Abstract
Metabolic reprogramming is associated with the adaptation of host cells to the disease environment, such as inflammation and cancer. However, little is known about microbial metabolic reprogramming or the role it plays in regulating the fitness of commensal and pathogenic bacteria in the gut. Here, we report that intestinal inflammation reprograms the metabolic pathways of Enterobacteriaceae, such as Escherichia coli LF82, in the gut to adapt to the inflammatory environment. We found that E. coli LF82 shifts its metabolism to catabolize l-serine in the inflamed gut in order to maximize its growth potential. However, l-serine catabolism has a minimal effect on its fitness in the healthy gut. In fact, the absence of genes involved in l-serine utilization reduces the competitive fitness of E. coli LF82 and Citrobacter rodentium only during inflammation. The concentration of luminal l-serine is largely dependent on dietary intake. Accordingly, withholding amino acids from the diet markedly reduces their availability in the gut lumen. Hence, inflammation-induced blooms of E. coli LF82 are significantly blunted when amino acids—particularly l-serine—are removed from the diet. Thus, the ability to catabolize l-serine increases bacterial fitness and provides Enterobacteriaceae with a growth advantage against competitors in the inflamed gut.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Source data for all figures and extended data figures are provided in the online version of the paper. The E. coli LF82 RNA-Seq data used in this study have been deposited in the Gene Expression Omnibus database repository under the accession number GSE106412. The metabolome data obtained in this study are available from the website of the NIH Common Fund’s Data Repository and Coordinating Center (supported by NIH grant U01-DK097430)—the Metabolomics Workbench (http://www.metabolomicsworkbench.org), under project ID PR000837.
References
Kamada, N. et al. Regulated virulence controls the ability of a pathogen to compete with the gut microbiota. Science 336, 1325–1329 (2012).
Conway, T. & Cohen, P. S. Commensal and pathogenic Escherichia coli metabolism in the gut. Microbiol. Spectr. 3, MBP-0006-2014 (2015).
Winter, S. E. et al. Host-derived nitrate boosts growth of E. coli in the inflamed gut. Science 339, 708–711 (2013).
Zhu, W. et al. Precision editing of the gut microbiota ameliorates colitis. Nature 553, 208–211 (2018).
Morgan, X. C. et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 13, R79 (2012).
Darfeuille-Michaud, A. et al. Presence of adherent Escherichia coli strains in ileal mucosa of patients with Crohn’s disease. Gastroenterology 115, 1405–1413 (1998).
Carvalho, F. A. et al. Crohn’s disease-associated Escherichia coli LF82 aggravates colitis in injured mouse colon via signaling by flagellin. Inflamm. Bowel Dis. 14, 1051–1060 (2008).
Carvalho, F. A. et al. Crohn’s disease adherent-invasive Escherichia coli colonize and induce strong gut inflammation in transgenic mice expressing human CEACAM. J. Exp. Med. 206, 2179–2189 (2009).
Chassaing, B., Koren, O., Carvalho, F. A., Ley, R. E. & Gewirtz, A. T. AIEC pathobiont instigates chronic colitis in susceptible hosts by altering microbiota composition. Gut 63, 1069–1080 (2014).
Small, C. L., Xing, L., McPhee, J. B., Law, H. T. & Coombes, B. K. Acute infectious gastroenteritis potentiates a Crohn’s disease pathobiont to fuel ongoing inflammation in the post-infectious period. PLoS Pathog. 12, e1005907 (2016).
Viladomiu, M. et al. IgA-coated E. coli enriched in Crohn’s disease spondyloarthritis promote TH17-dependent inflammation. Sci. Transl. Med. 9, eaaf9655 (2017).
Chang, D. E. et al. Carbon nutrition of Escherichia coli in the mouse intestine. Proc. Natl Acad. Sci. USA 101, 7427–7432 (2004).
Vijay-Kumar, M. et al. Deletion of TLR5 results in spontaneous colitis in mice. J. Clin. Invest. 117, 3909–3921 (2007).
Peekhaus, N. & Conway, T. What’s for dinner?: Entner–Doudoroff metabolism in Escherichia coli. J. Bacteriol. 180, 3495–3502 (1998).
Rigottier-Gois, L. Dysbiosis in inflammatory bowel diseases: the oxygen hypothesis. ISME J. 7, 1256–1261 (2013).
