Abstract
Bacterial adhesion is a general strategy for host–microbe and microbe–microbe interactions. Adhesive pili are essential for colonization, biofilm formation, virulence and pathogenesis of many environmental and pathogenic bacteria1,2. Members of the class Bacteroidia have unique type V pili, assembled by protease-mediated polymerization3. Porphyromonas gingivalis is the main contributor to periodontal disease and its type V pili are a key factor for its virulence4. However, the structure of the polymerized pilus and its assembly mechanism are unknown. Here we show structures of polymerized and monomeric states of FimA stalk pilin from P. gingivalis, determined by cryo-electron microscopy and crystallography. The atomic model of assembled FimA shows that the C-terminal strand of a donor subunit is inserted into a groove in the β-sheet of an acceptor subunit after N-terminal cleavage by the protease RgpB. The C terminus of the donor strand is essential for polymerization. We propose that type V pili assemble via a sequential polar assembly mechanism at the cell surface, involving protease-mediated strand exchange, employed by various Gram-negative species belonging to the class Bacteroidia. Our results reveal functional surfaces related to pathogenic properties of polymerized FimA. These insights may facilitate development of antibacterial drugs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The atomic coordinates of the crystal structures have been deposited into the RCSB Protein Data Bank under accession code 6JZK (FimA1) and 6JZJ (FimA2). The coordinates of the cryo-EM-based model have been deposited under accession code 6KMF. The cryo-EM map has been deposited into the Electron Microscopy Data Bank under accession code EMD-0724. Uncropped gels and western blot scans from Fig. 3d,f,h,j and Extended Data Figs. 2a,e and 5b and filament length data used to create Extended Data Fig. 2d are available with the paper. Additional raw data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Allen, W. J., Phan, G. & Waksman, G. Pilus biogenesis at the outer membrane of Gram-negative bacterial pathogens. Curr. Opin. Struct. Biol. 22, 500–506 (2012).
Hospenthal, M. K., Costa, T. R. D. & Waksman, G. A comprehensive guide to pilus biogenesis in Gram-negative bacteria. Nat. Rev. Microbiol. 15, 365–379 (2017).
Xu, Q. et al. A distinct type of pilus from the human microbiome. Cell 165, 690–703 (2016).
Lamont, R. J. & Jenkinson, H. F. Life below the gum line: pathogenic mechanisms of Porphyromonas gingivalis. Microbiol. Mol. Biol. Rev. 62, 1244–1263 (1998).
Wang, F. et al. Structure of microbial nanowires reveals stacked hemes that transport electrons over micrometers. Cell 177, 361–369 (2019).
Filman, D. J. et al. Cryo-EM reveals the structural basis of long-range electron transport in a cytochrome-based bacterial nanowire. Commun. Biol. 2, 219 (2019).
Lukaszczyk, M., Pradhan, B. & Remaut, H. The biosynthesis and structures of bacterial pili. Subcell. Biochem. 92, 369–413 (2019).
Remaut, H. et al. Donor-strand exchange in chaperone-assisted pilus assembly proceeds through a concerted β strand displacement mechanism. Mol. Cell 22, 831–842 (2006).
Holt, S. C. & Ebersole, J. L. Porphyromonas gingivalis, Treponema denticola, and Tannerella forsythia: the ‘red complex’, a prototype polybacterial pathogenic consortium in periodontitis. Periodontol. 2000 38, 72–122 (2005).
Demmer, R. T. & Desvarieux, M. Periodontal infections and cardiovascular disease: the heart of the matter. J. Am. Dent. Assoc. 137, 14S–20S (2006).
Michaud, D. S. et al. Plasma antibodies to oral bacteria and risk of pancreatic cancer in a large European prospective cohort study. Gut 62, 1764–1770 (2013).
Leech, M. T. & Bartold, P. M. The association between rheumatoid arthritis and periodontitis. Best Pr. Res. Clin. Rheumatol. 29, 189–201 (2015).
Dominy, S. S. et al. Porphyromonas gingivalis in Alzheimer’s disease brains: evidence for disease causation and treatment with small-molecule inhibitors. Sci. Adv. 5, eaau3333 (2019).
Yoshimura, F., Takahashi, K., Nodasaka, Y. & Suzuki, T. Purification and characterization of a novel type of fimbriae from the oral anaerobe Bacteroides gingivalis. J. Bacteriol. 160, 949–957 (1984).
