Abstract
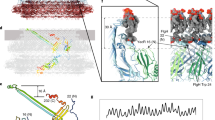
Motility structures are vital in all three domains of life. In Archaea, motility is mediated by the archaellum, a rotating type IV pilus-like structure that is a unique nanomachine for swimming motility in nature. Whereas periplasmic FlaF binds the surface layer (S-layer), the structure, assembly and roles of other periplasmic components remain enigmatic, limiting our knowledge of the archaellum’s functional interactions. Here, we find that the periplasmic protein FlaG and the association with its paralogue FlaF are essential for archaellation and motility. Therefore, we determine the crystal structure of Sulfolobus acidocaldarius soluble FlaG (sFlaG), which reveals a β-sandwich fold resembling the S-layer-interacting FlaF soluble domain (sFlaF). Furthermore, we solve the sFlaG2–sFlaF2 co-crystal structure, define its heterotetrameric complex in solution by small-angle X-ray scattering and find that mutations that disrupt the complex abolish motility. Interestingly, the sFlaF and sFlaG of Pyrococcus furiosus form a globular complex, whereas sFlaG alone forms a filament, indicating that FlaF can regulate FlaG filament assembly. Strikingly, Sulfolobus cells that lack the S-layer component bound by FlaF assemble archaella but cannot swim. These collective results support a model where a FlaG filament capped by a FlaG–FlaF complex anchors the archaellum to the S-layer to allow motility.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The X-ray diffraction data and coordinates of the sFlaG and sFlaG–sFlaF complex structures have been deposited with the PDB: 5TUH (sFlaG); 5TUG (sFlaGWT/sFlaFWT); and 6PBK (sFlaGV118K/sFlaFWT). The SAXS data have been deposited with the SASBDB: SASDEU7 (sFlaG); SASDES7 (sFlaGWT/sFlaFWT); SASDEV7 (sFlaGWT/sFlaFI96Y); and SASDET7 (sFlaGV118K/sFlaFWT). Source data for Fig. 4 and Supplementary Figs. 5, 7, 10 and 11 are provided with the paper.
References
Jarrell, K. F. & McBride, M. J. The surprisingly diverse ways that prokaryotes move. Nat. Rev. Microbiol. 6, 466–476 (2008).
Albers, S. V. & Jarrell, K. F. The archaellum: an update on the unique archaeal motility structure. Trends Microbiol. 26, 351–362 (2018).
Jarrell, K. F. & Albers, S. V. The archaellum: an old motility structure with a new name. Trends Microbiol. 20, 307–312 (2012).
Shahapure, R., Driessen, R. P. C., Haurat, M. F., Albers, S. V. & Dame, R. T. The archaellum: a rotating type IV pilus. Mol. Microbiol. 91, 716–723 (2014).
Kinosita, Y., Uchida, N., Nakane, D. & Nishizaka, T. Direct observation of rotation and steps of the archaellum in the swimming halophilic archaeon Halobacterium salinarum. Nat. Microbiol. 1, 16148 (2016).
Marwan, W., Alam, M. & Oesterhelt, D. Rotation and switching of the flagellar motor assembly in Halobacterium halobium. J. Bacteriol. 173, 1971–1977 (1991).
Lassak, K. et al. Molecular analysis of the crenarchaeal flagellum. Mol. Microbiol. 83, 110–124 (2012).
Chaban, B. et al. Systematic deletion analyses of the fla genes in the flagella operon identify several genes essential for proper assembly and function of flagella in the archaeon, Methanococcus maripaludis. Mol. Microbiol. 66, 596–609 (2007).
Tripepi, M., Imam, S. & Pohlschröder, M. Haloferax volcanii flagella are required for motility but are not involved in PibD-dependent surface adhesion. J. Bacteriol. 192, 3093–3102 (2010).
Thomas, N. A., Pawson, C. T. & Jarrell, K. F. Insertional inactivation of the flaH gene in the archaeon Methanococcus voltae results in non-flagellated cells. Mol. Genet. Genomics 265, 596–603 (2001).
Banerjee, A., Neiner, T., Tripp, P. & Albers, S.-V. Insights into subunit interactions in the Sulfolobus acidocaldarius archaellum cytoplasmic complex. FEBS J. 280, 6141–6149 (2013).
Ghosh, A. & Albers, S.-V. Assembly and function of the archaeal flagellum. Biochem. Soc. Trans. 39, 64–69 (2011).
Albers, S.-V., Szabó, Z. & Driessen, A. J. M. Protein secretion in the Archaea: multiple paths towards a unique cell surface. Nat. Rev. Microbiol. 4, 537–547 (2006).
Ng, S. Y. M., Chaban, B. & Jarrell, K. F. Archaeal flagella, bacterial flagella and type IV pili: a comparison of genes and posttranslational modifications. J. Mol. Microbiol. Biotechnol. 11, 167–191 (2006).
Takhar, H. K., Kemp, K., Kim, M., Howell, P. L. & Burrows, L. L. The platform protein is essential for type IV pilus biogenesis. J. Biol. Chem. 288, 9721–9728 (2013).
