Abstract
Viruses that infect microorganisms dominate marine microbial communities numerically, with impacts ranging from host evolution to global biogeochemical cycles1,2. However, virus community dynamics, necessary for conceptual and mechanistic model development, remains difficult to assess. Here, we describe the long-term stability of a viral community by analysing the metagenomes of near-surface 0.02–0.2 μm samples from the San Pedro Ocean Time-series3 that were sampled monthly over 5 years. Of 19,907 assembled viral contigs (>5 kb, mean 15 kb), 97% were found in each sample (by >98% ID metagenomic read recruitment) to have relative abundances that ranged over seven orders of magnitude, with limited temporal reordering of rank abundances along with little change in richness. Seasonal variations in viral community composition were superimposed on the overall stability; maximum community similarity occurred at 12-month intervals. Despite the stability of viral genotypic clusters that had 98% sequence identity, viral sequences showed transient variations in single-nucleotide polymorphisms (SNPs) and constant turnover of minor population variants, each rising and falling over a few months, reminiscent of Red Queen dynamics4. The rise and fall of variants within populations, interpreted through the perspective of known virus–host interactions5, is consistent with the hypothesis that fluctuating selection acts on a microdiverse cloud of strains, and this succession is associated with ever-shifting virus–host defences and counterdefences. This results in long-term virus–host coexistence that is facilitated by perpetually changing minor variants.
This is a preview of subscription content, access via your institution
Access options



Similar content being viewed by others
Data availability
All sequencing data are available at the JGI Genome portal under the proposal ID 2799 (to N.A. and J.A.F.). All data needed to evaluate the conclusions in the paper are provided in the paper or the Supplementary Information. Final cross-assembled sequences are deposited at NCBI under the BioProject ID PRJNA550983.
Code availability
Custom code is available at https://doi.org/10.6084/m9.figshare.8872796.v1.
References
Brum, J. R. et al. Patterns and ecological drivers of ocean viral communities. Science 348, 1261498 (2015).
Fuhrman, J. A. Marine viruses and their biogeochemical and ecological effects. Nature 399, 541–548 (1999).
Cram, J. A. et al. Seasonal and interannual variability of the marine bacterioplankton community throughout the water column over ten years. ISME J. 9, 563–580 (2015).
Brockhurst, M. A. et al. Running with the Red Queen: the role of biotic conflicts in evolution. Proc. R. Soc. B 281, 20141382 (2014).
Paterson, S. et al. Antagonistic coevolution accelerates molecular evolution. Nature 464, 275–278 (2010).
Roux, S. et al. Ecogenomics and potential biogeochemical impacts of globally abundant ocean viruses. Nature 537, 689–693 (2016).
Chow, C. E. T. et al. Temporal variability and coherence of euphotic zone bacterial communities over a decade in the Southern California Bight. ISME J. 7, 2259–2273 (2013).
Deng, L. et al. Viral tagging reveals discrete populations in Synechococcus viral genome sequence space. Nature 513, 242–245 (2014).
Gregory, A. C. et al. Marine DNA viral macro- and microdiversity from Pole to Pole. Cell 177, 1109–1123 (2019).
Aylward, F. O. et al. Diel cycling and long-term persistence of viruses in the ocean’s euphotic zone. Proc. Natl Acad. Sci. USA 114, 11446–11451 (2017).
Gregory, A. C. et al. Genomic differentiation among wild cyanophages despite widespread horizontal gene transfer. BMC Genom. 17, 930 (2016).
Chow, C. E. T. & Fuhrman, J. A. Seasonality and monthly dynamics of marine myovirus communities. Environ. Microbiol. 14, 2171–2183 (2012).
Lenski, R. E. Coevolution of bacteria and phage: are there endless cycles of bacterial defenses and phage counterdefenses?. J. Theor. Biol. 108, 319–325 (1984).
Hall, A. R., Scanlan, P. D., Morgan, A. D. & Buckling, A. Host-parasite coevolutionary arms races give way to fluctuating selection. Ecol. Lett. 14, 635–642 (2011).
Van Houte, S. et al. The diversity-generating benefits of a prokaryotic adaptive immune system. Nature 532, 385–388 (2016).
Weitz, J. S., Hartman, H. & Levin, S. A. Coevolutionary arms races between bacteria and bacteriophage. Proc. Natl Acad. Sci. USA 102, 9535–9540 (2005).
Thingstad, T. F., Pree, B., Giske, J. & Våge, S. What difference does it make if viruses are strain-, rather than species-specific? Front. Microbiol. 6, 320 (2015).