Lopez, C. A. et al. Virulence factors enhance Citrobacter rodentium expansion through aerobic respiration. Science 353, 1249–1253 (2016).
Pullan, S. T. et al. Nitric oxide in chemostat-cultured Escherichia coli is sensed by Fnr and other global regulators: unaltered methionine biosynthesis indicates lack of S nitrosation. J. Bacteriol. 189, 1845–1855 (2007).
Collins, J. W. et al. Citrobacter rodentium: infection, inflammation and the microbiota. Nat. Rev. Microbiol. 12, 612–623 (2014).
Sawers, G. The anaerobic degradation of l-serine and l-threonine in enterobacteria: networks of pathways and regulatory signals. Arch. Microbiol. 171, 1–5 (1998).
Reitzer, L. Nitrogen assimilation and global regulation in Escherichia coli. Annu. Rev. Microbiol. 57, 155–176 (2003).
Zinser, E. R. & Kolter, R. Mutations enhancing amino acid catabolism confer a growth advantage in stationary phase. J. Bacteriol. 181, 5800–5807 (1999).
Imai, J. et al. Flagellin-mediated activation of IL-33-ST2 signaling by a pathobiont promotes intestinal fibrosis. Mucosal Immunol. 12, 632–643 (2019).
Craven, M. et al. Inflammation drives dysbiosis and bacterial invasion in murine models of ileal Crohn’s disease. PLoS ONE 7, e41594 (2012).
Rasko, D. A. et al. The pangenome structure of Escherichia coli: comparative genomic analysis of E. coli commensal and pathogenic isolates. J. Bacteriol. 190, 6881–6893 (2008).
Miranda, R. L. et al. Glycolytic and gluconeogenic growth of Escherichia coli O157:H7 (EDL933) and E. coli K-12 (MG1655) in the mouse intestine. Infect. Immun. 72, 1666–1676 (2004).
Bloom, S. M. et al. Commensal Bacteroides species induce colitis in host-genotype-specific fashion in a mouse model of inflammatory bowel disease. Cell Host Microbe 9, 390–403 (2011).
Ralls, M. W. et al. Bacterial nutrient foraging in a mouse model of enteral nutrient deprivation: insight into the gut origin of sepsis. Am. J. Physiol. Gastrointest. Liver Physiol. 311, G734–G743 (2016).
Matsumoto, M. et al. Impact of intestinal microbiota on intestinal luminal metabolome. Sci. Rep. 2, 233 (2012).
Hashimoto, T. et al. ACE2 links amino acid malnutrition to microbial ecology and intestinal inflammation. Nature 487, 477–481 (2012).
Maddocks, O. D. et al. Serine starvation induces stress and p53-dependent metabolic remodelling in cancer cells. Nature 493, 542–546 (2013).
Pizer, L. I. & Potochny, M. L. Nutritional and regulatory aspects of serine metabolism in Escherichia coli. J. Bacteriol. 88, 611–619 (1964).
Nagao-Kitamoto, H. et al. Functional characterization of inflammatory bowel disease-associated gut dydbiosis in gnotobiotic mice. Cell. Mol. Gastroenterol. Hepatol. 2, 468–481 (2016).
Britton, G. J. et al. Microbiotas from humans with inflammatory bowel disease alter the balance of gut Th17 and RORγt+ regulatory T cells and exacerbate colitis in mice. Immunity 50, 212–224 (2019).
Matthews, R. G. & Neidhardt, F. C. Elevated serine catabolism is associated with the heat shock response in Escherichia coli. J. Bacteriol. 171, 2619–2625 (1989).
Sassone-Corsi, M. et al. Microcins mediate competition among Enterobacteriaceae in the inflamed gut. Nature 540, 280–283 (2016).
Velayudhan, J., Jones, M. A., Barrow, P. A. & Kelly, D. J. l-serine catabolism via an oxygen-labile l-serine dehydratase is essential for colonization of the avian gut by Campylobacter jejuni. Infect. Immun. 72, 260–268 (2004).
Hofreuter, D. et al. Contribution of amino acid catabolism to the tissue specific persistence of Campylobacter jejuni in a murine colonization model. PLoS ONE 7, e50699 (2012).
