Abstract
Pathogenomic evidence suggests that Mycobacterium tuberculosis (MTB) evolved from an environmental ancestor similar to Mycobacterium canettii, a rare human pathogen. Although the adaptations responsible for this transition are poorly characterized, the ability to persist in humans seems to be important. We set out to identify the adaptations contributing to the evolution of persistence in MTB. We performed an experimental evolution of eight M. canettii populations in mice; four populations were derived from the isolate STB-K (phylogenomically furthest from MTB) and four from STB-D (closest to MTB), which were monitored for 15 and 6 cycles, respectively. We selected M. canettii mutants with enhanced persistence in vivo compared with the parental strains, which were phenotypically closer to MTB. Genome sequencing of 140 mutants and complementation analysis revealed that mutations in two loci were responsible for enhanced persistence. Most of the tested mutants were more resistant than their parental strains to nitric oxide, an important effector of immunity. Modern MTB were similarly more resistant to nitric oxide than M. canettii. Our findings demonstrate phenotypic convergence during experimental evolution of M. canettii, which mirrors natural evolution of MTB. Furthermore, they indicate that the ability to withstand host-induced stresses was key for the emergence of persistent MTB.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The sequencing data were deposited in the NCBI repository under the project accession codes PRJNA662472, PRJNA664307 and PRJNA665366 for the Illumina-derived genome sequences of the clones and populations derived from STB-K and STB-D, and PacBio-derived genome sequence of STB-K, respectively. The genome sequences of the 6,167 clinical MTB isolates used in this study are from the NCBI genomes repository (https://www.ncbi.nlm.nih.gov/genome/166). All other data that support the findings of this study are available from the corresponding author on request. Source data are provided with this paper.
References
Behr, M. A., Edelstein, P. H. & Ramakrishnan, L. Revisiting the timetable of tuberculosis. Brit. Med. J. 362, k2738 (2018).
Global tuberculosis report (WHO, 2019).
Chiner-Oms, A. et al. Genomic determinants of speciation and spread of the Mycobacterium tuberculosis complex. Sci. Adv. 5, eaaw3307 (2019).
Blouin, Y. et al. Progenitor ‘Mycobacterium canettii’ clone responsible for lymph node tuberculosis epidemic, Djibouti. Emerg. Inf. Dis. 20, 21–28 (2014).
Koeck, J.-L. et al. Clinical characteristics of the smooth tubercle bacilli ‘Mycobacterium canettii’ infection suggest the existence of an environmental reservoir. Clin. Microbiol. Infect. 17, 1013–1019 (2011).
Fabre, M. et al. Molecular characteristics of ‘Mycobacterium canettii’ the smooth Mycobacterium tuberculosis bacilli. Infect. Genet. Evol. 10, 1165–1173 (2010).
Lopez, B. et al. A marked difference in pathogenesis and immune response induced by different Mycobacterium tuberculosis genotypes. Clin. Exp. Immunol. 133, 30–37 (2003).
Dormans, J. et al. Correlation of virulence, lung pathology, bacterial load and delayed type hypersensitivity responses after infection with different Mycobacterium tuberculosis genotypes in BALB/c mouse model. Clin. Exp. Immunol. 137, 460–468 (2004).
Supply, P. et al. Genomic analysis of smooth tubercle bacilli provides insights into ancestry and pathoadaptation of Mycobacterium tuberculosis. Nat. Genet. 45, 172–179 (2013).
Boritsch, E. C. et al. A glimpse into the past and predictions for the future: the molecular evolution of the tuberculosis agent. Mol. Microbiol. 93, 835–852 (2014).
Fabre, M. et al. High genetic diversity revealed by variable-number tandem repeat genotyping and analysis of hsp65 gene polymorphism in a large collection of ‘Mycobacterium canettii’ strains indicates that the M. tuberculosis complex is a recently emerged clone of ‘M. canettii’. J. Clin. Microbiol. 42, 3248–3255 (2004).
