Abstract
Bacteria respond to changes in their environment with specific transcriptional programmes, but even within genetically identical populations these programmes are not homogenously expressed1. Such transcriptional heterogeneity between individual bacteria allows genetically clonal communities to develop a complex array of phenotypes1, examples of which include persisters that resist antibiotic treatment and metabolically specialized cells that emerge under nutrient-limiting conditions2. Fluorescent reporter constructs have played a pivotal role in deciphering heterogeneous gene expression within bacterial populations3 but have been limited to recording the activity of single genes in a few genetically tractable model species, whereas the vast majority of bacteria remain difficult to engineer and/or even to cultivate. Single-cell transcriptomics is revolutionizing the analysis of phenotypic cell-to-cell variation in eukaryotes, but technical hurdles have prevented its robust application to prokaryotes. Here, using an improved poly(A)-independent single-cell RNA-sequencing protocol, we report the faithful capture of growth-dependent gene expression patterns in individual Salmonella and Pseudomonas bacteria across all RNA classes and genomic regions. These transcriptomes provide important reference points for single-cell RNA-sequencing of other bacterial species, mixed microbial communities and host–pathogen interactions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Code availability
All codes used to perform the analysis and reproduce the figures are available on GitHub: https://github.com/saliba-lab/single-bacterium-transcriptome-profiling.
References
Ackermann, M. A functional perspective on phenotypic heterogeneity in microorganisms. Nat. Rev. Microbiol. 13, 497–508 (2015).
Gollan, B., Grabe, G., Michaux, C. & Helaine, S. Bacterial persisters and infection: past, present, and progressing. Annu. Rev. Microbiol. 73, 359–385 (2019).
Kreibich, S. & Hardt, W. D. Experimental approaches to phenotypic diversity in infection. Curr. Opin. Microbiol. 27, 25–36 (2015).
Gasch, A. P. et al. Single-cell RNA sequencing reveals intrinsic and extrinsic regulatory heterogeneity in yeast responding to stress. PLoS Biol. 15, e2004050 (2017).
Muller, L. S. M. et al. Genome organization and DNA accessibility control antigenic variation in trypanosomes. Nature 563, 121–125 (2018).
Stubbington, M. J. T., Rozenblatt-Rosen, O., Regev, A. & Teichmann, S. A. Single-cell transcriptomics to explore the immune system in health and disease. Science 358, 58–63 (2017).
Kang, Y. et al. Transcript amplification from single bacterium for transcriptome analysis. Genome Res. 21, 925–935 (2011).
Wang, J., Chen, L., Chen, Z. & Zhang, W. RNA-seq based transcriptomic analysis of single bacterial cells. Integr. Biol. (Camb.) 7, 1466–1476 (2015).
Avital, G. et al. scDual-Seq: mapping the gene regulatory program of Salmonella infection by host and pathogen single-cell RNA-sequencing. Genome Biol. 18, 200 (2017).
Betin, V. et al. Hybridization-based capture of pathogen mRNA enables paired host–pathogen transcriptional analysis. Sci. Rep. 9, 19244 (2019).
Penaranda, C. & Hung, D. T. Single-cell RNA sequencing to understand host–pathogen interactions. ACS Infect. Dis. 5, 336–344 (2019).
Milo, R. & Phillips, R. Cell Biology by the Numbers (Garland Science, 2015).
Bagnoli, J. W. et al. Sensitive and powerful single-cell RNA sequencing using mcSCRB-seq. Nat. Commun. 9, 2937 (2018).
Taniguchi, Y. et al. Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 329, 533–538 (2010).
Sheng, K., Cao, W., Niu, Y., Deng, Q. & Zong, C. Effective detection of variation in single-cell transcriptomes using MATQ-seq. Nat. Methods 14, 267–270 (2017).
Kroger, C. et al. An infection-relevant transcriptomic compendium for Salmonella enterica serovar Typhimurium. Cell Host Microbe 14, 683–695 (2013).
Smirnov, A. et al. Grad-seq guides the discovery of ProQ as a major small RNA-binding protein. Proc. Natl Acad. Sci. USA 113, 11591–11596 (2016).
Chao, Y. et al. In vivo cleavage map illuminates the central role of RNase E in coding and non-coding RNA pathways. Mol. Cell 65, 39–51 (2017).
Hor, J., Matera, G., Vogel, J., Gottesman, S. & Storz, G. Trans-acting small RNAs and their effects on gene expression in Escherichia coli and Salmonella enterica. EcoSal Plus https://doi.org/10.1128/ecosalplus.ESP-0030-2019 (2020).
Westermann, A. J. et al. Dual RNA-seq unveils noncoding RNA functions in host–pathogen interactions. Nature 529, 496–501 (2016).
Westermann, A. J. & Vogel, J. Host–pathogen transcriptomics by dual RNA-seq. Methods Mol. Biol. 1737, 59–75 (2018).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013).
Haas, B. J., Chin, M., Nusbaum, C., Birren, B. W. & Livny, J. How deep is deep enough for RNA-Seq profiling of bacterial transcriptomes? BMC Genomics 13, 734 (2012).
Blattman, S. B., Jiang, W., Oikonomou, P. & Tavazoie, S. Prokaryotic single-cell RNA sequencing by in situ combinatorial indexing. Nat. Microbiol. https://doi.org/10.1038/s41564-020-0729-6 (2020).
Barczak, A. K. et al. RNA signatures allow rapid identification of pathogens and antibiotic susceptibilities. Proc. Natl Acad. Sci. USA 109, 6217–6222 (2012).
Gu, W. et al. Depletion of Abundant Sequences by Hybridization (DASH): using Cas9 to remove unwanted high-abundance species in sequencing libraries and molecular counting applications. Genome Biol. 17, 41 (2016).