Hamada, N., Sojar, H. T., Cho, M. I. & Genco, R. J. Isolation and characterization of a minor fimbria from Porphyromonas gingivalis. Infect. Immun. 64, 4788–4794 (1996).
Hajishengallis, G., Shakhatreh, M.-A. K., Wang, M. & Liang, S. Complement receptor 3 blockade promotes IL-12-mediated clearance of Porphyromonas gingivalis and negates its virulence in vivo. J. Immunol. 179, 2359–2367 (2007).
Amano, A. Bacterial adhesins to host components in periodontitis. Periodontol. 2000 52, 12–37 (2010).
Park, Y. et al. Short fimbriae of Porphyromonas gingivalis and their role in coadhesion with Streptococcus gordonii. Infect. Immun. 73, 3983–3989 (2005).
Sojar, H. T., Hamada, N. & Genco, R. J. Isolation and characterization of fimbriae from a sparsely fimbriated strain of Porphyromonas gingivalis. Appl. Environ. Microbiol. 63, 2318–2323 (1997).
Hasegawa, Y. et al. Anchoring and length regulation of Porphyromonas gingivalis Mfa1 fimbriae by the downstream gene product Mfa2. Microbiology 155, 3333–3347 (2009).
Nagano, K., Hasegawa, Y., Murakami, Y., Nishiyama, S. & Yoshimura, F. FimB regulates FimA fimbriation in Porphyromonas gingivalis. J. Dent. Res. 89, 903–908 (2010).
Hamada, S. et al. Molecular and immunological characterization of the fimbriae of Porphyromonas gingivalis. Microbiol. Immunol. 38, 921–930 (1994).
Fujiwara, T., Nakagawa, I., Morishima, S., Takahashi, I. & Hamada, S. Inconsistency between the fimbrilin gene and the antigenicity of lipopolysaccharides in selected strains of Porphyromonas gingivalis. FEMS Microbiol. Lett. 124, 333–341 (1994).
Nakagawa, I. et al. Distribution and molecular characterization of Porphyromonas gingivalis carrying a new type of fimA gene. J. Clin. Microbiol. 38, 1909–1914 (2000).
Nakagawa, I. et al. Identification of a new variant of fimA gene of Porphyromonas gingivalis and its distribution in adults and disabled populations with periodontitis. J. Periodontal Res . 37, 425–432 (2002).
Okuda, S. & Tokuda, H. Lipoprotein sorting in bacteria. Annu. Rev. Microbiol. 65, 239–259 (2011).
Korotkov, K. V. et al. Calcium is essential for the major pseudopilin in the type 2 secretion system. J. Biol. Chem. 284, 25466–25470 (2009).
Shoji, M. et al. The major structural components of two cell surface filaments of Porphyromonas gingivalis are matured through lipoprotein precursors. Mol. Microbiol. 52, 1513–1525 (2004).
Konovalova, A. & Silhavy, T. J. Outer membrane lipoprotein biogenesis: Lol is not the end. Phil. Trans. R. Soc. B 370, 20150030 (2015).
Nakayama, K., Yoshimura, F., Kadowaki, T. & Yamamoto, K. Involvement of arginine-specific cysteine proteinase (Arg-gingipain) in fimbriation of Porphyromonas gingivalis. J. Bacteriol. 178, 2818–2824 (1996).
Shoji, M. et al. Recombinant Porphyromonas gingivalis FimA preproprotein expressed in Escherichia coli is lipidated and the mature or processed recombinant FimA protein forms a short filament in vitro. Can. J. Microbiol. 56, 959–967 (2010).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Lee, J. Y. et al. Maturation of the Mfa1 fimbriae in the oral pathogen Porphyromonas gingivalis. Front. Cell Infect. Microbiol. 8, 137 (2018).
Nishiyama, M., Ishikawa, T., Rechsteiner, H. & Glockshuber, R. Reconstitution of pilus assembly reveals a bacterial outer membrane catalyst. Science 320, 376–379 (2008).
Nishiyama, S. et al. Involvement of minor components associated with the FimA fimbriae of Porphyromonas gingivalis in adhesive functions. Microbiology 153, 1916–1925 (2007).
Hajishengallis, G., Ratti, P. & Harokopakis, E. Peptide mapping of bacterial fimbrial epitopes interacting with pattern recognition receptors. J. Biol. Chem. 280, 38902–38913 (2005).
Nakano, K. et al. Comparison of inflammatory changes caused by Porphyromonas gingivalis with distinct fimA genotypes in a mouse abscess model. Oral Microbiol. Immunol. 19, 205–209 (2004).
