Abstract
Although much research has been done on the diversity of the gut microbiome, little is known about how it influences intestinal homeostasis under normal and pathogenic conditions. Epigenetic mechanisms have recently been suggested to operate at the interface between the microbiota and the intestinal epithelium. We performed whole-genome bisulfite sequencing on conventionally raised and germ-free mice, and discovered that exposure to commensal microbiota induced localized DNA methylation changes at regulatory elements, which are TET2/3-dependent. This culminated in the activation of a set of ‘early sentinel’ response genes to maintain intestinal homeostasis. Furthermore, we demonstrated that exposure to the microbiota in dextran sodium sulfate-induced acute inflammation results in profound DNA methylation and chromatin accessibility changes at regulatory elements, leading to alterations in gene expression programs enriched in colitis- and colon-cancer-associated functions. Finally, by employing genetic interventions, we show that microbiota-induced epigenetic programming is necessary for proper intestinal homeostasis in vivo.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data to support the findings of this study are available from the corresponding authors upon reasonable request. All sequencing data are available from the GEO database under accession number GSE137037. Source data for Figs. 2a–e, 4a,b,g and 5a–g and Extended Data Figs. 2e, 4b–g,i and 5a,b are included in this article.
References
Yin, Y. et al. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science 356, eaaj2239 (2017).
Alenghat, T. et al. Histone deacetylase 3 coordinates commensal-bacteria-dependent intestinal homeostasis. Nature 504, 153–157 (2013).
Yu, D. H. et al. Postnatal epigenetic regulation of intestinal stem cells requires DNA methylation and is guided by the microbiome. Genome Biol. 16, 211 (2015).
Fellows, R. et al. Microbiota derived short chain fatty acids promote histone crotonylation in the colon through histone deacetylases. Nat. Commun. 9, 105 (2018).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Khor, B., Gardet, A. & Xavier, R. J. Genetics and pathogenesis of inflammatory bowel disease. Nature 474, 307–317 (2011).
Elliott, E. N., Sheaffer, K. L., Schug, J., Stappenbeck, T. S. & Kaestner, K. H. Dnmt1 is essential to maintain progenitors in the perinatal intestinal epithelium. Development 142, 2163–2172 (2015).
Kim, R., Sheaffer, K. L., Choi, I., Won, K. J. & Kaestner, K. H. Epigenetic regulation of intestinal stem cells by Tet1-mediated DNA hydroxymethylation. Genes Dev. 30, 2433–2442 (2016).
Jenke, A. C. & Zilbauer, M. Epigenetics in inflammatory bowel disease. Curr. Opin. Gastroenterol. 28, 577–584 (2012).
Thaiss, C. A. et al. Microbiota diurnal rhythmicity programs host transcriptome oscillations. Cell 167, 1495–1510 (2016).
Gury-BenAri, M. et al. The spectrum and regulatory landscape of intestinal innate lymphoid cells are shaped by the microbiome. Cell 166, 1231–1246 e13 (2016).
Gordon, H. A. & Pesti, L. The gnotobiotic animal as a tool in the study of host microbial relationships. Bacteriol. Rev. 35, 390–429 (1971).
Park, J. H. et al. Promotion of intestinal epithelial cell turnover by commensal bacteria: role of short-chain fatty acids. PLoS ONE 11, e0156334 (2016).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011).
Orlanski, S. et al. Tissue-specific DNA demethylation is required for proper B-cell differentiation and function. Proc. Natl Acad. Sci. USA 113, 5018–5023 (2016).
Ye, D. Z. & Kaestner, K. H. Foxa1 and Foxa2 control the differentiation of goblet and enteroendocrine L- and D-cells in mice. Gastroenterology 137, 2052–2062 (2009).
Gosalia, N., Yang, R., Kerschner, J. L. & Harris, A. FOXA2 regulates a network of genes involved in critical functions of human intestinal epithelial cells. Physiol. Genomics 47, 290–297 (2015).
Yu, T. et al. Kruppel-like factor 4 regulates intestinal epithelial cell morphology and polarity. PLoS ONE 7, e32492 (2012).
Schonthaler, H. B., Guinea-Viniegra, J. & Wagner, E. F. Targeting inflammation by modulating the Jun/AP-1 pathway. Ann. Rheum. Dis. 70(Suppl 1), i109–i112 (2011).
