Abstract
Sensing of microbes activates the innate immune system, depending on functional mitochondria. However, pathogenic bacteria inhibit mitochondrial activity by delivering toxins via outer membrane vesicles (OMVs). How macrophages respond to pathogenic microbes that target mitochondria remains unclear. Here, we show that macrophages exposed to OMVs from Neisseria gonorrhoeae, uropathogenic Escherichia coli and Pseudomonas aeruginosa induce mitochondrial apoptosis and NLRP3 inflammasome activation. OMVs and toxins that cause mitochondrial dysfunction trigger inhibition of host protein synthesis, which depletes the unstable BCL-2 family member MCL-1 and induces BAK-dependent mitochondrial apoptosis. In parallel with caspase-11-mediated pyroptosis, mitochondrial apoptosis and potassium ion efflux activate the NLRP3 inflammasome after OMV exposure in vitro. Importantly, in the in vivo setting, the activation and release of interleukin-1β in response to N. gonorrhoeae OMVs is regulated by mitochondrial apoptosis. Our data highlight how innate immune cells sense infections by monitoring mitochondrial health.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Code availability
The MetaMorph journals, which are command scripts used for the quantification of live cell imaging data, are freely available at https://cloudstor.aarnet.edu.au/plus/s/8XGL750dIgwLYDn.
References
Kaparakis-Liaskos, M. & Ferrero, R. L. Immune modulation by bacterial outer membrane vesicles. Nat. Rev. Immunol. 15, 375–387 (2015).
Schwechheimer, C. & Kuehn, M. J. Outer-membrane vesicles from Gram-negative bacteria: biogenesis and functions. Nat. Rev. Microbiol. 13, 605–619 (2015).
Toyofuku, M., Nomura, N. & Eberl, L. Types and origins of bacterial membrane vesicles. Nat. Rev. Microbiol. 17, 13–24 (2019).
Vanaja, S. K. et al. Bacterial outer membrane vesicles mediate cytosolic localization of LPS and caspase-11 activation. Cell 165, 1106–1119 (2016).
Finethy, R. et al. Inflammasome activation by bacterial outer membrane vesicles requires guanylate binding proteins. mBio 8, e01188-17 (2017).
Santos, J. C. et al. LPS targets host guanylate-binding proteins to the bacterial outer membrane for non-canonical inflammasome activation. EMBO J. 37, e98089 (2018).
Kayagaki, N. et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signalling. Nature 526, 666–671 (2015).
Ding, J. et al. Pore-forming activity and structural autoinhibition of the gasdermin family. Nature 535, 111–116 (2016).
Mulvihill, E. et al. Mechanism of membrane pore formation by human gasdermin-D. EMBO J. 37, e98321 (2018).
Kayagaki, N. et al. Non-canonical inflammasome activation targets caspase-11. Nature 479, 117–121 (2011).
Rathinam, V. A. et al. TRIF licenses caspase-11-dependent NLRP3 inflammasome activation by Gram-negative bacteria. Cell 150, 606–619 (2012).
Hagar, J. A., Powell, D. A., Aachoui, Y., Ernst, R. K. & Miao, E. A. Cytoplasmic LPS activates caspase-11: implications in TLR4-independent endotoxic shock. Science 341, 1250–1253 (2013).
He, W. T. et al. Gasdermin D is an executor of pyroptosis and required for interleukin-1β secretion. Cell Res. 25, 1285–1298 (2015).
Aachoui, Y. et al. Caspase-11 protects against bacteria that escape the vacuole. Science 339, 975–978 (2013).
Shi, J. et al. Inflammatory caspases are innate immune receptors for intracellular LPS. Nature 514, 187–192 (2014).
Shi, J. et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–665 (2015).
Bitto, N. J. et al. Membrane vesicles from Pseudomonas aeruginosa activate the noncanonical inflammasome through caspase-5 in human monocytes. Immunol. Cell Biol. 10, 1120–1130 (2018).
Deo, P. et al. Outer membrane vesicles from Neisseria gonorrhoeae target PorB to mitochondria and induce apoptosis. PLoS Pathog. 14, e1006945 (2018).