Gutierrez, M. C. et al. Ancient orgin and gene mosaicism of the progenitor of Mycobacterium tuberculosis. PLoS Pathog. 1, e5 (2005).
Thomson, M. et al. Modulation of the cAMP levels with a conserved actinobacteria phosphodiesterase enzyme reduces antimicrobial tolerance in mycobacteria. Preprint at bioRxiv https://doi.org/10.1101/2020.08.26.267864 (2020).
Kramnik, I., Demant, P. & Bloom, B. B. in Genetics and Tuberculosis: Novartis Foundation Symposium 217 (eds Chadwick, D. J. & Cardew, G.) Ch. 9 (Novartis, 1998).
Kramnik, I., Dietrich, W. F., Demant, P. & Bloom, B. R. Genetic control of resistance to experimental infection with virulent Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 97, 8560–8565 (2000).
Lanoix, J. P., Lenaerts, A. J. & Nuermberger, E. L. Heterogeneous disease progression and treatment response in a C3HeB/FeJ mouse model of tuberculosis. Dis. Mod. Mech. 8, 603–610 (2015).
Harper, J. et al. Mouse model of necrotic tuberculosis granulomas develops hypoxic lesions. J. Inf. Dis. 205, 595–602 (2012).
Singh, A. et al. Mycobacterium tuberculosis WhiB3 maintains redox homeostasis by regulating virulence lipid anabolism to modulate macrophage response. PLoS Pathog. 5, e1000545 (2009).
Vandal, O. H., Pierini, L. M., Schnappinger, D., Nathan, C. F. & Ehrt, S. A membrane protein preserves intrabacterial pH in intraphagosomal Mycobacterium tuberculosis. Nat. Med. 14, 849–854 (2008).
Darwin, K. H., Ehrt, S., Gutierrez-Ramos, J. C., Weich, N. & Nathan, C. F. The proteasome of Mycobacterium tuberculosis is required for resistance to nitric oxide. Science 302, 1963–1966 (2003).
Albrethsen, J. et al. Proteomic profiling of Mycobacterium tuberculosis identifies nutrient-starvation-responsive toxin-antitoxin systems. Mol. Cell. Prot. 12, 1180–1191 (2013).
Schaible, U. E., Sturgill-Koszycki, S., Schlesinger, P. H. & Russell, D. G. Cytokine activation leads to acidification and increases maturation of Mycobacterium avium-containing phagosomes in murine macrophages. J. Immunol. 160, 1290–1296 (1998).
MacMicking, J. D., Taylor, G. A. & McKinney, J. D. Immune control of tuberculosis by IFN-γ-inducible LRG-47. Science 302, 654–659 (2003).
Neyrolles, O., Wolschendorf, F., Mitra, A. & Niederweis, M. Mycobacteria, metals, and the macrophage. Immunol. Rev. 264, 249–263 (2015).
Rifat, D., Bishai, W. R. & Karakousis, P. C. Phosphate depletion: a novel trigger for Mycobacterium tuberculosis persistence. J. Inf. Dis. 200, 1126–1135 (2009).
Rengarajan, J., Bloom, B. R. & Rubin, E. J. Genome-wide requirements for Mycobacterium tuberculosis adaptation and survival in macrophages. Proc. Natl Acad. Sci. USA 102, 8327–8332 (2005).
Wolschendorf, F. et al. Copper resistance is essential for virulence of Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 108, 1621–1626 (2011).
Botella, H. et al. Mycobacterial P1-type ATPases mediate resistance to zinc poisoning in human macrophages. Cell Host Microbe 10, 248–259 (2011).
MacMicking, J. D. et al. Identification of nitric oxide synthase as a protective locus against tuberculosis. Proc. Natl Acad. Sci. USA 94, 5243–5248 (1997).
Nicholson, S. et al. Inducible nitric oxide synthase in pulmonary alveolar macrophages from patients with tuberculosis. J. Exp. Med. 183, 2293–2302 (1996).