Alteber, Z. et al. The anti-inflammatory IFITM genes ameliorate colitis and partially protect from tumorigenesis by changing immunity and microbiota. Immunol. Cell. Biol. 96, 284–297 (2018).
Okita, Y. et al. Interleukin-22-induced antimicrobial phospholipase A2 group IIA mediates protective innate immunity of nonhematopoietic cells against listeria monocytogenes. Infect. Immun. 84, 573–579 (2016).
Muhl, H., Bachmann, M. & Pfeilschifter, J. Inducible NO synthase and antibacterial host defence in times of Th17/Th22/T22 immunity. Cell. Microbiol. 13, 340–348 (2011).
Johansson, M. E. et al. Bacteria penetrate the inner mucus layer before inflammation in the dextran sulfate colitis model. PLoS ONE 5, e12238 (2010).
Berman, B. P. et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina-associated domains. Nat. Genet. 44, 40–46 (2012).
Abu-Remaileh, M. et al. Chronic inflammation induces a novel epigenetic program that is conserved in intestinal adenomas and in colorectal cancer. Cancer Res. 75, 2120–2130 (2015).
Elinav, E. et al. Inflammation-induced cancer: crosstalk between tumours, immune cells and microorganisms. Nat. Rev. Cancer 13, 759–771 (2013).
Sheaffer, K. L. et al. DNA methylation is required for the control of stem cell differentiation in the small intestine. Genes Dev. 28, 652–664 (2014).
Lawrence, T. The nuclear factor NF-κB pathway in inflammation. Cold Spring Harb. Perspect. Biol. 1, a001651 (2009).
Kirillov, A. et al. A role for nuclear NF-κB in B-cell-specific demethylation of the Igkappa locus. Nat. Genet. 13, 435–441 (1996).
Kagan, J. C. & Medzhitov, R. Phosphoinositide-mediated adaptor recruitment controls Toll-like receptor signaling. Cell 125, 943–955 (2006).
Gu, T. P. et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature 477, 606–610 (2011).
Wossidlo, M. et al. 5-Hydroxymethylcytosine in the mammalian zygote is linked with epigenetic reprogramming. Nat. Commun. 2, 241 (2011).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Koh, K. P. et al. Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 8, 200–213 (2011).
Vincent, J. J. et al. Stage-specific roles for tet1 and tet2 in DNA demethylation in primordial germ cells. Cell Stem Cell 12, 470–478 (2013).
Dawlaty, M. M. et al. Loss of Tet enzymes compromises proper differentiation of embryonic stem cells. Dev. Cell 29, 102–111 (2014).
Ko, M. et al. Ten-Eleven-Translocation 2 (TET2) negatively regulates homeostasis and differentiation of hematopoietic stem cells in mice. Proc. Natl Acad. Sci. USA 108, 14566–14571 (2011).
Ko, M. et al. TET proteins and 5-methylcytosine oxidation in hematological cancers. Immunol. Rev. 263, 6–21 (2015).
Madison, B. B. et al. Cis elements of the villin gene control expression in restricted domains of the vertical (crypt) and horizontal (duodenum, cecum) axes of the intestine. J. Biol. Chem. 277, 33275–33283 (2002).
el Marjou, F. et al. Tissue-specific and inducible Cre-mediated recombination in the gut epithelium. Genesis 39, 186–193 (2004).
Elliott, E. N., Sheaffer, K. L. & Kaestner, K. H. The ‘de novo’ DNA methyltransferase Dnmt3b compensates the Dnmt1-deficient intestinal epithelium. eLife 5, e12975 (2016).
Pan, W. H. et al. Exposure to the gut microbiota drives distinct methylome and transcriptome changes in intestinal epithelial cells during postnatal development. Genome Med. 10, 27 (2018).
Cheng, J., Palva, A. M., de Vos, W. M. & Satokari, R. Contribution of the intestinal microbiota to human health: from birth to 100 years of age. Curr. Top. Microbiol. Immunol. 358, 323–346 (2013).
Davison, J. M. et al. Microbiota regulate intestinal epithelial gene expression by suppressing the transcription factor Hepatocyte nuclear factor 4 alpha. Genome Res. 27, 1195–1206 (2017).
Camp, J. G. et al. Microbiota modulate transcription in the intestinal epithelium without remodeling the accessible chromatin landscape. Genome Res. 24, 1504–1516 (2014).
Golson, M. L. & Kaestner, K. H. Fox transcription factors: from development to disease. Development 143, 4558–4570 (2016).