Sugano, N. et al. Differential cytokine induction by two types of Porphyromonas gingivalis. Oral Microbiol. Immunol. 19, 121–123 (2004).
Bodet, C., Chandad, F. & Grenier, D. Porphyromonas gingivalis-induced inflammatory mediator profile in an ex vivo human whole blood model. Clin. Exp. Immunol. 143, 50–57 (2006).
Nagano, K. et al. Porphyromonas gingivalis FimA fimbriae: fimbrial assembly by fimA alone in the fim gene cluster and differential antigenicity among fimA genotypes. PLoS ONE 7, e43722 (2012).
Alaei, S. R., Park, J. H., Walker, S. G. & Thanassi, D. G. Peptide-based inhibitors of fimbrial biogenesis in Porphyromonas gingivalis. Infect. Immun. 87, e00750–18 (2019).
Veillard, F. et al. Purification and characterisation of recombinant His-tagged RgpB gingipain from Porphymonas gingivalis. Biol. Chem. 396, 377–384 (2015).
Battye, T. G. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. W. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D 67, 271–281 (2011).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, T., Nakayama, K., Belliveau, L. & Duncan, M. J. Porphyromonas gingivalis gingipains and adhesion to epithelial cells. Infect. Immun. 69, 3048–3056 (2001).
Kikuchi, Y. et al. Novel stationary-phase-upregulated protein of Porphyromonas gingivalis influences production of superoxide dismutase, thiol peroxidase and thioredoxin. Microbiology 151, 841–853 (2005).
Shoji, M. et al. Characterization of hemin-binding protein 35 (HBP35) in Porphyromonas gingivalis: its cellular distribution, thioredoxin activity and role in heme utilization. BMC Microbiol. 10, 152 (2010).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Scheres, S. H. W. Processing of structurally heterogeneous Cryo-EM data in RELION. Methods Enzymol. 579, 125–157 (2016).
Grant, T., Rohou, A. & Grigorieff, N. cisTEM, user-friendly software for single-particle image processing. eLife 7, e35383 (2018).
Frank, J. et al. SPIDER and WEB: Processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Pettersen, E. F. et al. UCSF chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Kyte, J. & Doolittle, R. F. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 157, 105–132 (1982).
Acknowledgements
We thank J. Potempa for providing the His-tagged RgpB-expressing P. gingivalis strain, S. Aizawa for critically reading the manuscript and S. D. Aird for technical editing. X-ray diffraction data were collected at synchrotron beamlines BL32XU and BL41XU in SPring-8 (Harima, Japan) with the approval of the Japan Synchrotron Radiation Research Institute (JASRI) (proposal 2014B1478, 2016A2539, 2017A2588 and 2018A2569). This work was supported by the Platform Project for Supporting Drug Discovery and Life Science Research (BINDS) from AMED under grant no. JP18am0101076 (to M.W.), by JSPS KAKENHI grant nos. JP17K17085 and JP19K10083 (to S.S.), by JSPS KAKENHI grant no. JP16H05504 (to K.N., K.I. and M.S.), and by JSPS KAKENHI grant no. JP17K07318 (to H.M.). M.W. was supported by direct funding from Okinawa Institute of Science and Technology Graduate University.
Author information
Authors and Affiliations
Contributions
S.S., M.S., K.I., K.N. and M.W. conceived and designed experiments. M.S., K.O. and S.S. performed molecular cloning and protein purification. M.S. created mutants. S.S., H.M., M.M.M. and M.W. carried out cryo-EM experiments and image processing. H.M. and M.M.M. built atomic models into cryo-EM maps. K.O. and K.I. performed crystallization experiments and analysis. S.S. wrote the first draft. All authors analysed results and contributed to writing the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Comparison of FimA subtypes and mapping of functional regions and immune epitopes onto the polymerized FimA1.