Rich, E. A. et al. Mycobacterium tuberculosis (MTB)-stimulated production of nitric oxide by human alveolar macrophages and relationship of nitric oxide production to growth inhibition of MTB. Tuber. Lung Dis. 78, 247–255 (1997).
Martineau, A. R. et al. Neutrophil-mediated innate immune resistance to mycobacteria. J. Clin. Invest. 117, 1988–1994 (2007).
Martineau, A. R. et al. IFN-γ- and TNF-independent vitamin D-inducible human suppression of mycobacteria: the role of cathelicidin LL-37. J. Immunol. 178, 7190–7198 (2007).
Liu, P. T., Stenger, S., Tang, D. H. & Modlin, R. L. Cutting edge: vitamin D-mediated human antimicrobial activity against Mycobacterium tuberculosis is dependent on the induction of cathelicidin. J. Immunol. 179, 2060–2063 (2007).
Stenger, S. et al. An antimicrobial activity of cytolytic T cells mediated by granulysin. Science 282, 121–125 (1998).
Buchmeier, N. et al. A parallel intraphagosomal survival strategy shared by Mycobacterium tuberculosis and Salmonella enterica. Mol. Microbiol. 35, 1375–1382 (2000).
Ng, V. H., Cox, J. S., Sousa, A. O., MacMicking, J. D. & McKinney, J. D. Role of KatG catalase-peroxidase in mycobacterial pathogenesis: countering the phagocyte oxidative burst. Mol. Microbiol. 52, 1291–1302 (2004).
Primm, T. P. et al. The stringent response of Mycobacterium tuberculosis is required for long-term survival. J. Bacteriol. 182, 4889–4898 (2000).
Lieberman, T. D. et al. Parallel bacterial evolution within multiple patients identifies candidate pathogenicity genes. Nat. Genet. 43, 1275–1280 (2011).
Lenski, R. E. Chance and necessity in the evolution of a bacterial pathogen. Nat. Genet. 43, 1174–1176 (2011).
Stern, D. L. The genetic causes of convergent evolution. Nat. Rev. Genet. 14, 751–764 (2013).
Didelot, X., Walker, A. S., Peto, T. E., Crook, D. W. & Wilson, D. J. Within-host evolution of bacterial pathogens. Nat. Rev. Microbiol. 14, 150–162 (2016).
Wirth, T. et al. Niche specialization and spread of Staphylococcus capitis involved in neonatal sepsis. Nat. Microbiol. 5, 735–745 (2020).
Chan, J., Xing, Y., Magliozzo, R. S. & Bloom, B. R. Killing of virulent Mycobacterium tuberculosis by reactive nitrogen intermediates produced by activated murine macrophages. J. Exp. Med. 175, 1111–1122 (1992).
Flesch, I. E. & Kaufmann, S. H. Mechanisms involved in mycobacterial growth inhibition by gamma interferon-activated bone marrow macrophages: role of reactive nitrogen intermediates. Infect. Immun. 59, 3213–3218 (1991).
Long, R., Light, B. & Talbot, J. A. Mycobacteriocidal action of exogenous nitric oxide. Antimicrob. Agents Chemother. 43, 403–405 (1999).
Nozaki, Y., Hasegawa, Y., Ichiyama, S., Nakashima, I. & Shimokata, K. Mechanism of nitric oxide-dependent killing of Mycobacterium bovis BCG in human alveolar macrophages. Infect. Immun. 65, 3644–3647 (1997).
Mishra, B. B. et al. Nitric oxide prevents a pathogen-permissive granulocytic inflammation during tuberculosis. Nat. Microbiol. 2, 17072 (2017).
Mishra, B. B. et al. Nitric oxide controls the immunopathology of tuberculosis by inhibiting NLRP3 inflammasome-dependent processing of IL-1β. Nat. Immunol. 14, 52–60 (2013).
Chandrasekera, N. S. et al. Improved phenoxyalkylbenzimidazoles with activity against Mycobacterium tuberculosis appear to target QcrB. ACS Infect. Dis. 3, 898–916 (2017).