Ma, E. H. et al. Serine is an essential metabolite for effector T cell expansion. Cell Metab. 25, 345–357 (2017).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Szklarczyk, D. et al. STRINGv10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Hirayama, A. et al. Metabolic profiling reveals new serum biomarkers for differentiating diabetic nephropathy. Anal. Bioanal. Chem. 404, 3101–3109 (2012).
Sugimoto, M., Wong, D. T., Hirayama, A., Soga, T. & Tomita, M. Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics 6, 78–95 (2010).
Schauer, D. B. & Falkow, S. Attaching and effacing locus of a Citrobacter freundii biotype that causes transmissible murine colonic hyperplasia. Infect. Immun. 61, 2486–2492 (1993).
Acknowledgements
The authors thank the University of Michigan Center for Gastrointestinal Research (NIH 5P30DK034933), Host Microbiome Initiative, Germ-Free Animal Facility, DNA Sequencing Core, Bioinformatics Core for research support, and In Vivo Animal Core at the University of Michigan Unit for Laboratory Animal Medicine for performing the pathology assessment. We also thank S. Yamada, J. Imai, Y.-G. Kim and E. C. Martens for technical assistance, and M. Y. Zeng for critical reading of the manuscript. This work was supported by a Kenneth Rainin Foundation Innovator Award (to N.K.), National Institute of Health grants DK110146, DK108901 and DK119219 (to N.K.), T32 grant DK094775 (to T.L.M.), the Crohn’s and Colitis Foundation of America (to N.K. and H.N.-K.), a JSPS Postdoctoral Fellowship for Research Abroad (to S.K., H.N.-K. and K.S.), the Uehara Memorial Foundation Postdoctoral Fellowship Award (to S.K. and K.S.), the University of Michigan Clinical and Translational Science Awards Program (to S.K.), the Prevent Cancer Foundation (to S.K.), JSPS KAKENHI grants 16H04901, 17H05654 and 18H04805 (to S.F.), JST PRESTO award JPMJPR1537 (to S.F.), JST ERATO JPMJER1902 (to S.F.), AMED-CREST grant JP19gm1010009 (to S.F.), the Takeda Science Foundation (to S.F.), the Food Science Institute Foundation (to S.F.), Université Clermont Auvergne (to N.B.), Inserm U1071 (to N.B.) and INRA USC-2018 (to N.B.).
Author information
Authors and Affiliations
Contributions
S.K. and N.K. conceived and designed the experiments. S.K. conducted most of the experiments with help from C.J.A., M.R., H.N.-K., K.S., S.D.H., M.B., M.M., T.N., A.Hayashi, T.L.M. and P.K. C.I., A.Hirayama and S.F. performed the metabolome analysis. N.I. assisted with the bacterial RNA-Seq analysis. K.A.E. helped with the germ-free animal experiments. H.G., M.E.-Z., S.B., H.L.T.M., J.Y.K. and N.B. provided advice and discussion. B.D. and K.W.S. provided critical materials. S.K., C.J.A., S.F. and N.K. analysed the data. S.K. and N.K. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 L-serine catabolism mutant LF82 ΔtdcΔsda has serine-dependent growth defect.
Diagram of tdc gene operon (a) and L-serine catabolism pathway in E. coli (b). (c, d) AIEC LF82 WT, Δtdc (ΔT), Δsda (ΔS), and ΔtdcΔsda (ΔTS) mutant strains were cultured in DMEM (0.45% glucose) or a minimal medium (0.1% glucose) supplemented with 1 mM serine for 8 hours at 37 °C with 20% O2 and 5% CO2. Growth kinetics (O.D600) are shown. Data represent mean ± s.d. (N=5, technical replicates). Results are representative of 2 biologically independent experiments. ****; P < 0.0001 by 2-Way ANOVA (∆TS vs WT, ∆T or ∆S). (d) Bacterial CFUs at 8 h were measured by plating on LB agar. Each number inside the bar indicates percent growth to WT LF82. (e) Luminal contents from ceca were collected from SPF C57BL/6 WT mice. The debris and bacteria were removed by centrifugation followed by filtration (0.2 μm). AIEC LF82 WT or ∆TS mutant were cultured in 20% sterilized luminal content for 8 h at 37 °C with 20% O2 and 5% CO2. Bacterial CFUs were measured by plating on LB agar. (d, e) Data represent geometric mean ± s.d. (N = 4-5, biological replicates). N.S.; not significant, ***; P < 0.001 by Mann–Whitney U test (two-sided) (e) or 1-Way ANOVA followed by Bonferroni post-hoc test (d).