a, Sequence alignment of five types of FimA: FimA1 from strain ATCC 33277, FimA2 from strain TDC60, FimA3 from strain 6/26, FimA4 from strain W83, and FimA5 from strain HNA99. Non-conserved regions (as assigned by ClustalW (Thompson et al, 1994)) are coloured red. Sequences containing functional regions are marked with coloured lines, highlighting regions binding to salivary protein (312-332, cyan), PRP1 (372-383, yellow), Statherin (339-352 and 353-363, orange), and a shared PRP1 and Statherin-binding site (364-372, red). Sequences containing immune epitopes are marked with coloured lines, highlighting regions binding to CD14 (115-136, dark blue), CD11b/CD18 binding epitope (212-231, dark green), and the latter part of the CD11b/CD18 epitope (252-271, bright green). Partial sequences surrounded by dashed line boxes correspond to labeled loop regions in (b-c). b, Structure alignment of the crystal structures of FimA subtypes FimA1 (red) and FimA2 (blue), root mean square deviation (RMSD) 0.75 Å; c, FimA1 (red) and FimA4 PDB 4q98 (yellow), RMSD 0.93 Å; d, FimA2 (blue) and FimA4 (yellow), RMSD 1.00 Å. Loop regions that show major structural divergence are marked with dashed circles. e, Distribution of non-conserved variable amino acid residues (pink) mapped onto the molecular surface of the FimA1 filament after identification by multiple sequence alignment of FimA1-5 subtypes (a). f, Regions mapped onto the molecular surface of the FimA1 filament are highlighted in colour: Functional binding regions of salivary protein (312-332, cyan), PRP1 (372-383, yellow), Statherin (339-352 and 353-363, orange), and a shared PRP1 and Statherin-binding region (364-372, red); Immune epitopes binding to CD14 (115-136, dark blue), CD11b/CD18 (212-231, dark green), and the latter part of CD11b/CD18 (252-271, bright green). Right: ribbon representation.
Extended Data Fig. 2 In vitro assembly rate of rFimA filament.
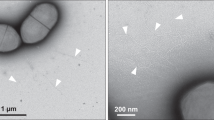
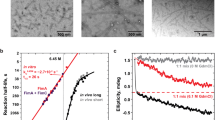
a, SDS-PAGE result shows that 30 min after addition of RgpB almost all rFimA monomers have been cleaved. Samples were heated at 100 °C for 10 min. Lanes: (i) rFimA without RgpB, (ii) after 10 min incubation with RgpB, (iii) after 30 min incubation with RgpB, (iv) after 60 min incubation with RgpB. Single asterisk indicates full length rFimA, double asterisk indicates cleaved rFimA. b, Negatively stained TEM image of sample from lane iv (after 60 min incubation with RgpB). Although the sample contains an excess of cleaved rFimA monomer, the efficiency of in vitro polymerization is low. Scale bar, 100 nm. c, TEM image of negatively stained purified filaments from (b) using by PEG precipitation. Scale bar, 100 nm. d, Relative frequency histogram of filament length distribution measured from images of purified filaments after 30 min incubation with RgpB (white columns, mean = 249 ± 123 nm) and after 60 min incubation (black columns, mean = 383 ± 262 nm) at 37 °C. 100 filaments were picked from randomly imaged areas. Experiments were carried out in triplicate. Histogram and error bars (standard deviation) include data from all 3 experiments (total of n = 300 observations). Filaments shorter than 100 nm could not be separated by precipitation. The longest filament measured 740 nm after 30 min, and 1249 nm after 60 min, corresponding to a maximum observed growth rate of ~17 nm/min or ~2.5 subunits/min (helical rise of the rFimA filament is 66.7 Å). Under these conditions, polymerization rate is rate-limiting, not the cleavage rate. e, Western blot of polymerizing rFimA after RgpB cleavage. Samples were heated at 80 °C for 10 min. Primary antibodies against FimA. Lanes: (i) Absence of RgpB, (ii) 1 min after, (iii) 3 min, (iv) 5 min, (v) 10 min, (vi) 30 min, (vii) 60 min. Ladder patterns indicate FimA polymerization. Asterisk: FimA dimer. Experiments for (a-c, e) were carried out in triplicate.
Extended Data Fig. 3 Resolution estimation of the cryo-EM map.
a, The best six 2D class-average images (from a total of 50 class averages calculated from 831,459 aligned image segments selected from 1,153 digital micrographs). b, Map resolution was estimated from two independently refined half-set maps by Fourier shell correlation (FSC). The average map resolution was 3.6 Å (FSC=0.143). c, Angular distribution plot of particle orientations (Euler angles). The coloured bar indicates the number of particles per orientation. Note that blue colour indicates non-zero values. Hence, all views are represented by at least one particle image.
Extended Data Fig. 4 Comparison of cryo-EM reconstructions of rFimA filament and Fim pilus purified from the mfa1 mutant.