O’Malley, T. et al. Imidazopyridine compounds inhibit mycobacterial growth by depleting ATP levels. Antimicrob. Agents Chemother. 62, e02439-17 (2018).
Nazarova, E. V. et al. The genetic requirements of fatty acid import by Mycobacterium tuberculosis within macrophages. eLife 8, e4362 (2019).
Karboul, A. et al. Frequent homologous recombination events in Mycobacterium tuberculosis PE/PPE multigene families: potential role in antigenic variability. J. Bacteriol. 190, 7838–7846 (2008).
Ates, L. S. et al. Mutations in ppe38 block PE_PGRS secretion and increase virulence of Mycobacterium tuberculosis. Nat. Microbiol. 3, 181–188 (2018).
Wang, Q. et al. PE/PPE proteins mediate nutrient transport across the outer membrane of Mycobacterium tuberculosis. Science 367, 1147–1151 (2020).
Helsen, J. et al. Gene loss predictably drives evolutionary adaptation. Mol. Biol. Evol. 37, 2989–3002 (2020).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
van Kessel, J. C. & Hatfull, G. F. Mycobacterial recombineering. Methods Mol. Biol. 435, 203–215 (2008).
Pelicic, V., Reyrat, J.-M. & Gicquel, B. Generation of unmarked directed mutations in mycobacteria, using sucrose counterselectable suicide vectors. Mol. Microbiol. 20, 919–925 (1996).
Stover, C. K. et al. New use of BCG for recombinant vaccines. Nature 351, 456–460 (1991).
Le Dantec, C., Winter, N., Gicquel, B., Vincent, V. & Picardeau, M. Genomic sequence and transcriptional analysis of a 23-kb mycobacterial linear plasmid: evidence for horizontal transfer and identification of plasmid maintenance systems. J. Bacteriol. 183, 2157–2164 (2001).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Rodrigues, L., Viveiros, M. & Ainsa, J. A. Measuring efflux and permeability in mycobacteria. Methods Mol. Biol. 1285, 227–239 (2015).
Huson, D. H. & Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 23, 254–267 (2006).
Comas, I. et al. Human T cell epitopes of Mycobacterium tuberculosis are evolutionarily hyperconserved. Nat. Genet. 42, 498–503 (2010).
Coll, F. et al. A robust SNP barcode for typing Mycobacterium tuberculosis complex strains. Nat. Commun. 5, 4812 (2014).
Merker, M. et al. Phylogenetically informative mutations in genes implicated in antibiotic resistance in Mycobacterium tuberculosis complex. Genome Med. 12, 27 (2020).
Acknowledgements
The authors thank T. Seemann for their initial help with the network phylogeny analysis. The authors thank E. Russo and P. Bousso (Institut Pasteur, France) for the gift of NOS2−/− mice. The authors acknowledge support from the GenoToul (Toulouse, France), Bioinformatics and GeT platforms for sequencing, computing and storage resources, TRI platform for image analyses and the Anexplo platform for animal experimentation. The authors thank J. Korlach, P. Baybayan and J. Harting from Pacific Biosciences for sequencing the parental STB-K strain. The authors thank S. Milia for technical assistance at the Experimental Histopathology Facility of INSERM/UPS US006 CREFRE, Toulouse Purpan, France. The authors also thank Life Science Editors for their professional editing assistance during the preparation of the manuscript. This project received funding from the European Respiratory Society and the European Union’s H2020 research and innovation programme under the Marie Sklodowska–Curie grant agreement no. 713406 awarded to A.C.A. This work was also supported by grants from the French National Research Agency (TBemerg contract no. ANR-16-CE35-0009 awarded to R.B., P.S. and C. Guilhot; Equipex ANINFIMIP ANR-11-EQUIPEX-0003 to C. Guilhot; Labex ANR-10-LABX-62-IBEID awarded to R.B.), the European Union (PathoNgenTrace contract no. FP7-278864 awarded to P.S.); and the Fondation pour la Recherche Médicale (Equipe FRM 2016 DEQ20160334879), the Fondation Bettencourt Schueller and the Fondation MSDAvenir (Fight-TB project) to C. Guilhot.