Chaudhury, P. et al. The nucleotide‐dependent interaction of FlaH and FlaI is essential for assembly and function of the archaellum motor. Mol. Microbiol. 99, 674–685 (2016).
Reindl, S. et al. Insights into FlaI functions in archaeal motor assembly and motility from structures, conformations, and genetics. Mol. Cell 49, 1069–1082 (2013).
Ghosh, A., Hartung, S., van der Does, C., Tainer, J. A. & Albers, S.-V. Archaeal flagellar ATPase motor shows ATP-dependent hexameric assembly and activity stimulation by specific lipid binding. Biochem. J. 437, 43–52 (2011).
Jakovljevic, V., Leonardy, S., Hoppert, M. & Søgaard-Andersen, L. PilB and PilT are ATPases acting antagonistically in type IV pilus function in Myxococcus xanthus. J. Bacteriol. 190, 2411–2421 (2008).
Mancl, J. M., Black, W. P., Robinson, H., Yang, Z. & Schubot, F. D. Crystal structure of a type IV pilus assembly ATPase: insights into the molecular mechanism of PilB from Thermus thermophilus. Structure 24, 1886–1897 (2016).
Tsai, C. L. & Tainer, J. A. The ATPase motor turns for type IV pilus assembly. Structure 24, 1857–1859 (2016).
Banerjee, A. et al. FlaX, a unique component of the crenarchaeal archaellum, forms oligomeric ring-shaped structures and interacts with the motor ATPase FlaI. J. Biol. Chem. 287, 43322–43330 (2012).
Banerjee, A. et al. FlaF is a β-sandwich protein that anchors the archaellum in the archaeal cell envelope by binding the S-layer protein. Structure 23, 863–872 (2015).
Szabó, Z. et al. Identification of diverse archaeal proteins with class III signal peptides cleaved by distinct archaeal prepilin peptidases. J. Bacteriol. 189, 772–778 (2007).
Albers, S. V., Szabó, Z. & Driessen, A. J. M. Archaeal homolog of bacterial type IV prepilin signal peptidases with broad substrate specificity. in. J. Bacteriol. 185, 3918–3925 (2003).
Brock, T. D., Brock, K. M., Belly, R. T. & Weiss, R. L. Sulfolobus: a new genus of sulfur-oxidizing bacteria living at low pH and high temperature. Arch. Mikrobiol. 84, 54–68 (1972).
Rambo, R. P. & Tainer, J. A. Characterizing flexible and intrinsically unstructured biological macromolecules by SAS using the Porod-Debye law. Biopolymers 95, 559–571 (2011).
Kikhney, A. G. & Svergun, D. I. A practical guide to small angle X-ray scattering (SAXS) of flexible and intrinsically disordered proteins. FEBS Lett. 589, 2570–2577 (2015).
Burger, V. M., Arenas, D. J. & Stultz, C. M. A structure-free method for quantifying conformational flexibility in proteins. Sci. Rep. 6, 29040 (2016).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Zhang, C., Phillips, A. P. R., Wipfler, R. L., Olsen, G. J. & Whitaker, R. J. The essential genome of the crenarchaeal model Sulfolobus islandicus. Nat. Commun. 9, 4908 (2018).
Zhang, C. et al. Cell structure changes in the hyperthermophilic crenarchaeon Sulfolobus islandicus lacking the S-Layer. mBio 10, e01589-19 (2019).
Jarrell, K. F., Ding, Y., Nair, D. B. & Siu, S. Surface appendages of Archaea: structure, function, genetics and assembly. Life 3, 86–117 (2013).
Briegel, A. et al. Morphology of the archaellar motor and associated cytoplasmic cone in Thermococcus kodakaraensis. EMBO Rep. 18, 1660–1670 (2017).
Daum, B. et al. Structure and in situ organisation of the Pyrococcus furiosus archaellum machinery. eLife 6, e27470 (2017).
Craig, L. et al. Type IV pilus structure by cryo-electron microscopy and crystallography: implications for pilus assembly and functions. Mol. Cell 23, 651–662 (2006).
Meshcheryakov, V. A. et al. High‐resolution archaellum structure reveals a conserved metal‐binding site. EMBO Rep. 20, e46340 (2019).
Van Way, S. M., Hosking, E. R., Braun, T. F. & Manson, M. D. Mot protein assembly into the bacterial flagellum: a model based on mutational analysis of the motB gene. J. Mol. Biol. 297, 7–24 (2000).
Chaban, B., Coleman, I. & Beeby, M. Evolution of higher torque in Campylobacter-type bacterial flagellar motors. Sci. Rep. 8, 97 (2018).
Hura, G. L. et al. Comprehensive macromolecular conformations mapped by quantitative SAXS analyses. Nat. Methods 10, 453–454 (2013).
Classen, S. et al. Implementation and performance of SIBYLS: A dual endstation small-angle X-ray scattering and macromolecular crystallography beamline at the Advanced Light Source. J. Appl. Crystallogr. 46, 1–13 (2013).
