-20 (2020).
Walls, A. C. et al. Tectonic conformational changes of a coronavirus spike glycoprotein promote membrane fusion. Proc. Natl Acad. Sci. USA 114, 11157–11162 (2017).
Jiang, L. et al. Potent neutralization of MERS-CoV by human neutralizing monoclonal antibodies to the viral spike glycoprotein. Sci. Transl. Med. 6, 234–259 (2014).
Giang, E. et al. Human broadly neutralizing antibodies to the envelope glycoprotein complex of hepatitis C virus. Proc. Natl Acad. Sci. USA 109, 6205–6210 (2012).
Ge, J. et al. Antibody neutralization of SARS-CoV-2 through ACE2 receptor mimicry. Nat. Commun. 12, 250 (2021).
Ren, H. & Zhou, P. Epitope-focused vaccine design against influenza A and B viruses. Curr. Opin. Immunol. 42, 83–90 (2016).
Correia, B. E. et al. Proof of principle for epitope-focused vaccine design. Nature 507, 201–206 (2014).
Yi, C. et al. Key residues of the receptor binding motif in the spike protein of SARS-CoV-2 that interact with ACE2 and neutralizing antibodies. Cell. Mol. Immunol. 17, 621–630 (2020).
Jacomy, H. & Talbot, P. J. Vacuolating encephalitis in mice infected by human coronavirus OC43. Virology 315, 20–33 (2003).
Xia, S. et al. A pan-coronavirus fusion inhibitor targeting the HR1 domain of human coronavirus spike. Sci. Adv. 5, eaav4580 (2019).
Xia, S. et al. Inhibition of SARS-CoV-2 (previously 2019-nCoV) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion. Cell Res. 30, 343–355 (2020).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Tan, J. et al. A public antibody lineage that potently inhibits malaria infection through dual binding to the circumsporozoite protein. Nat. Med. 24, 401–407 (2018).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
McNicholas, S., Potterton, E., Wilson, K. S. & Noble, M. E. M. Presenting your structures: the CCP4mg molecular-graphics software. Acta Crystallogr. D D67, 386–394 (2011).
The PyMOL Molecular Graphics System, Version 2.4.1 (Schrodinger LLC, 2015).
Sun, X. et al. Unique binding pattern for a lineage of human antibodies with broad reactivity against influenza A virus. Nat. Commun. 13, 2378 (2022).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Acknowledgements
We thank the Core Facilities of Chemical Biology, Cell Biology and Molecular Biology of Shanghai Institute of Biochemistry and Cell Biology, Center for Excellence in Molecular Cell Science for their help. This work was supported by the Ministry of Science and Technology of China (2018YFA0507402 to B.S.), the Key International Partnership Program of the Chinese Academy of Sciences (153D31KYSB20180055 to B.S.), the National Natural Science Foundation of China (81971921 to Y.X. and 81761128007 to J.X., 32100751 to X.S. and 32100123 to C.Y.), the National Key Research and Development Program of China (2021YFC2300703 to L.L.), the Shanghai Municipal Science and Technology Major Project (ZD2021CY001 to Y.X. and L.L.), the National Major Science and Technology Projects of China (2018ZX10301403 to L.D. and 2018ZX10301208 to Y.X.), and Shanghai Science and Technology Commission (21DZ2291300 to X. Zhang).
Author information
Authors and Affiliations
Contributions
B.S., Y.X., J.X., L.L., J.D. and Z.L. initiated and supervised the study. X.S. and C.Y. designed and performed most of the experiments. Y. Zhu, S.X., S.D. and G.H. performed all the animal studies. X.C. and T.Z. performed structural analysis. L.D., X. Lu, L.D. and C.Q. contributed to the isolation of spike-specific memory B cells. C.G. and M.L. contributed to the authentic virus neutralization experiments. W.F., S.Y., C.Z. and X. Zhang collected the blood sample. A.Q., X. Zhou, X. Li, R.Y., X. Lu, S.X., Y. Zhang, L.M., G.W. Y.F., S.C., Z.Y., Y.H. and W.G. provided materials and helped with some experiments during this study. Q.Z., H.L. and Q.D. provided valuable suggestions. X.S. and X.C. wrote the paper. C.Y., B.S., Y.X., J.X., L.L., Z.L. and J.D. revised the paper.
Corresponding authors
Ethics declarations
Competing interests
B.S., X.S., C.Y., Z.L. and Y. Zhang are inventors on the patent applications (202110008031.1, China, 2021, and PCT/CN2022/070309, China, 2022) filed by Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences, for antibody 76E1. The other authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cross-reactivity of COVID-19 convalescent plasmas against multiple human coronaviruses.