Bielaszewska, M. et al. Enterohemorrhagic Escherichia coli hemolysin employs outer membrane vesicles to target mitochondria and cause endothelial and epithelial apoptosis. PLoS Pathog. 9, e1003797 (2013).
Czabotar, P. E., Lessene, G., Strasser, A. & Adams, J. M. Control of apoptosis by the BCL-2 protein family: implications for physiology and therapy. Nat. Rev. Mol. Cell Biol. 15, 49–63 (2014).
Speir, M. et al. Eliminating Legionella by inhibiting BCL-XL to induce macrophage apoptosis. Nat. Microbiol. 1, 15034 (2016).
Zhou, X. et al. Hexa-acylated lipid A is required for host inflammatory response to Neisseria gonorrhoeae in experimental gonorrhea. Infect. Immun. 82, 184–192 (2014).
Duncan, J. A. et al. Neisseria gonorrhoeae activates the proteinase cathepsin B to mediate the signaling activities of the NLRP3 and ASC-containing inflammasome. J. Immunol. 182, 6460–6469 (2009).
Hobbs, M. M. et al. Lipid A’s structure mediates Neisseria gonorrhoeae fitness during experimental infection of mice and men. mBio 4, e00892-13 (2013).
Kahler, C. M., Nawrocki, K. L., Anandan, A., Vrielink, A. & Shafer, W. M. Structure–function relationships of the neisserial EptA enzyme responsible for phosphoethanolamine decoration of lipid A: rationale for drug targeting. Front. Microbiol. 9, 1922 (2018).
Bomberger, J. M. et al. Long-distance delivery of bacterial virulence factors by Pseudomonas aeruginosa outer membrane vesicles. PLoS Pathog. 5, e1000382 (2009).
Goncalves, A. V. et al. Gasdermin-D and caspase-7 are the key caspase-1/8 substrates downstream of the NAIP5/NLRC4 inflammasome required for restriction of Legionella pneumophila. PLoS Pathog. 15, e1007886 (2019).
Akhter, A. et al. Caspase-7 activation by the Nlrc4/Ipaf inflammasome restricts Legionella pneumophila infection. PLoS Pathog. 5, e1000361 (2009).
Rueda, F. et al. Production of functional inclusion bodies in endotoxin-free Escherichia coli. Appl. Microbiol. Biotechnol. 98, 9229–9238 (2014).
Shpilka, T. & Haynes, C. M. The mitochondrial UPR: mechanisms, physiological functions and implications in ageing. Nat. Rev. Mol. Cell Biol. 19, 109–120 (2018).
Vince, J. E. et al. The mitochondrial apoptotic effectors BAX/BAK activate caspase-3 and -7 to trigger NLRP3 inflammasome and caspase-8 driven IL-1β activation. Cell Rep. 25, 2339–2353 (2018).
Morciano, G. et al. Mcl-1 involvement in mitochondrial dynamics is associated with apoptotic cell death. Mol. Biol. Cell 27, 20–34 (2016).
Okamoto, T. et al. Enhanced stability of Mcl1, a prosurvival Bcl2 relative, blunts stress-induced apoptosis, causes male sterility, and promotes tumorigenesis. Proc. Natl Acad. Sci. USA 111, 261–266 (2014).
Bae, J., Leo, C. P., Hsu, S. Y. & Hsueh, A. J. MCL-1S, a splicing variant of the antiapoptotic BCL-2 family member MCL-1, encodes a proapoptotic protein possessing only the BH3 domain. J. Biol. Chem. 275, 25255–25261 (2000).
Chauhan, D. et al. BAX/BAK-induced apoptosis results in caspase-8-dependent IL-1β maturation in macrophages. Cell Rep. 25, 2354–2368 (2018).
Chen, K. W. et al. Extrinsic and intrinsic apoptosis activate pannexin-1 to drive NLRP3 inflammasome assembly. EMBO J. 15, e101638 (2019).
Rudel, T., Kepp, O. & Kozjak-Pavlovic, V. Interactions between bacterial pathogens and mitochondrial cell death pathways. Nat. Rev. Microbiol. 8, 693–705 (2010).
Pellegrino, M. W. et al. Mitochondrial UPR-regulated innate immunity provides resistance to pathogen infection. Nature 516, 414–417 (2014).