Ghaleb, A. M., McConnell, B. B., Kaestner, K. H. & Yang, V. W. Altered intestinal epithelial homeostasis in mice with intestine-specific deletion of the Kruppel-like factor 4 gene. Dev. Biol. 349, 310–320 (2011).
Liu, Y., Chidgey, M., Yang, V. W. & Bialkowska, A. B. Kruppel-like factor 5 is essential for maintenance of barrier function in mouse colon. Am. J. Physiol. Gastrointest. Liver Physiol. 313, G478–G491 (2017).
Sardina, J. L. et al. Transcription factors drive Tet2-mediated enhancer demethylation to reprogram cell fate. Cell Stem Cell 23, 727–741 (2018).
Macpherson, A. J. & Harris, N. L. Interactions between commensal intestinal bacteria and the immune system. Nat. Rev. Immunol. 4, 478–485 (2004).
Gensollen, T., Iyer, S. S., Kasper, D. L. & Blumberg, R. S. How colonization by microbiota in early life shapes the immune system. Science 352, 539–544 (2016).
Bergman, Y. & Cedar, H. DNA methylation dynamics in health and disease. Nat. Struct. Mol. Biol. 20, 274–281 (2013).
Ben-Neriah, Y. & Karin, M. Inflammation meets cancer, with NF-κB as the matchmaker. Nat. Immunol. 12, 715–723 (2011).
Hecht, G. et al. A simple cage-autonomous method for the maintenance of the barrier status of germ-free mice during experimentation. Lab Anim. 48, 292–297 (2014).
Thaiss, C. A. et al. Persistent microbiome alterations modulate the rate of post-dieting weight regain. Nature 540, 544–551 (2016).
Sato, T. & Clevers, H. Primary mouse small intestinal epithelial cell cultures. Methods Mol. Biol. 945, 319–328 (2013).
Haber, A. L. et al. A single-cell survey of the small intestinal epithelium. Nature 551, 333–339 (2017).
Erben, U. et al. A guide to histomorphological evaluation of intestinal inflammation in mouse models. Int. J. Clin. Exp. Pathol. 7, 4557–4576 (2014).
Raddatz, G., Gao, Q., Bender, S., Jaenisch, R. & Lyko, F. Dnmt3a protects active chromosome domains against cancer-associated hypomethylation. PLoS Genet. 8, e1003146 (2012).
Xi, Y. & Li, W. BSMAP: whole genome bisulfite sequence MAPping program. BMC Bioinformatics 10, 232 (2009).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank all members of our groups for helpful discussions. This work was supported by research grants from the Israel Academy of Sciences (grant 734/13 Y.B.), the Israel Cancer Research Foundation (grant 211410 to Y.B.), The Emanuel Rubin Chair in Medical Sciences (Y.B.), the Israel Center of Excellence Program (grant 1796/12 to Y.B.), the Helmholtz-Israel-Cooperation in Personalized Medicine (to Y.B. and F.L.), the Helmholtz program ‘Aging and Metabolic Programming’ (AMPro, to F.L.) and the German-Israeli Foundation (grant 1424 to Y.B. and F.L.).
Author information
Authors and Affiliations
Contributions
I.A. conceived and carried out most of the experiments, and analysed and interpreted the results. M.R. prepared the samples, targeted bisulfite, qPCR and ATAC-seq analyses. D.C. performed targeted bisulfite analyses. M.A.-R. initiated the acute inflammation experiments. T.T performed and analysed the FACS experiments. H.S. conducted experiments with germ free mice. G.R., J.G. and I.A. analysed and interpreted the genome-wide data. E.P. evaluated all histological samples. E.E., E.P., F.L. and Y.B. designed and supervised this study. I.A, F.L. and Y.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Microbiota induces transcriptional alterations.
a, Pie chart showing the number of significantly differentially expressed genes (1182) with a fold-change of ≥2, relative to germ free (GF). Conventional (CNV) upregulated genes are shown in red and downregulated genes are shown in green. b, Ingenuity pathway analysis of the 358 downregulated genes from (a). The highly enriched functions from the most highly enriched categories are shown. c, Gene Ontology (GO) analysis of the CNV 824 upregulated genes from (a). The highly enriched biological processes from the enriched categories are shown. P values (b,c) were calculated using two-tailed Fisher’s exact test. d, Expression levels of proliferation genes (Mki67 and Top2a) in GF (n = 3) and CNV (n = 3), data were extracted from RNA-seq analysis. e, Mki67 staining on distal colon specimens from GF (n = 3) and CNV (n = 3) mice. Scale bar 100 µm. Quantification of Mki67-positive cells is also shown. Significance (d,e) was determined using two-sided t-test and is expressed as the mean ± SEM.