Author information
Authors and Affiliations
Contributions
C. Guilhot, R.B. and P.S. planned the project. A.C.A., C.A.-D. and C. Guilhot designed the experiments. A.C.A., W. M., C. Gaudin, F.M., A.H., C.A.-D., A. Peixoto, R.A., A. Pawlik, W.F., C.B. and C. Guilhot performed the experiments. A.C.A., W.M., C. Gaudin, A.V., A.H., C.A.-D., A. Peixoto, R.A., A. Pawlik, R.B., P.S. and C. Guilhot analysed the data. A.C.A., C.A.-D., R.B., P.S. and C. Guilhot contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Luiz Pedro de Carvalho and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phylogenetic tree showing the distribution of M. canettii and MTB strains.
Network phylogeny inferred among 28 genomes by NeighborNet analysis, based on pairwise alignments of whole-genome SNP data from a selection of strains (17 M. canettii, 8 MTB from various phylogenetic lineages and 3 strains from the M. africanum and M. bovis lineages), depicted by the SplitsTree software64. The whole SNP dataset was originally generated for Supplementary Fig. 1 of reference10, using genome data described by4,9,65. Strains used in experimental evolution experiments are coloured in red.
Extended Data Fig. 2 Experimental evolution in mice selects M. canettii clones with enhanced persistence.
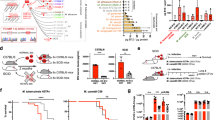
Persistence phenotypes of individual STB-K clones recovered from STB-K evolution lineages A-D after cycles a) 6, b) 12 and c) 15. Mice were i.n. infected with 103 c.f.u. of a) KC6A1, KC6A2, KC6B1, KC6B2, KC6C1, KC6C2, KC6D1, and KC6D2, b) KC12A1, KC12B1, KC12C1, and KC12D1, or c) KC15A1, KC15B1, KC15C1, and KC15D1. Parental STB-K and control MTB H37Rv strains were included in all three experiments. The bacterial burden in the lungs and spleen was assessed at various intervals post infection by plating serial dilutions of organ homogenates on 7H11 agar plates. Data are shown as log10 c.f.u. for individual mice and the mean. n = 4 individual mice for each time point and each strain over one experiment (when only 3 values are visible in the figure, it indicates that an animal died or had to be euthanized due to health issues prior to the end point). When bacteria were undetectable, the c.f.u. value was set at the limit of detection (log10 1.7; indicated by the broken horizontal line).
Extended Data Fig. 3 KC9A1 and KC9C1 display similar bacterial burden to STB-K during the acute phase of infection.
C3HeB/FeJ mice were i.v. infected with approximately 103 c.f.u. of STB-K, KC9A1, KC9C1, or MTB H37Rv, and c.f.u. counts in the a) lung and b) spleen were quantified at 21 days post infection by plating dilutions of the organ homogenate on 7H11 agar plates. Results are shown as the log10 c.f.u. for individual mice (5 per group) and the mean, and are representative of two independent experiments. Non-significant by one-way ANOVA with the Bonferroni post-hoc test. When bacteria were undetectable, the c.f.u. value was set at the limit of detection (log10 1.7; indicated by the broken horizontal line).
Extended Data Fig. 4 KC9A1 and KC9C1 do not induce a greater inflammatory response in the lungs of mice than STB-K.
Lungs were isolated from C3HeB/FeJ mice 76 days after i.n. infection with approximately 200 c.f.u. of STB-K, KC9A1, KC9C1, or H37Rv for histopathology. Lung samples were fixed and stained with haematoxylin and eosin, and analysed using Pannoramic Viewer. For each mouse, one entire lung section was analysed and the images shown are representative. n = 5 individual mice for each strain over one experiment (except for KC9C1 where n = 4 mice as one had to be euthanized due to health issues prior to the end point). The bars correspond to 2 mm for 0.5x images and 50 µm for 20x images.