Dzhagalov, I., St John, A. & He, Y. W. The antiapoptotic protein Mcl-1 is essential for the survival of neutrophils but not macrophages. Blood 109, 1620–1626 (2007).
Suzuki, T. et al. Infection with flaviviruses requires BCLXL for cell survival. PLoS Pathog. 14, e1007299 (2018).
Lemaitre, B. & Girardin, S. E. Translation inhibition and metabolic stress pathways in the host response to bacterial pathogens. Nat. Rev. Microbiol. 11, 365–369 (2013).
Vikstrom, I. et al. Mcl-1 is essential for germinal center formation and B cell memory. Science 330, 1095–1099 (2010).
Glaser, S. P. et al. Anti-apoptotic Mcl-1 is essential for the development and sustained growth of acute myeloid leukemia. Genes Dev. 26, 120–125 (2012).
Martinon, F., Petrilli, V., Mayor, A., Tardivel, A. & Tschopp, J. Gout-associated uric acid crystals activate the NALP3 inflammasome. Nature 440, 237–241 (2006).
Lawlor, K. E. et al. RIPK3 promotes cell death and NLRP3 inflammasome activation in the absence of MLKL. Nat. Commun. 6, 6282 (2015).
Vince, J. E. et al. Inhibitor of apoptosis proteins limit RIP3 kinase-dependent interleukin-1 activation. Immunity 36, 215–227 (2012).
Knudson, C. M., Tung, K. S., Tourtellotte, W. G., Brown, G. A. & Korsmeyer, S. J. Bax-deficient mice with lymphoid hyperplasia and male germ cell death. Science 270, 96–99 (1995).
Ke, F. F. S. et al. Embryogenesis and adult life in the absence of intrinsic apoptosis effectors BAX, BAK, and BOK. Cell 173, 1217–1230.e17 (2018).
Speir, M. et al. Legionella pneumophila strain 130b evades macrophage cell death independent of the effector SidF in the absence of flagellin. Front. Cell. Infect. Microbiol. 7, 35 (2017).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
We thank A. Strasser, D. Huang and S. Masters (Walter and Eliza Hall Institute of Medical Research) and V. Dixit (Genentech) for providing the genetically modified mice; I. D. Hay (University of Auckland) for the ClearColi and uropathogenic E. coli bacterial strains; M. Ryan (Monash University) for the antibodies; D. Creek (Monash Proteomics and Metabolomics Facility) for liquid chromatography–mass spectrometry support; the Monash Micro Imaging Facility for expert advice on live cell imaging microscopy; and the Monash Animal Research Platform for providing mice. This research was supported by grants from the National Health and Medical Research Council (grant nos. 1145788, 1183070 and 1141466 to J.E.V.; 1162765 and 1181089 to K.E.L.; and 1183848 and 1163556 to T.N.). J.L. and J.E.V. are Australian National Health and Medical Research Council Principal Research and Career Development fellows. K.E.L. and T.N. are Australian Research Council (ARC) Future fellows (FT190100266 and FT170100313).
Author information
Authors and Affiliations
Contributions
P.D. performed and analysed the vesicle purification, live cell imaging, ELISA, immunoblot, proteomics and mouse experiments. S.H.C. performed the radiography, ELISA and live cell imaging experiments. M.-L.H. and J.L. performed the lipid mass spectrometry analysis. C.H. and R.B.S. performed the protein mass spectrometry analysis. M.S., J.E. and K.E.L. performed and analysed the ELISA and mouse experiments. S.D. performed the ELISA and live cell imaging experiments. B.T.K., J.E.V. and K.E.L. generated the genetically modified mice. J.E.V., K.E.L. and T.N. supervised the project.
Corresponding author
Ethics declarations
Competing interests
B.T.K., J.E.V. and K.E.L. are currently or have been employed at the Walter and Eliza Hall Institute, which received research funding and milestone payments from AbbVie and Genentech regarding ABT-199, which was developed based on ABT-737. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Lipid A modification of N. gonorrhoeae OMVs.
a, b, The mass spectrum (m/z) of lipid A purified from N. gonorrhoeae (a) and NOMVs (b) and identification of the major lipid species (M1, M2, M3 and M4 with their measured and predicted molecular weight to charge ratios, m/z) that contain phosphate (blue) or phosphoethanolamine (PEA, red) headgroups as indicated in (c). Representative of at least four independent experiments.