Martiny, J. B. H., Riemann, L., Marston, M. F. & Middelboe, M. Antagonistic coevolution of marine planktonic viruses and their hosts. Ann. Rev. Mar. Sci. 6, 393–414 (2013).
Waterbury, J. B. & Valois, F. W. Resistance to co-occurring phages enables marine Synechococcus communities to coexist with cyanophages abundant in seawater. Appl. Environ. Microbiol. 59, 3393–3399 (1993).
Arkhipova, K. et al. Temporal dynamics of uncultured viruses: a new dimension in viral diversity. ISME J. 12, 199–211 (2018).
Rodriguez-Brito, B. et al. Viral and microbial community dynamics in four aquatic environments. ISME J. 4, 739–751 (2010).
Schloissnig, S. et al. Genomic variation landscape of the human gut microbiome. Nature 493, 45–50 (2013).
Martinez-Hernandez, F. et al. Single-virus genomics reveals hidden cosmopolitan and abundant viruses. Nat. Commun. 8, 15892 (2017).
Bendall, M. L. et al. Genome-wide selective sweeps and gene-specific sweeps in natural bacterial populations. ISME J. 10, 1589–1602 (2016).
Minot, S. et al. Rapid evolution of the human gut virome. Proc. Natl Acad. Sci. USA 110, 12450–12455 (2013).
Hickey, B. M. Circulation over the Santa Monica-San Pedro Basin and Shelf. Prog. Oceanogr. 30, 37–115 (1992).
Marston, M. F. et al. Rapid diversification of coevolving marine Synechococcus and a virus. Proc. Natl Acad. Sci. USA 109, 4544–4549 (2012).
Breitbart, M. & Rohwer, F. Here a virus, there a virus, everywhere the same virus? Trend. Microbiol. 13, 278–284 (2005).
Woolhouse, M. E. J., Webster, J. P., Domingo, E., Charlesworth, B. & Levin, B. R. Biological and biomedical implications of the co-evolution of pathogens and their hosts. Nat. Genet. 32, 569–577 (2002).
Betts, A., Gray, C., Zelek, M., MacLean, R. C. & King, K. C. High parasite diversity accelerates host adaptation and diversification. Science 360, 907–911 (2018).
Steward, G. F. & Culley, A. I. in Manual of Aquatic Viral Ecology (eds. Wilhelm, S. W. et al.) Ch. 16 (2010).
Li, H. BFC: correcting Illumina sequencing errors. Bioinformatics 31, 2885–2887 (2015).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Sommer, D. D., Delcher, A. L., Salzberg, S. L. & Pop, M. Minimus: a fast, lightweight genome assembler. BMC Bioinform. 8, 64 (2007).
Sieradzki, E. T., Ignacio-Espinoza, J. C., Needham, D. M., Fichot, E. B. & Fuhrman, J. A. Dynamic marine viral infections and major contribution to photosynthetic processes shown by spatiotemporal picoplankton metatranscriptomes. Nat. Commun. 10, 1169 (2019).
Roux, S., Enault, F., Hurwitz, B. L. & Sullivan, M. B. VirSorter: mining viral signal from microbial genomic data. PeerJ 3, e985 (2015).
Ren, J., Ahlgren, N. A., Lu, Y. Y., Fuhrman, J. A. & Sun, F. VirFinder: a novel k-mer based tool for identifying viral sequences from assembled metagenomic data. Microbiome 5, 69 (2017).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 11, 119 (2010).
Nishimura, Y. et al. Environmental viral genomes shed new light on virus-host interactions in the ocean. mSphere 2, e00359-16 (2017).
Mizuno, C. M., Rodriguez-Valera, F., Kimes, N. E. & Ghai, R. Expanding the marine virosphere using metagenomics. PLoS Genet. 9, e1003987 (2013).
Ahlgren, N. A., Ren, J., Lu, Y. Y., Fuhrman, J. A. & Sun, F. Alignment-free d2* oligonucleotide frequency dissimilarity measure improves prediction of hosts from metagenomically-derived viral sequences. Nucleic Acids Res. 45, 39–53 (2017).
Uchiyama, I., Mihara, M., Nishide, H. & Chiba, H. MBGD update 2015: microbial genome database for flexible ortholog analysis utilizing a diverse set of genomic data. Nucleic Acids Res. 43, D270–D276 (2015).
Parks, D. H. et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 36, 996–1004 (2018).