Extended Data Fig. 5 STB-K and KC9A1 induce a similar recruitment of immune cells to the lungs whereas the pattern for KC9C1 is closer to that of H37Rv.
Lung samples from C3HeB/FeJ mice 76 days after i.n. infection with approximately 200 c.f.u. of STB-K, KC9A1, KC9C1, or H37Rv were fixed, and analysed by immunohistochemistry. Serial lung sections, one per infected mouse and per primary antibody, were labelled with antibodies against a) B lymphocytes (B220), b) T lymphocytes (CD3), c) macrophages (F4/80), and d) neutrophils (neutrophil marker). At least four sections from 4 mice were analysed with each antibody. The percentage of each cell type in the various lesions was quantified. *P < 0.05, **P < 0.01, ***P < 0.001, by Brown–Forsythe and Welch one-way ANOVA with the Games-Howell post-hoc test. a) H37Rv vs STB-K; P < 0.0001, STB-K vs KC9C1; P = 0.0020, b) H37Rv vs STB-K; P = 0.0462, STB-K vs KC9C1; P < 0.0001, c) H37Rv vs STB-K; P = 0.0026, STB-K vs KC9A1; P = 0.0125, d) H37Rv vs STB-K; P < 0.0001, STB-K vs KC9C1; P < 0.0001.
Extended Data Fig. 6 The persistent phenotypes of KC9A1 and KC9C1 are not associated with a change in cytokine production.
C3HeB/FeJ mice were i.v. infected with approximately 103 c.f.u. of STB-K, KC9A1, KC9C1, or MTB H37Rv, and IL-10, IFN-γ and TNF concentrations in the lung and spleen homogenates were quantified at a) 21 days and b) 76 days post infection by ELISA. Mean cytokine concentrations are shown, in addition to the concentrations for individual mice. n = 5 individual mice for each time point and each strain, except for H37Rv which had n = 4 individual mice at day 70 as one mouse had to be euthanized due to health issues prior to the end point. *P < 0.05 by one-way ANOVA with the Bonferroni post-hoc test. a) STB-K vs KC9A1; P = 0.0102.
Extended Data Fig. 7 The pe18-ppe26 deletion does not confer enhanced persistence to STB-D.
BALB/c mice were i.n. infected with approximately 103 c.f.u. of M. canettii D::EV, DΔppe25–ppe27::EV,::D,::K, or::C1, and c.f.u. counts in the lung were quantified at 1 and 70 days post infection by plating dilutions of the organ homogenate on 7H11 agar plates. Results are shown as a) the log10 c.f.u. per lung for individual mice and the mean, and b) normalized values (log10 c.f.u. D77 – log10 c.f.u. D1), from an experiment performed with 4 mice per group. Differences between DΔppe25–ppe27::EV,::D,::K and::C1 were deemed non-significant by a) two-way and b) one-way ANOVA with the Bonferroni post-hoc test. The statistical difference between D::EV and the other groups is not shown. When bacteria were undetectable, the c.f.u. value was set at the limit of detection (log10 1.7; indicated by the broken line) for the purposes of statistical analysis.
Extended Data Fig. 8 STB-K and the two persistent clones KC9A1 and KC9C1 display similar resistance to H2O2 and grow similarly in phosphate-limited medium or at acidic pH.