Prezza, G. et al. Improved bacterial RNA-seq by Cas9-based depletion of ribosomal RNA reads. RNA 26, 1069–1078 (2020).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank S. Gorski for critical comments on the manuscript; K. Secener and X. Hartmann for performing preliminary experiments; C. Michaux for advice on bacterial culture; L. Barquist and F. Erhard for advice on statistical analysis; and P. Arampatzi, T. Heckel (SysMed Würzburg) and R. Geffers (Genome Analytics, HZI) for conducting the sequencing.
Author information
Authors and Affiliations
Contributions
F.I. and C.H. conducted experiments. E.V. performed data analysis. A.-E.S. designed research. J.V and A.-E.S. directed research. A.-E.S. and J.V. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Single-bacteria sorting.
Evaluation of single-bacteria sorting efficiency. On a LB agar molded in a 96 well format dish, GFP-fluorescent and non-fluorescent Salmonella 100, 10, 2 and 1 bacteria are sorted systematically in alternative manner. After overnight growth colonies are observed (upper bright field image) and absent colonies are indicated with a red circle. Observing the plate under fluorescent reader allows to count the sorting mismatches (indicated by an orange square) and define the sorting precision.
Extended Data Fig. 2 Comparison of library size and number of detected genes between different growth conditions.
a, Violin plots represent the size of libraries cultured in the different growth conditions for 10-pooled and single bacteria. b, Scatter plots depicting the relation between the number of detected genes and library size. All libraries labelled as outliers have been removed from the analysis (Methods, Supplementary Table 1).
Extended Data Fig. 3 Coverage plot and read densities.
a, Read densities of structural genes with a high expression in 10- and single-cell libraries. b, Gene coverage of selected differentially expressed genes in the respective conditions in 10-cell libraries. c, Read densities of csrA in 10-pooled libraries and respective conditions overlapping to 5’ UTR. Libraries have been automatically log scaled by Integrative Genomics Viewer.
Extended Data Fig. 4 Technical noise assessment of 10-pooled bacteria and single‐bacteria RNA‐seq data.
a, b, Coefficient of variation is plotted against the log2 (average of normalized read count) for (a) 10-pooled bacteria and (b) single bacteria conditions. Color code refers to salt (NaCl) shock (green), anaerobic shock (red) and late stationary phase (blue). Such analysis are routinely conducted when analyzing mammalian single-cell RNA-seq data (see ref. 23). Note that upper bound corresponds to CV=√n with n the number of samples analyzed.
Extended Data Fig. 5 Correlations between matching growth conditions.
a, Scatter plots show the correlation between the matching pooled (10-pooled) and single bacteria in the respective conditions with the associated Spearman’s correlation coefficient p<2.2e10-16 in all three growth conditions. b, Scatter plots show the correlation between 10-pooled and single bacteria (this study) and bulk RNA-seq in the respective conditions with the associated Spearman’s correlation coefficient (ρ) with the associated p-values.
Extended Data Fig. 6 Technical parameters associated to the Principal Component Analysis (PCA) of 10-pooled and single bacteria transcriptomes.
a, b, For single-bacteria (a) and 10-pooled bacteria transcriptomes (b) library size (left), number of detected genes (middle) and genes associated with PC loading 1 and 2 are overlaid on top of the PCA. The top 15 genes with the highest contribution to PC were selected and their related loading vectors are shown on the PCA plot. The vectors show how the original variables contribute to creating the principal component. c, Scree plots give the variance associated to each PC loading.
Extended Data Fig. 7 Application of MATQ-seq to Pseudomonas aeruginosa.
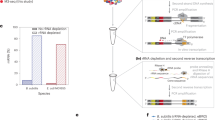
a, Scatter plots depicting the relation between the number of detected genes versus library size for single bacteria libraries (blue) and 10-pooled bacteria libraries (red). b, Violin plots representing the number genes detected for 10-pooled and single bacteria. c, Average proportions of transcript categories after removal of unmapped reads obtained for 10-pooled and single bacteria. CDS: coding sequences; IGR, intergenic region; ncRNA: non-coding RNA; sRNA: small RNA; other: all other RNA classes (Supplementary Tables 7 and 8). d, Representative reads aligning to the reference sequence of tmRNA-encoding ssrA gene (±300 bp upstream and downstream the CDS) across 10-pooled and single bacteria.
Supplementary information
Source data
Source Data Fig. 2
Raw source data.
Source Data Fig. 3
Raw source data.
Source Data Extended Data Fig. 2
Raw source data.
Source Data Extended Data Fig. 4
Raw source data.
Source Data Extended Data Fig. 5
Raw source data.
Source Data Extended Data Fig. 6
Raw source data.
Source Data Extended Data Fig. 7
Raw source data.
Rights and permissions
About this article
Cite this article
Imdahl, F., Vafadarnejad, E., Homberger, C. et al. Single-cell RNA-sequencing reports growth-condition-specific global transcriptomes of individual bacteria. Nat Microbiol 5, 1202–1206 (2020). https://doi.org/10.1038/s41564-020-0774-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-0774-1
This article is cited by
-
Transcription–replication interactions reveal bacterial genome regulation
Nature (2024)
-
Co-transcriptional gene regulation in eukaryotes and prokaryotes
Nature Reviews Molecular Cell Biology (2024)
-
Emerging tools for uncovering genetic and transcriptomic heterogeneities in bacteria
Biophysical Reviews (2024)
-
Plant biotechnology research with single-cell transcriptome: recent advancements and prospects
Plant Cell Reports (2024)
-
Multidimensional specialization and generalization are pervasive in soil prokaryotes
Nature Ecology & Evolution (2023)