Langmead, B., Wilks, C., Antonescu, V. & Charles, R. Scaling read aligners to hundreds of threads on general-purpose processors. Bioinformatics 35, 421–432 (2019).
Zhao, Y. et al. Abundant SAR11 viruses in the ocean. Nature 494, 357–360 (2013).
Tithi, S. S., Aylward, F. O., Jensen, R. V. & Zhang, L. FastViromeExplorer: a pipeline for virus and phage identification and abundance profiling in metagenomics data. PeerJ 6, e4227 (2018).
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Acknowledgements
We thank the directors and staff of the USC Wrigley Institute for Environmental Studies for supporting the SPOT, T. Gundersen and Fuhrman laboratory members, and the crew of the R/V Yellowfin for their help during our monthly sampling. We also thank J. McNichol for curating the local genome database and D. Needham, E. Sieradzki and S. Hou for providing metagenomically assembled genomes. This work was supported by NSF grant no. 1737409, the NIH (1R01GM120624-01A1), the Gordon and Betty Moore Foundation Marine Microbiology Initiative grant no. 3779 to J.A.F., and the Simons Collaboration on Computational Biogeochemical Modeling of Marine Ecosystems (CBIOMES; grant ID 549943) to J.A.F.
Author information
Authors and Affiliations
Contributions
J.C.I.-E., N.A. and J.A.F. designed the study and wrote the manuscript. N.A. initiated the project and performed the DNA extractions. J.C.I.-E. performed the bioinformatic analyses related to the identification, annotation, read mapping, variant calling and population genetics of the viral contigs.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
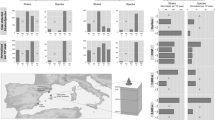
Extended Data Fig. 1 Rank abundance and placement of identifiable viral populations.
Rank abundance and placement of identifiable viral populations. From right to left: Right-most column is a rank abundance plot, where the most abundant viral contigs are at the top and the least abundant at the bottom. Cumulative abundances across all time points are shown, there are the product of normalization as stated in the methods. Middle columns show (i) In black, the positions of contigs with high (> 95% ID over 5kb) identity to previously identified virus sequences in databases, n = 746 (Details in Supplementary Tables). Due to constraints on visible line thickness, they appear to represent a majority of contigs, but note they are only 746 out of 19,907 total contigs (See details on Supplementary Tables); (ii) Green lines show viral contigs identified as cyanophage n = 73 (Supplementary Tables). (iii) Maroon lines show viral contigs identified as Pelagibacter phages n = 68 (Supplementary Tables). (iv) Red lines show viral contigs that are nearly identical, > 99% ID over at least 40 kb at the nucleotide level to previously described viruses (Supplementary Tables). Left-most panel shows genomic diagrams from selected contigs and their identified hit in databases (>99% identity), all drawn to the same scale. Remarkably, all these illustrations represent Synechococcus phages isolated off the coast of California.
Extended Data Fig. 2 Relative abundance of viral contigs on a linear scale.
Heat Maps otherwise similar to Fig. 1b (which has a 7-decade log scale), showing relative abundance of the 19,907 viral contigs, one contig per row, during monthly sampling. Contigs are ordered by average abundance (over all months), highest at the top. White columns represent months with missing data (all data are in Supplementary Tables). Each panel has a different range, where abundances at or exceeding the maximum value appear as red. This display better allows visualization of temporal changes in contig abundance within orders of magnitude, compared to Fig. 1b where colors change little within each order of magnitude. Different subpanels are needed to visualize the ranges of all the contigs. “Zooming” the image optimizes the ability to visualize details.
Extended Data Fig. 3 Seasonality is driven by medium range abundance contigs.
Similar to Fig. 2, these depict the Bray-Curtis distance vs time lag between samples, but here divided into different fractions of the rank abundance curve. The left column of graphs starts from the bottom (rare) part of the curve, with an increasing fraction of the contigs included in graphs displayed from top to bottom. The right column of graphs starts with the top most abundant contigs, with an increasing fraction of the curve included in graphs from top to bottom. Note on the right that as more members are included from the long rare tail of the rank-abundance curve, consistent seasonality increases and average similarity decreases. Generally, the rarer contigs show stronger seasonality than the most abundant ones. Because Bray Curtis similarity is proportionately more affected by the more abundant organisms in general, these indicate that middle-high percentiles (top 50th-75th) may dominate the collective community seasonality. Bottom 5% n = 4587 viral contigs, bottom 10 % n = 7092, bottom 25 % n = 12031, bottom 50% n = 16741, bottom 75% n=19093. Top 5% n = 61, Top 10% n=168, Top 25% n= 814, top 50% n = 3166, top 75% n = 7876.