a, Flasks of 7H9/ADC/Tween at pH 6.8 were inoculated with STB-K, KC9A1, KC9C1, and MTB H37Rv at a final OD600nm of approximately 0.05 and growth was evaluated by monitoring the OD600nm. Data are represented as the mean and standard deviation from three independent experiments. b, STB-K, KC9A1, KC9C1, and H37Rv were inoculated to a final OD600nm of approximately 0.05 into flasks containing 7H9 supplemented with ADC and 0.05% Tyloxapol adjusted to pH 5, 5.5, 6 or 6.6. Strain growth was evaluated by monitoring the OD600nm. Data are represented as the mean and standard deviation from three independent experiments. c, STB-K, KC9A1, KC9C1, and H37Rv were inoculated to a final OD600nm of approximately 0.05 into flasks containing Sauton’s medium where the final concentration of phosphate had been adjusted to either 0 µM, 50 µM or 2870 µM supplemented with 0.05% Tween 80. Strain growth was evaluated by monitoring the OD600nm. Data are represented as the mean and standard deviation from three independent experiments. d, Flasks of 7H9/ADC/Tween + 10 mM H2O2 were inoculated with 105 c.f.u. of STB-K, KC9A1, KC9C1 and H37Rv. Bacterial survival was monitored by evaluating c.f.u. numbers at 0, 16 and 24 h post treatment with H2O2. Results are shown as log10 c.f.u./ml for individual data points from eight independent experiments and the mean. When bacteria were undetectable, the c.f.u. value was set at the limit of detection (log10 1.3; indicated by the horizontal broken line) for the purposes of statistical analysis. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001 by two-way ANOVA with the Bonferroni post-hoc test. T0: KC9C1 vs H37Rv; P = 0.0264. 16 h: STB-K vs KC9C1; P = 0.0237, STB-K vs H37Rv; P < 0.0001, KC9A1 vs H37Rv; P = 0.0022, KC9C1 vs H37Rv; P = 0.0319. 24 h: STB-K vs H37Rv; P < 0.0001, KC9A1 vs H37Rv; P = 0.0002, KC9C1 vs H37Rv; P = 0.0001.
Extended Data Fig. 9 The persistence phenotype of the rv1339 STB-K mutant (KC9A1) is due in part to enhanced resistance to NO.
NOS2-/- mice or C57BL/6 wild-type controls (WT) were infected via aerosol with approximately 100 c.f.u. of STB-K or KC9A1 and bacterial burden in the a) lung and b) spleen was quantified at 56 days post infection by c.f.u. plating on solid medium. Results are shown as the log10 c.f.u. for individual mice and the mean. n = 5 individual mice for each time point and each strain (except for KC9A1 which had n = 6 NOS2-/- individual mice) over one experiment. When bacteria were undetectable, the c.f.u. value was set at the limit of detection (log10 2; indicated by the broken horizontal line) for the purposes of statistical analysis. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001 by one-way ANOVA with the Bonferroni post-hoc test. a, STB-K (WT) vs KC9A1 (WT); P = 0.0015, STB-K (WT) vs KC9A1 (NOS2-/-); P < 0.0001, KC9A1 (WT) vs STB-K (NOS2-/-); P = 0.0037; STB-K (NOS2-/-) vs KC9A1 (NOS2-/-); P = 0.0001, b) STB-K (WT) vs KC9A1 (WT); P = 0.0005, STB-K (WT) vs KC9A1 (NOS2-/-); P < 0.0001, KC9A1 (WT) vs STB-K (NOS2-/-); P = 0.0357, KC9A1 (WT) vs KC9A1 (NOS2-/-); P < 0.0001, STB-K (NOS2-/-) vs KC9A1 (NOS2-/-); P < 0.0001. Using this route of infection and the C57BL/6 mouse strain, the bacterial burden in the lungs and spleen at day 56 post infection was consistent with the enhanced persistence phenotype of KC9A1 when compared with STB-K. In NOS2-/- mice, the persistence of STB-K in the spleen and KC9A1 in both organs were increased showing that NO is an important factor reducing STB-K and KC9A1 persistence. However, the difference between STB-K and KC9A1 was maintained in NOS2-/- mice, and even increased in comparison to C57BL/6 mice, indicating that the rv1339 mutation also confers strong resistance to NOS2-/- independent mechanisms. c, BMDMs from BALB/c mice were either unstimulated, or stimulated with IFN-γ (100 U/ml) + LPS (100 ng/ml), or stimulated with IFN-γ + LPS and treated with the inducible nitric oxide synthase (iNOS) inhibitor L-NAME (4 mM). BMDMs (2.5 ×105 cells/well) were infected at a multiplicity of infection (MOI) of 0.5 bacteria/cell with STB-K, KC9A1, KC9C1 or MTB H37Rv and the number of surviving bacteria were evaluated by plating on 7H11 solid medium. The indicated value are the Log10 (ratio of bacterial load at day 3 over bacterial load at day 0 in the same experiment). *P < 0.05 by one-way repeated measure ANOVA with the Bonferroni post-hoc test. c) STB-K (IFN-γ + LPS) vs KC9A1 (IFN-γ + LPS); P = 0.0225, STB-K (IFN-γ + LPS) vs KC9C1 (IFN-γ + LPS); P = 0.0437. In these experiments, stimulation of BMDMs with IFN-γ and LPS reduced the survival of the 4 strains and this effect is partially suppressed by inhibiting iNOS.