Extended Data Fig. 2 Inflammation-associated milieu may regulate the expression of tdcA in E. col.
(a) Mucin broth was prepared by dissolving 0.5 % mucin in None-Carbon E medium (NCE) supplemented with trace elements. Sodium nitrate, dimethyl sulfoxide (DMSO), and trimethylamine N-oxide (TMAO) were added to a final concentration of 40 mM. Mucin broth without any supplementation (None) was used as a control. Each E. coli strain (MG1655 (MG), LF82 (LF), and LF82∆tdcΔsda double mutant (TS)) was inoculated (1 x 104 CFU/ml) in the medium and incubated anaerobically for 24 h at 37 °C. Bacterial numbers were determined by spreading dilutions on selective LB agar plates. The fold increase was calculated by normalizing the CFU at 24 hrs to the respective CFU at 0 hrs. Data represent geometric mean ± s.d. (N=3, biologically independent samples). N.S.; not significant by 1-Way ANOVA followed by Bonferroni post-hoc test. (b) AIEC LF82 was cultured in vitro for 8 hrs at 37 °C. The expression of tdcA mRNA was assessed by qPCR. Fold expression of tdcA in steady state-like conditions (DMEM supplemented with 0.1% glucose, cultured in 0% oxygen) is shown. Different concentrations of glucose (0.1% glucose or 0% glucose) and oxygen (0% oxygen or 2% oxygen) were tested. 3-[2-hydroxy- 1-(1-methylethyl)-2-nitrosohydrazino]-1-propanamine (NOC- 5) (10 mM) was added to mimic the presence of nitric oxide (NO). Data are represented as mean ± s.d.. Dots indicate individual biological replicates. ***; P < 0.001, ****; P < 0.0001 by Dunnett test. (c) SPF C57BL/6 mice WT (Ctrl) and Cybb−/− mice were treated with 3% DSS for 5 days. At day 5 post DSS treatment, mice were co-inoculated with LF82 WT and ∆tdc∆sda (∆TS) mutant (1 x 109 CFU each/mouse). Fecal samples were collected 24 hrs post E. coli inoculation. Fecal lipocalin-2 (Lcn2) levels (a) and the competitive index (LF82 WT/LF82 ΔTS) (d) are shown. Bars represent geometric mean ± s.d. Dots indicate individual mice. (N=4-5, biologically independent animals). N.S.; not significant, * P < 0.05 by Mann-Whitney U test (two-sided).
Extended Data Fig. 3 L-serine metabolism pathways are not required for AIEC virulence.
(a) Experimental procedure: BMDMs (2 x 105 cells/well/48 well plate in 200 μl) were stimulated with MG1655, AIEC LF82 WT or LF82 ∆tdcDsda double mutant (ΔTS) strains at a MOI=5 for 3 hrs, followed by 15 hrs of additional culture in the presence of gentamycin (100 μg/ml) to prevent bacterial overgrowth. (b) Bacterial CFU was measured before gentamicin treatment (3 hrs). Data represents geometric mean ± s.d. Dots indicate individual biological replicate (N=3). N.S.; not significant. (c) Culture supernatants were harvested, and cytokines were measured by ELISA. Data represents mean ± s.d. Dots indicate individual biological replicate (N=3). N.S.; not significant, ***; P < 0.001, ****; P < 0.0001 by 1-Way ANOVA followed by Bonferroni post-hoc test. (d) T84 intestinal epithelial cells (2 x 105 cells/well/24 well plate in 500 μl) were grown for 3 weeks and infected with each E. coli strain (2 x 106 cells/well). After 3 hrs, cells were washed, and extracellular bacteria were killed by gentamycin (100 μg/ml). Intracellular bacteria were then quantified by culture on LB agar plates. The percentage of intracellular bacteria was calculated. Data represents mean ± s.d. Dots indicate individual biological replicate (N=3). N.S.; not significant, **; P < 0.01 by 1-Way ANOVA followed by Bonferroni post-hoc test.