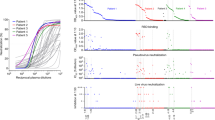
a, Amino acid sequence alignment of the S2 regions of seven human coronaviruses. The sequence identities of the human coronaviruses to SARS-CoV-2 are shown at right. b-e, Results from ELISA assessing antibody endpoint titers of plasmas from five COVID-19 convalescent patients (76, 93, 111, 125 and 555) and four healthy participants (DLF, HXC, ZC and SP), whose plasma samples were collected before the emergence of COVID-19. Antibody endpoint titers were assessed against S1, RBD, S2 and S-ECD proteins for SARS-CoV-2 (b), SARS-CoV (c), MERS-CoV (d) and S-ECD proteins for the common viruses HCoV-NL63, HCoV-OC43, HCoV-HKU1 and HCoV-229E (e). Antibody endpoint titer represents the maximum dilution of the sera that can still bind the antigen. The antibody endpoint titers were evaluated at a top dilution of 100-fold and seven additional 3-fold serial dilutions. The wells with no plasma incubation were used as controls, and the cut-off values were calculated as the OD450 of the control × 2.1. Representative data from two independent experiments are shown.
Extended Data Fig. 2 Broad binding affinity of 76E1.
a, Schematic of mutations landscape in SARS-CoV-2 VOCs, variants of concern. del, deletion; ins, insertion. b, Binding kinetics of 76E1 to multiple soluble S-ECD proteins of SARS-CoV-2 variants as measured by BLI. 76E1 was loaded on Octet AHC sensor and tested for real-time association and dissociation of the serial diluted S-ECD proteins from SARS-CoV-2 variants as determined by BLI. 76E1 retained higher affinity to all the variants tested. c, Binding kinetics of 76E1 to multiple soluble S-ECD proteins of α- and β-coronaviruses as measured by BLI. Serial diluted S proteins were immersed into the immortalized 76E1 on AHC sensors, the association and disassociation kinetics were recorded and shown. d, Binding curves of 76E1 to a panel of S-ECD proteins representative of α- and β-coronaviruses as measured by ELISA. e, 76E1 recognize a continuous epitope within spike protein. ELISA-based binding curves of 76E1 to SARS-CoV-2 S-ECD protein that was 100 °C-denatured in the presence of DTT and SDS versus untreated S-ECD. A SARS-CoV-2 RBD antibody 76F6 was used as a control. Data are shown as the mean values of duplicates (d,e). Representative data from two independent experiments were shown.
Extended Data Fig. 3 Prophylatical treatment of 76E1 protects mice from SARS-CoV-2-induced disease.
a, Study outline: A single dose of 150 mg/kg 76E1 (n = 5), 50 mg/kg 76E1 (n = 4) or PBS (n = 6) treatment was delivered intraperitoneally 24 hours before challenge the hACE2 transgenic mice with 3.4 x 104 PFU of SARS-CoV-2. b, The weight loss was monitored for 3 days. c, Lung virus titers were measured 3 days post-infection (d.p.i.) and are shown as log10 RNA copies per gram of lung tissue. The data were representative of two independent experiments. d, H&E staining of lung tissue sections at 3 d.p.i. Representative images are shown with low (top, scale bars, 500 µm) and high power (below, scale bars, 50 µm) from two independent experiments. Results are depicted as the mean ± s.d. The P values are analyzed by two-sided unpaired t tests. ****P < 0.0001, ns, not significant (b,c).
Extended Data Fig. 4 Sequence alignment of the S proteins of seven human coronaviruses indicates seven conserved sequences in S2.
100% Identical residues are denoted in red background. The blue line indicates fusion peptide. The 7 conserved sequences are labeled with black dotted lines. The S1/S2 and S2’ cleavage sites of SARS-CoV-2 are annotated above the site. The sequence alignment was conducted by ESPript54. The sequences used in the alignment are: SARS-CoV-2 (NCBI GenBank: MT121215.1), SARS-CoV (NCBI GenBank: ABF65836.1), MERS-CoV (NCBI GenBank: AKN11072.1), HCoV-229E (NCBI GenBank: NP_073551.1), HCoV-NL63 (NCBI GenBank: YP_003767.1), HCoV-OC43 (NCBI GenBank: YP_009555241.1), HCoV-HKU1 (NCBI GenBank: ABD75513.1).
Extended Data Fig. 5 76E1 showed decreased binding activity to peptide809-823 than peptide809-828 and peptide809-833.
Isothermal titration calorimetry (ITC) analysis of the interactions between the 76E1 Fab and the SARS-CoV-2 S peptides 809-823 (a), 809-828 (b) and 809-833 (c).
Extended Data Fig. 6 76E1 is broadly resilient to natural mutations within its epitopes.
a, Frequency of mutants within the 76E1 epitope based on 5,506,473 sequences of SARS-CoV-2 available on the National Genomics Data Center (NGDC) of China National Center for Bioinformation (CNCB) as of November 19, 2021. The mutation counts more than 15 for each substitution were shown with black and the others are shown in gray. 8 mutants were identified within the 76E1 epitope residues. b, The infectivity of SARS-CoV-2 pseudoviruses with each mutation as compared with the wide type pseudovirus. n = 6. The P values are analysed by two-sided unpaired t tests. ****P < 0.0001, ns, not significant. c, Neutralizing curves of 76E1 against SARS-CoV-2 pseudoviruses with mutations. n = 3. d, The mutation numbers and IC50 valves of 76E1 to each mutant. n.d., not detectable due to the lower virus infectivity. Data are shown as the mean ± s.d. (b,c). Representative data are shown from two independent experiments (b,c).