Extended Data Fig. 10 Distribution of mutations within the rv1339 orthologue among MTB complex lineages.
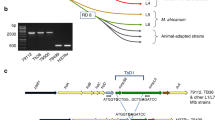
a, Wild-type sequences (WT), as well as fixed synonymous and non-synonymous mutations were identified in a collection of genome assemblies from 6167 isolates, which were classified in lineages or main sub-lineages based on phylogenetic SNPs and the conventional nomenclature66,67. b, Nucleotide (left) and corresponding amino acid (right) changes or combined changes detected in isolates from different (sub-) lineages are shown. The graph in the center is a zoom-in that represents mutations detected in 8 isolates or less. The G57A mutation detected in >400 isolates (bottom) is a synonymous SNP shared by basal lineages of the MTB complex (for example lineages 1, 5, 6 and animal lineages). SM, synonymous; FS, frameshift. These analyses identified about 30 different non-synonymous mutations in rv1339 in this dataset, which were phylogenetically linked in all cases, that is found in one (sub)lineage only. Of note, two frameshift-causing indels and one non-sense mutation were detected, which were however represented only in 2 isolates at most. These results indicate a strong conservation of the rv1339 gene among MTB isolates.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3, Supplementary Tables 1–5, uncropped pictures of gels for Supplementary Figs. 1–3.
Supplementary Tables
Supplementary Table 1. Mutations identified in STB-K. Supplementary Table 2. Mutations identified in STB-D. Supplementary Table 3. M. canettii, MTB and M. smegmatis strains used in this study. Supplementary Table 4. Plasmids used in this study. Supplementary Table 5. Primers used in this study.
Source data
Source Data Fig. 1
Numerical data.
Source Data Fig. 2
Numerical data.
Source Data Fig. 3
Numerical data.
Source Data Fig. 4
Numerical data.
Source Data Fig. 5
Numerical data.
Source Data Fig. 6
Numerical data.
Source Data Extended Data Fig. 2
Numerical data.
Source Data Extended Data Fig. 3
Numerical data.
Source Data Extended Data Fig. 5
Numerical data.
Source Data Extended Data Fig. 6
Numerical data.
Source Data Extended Data Fig. 7
Numerical data.
Source Data Extended Data Fig. 8
Numerical data.
Source Data Extended Data Fig. 9
Numerical data.
Source Data Extended Data Fig. 10
Numerical data.
Rights and permissions
About this article
Cite this article
Allen, A.C., Malaga, W., Gaudin, C. et al. Parallel in vivo experimental evolution reveals that increased stress resistance was key for the emergence of persistent tuberculosis bacilli. Nat Microbiol 6, 1082–1093 (2021). https://doi.org/10.1038/s41564-021-00938-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-021-00938-4
This article is cited by
-
A smooth tubercle bacillus from Ethiopia phylogenetically close to the Mycobacterium tuberculosis complex
Nature Communications (2023)
-
Emergence of persistent tubercle bacilli
Nature Microbiology (2021)
-
2021 in review
Nature Microbiology (2021)