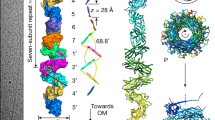
Iso-electron potential surface representations of cryo-EM reconstructions. a, rFimA (3.6 Å resolution, contoured at 5.5σ above average). The rFimA filament formed a right-handed helix with a helical twist of 71.0° (5.1 subunits per turn) and rise 66.7 Å (pitch 339 Å per turn). b, Purified Fim pilus from the mfa1 mutant (7.2 Å resolution, contoured at 4.3σ). The map was sharpened with a B-factor of -150 Å-1. c, Same as (b), including 3 fitted copies of the rFimA atomic model. The Fim pilus formed a right-handed helical filament with a helical twist of 70.0° (5.1 subunits per turn) and a rise of 67.6 Å (pitch 345 Å per turn). Scale bar, 20Å.
Extended Data Fig. 5 New interactions after strand exchange stabilize the filament.
a, Amino acid pairs within hydrogen-bonding distance in the β-sheets of the self-complementing form (monomer) and the donor-strand exchanged form (assembled). Labels indicate participating amino acid residues. Interactions involving only main chain atoms are shown as dashed lines. Interactions involving at least one side chain atom are shown as solid lines. b, Western blot of pilus-defective (fimA mfa1 mfa2) strains expressing (i), FimA(Q379A); (ii), FimA(Δ383); (iii), FimA+. Samples were heated at 80 °C for 10 min. Ladder patterns indicate FimA polymerization. Substitution of Gln379 by Ala does not affect FimA polymerization. Experiments were carried out in duplicate. c, Electron micrograph of negatively stained rFimA(Q379A) filaments after RgpB treatment. d, Electron micrograph of negatively stained rFimA(Δ383) monomers after RgpB treatment. Scale bars (c, d): 100 nm. Each image in (c, d) is representative for observations from at least 10 grid areas, repeated twice. e, The donor-acceptor pilin interface. f, Amino acid pairs within hydrogen-bonding distance at the interface (Tyr186 side chain with backbone carbonyl of Gln70, and Gln70 side chain with backbone carbonyl of Ala185).
Extended Data Fig. 6 The N-terminal strand remains in the subunit at the base of the rFimA filament after cleavage.
a, Two electron micrographs of negatively stained rFimA filaments labeled with secondary anti-mouse IgG antibody-conjugated gold (negative control). b, Four sample electron micrographs of rFimA filaments prepared and imaged as (a), but using primary anti-His tag antibody. Scale bar, 100 nm. Images of samples at each condition (a, b) are representative for observations from at least 10 grid areas, repeated 3 times.
Extended Data Fig. 7
X-ray data collection and refinement statistics; Cryo-EM data collection, refinement and validation statistics.
Supplementary information
Source data
Source Data Fig. 3
Unprocessed western blot for Fig. 3d, unprocessed Coomassie brilliant blue (CBB)-stained gel and western blot for Fig. 3f, unprocessed immunodot blot for Fig. 3h, and unprocessed CBB-stained gel and western blot for Fig. 3h.
Source Data Extended Data Fig. 2
Unprocessed CBB stained gel for Extended Data Fig. 2a, and unprocessed CBB-stained gel and western blot for Extended Data Fig. 2e.
Source Data Extended Data Fig. 2d
Filament length data for Extended Data Fig. 2d.
Source Data Extended Data Fig. 5
Unprocessed western blot for Extended Data Fig. 5b.
Rights and permissions
About this article
Cite this article
Shibata, S., Shoji, M., Okada, K. et al. Structure of polymerized type V pilin reveals assembly mechanism involving protease-mediated strand exchange. Nat Microbiol 5, 830–837 (2020). https://doi.org/10.1038/s41564-020-0705-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-0705-1
This article is cited by
-
Serine peptidase Vpr forms enzymatically active fibrils outside Bacillus bacteria revealed by cryo-EM
Nature Communications (2023)
-
Structure based High-Throughput Virtual Screening, Molecular Docking and Molecular Dynamics Study of anticancer natural compounds against fimbriae (FimA) protein of Porphyromonas gingivalis in oral squamous cell carcinoma
Molecular Diversity (2023)
-
Electron cryo-microscopy reveals the structure of the archaeal thread filament
Nature Communications (2022)
-
Donor-strand exchange drives assembly of the TasA scaffold in Bacillus subtilis biofilms
Nature Communications (2022)
-
Heads or tails for type V pilus assembly
Nature Microbiology (2020)