Extended Data Fig. 4 LF82 but not MG1655 employs L-serine for its growth.
(a) AIEC LF82 and commensal E. coli MG1655 were cultured in DMEM (0.45% glucose) or in a minimal medium (0.1% glucose) supplemented with a single L-amino acid (final concentration was 1mM) for 8 h at 37 °C in 20% O2 and 5% CO2. Bacterial proliferation (O.D600) is shown. Data represent mean ± s.d. (N=2, technical replicates). Results are representative of 2 biologically independent experiments. SPF C57BL/6 mice were pretreated with streptomycin (800mg/kg, p.o.) 1 day prior to the treatment with 3.0% DSS. Control (Ctrl) and DSS-treated mice (day 5 post 3.0% treatment) were then co-inoculated with MG1655 WT and MG1655 ΔtdcΔsda (ΔTS) mutant (1 x 109 CFU each/mouse). Fecal samples were collected 24 hrs post E. coli inoculation. Bacterial CFUs in feces (b) and the competitive index (WT/ΔTS) (c) are shown. Bars represent geometric mean ± s.d. Dots indicate individual mice (N=5, biologically independent animals). N.S.; not significant by Mann-Whitney U test (two-sided) or 1-Way ANOVA followed by Bonferroni post-hoc test.
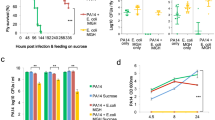
Extended Data Fig. 5 Severity of DSS colitis in gnotobiotic mice.
Germ-free (GF) C57BL/6 mice (N=4, biologically independent animals) were mono-colonized either with MG1655 or LF82, or co-colonized with those two strains (1 x 109 CFU each/mouse) for 10 days. On day 10, colitis was induced by 1.5% DSS (for 5 days). Body weight change at day 15 (% of initial (day 10)) (a) and disease activity index (DAI) (day 10 and day 15) (b) are shown. Data represent mean ± s.e.m. Dots indicate individual mice. **; P < 0.01, **** P < 0.0001 by Mann-Whitney U test (two-sided).
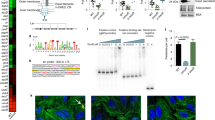
Extended Data Fig. 6 Intestinal inflammation alters the luminal metabolome.
SPF C57BL/6 mice (N=5, biologically independent animals) were treated with 1.5% DSS for 5 days. Cecal samples were harvested and luminal metabolic profiles were analyzed by capillary electrophoresis time-of-flight mass spectrometry (CE- TOF/MS). (a) A heatmap of the quantified luminal metabolites. The concentrations of metabolites were transformed into Z-scores and clustered according to their Euclidean distance. Gray areas in the heatmap indicate that respective metabolites were not detected. (b) Principal component analysis (PCA) of the luminal metabolome data. The ellipse denotes the 95% significance limit of the model, as defined by Hotelling’s t-test. (c) A loading scatter plot of the PCA. (d) The bar graphs showing the selected metabolites whose concentrations were altered significantly during DSS colitis. Data are presented as mean ± s.d. Dots represents individual mice (N=5, biologically independent animals). N.D.; not detected, *; P < 0.05, **; P < 0.01 by Mann–Whitney U test (two-sided).
Extended Data Fig. 7 Effect of dietary amino acid modification on mice.
(a, b) SPF C57BL/6 mice were fed a control amino acid defined diet (Ctrl), protein-free diet (PFD), L-serine-L-glycine-deficient diet (SDD), or L-aspartic acid-deficient diet (DDD) for 7 days. Food consumption (a) and body weight change (b) were monitored at indicated time points. Four individual cages were used for each diet. Each cage contains 2-5 biologically independent mice. Food consumption amount per mouse in each cage was calculated. Data represent mean ± s.e.m. (N=4, biologically independent experiments). N.S.; not significant, *; P < 0.05, *** P < 0.001, **** P < 0.0001 by two-way ANOVA followed by Bonferroni post-hoc test (Ctrl vs PFD). (c) SPF C57BL/6 mice were fed the Ctrl, PFD, SDD, or DDD for 3 days. On day 3, fecal samples were collected from each mouse. Capillary electrophoresis time-of-flight mass spectrometry (CE-TOF/MS) was used to measure the concentration of luminal L-amino acids. Data represent mean ± s.e.m. Dots indicate individual mice (N=5-6, biologically independent animals). N.S.; not significant, *; P < 0.05, **; P < 0.01, ***; P < 0.001, **** P < 0.0001 by one-way ANOVA followed by Bonferroni post-hoc test or Mann-Whitney U test (two-sided).