Pettersen, E. F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Trewhella, J. et al. 2017 publication guidelines for structural modelling of small-angle scattering data from biomolecules in solution: an update. Acta Crystallogr. D 73, 710–728 (2017).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinformatics 54, 5.6.1–5.6.37 (2016).
Schneidman-Duhovny, D., Hammel, M., Tainer, J. A. & Sali, A. FoXS, FoXSDock and MultiFoXS: single-state and multi-state structural modeling of proteins and their complexes based on SAXS profiles. Nucleic Acids Res. 44, W424–W429 (2016).
Schneidman-Duhovny, D., Hammel, M., Tainer, J. A. & Sali, A. Accurate SAXS profile computation and its assessment by contrast variation experiments. Biophys. J. 105, 962–974 (2013).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Mastronarde, D. N. Dual-axis tomography: an approach with alignment methods that preserve resolution. J. Struct. Biol. 120, 343–352 (1997).
de la Rosa-Trevín, J. M. et al. Scipion: a software framework toward integration, reproducibility and validation in 3D electron microscopy. J. Struct. Biol. 195, 93–99 (2016).
de la Rosa-Trevín, J. M. et al. Xmipp 3.0: an improved software suite for image processing in electron microscopy. J. Struct. Biol. 184, 321–328 (2013).
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Scheres, S. H. W. RELION: Implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Acknowledgements
P.T. received funding from the European Union’s Horizon 2020 research and innovation program under grant agreement no. 686647. S.-V.A. was funded by the Deutsche Forschungsgemeinschaft (German Research Foundation) under project no. 403222702-SFB 1381. The work was conducted at ALS, a national user facility operated by the Lawrence Berkeley National Laboratory on behalf of the US Department of Energy’s Office of Basic Energy Sciences, through the Integrated Diffraction Analysis Technologies program, supported by US Department of Energy Office of Biological and Environmental Research. Additional support comes from the National Institutes of Health (NIH) grant no. P30GM124169. J.A.T. acknowledges start-up funds from the University of Texas Safety Tracking and Reporting System (NIH grant no. R35CA220430) and the Robert A. Welch Chair in Chemistry (grant no. CFS127800-80-101399-50). C.Z., R.L.W. and R.J.W. are supported by a grant from NASA (National Aeronautics and Space Administration) through the NASA Astrobiology Institute under cooperative agreement no. NNA13AA91A, issued through the Science Mission Directorate. We thank S. Robinson from the Microscopy Suite at the Beckman Institute for Advance Science and Technology, University of Illinois at Urbana-Champaign, for providing TEM assistance. We especially thank K. Burnett for her assistance on the SAXS data collection of the sFlaG and sFlaG–sFlaF complexes, G. Hura for the SAXS similarity calculation suggestion and S. Classen for his assistance on the X-ray crystallography data collection at the SIBYLS Beamline (BL12.3.1). We thank A. Ghosh for technical help; B. Tutt at the Department of Scientific Publications at the MD Anderson Cancer Center for proofreading and editing our manuscript; and we thank P. Simpson for technical assistance with imaging the P. furiosus FlaF and FlaG assemblies. The TEM is operated by the University of Freiburg, Faculty of Biology, as a partner unit within the Microscopy and Image Analysis Platform, Freiburg.
Author information
Authors and Affiliations
Contributions
C.-L.T, P.T., J.A.T. and S.-V.A. designed the experiments, analysed the data and wrote the manuscript. C.-L.T. and J.A.T. contributed to protein crystallization, SAXS, X-ray data collection, structure determination, data analyses, the initial EM model reconstruction and the figures. P.T., S.S., M.R.-F., A.B., P.C., M.B. and S.-V.A. contributed to protein production and purification, SEC and reconstitution, microscale thermophoresis, cell motility, cell fractionation, data analysis, TEM analysis and the figures. C.Z., R.L.W. and R.J.W. contributed to the S-layer protein knockout, TEM analysis and the figures.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Results, Supplementary Figs. 1–11, Supplementary Tables 1–3 and Supplementary References.
Supplementary Data 1
Unprocessed gels.
Supplementary Data 2
Unprocessed western blots and gels.
Supplementary Data 3
Unprocessed western blots and gels.
Supplementary Data 4
Unprocessed western blots and gels.
Source data
Source Data Fig. 4
Unprocessed western blots and gels.
Rights and permissions
About this article
Cite this article
Tsai, CL., Tripp, P., Sivabalasarma, S. et al. The structure of the periplasmic FlaG–FlaF complex and its essential role for archaellar swimming motility. Nat Microbiol 5, 216–225 (2020). https://doi.org/10.1038/s41564-019-0622-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0622-3
This article is cited by
-
An archaellum filament composed of two alternating subunits
Nature Communications (2022)
-
The cell biology of archaea
Nature Microbiology (2022)
-
Electron cryo-microscopy reveals the structure of the archaeal thread filament
Nature Communications (2022)
-
Wie ein rotierender Typ-IV-Pilus Archaeen das Schwimmen beibrachte
BIOspektrum (2020)