Extended Data Fig. 7 The epitope of 76E1 is partially buried in the prefusion S trimer.
a, Structural display of peptide814-825 (orange) on the prefusion S trimer (gray) (PDB code 6XR8) of SARS-CoV-2. The epitope residues of 76E1 were shown. b, The solvent assessable surface area (SASA) and the buried surface area (BSA) of residues 814-825 in the structure of the SARS-CoV-2 S trimer in the prefusion state (PDB code 6XR8).
Extended Data Fig. 8 76E1 is synergistic with RBD antibodies to neutralize SARS-CoV-2 pseudovirus.
a, RBD antibodies CB6, P2C1F11 and 28-8 L facilitated 76E1 binding to the cell surface-expressed S proteins of SARS-CoV-2, as determined by flow cytometry, showing as Mean fluorescence intensity (MFI) values. The data are shown as the mean value of duplicates. b-j, The synergistic effect of 76E1 and RBD antibody in neutralizing SARS-CoV-2 pseudovirus. b, e, h, Neutralizing curves of 76E1 and RBD antibodies CB6 (b), P2C1F11 (e) and 28-8 L (h) alone and in combination at a constant ratio with serial 3-fold-diluted concentration from 81 times IC50. A dose of 1 represents the concentration at IC50. The constant ratio indicates the ratio between 76E1 and RBD antibody at IC50. Results are depicted as the mean ± s.d. from three replicates. c, f, i, CI (combination index) values of CB6 (c), P2C1F11 (f) and 28-8 L (i) at ED50 and ED75 (50% and 75% effective dose). The CI values was calculated using the CompuSyn program. The dotted line in red represents CI = 1 below of which was defined as synergism versus top of which as antagonism. d, g, j Summary of the synergistic effect parameters of 76E1 and CB6 (d), P2C1F11 (g) and 28-8 L (j). Dose reduction index (DRI) was calculated by comparing the dose required to reach 50% or 75% virus neutralization when 76E1 or RBD antibody was used alone and in combination. α, Concentrations of 76E1 and RBD antibody in the mixture, respectively. β, DRI of 76E1 and RBD antibody in the mixture, respectively. For all data, representative data are shown from two independent experiments.
Extended Data Fig. 9 Antibodies toward the peptide809-833 are elicited during natural SARS-CoV-2 infection.
a, ELISA-based reactivity to the peptide809-833 of plasmas from 36 COVID-19 convalescent patients and 12 healthy people whose plasma samples were collected before the emergence of COVID-19, showing as the antibody endpoint titers. b, Results from ELISA to assess the competing activity of the plasmas to 76E1. The plasmas from 36 COVID-19 convalescent patients competed 76E1 at different level, but not for the healthy people. Results are depicted as the mean value of duplicates. Representative data were shown from two independent experiments.
Extended Data Fig. 10 Peptide809-833 binding mAbs 555D6 and 125C1 could not neutralize SARS-CoV-2 pseudovirus.
a, ELISA binding activities of 9 isolated S2 targeting antibodies to peptide809-833. Data are shown as mean value of duplicates. b-c, Neutralizing activities of 555D6 (b) and 125C1 (c) against SARS-CoV-2 pseudovirus. Results are depicted as the mean ± s.d. n = 3. d-e, ELISA-based epitope alanine scanning of 555D6 and 125C1 on the peptide809-833 of SARS-CoV-2, shown as EC50 (ng/ml). Non-binding EC50 were set to over 10,000 ng/ml. f, The EC50 fold change of 555D6 and 125C1. The EC50 fold change was the ratio of EC50 of antibodies against mutants compared with wildtype peptide. g, 125C1 and 555D6 could not block the trypsin-induced S2’ cleavage. 76E1 was included as a positive control. Representative data were shown from two independent experiments (a-g).
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Sun, X., Yi, C., Zhu, Y. et al. Neutralization mechanism of a human antibody with pan-coronavirus reactivity including SARS-CoV-2. Nat Microbiol 7, 1063–1074 (2022). https://doi.org/10.1038/s41564-022-01155-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01155-3
This article is cited by
-
MVA-based vaccine candidates encoding the native or prefusion-stabilized SARS-CoV-2 spike reveal differential immunogenicity in humans
npj Vaccines (2024)
-
Human coronavirus OC43-elicited CD4+ T cells protect against SARS-CoV-2 in HLA transgenic mice
Nature Communications (2024)
-
Targetable elements in SARS-CoV-2 S2 subunit for the design of pan-coronavirus fusion inhibitors and vaccines
Signal Transduction and Targeted Therapy (2023)
-
Broadly neutralizing antibodies to SARS-CoV-2 and other human coronaviruses
Nature Reviews Immunology (2023)
-
Repeated vaccination of inactivated SARS-CoV-2 vaccine dampens neutralizing antibodies against Omicron variants in breakthrough infection
Cell Research (2023)