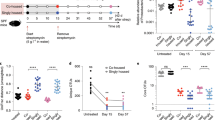
Extended Data Fig. 8 Dietary L-serine regulates intraspecific competition between E. coli in the inflamed gut.
(a) Germ-free (GF) C57BL/6 mice were co-inoculated with LF82 and MG1655 for 7 days. All mice were fed the control amino acid-defined (Ctrl) diet during this period. On day 7, the diet was switched to protein-free diet (PFD) or L-serine- deficient diet (SDD). The control group stayed on the ctrl diet. Three days after switching diets, colitis was induced by 1.5% DSS (5-day treatment). (Left) Bacterial CFUs and the (Right) competitive index of LF82/MG1655 were analyzed at indicated time points. Data are represented as geometric mean ± s.d. (N=5-8, biologically independent animals). *; P<0.05, ****; P< 0.0001: 2-Way ANOVA followed by Bonferroni post-hoc test (Left: Ctrl diet + MG1655 vs. SDD + MG1655, Right: Ctrl Diet vs SDD). (b) GF C57BL/6 mice were fed Ctrl diet or SDD. After three days, the mice were co- inoculated with LF82 and MG1655. Bacterial CFUs and the competitive index of LF82/MG1655 were analyzed at indicated time points. Data are represented as geometric mean ± s.d. (N=5, biologically independent animals).
Extended Data Fig. 9 Deprivation of dietary L- serine does not influence host anti- microbial immunity.
SPF C57BL/6 mice (N=5, biologically independent animals) were fed a control amino acid-defined diet (Ctrl), protein-free diet (PFD), or L-serine and L-glycine-deficient diet (SDD) for 14 days. Colonic mucosa was isolated at day 14 and the expression of host anti-microbial genes was analyzed by qPCR. Data are represented as mean ± s.d. Dots indicate individual mice. N.S.; not significant, by 1-Way ANOVA followed by Bonferroni post-hoc test.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Tables 3 and 4.
Supplementary Tables 1 and 2
Gene expression profile of E. coli LF82 in the inflamed gut. Supplementary Table 2: luminal metabolome changes during intestinal inflammation.
Source data
Source Data Fig. 1
Statistical Source Data to Fig. 1.
Source Data Fig. 2
Statistical Source Data to Fig. 2.
Source Data Fig. 3
Statistical Source Data to Fig. 3.
Source Data Fig. 4
Statistical Source Data to Fig. 4.
Source Data Fig. 5
Statistical Source Data to Fig. 5.
Source Data Extended Data Fig. 1
Statistical Source Data to Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Statistical Source Data to Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Statistical Source Data to Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Statistical Source Data to Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Statistical Source Data to Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Statistical Source Data to Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Statistical Source Data to Extended Data Fig. 7.
Source Data Extended Data Fig. 8
Statistical Source Data to Extended Data Fig. 8.
Source Data Extended Data Fig. 9
Statistical Source Data to Extended Data Fig. 9.
Rights and permissions
About this article
Cite this article
Kitamoto, S., Alteri, C.J., Rodrigues, M. et al. Dietary l-serine confers a competitive fitness advantage to Enterobacteriaceae in the inflamed gut. Nat Microbiol 5, 116–125 (2020). https://doi.org/10.1038/s41564-019-0591-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0591-6
This article is cited by
-
Metabolic network of the gut microbiota in inflammatory bowel disease
Inflammation and Regeneration (2024)
-
A consortia of clinical E. coli strains with distinct in vitro adherent/invasive properties establish their own co-colonization niche and shape the intestinal microbiota in inflammation-susceptible mice
Microbiome (2023)
-
Effect of environmental variance-based resilience selection on the gut metabolome of rabbits
Genetics Selection Evolution (2023)
-
Genetic evidence strengthens the bidirectional connection between gut microbiota and periodontitis: insights from a two-sample Mendelian randomization study
Journal of Translational Medicine (2023)
-
The oral-gut axis: a missing piece in the IBD puzzle
Inflammation and Regeneration (2023)