Extended Data Fig. 2 OMVs derived from N. gonorrhoeae induce inflammation in mice independent of caspase-1 and 11.
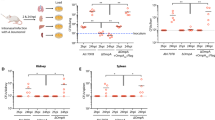
WT and caspase-1/11 deficient (Casp1/11-/-) mice were intraperitoneally (i.p) injected with N. gonorrhoeae OMVs (NOMV, 100µg) or PBS. a, Change in total body weight over time. Mean and SD, n = 2 PBS and 5 OMV-treated mice. The statistical significance was determined by two-way ANOVA followed by Sidak’s multiple comparison test. b, Images of Casp1/11-/- mice i.p injected with PBS or NOMV 24 hours post injection. c, Serum IL-1β levels 24 hours post treatment. Mean and SEM (n = 5 mice/group). The statistical significance was determined by two-tailed unpaired t test.
Extended Data Fig. 3 BAK-mediated cell death in OMV-treated macrophages.
Live-cell imaging to monitor the cell death (DRAQ7 staining) over time (a–h) of BAK (Bak-/-) and BAX (Bax-/-) deficient bone-marrow derived macrophages (BMDMs) treated with (a) N. gonorrhoeae OMVs (NOMV), (b) PBS, (d) uropathogenic E. coli derived OMVs (UOMV), (e) P. aeruginosa (POMV), (f) E.coli BW25113 (EOMVs) and (h) OMVs derived from LPS-deficient clear E. coli BL21 (COMV). c, Cell death (DRAQ7 staining) of WT and Casp8/Ripk3-/- deficient BMDMs treated with PBS or NOMV determined by live-cell imaging. g, Cell death (DRAQ7 staining) of WT BMDMs treated with PBS or E. coli OMVs (EOMV) derived from strain BW25113, BL21 and clear E. coli (COMV) using live-cell imaging. Mean and SD from three biological samples. Representative of at least two independent experiments.
Extended Data Fig. 4 Lipid A modifications in OMVs derived from UPEC and P. aeruginosa.
a, c, The mass spectrum (m/z) of lipid A purified from uropathogenic E. coli (UPEC) (a) and (c) P. aeruginosa bacterial cells and (b, d) their respective OMVs. e, f, Identification and delineation of the major lipid A species (M5, M6, M7, M11, M12 & M13 and their modifications M8, M9, M10, M14, M15, M16 and M17). Both unmodified and 4-amino-4-deoxy-L-arabinose-modified (L-Ara4N-modified) or palmitate-modified (C16:0) forms are shown. Blue colour shows unmodified lipid A, red colour shows (PEA and L-Ara4N-modified lipid A, and brown colour shows C16:0 modified lipid A. A minor peak at m/z 1445.86 indicates the lipid A at m/z 1615.99 losing a fatty acyl chain at position 3. Representative of three independent experiments.
Extended Data Fig. 5 N. gonorrhoeae OMVs induce metabolic reprogramming in macrophages.
a, b, Bone marrow-derived macrophages (BMDMs) treated with PBS or N. gonorrhoeae OMVs (NOMV) were analysed by proteomics to identify up (red) and down (blue) -regulated proteins involved in (a) cell death signalling and (b) metabolism. Data from three (1, 2 and 3) independent experiments. AA, amino acid; Glyc, glycolysis; FA fatty acid; NT, nucleotide; PTM, post-translational modification; DT, detoxification; RA, retinoic acid.
Extended Data Fig. 6 Imaging mitochondrial membrane potential, caspase-3/7 activation and cell death in OMV treated macrophages.
WT BMDMs treated with (a) N. gonorrhoeae OMVs (NOMV, 50 µg/mL) or (b) cycloheximide (CHX, 1 µg/mL) were simultaneously analysed for mitochondrial membrane potential (ΔΨm) using TMRM staining, caspase-3/7 activity (CellEvent) and cell death (DRAQ7) at the single cell level using live-cell imaging. Mean and SEM from n = 3 independent experiments. c, Bright field, TMRM, caspase-3/7 activity and DRAQ7 staining, including merged images, of WT BMDMs treated with NOMVs at indicated time points. Yellow arrow indicates a cell that transiently regains TMRM staining before caspase-3/7 activation and cell death. Scale bar 50 µm. Representative images from three independent experiments.