Extended Data Fig. 2 Microbiota induces DNA methylation changes.
a, Average DNA methylation ratios of various intragenic sub-segments are shown for germ free (GF) (n = 2, yellow and green) and conventional (CNV) (n = 2, blue and red) mice. b, Average methylation profiles of all promoters in all 4 mice that were analyzed by whole-genome bisulfite sequencing. c, Average methylation profiles of all canyons in all 4 mice that were analyzed by whole-genome bisulfite sequencing. d, Number of unmethylated regions (UMRs) and low methylated regions (LMRs) in GF and CNV samples. e, Bisulfite sequencing results for LMRs defined by comparing CNV versus GF mice. The heatmap shows average methylation ratios of 4 LMR amplicons from GF (sorted intestinal epithelial cells (IECs) n = 4 and crypts n = 3) and CNV (sorted IECs n = 3 and crypts n = 4). P values were calculated using two-sided t-test for GF versus CNV IECs and for GF versus CNV crypts. The precise P values can be found in Source Data.
Extended Data Fig. 3 Acute inflammation in conventional (CNV) mice provokes DNA methylation changes.
a, A schematic diagram of the protocol used to induce acute inflammation. Briefly, acute inflammation was induced by administration of 2% DSS in the drinking water for 5 days followed by regular drinking water for an additional 16 days. b, Average global DNA methylation ratios are shown for CNV (n = 2, blue) and DSS-treated CNV (CNV/DSS) (n = 2, orange) mice, respectively. c, Average DNA methylation ratios of various intragenic sub-segments are shown for CNV (n = 2, yellow and green) and CNV/DSS (n = 2, blue and red) mice. d, Methylation and lamina-associated domain (LAD) tracks of mouse chromosome 4 (blue CNV, red CNV/DSS). e, Average methylation profiles of all canyons in all 4 mice that were analyzed by whole-genome bisulfite sequencing. f, Bar graph showing the number of up- and down- regulated genes in CNV/DSS versus CNV samples associated with hyper- and hypo- methylated promoters. g, Gene Ontology (GO) analysis of the upregulated genes (n = 185) associated with hypomethylated promoters in CNV/DSS compared to CNV. The highly enriched processes are shown (P values were calculated using two-tailed Fisher’s exact test). h, Diseases-related with the upregulated genes (n = 185) associated with hypomethylated promoter in CNV/DSS compared to CNV samples (P values were calculated using two-tailed Fisher’s exact test, adjusted p-value calculated using the Benjamini-Hochberg method for correction for multiple hypotheses testing).
Extended Data Fig. 4 Validation analyses of LMRs induced by acute inflammation.
a, Comparison of average LMR methylation levels in conventional (CNV) (n = 2) and DSS-treated CNV (CNV/DSS) (n = 2) mice. The upper (lower) line indicates the positions in the plot where CNV/DSS is exactly 0.1 hypermethylated (hypomethylated) compared to CNV. There are 20061 (14585) LMRs which are more than 0.1 hypermethylated (hypomethylated) in CNV/DSS versus CNV. b, Bisulfite sequencing results for LMRs defined in CNV versus CNV/DSS mice. The heatmap shows average methylation ratios of 5 LMR amplicons from CNV sorted intestinal epithelial cells (IECs) (n = 5) and CNV/DSS IECs (n = 5) mice. c, Changes in body weight of CNV (n = 5) and CNV/DSS (n = 7) mice and (d) disease activity index (DAI) were monitored daily. e,f, on day 21, mice weight and colon length (respectively) were measured. g, Histological score shows the combined score of inflammatory cell infiltration and tissue damage. h, Hematoxylin and eosin (H&E)-stained histologic images of the colon from CNV and CNV/DSS mice. Scale bar 100 µm. i, Bisulfite sequencing results for indicated LMRs defined in CNV versus CNV/DSS mice. The heatmap shows average methylation ratios of 5 LMR amplicons from CNV crypts (n = 5) and CNV/DSS crypts (n = 5) isolated from mice raised in a different animal facility than in (b). j, Average methylation profiles of hypomethylated LMRs-containing NF-κB and AP-1 binding sites, respectively. Significance (b-g and i) was determined using two-sided t-test and is expressed as the mean ± SEM. The exact P values (b-d and i) can be found in Source Data. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Extended Data Fig. 5 Validation analyses of LMR methylation changes in sterile inflammation.