Extended Data Fig. 4 Distribution of polymorphic sites along three selected genomes.
The three longest T4-like genome fragments were chosen (ranked 9433rd, 10382nd and 13113rd) because T4-likes are the most extensively studied group and have the best annotations. Each diagram shows all predicted open reading frames and their sense direction as depicted by an arrow. The y-axis on the left side (for thin black bars) shows the number of months that each location exhibited a SNP. The position along the x axis corresponds to their position along the genome. Y axis on the right (red line) shows the average number of SNPs per basepair on a 500 bp moving average. Note that only about ~5% of the sites are polymorphic although this is hard to visualize. For details, please refer to annotations and the per gene density of 20 representative (including the ones shown here) T4-like viruses is included in Supplementary Tables under the tab “SNPsAmongT4LikeViruses” Although not selected for this reason, these three show strikingly different patterns in SNP distributions, from relatively uniform (top panel) to very patchy with a few hotspots (bottom panel). Note y axes are scaled for each panel, and the top and middle ones have similar SNP densities to each other over most of their lengths.
Extended Data Fig. 5 Post hoc evaluation of our cross-assembly strategy.
Cross assembly merged contig sequences from different months when overall identity of overlapping regions greater than 1000 bp in length exceeded 95%. We evaluated how often these merged overlaps occurred at different percent identities to assess how much variation was combined, and also examined other useful statistics. a) Distribution of percent identity of all alignments used to merge contigs during our cross-assembly step, dotted lines represent the percent of alignments covered to the right of the line. Note that 92% of merges had >98% sequence identity b). Distribution of lengths of all alignments used to merge contigs during our cross-assembly step. Note most merged regions were 5,000–10,000 bp in length. c) Distribution of the fractions of the contig used during our merging step (that is length of the alignment divided by the contig length). Note that the vast majority of merges occurred over almost the entire lengths (90–100%) of the contigs d). Distribution of the number of contigs that were merged into a single contig during cross- assembly. Note that the vast majority of merged contigs came from 2 or three individual contigs. All panels taken together show that while merging occurred, the vast majority (86%) was between almost completely overlapping (including nested) and >98% identical sequence contigs, rather than bridging between long contigs with short overlaps.
Extended Data Fig. 6 Read recruitment to fully sequenced isolates reveals identical patterns of succession.
Dynamics of SNP profiles calculated from reads recruited (within 98% ID) to two reference genomes. These profiles were generated as those shown in Fig. 3 and as described in the methods. NOTE: The absolute number of polymorphic sites is bigger since a full genome is being considered.
Supplementary information
Supplementary Information
Supplementary Discussion and Supplementary Figs. 1–3.
Supplementary Tables
Raw data and counts: (1) normalized virus abundance per sampled month, calculated as described in the Methods, sorted from most to least abundant. (2) Cumulative abundance and sizes of all contigs, sorted as in (1). (3) Taxonomy for each contig, from genome-wide nucleotide-level identity and from other computational methods. (4) Distribution of polymorphic sites along selected T4-like viral contigs. (5) Accession numbers genome and genome names of the sequences used to build our custom database for bioinformatic analyses. (6) Summary of data.
Supplementary Data
Concatenated GenBank-like files from all 19,907 contigs identified as viral. Annotations and predicted probable host assignments were performed as described in the Methods.
Rights and permissions
About this article
Cite this article
Ignacio-Espinoza, J.C., Ahlgren, N.A. & Fuhrman, J.A. Long-term stability and Red Queen-like strain dynamics in marine viruses. Nat Microbiol 5, 265–271 (2020). https://doi.org/10.1038/s41564-019-0628-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0628-x
This article is cited by
-
Eco-evolutionary dynamics of gut phageome in wild gibbons (Hoolock tianxing) with seasonal diet variations
Nature Communications (2024)
-
A subset of viruses thrives following microbial resuscitation during rewetting of a seasonally dry California grassland soil
Nature Communications (2023)
-
Highly diverse flavobacterial phages isolated from North Sea spring blooms
The ISME Journal (2022)
-
Unexpected myriad of co-occurring viral strains and species in one of the most abundant and microdiverse viruses on Earth
The ISME Journal (2022)
-
The OceanDNA MAG catalog contains over 50,000 prokaryotic genomes originated from various marine environments
Scientific Data (2022)