Extended Data Fig. 7 OMV treated macrophages transiently regain mitochondrial membrane potential.
Bak+/- control BMDMs treated with N. gonorrhoeae OMVs (NOMV, 50 µg/mL) were stained with TMRM (ΔΨm, red), caspase-3/7 substrate (green) and DRAQ7 (cell death, magenta) at indicated time points using live-cell imaging. Scale bar 50 µm. Representative images from three independent experiments.
Extended Data Fig. 8 OMV-treated macrophages show reduced protein synthesis.
a, 35S methionine labelled proteins from bone marrow-derived macrophages (BMDMs) exposed to two independent OMVs from N. gonorrhoeae (NOMV1, NOMV2), E. coli (EOMV), uropathogenic E. coli (UOMV) and LPS-deficient clear E. coli (COMV). Cycloheximide (CHX, 1 µg/mL), no 35S methionine (no35S) and Coomassie stained gel are controls. Representative of two independent experiments. b, 35S methionine labelled proteins from BMDMs treated with oligomycin (0.5 µM), antimycin A (0.5 µg/mL), FCCP (0.5 µM), NOMV (50 µg/mL), cycloheximide (CHX, 1 µg/mL) or PBS. Representative of two independent experiments. c, Dead (DRAQ7 positive) BMDMs treated with oligomycin (0.5 µM), antimycin (0.5 µg/mL) or DMSO. Data represent mean and SD from n = 3 independent experiments. d, BMDMs treated with oligomycin (0.5 µM), antimycin (0.5 µg/mL) or PBS with or without ABT-737 (1 µM) for 24 hours. Immunoblot of MCL-1, BCL-XL and cleaved caspase-3. Ponceau staining is loading control. Representative of two independent experiments. e, Immunoblot analysis of MCL-1 (L, long isoform; S, small isoform; fN, floxed N-terminus), BCL-XL and tubulin after treatment with N. gonorrhoeae OMV (NOMV, 50 µg/mL) or cycloheximide (CHX, 1 µg/mL) four hours. Representative of three independent experiments. f, WT, Bcl-xL-/- and Mcl-1-fN BMDMs exposed to NOMVs or PBS were probed for MCL-1, BCL-XL, cleaved caspase-3 at 24 hours post treatment. Tubulin is shown as loading control. Representative of two independent experiments. g, Dead (DRAQ7 stained) WT and BCL-XL deficient (Bcl-xL-/-) BMDMs treated with E. coli OMVs (EOMVs). Mean SD, n = 3 biological samples. Representative of two independent experiments.
Extended Data Fig. 9 OMV-mediated secretion of IL-1β.
a, Dead (DRAQ7 stained) wild-type (WT) bone-marrow derived macrophages (BMDMs) treated with N. gonorrhoeae OMVs (NOMV) with or without NLRP3 inhibitor (MCC950) were analysed by live-cell imaging. Mean, SD, n = 3 biological samples. Representative of two independent experiments. b, Dead WT and Nlrp3-/- BMDMs. Mean and SEM from n = 6 independent experiments. c, Secreted IL-1β levels from WT BMDMs unprimed or primed with LPS (50 ng/mL) and treated with NOMV (50 µg/mL) or nigericin (10 µM) at indicated time points. Mean, SEM, n = 3 independent experiments. d, e and f, Secreted IL-1β levels from (d) Bak+/-, Bak-/- and Bak-/-/Baxfl/fl,LysMCre (Bax/Bak-/-), (e) Casp1/11-/-, Casp-11-/- and (f) Gsdmd-/- BMDMs exposed to NOMV, E. coli (EOMV), LPS deficient clear E. coli (COMV), uropathogenic E. coli (UOMV) and P. aeruginosa (POMV) 24 hours post-treatment. Mean, SD from n = 3-4 independent experiments. g, Immunoblot analysis of full length (FL) and cleaved GSDMD (p30). Ponceau staining is loading control. Molecular weight markers (kDa) on the left. Representative of two independent experiments. h, Serum IL-1β levels after N. gonorrhoeae OMVs (100 µg) peritoneal injection 6 hours post-treatment. Data represent individual mice, mean and SEM (n = 8 mice per group). Two Bcl-xL-/- mice were excluded because of morbidity. (i) Serum IL-1β and (j) TNF-α 6 hours post E. coli LPS i.p. injection. Data represent individual mice, mean and SEM (n = 5). (k) Images of Bax/Bak-/- deficient mice i.p injected with NOMV 24 hours post treatment. Representative image of five mice.