a, Bisulfite sequencing results for indicated LMRs defined in germ free (GF) versus DSS-treated GF (GF/DSS) mice. The heatmap shows average methylation ratios of LMRs from GF crypt IECs (n = 5) and CNV/DSS crypt IECs (n = 5). b, Bisulfite sequencing results for indicated LMRs defined in GF versus GF/DSS mice. The heatmap shows average methylation ratios of LMRs from GF FACS-sorted intestinal epithelial cells (IECs) (n = 5) and CNV/DSS FACS-sorted IECs (n = 5). P values (a,b) were calculated using two-sided t-test. The exact P values can be found in Source Data. c, Pie chart indicating the number of up- and down- regulated genes that are associated with hypermethylated- and hypomethylated- LMRs in GF/DSS compared to GF mice.
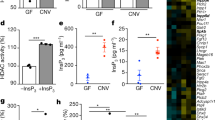
Extended Data Fig. 6 TET2/3 play a key role in microbiota-induced DNA demethylation.
a, Expression levels of TET genes in conventional (CNV) (n = 3) and DSS-treated CNV (CNV/DSS) (n = 3) mice, data extracted from RNA-seq analysis. b, Normalized expression levels of TET2 and TET3 genes from colonic crypts isolated from germ free (GF) (n = 5), CNV (n = 5) and antibiotics (Abx)-treated (n = 5) mice. c, Normalized expression levels of TET2 and TET3 genes from WT (n = 4) and LPS-treated (n = 4) organoids. Significance (a-c) was determined using two-sided t-test and is expressed as the mean ± SEM. d, Average global DNA methylation ratios are shown for Tet2/3 fl/fl (WT, n = 3) and 3 Tet2/3 fl/fl VillinCre (KO, n = 3) mice, respectively. Significance (d) was determined using two-sided Welch two-sample t-test and is expressed as the mean ± SEM. e, Comparison of average LMR methylation levels in Tet2/3 fl/fl and Tet2/3 fl/fl VillinCre mice. The upper (lower) line indicates the positions in the plot where TET2/3 KO is exactly 0.1 hypermethylated (hypomethylated) compared to TET2/3 WT. f, Bar plot of all LMRs identified in TET2/3 WT and KO mice, emphasizing the hypermethylation in KO mice.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and tables.
Supplementary Table 1
Supplementary Tables 3 and 8.
Source data
Source Data Fig. 2
Source data of the in vivo experiments and statistical analyses.
Source Data Fig. 4
Source data of the in vivo experiments and statistical analyses.
Source Data Fig. 5
Source data of the in vivo experiments and statistical analyses.
Source Data Extended Data Fig. 2
Source data of the in vivo experiments and statistical analyses.
Source Data Extended Data Fig. 4
Source data of the in vivo experiments and statistical analyses.
Source Data Extended Data Fig. 5
Source data of the in vivo experiments and statistical analyses.
Rights and permissions
About this article
Cite this article
Ansari, I., Raddatz, G., Gutekunst, J. et al. The microbiota programs DNA methylation to control intestinal homeostasis and inflammation. Nat Microbiol 5, 610–619 (2020). https://doi.org/10.1038/s41564-019-0659-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0659-3
This article is cited by
-
Increased CpG methylation at the CDH1 locus in inflamed ileal mucosa of patients with Crohn disease
Clinical Epigenetics (2024)
-
Integrative analysis reveals associations between oral microbiota dysbiosis and host genetic and epigenetic aberrations in oral cavity squamous cell carcinoma
npj Biofilms and Microbiomes (2024)
-
Perinatal foodborne titanium dioxide exposure-mediated dysbiosis predisposes mice to develop colitis through life
Particle and Fibre Toxicology (2023)
-
Targeting the epigenome to reinvigorate T cells for cancer immunotherapy
Military Medical Research (2023)
-
Polyphenol-rich diet mediates interplay between macrophage-neutrophil and gut microbiota to alleviate intestinal inflammation
Cell Death & Disease (2023)