Extended Data Fig. 10 Model of the role of mitochondrial apoptotic factors activated by OMVs.
The outer membrane vesicle (OMV) released by Gram-negative bacteria contains lipopolysaccharide (LPS) which activates cytosolic caspase-11, triggering gasdermin-D cleavage and pyroptosis in macrophages in vitro. Pore-formation by gasdermin-D causes potassium (K+) efflux which is sensed by the NLRP3/caspase-1 inflammasome to secrete mature IL-1β. OMVs also traffic pore-forming toxins to mitochondria and disrupt organellar homeostasis. Mitochondrial stress eventually prevents host protein synthesis which depletes pro-survival factor MCL-1. MCL-1 together with BCL-XL inhibit pro-death factor BAK and activation of apoptotic caspases, such as caspase-3 and -7. Apoptosis leads to potassium efflux, NLRP3/caspase-1 activation and secretion of IL-1β. Caspase-3/7 can also activate caspase-8 which can trigger IL-1β secretion in an NLRP3-dependent and independent manner. Activation of NLRP3 further contributes to cell death after OMV exposure, likely independent of caspase-1, but perhaps engaging caspase-8 or unknown factors. In vivo, IL-1β secretion and inflammation are primarily regulated by BCL-XL, BAK and BAX, suggesting N. gonorrhoeae OMVs activate additional host apoptotic factors and evade caspase-11 sensing.
Supplementary information
Supplementary Table 1
WT BMDMs treated with NOMVs or PBS for 24 h were analysed by mass spectrometry to identify up- and downregulated proteins in three independent samples.
Supplementary Video 1
Bright-field and DRAQ7 staining of WT BMDMs treated with EOMVs acquired every 30 min over 48 h. The results are representative of three independent experiments.
Supplementary Video 2
Bright-field and DRAQ7 staining of caspase-11-deficient (Casp11−/−) BMDMs treated with EOMVs acquired every 30 min over 48 h. The results are representative of three independent experiments.
Supplementary Video 3
Bright-field and DRAQ7 staining of WT BMDMs treated with NOMVs acquired every 30 min over 48 h. The results are representative of three independent experiments.
Supplementary Video 4
Bright-field and DRAQ7 staining of caspase-1- and caspase-11-deficient (Casp1/11−/−) BMDMs treated with NOMVs acquired every 30 min over 48 h. The results are representative of three independent experiments.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed autoradiograph, Coomassie gel and western blot.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Western blot and Ponceau stain of membrane.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed autoradiograph, Coomassie gel, western blot and Ponceau stain of membrane.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 9
Western blot and Ponceau stain of membrane.
Rights and permissions
About this article
Cite this article
Deo, P., Chow, S.H., Han, ML. et al. Mitochondrial dysfunction caused by outer membrane vesicles from Gram-negative bacteria activates intrinsic apoptosis and inflammation. Nat Microbiol 5, 1418–1427 (2020). https://doi.org/10.1038/s41564-020-0773-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-0773-2
This article is cited by
-
Gonococcal OMV-delivered PorB induces epithelial cell mitophagy
Nature Communications (2024)
-
Identifying pyroptosis- and inflammation-related genes in intracranial aneurysms based on bioinformatics analysis
Biological Research (2023)
-
Extracellular vesicles produced by avian pathogenic Escherichia coli (APEC) activate macrophage proinflammatory response and neutrophil extracellular trap (NET) formation through TLR4 signaling
Microbial Cell Factories (2023)
-
Emerging role of bacterial outer membrane vesicle in gastrointestinal tract
Gut Pathogens (2023)
-
A critical role of outer membrane vesicles in antibiotic resistance in carbapenem-resistant Klebsiella pneumoniae
Annals of Clinical Microbiology and Antimicrobials (2023)