Abstract
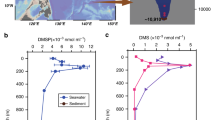
Dimethylsulfoniopropionate (DMSP) and its catabolite dimethyl sulfide (DMS) are key marine nutrients1,2 that have roles in global sulfur cycling2, atmospheric chemistry3, signalling4,5 and, potentially, climate regulation6,7. The production of DMSP was previously thought to be an oxic and photic process that is mainly confined to the surface oceans. However, here we show that DMSP concentrations and/or rates of DMSP and DMS synthesis are higher in surface sediment from, for example, saltmarsh ponds, estuaries and the deep ocean than in the overlying seawater. A quarter of bacterial strains isolated from saltmarsh sediment produced DMSP (up to 73 mM), and we identified several previously unknown producers of DMSP. Most DMSP-producing isolates contained dsyB8, but some alphaproteobacteria, gammaproteobacteria and actinobacteria used a methionine methylation pathway independent of DsyB that was previously only associated with higher plants. These bacteria contained a methionine methyltransferase gene (mmtN)—a marker for bacterial synthesis of DMSP through this pathway. DMSP-producing bacteria and their dsyB and/or mmtN transcripts were present in all of the tested seawater samples and Tara Oceans bacterioplankton datasets, but were much more abundant in marine surface sediment. Approximately 1 × 108 bacteria g−1 of surface marine sediment are predicted to produce DMSP, and their contribution to this process should be included in future models of global DMSP production. We propose that coastal and marine sediments, which cover a large part of the Earth’s surface, are environments with high levels of DMSP and DMS productivity, and that bacteria are important producers of DMSP and DMS within these environments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The 16S rRNA gene amplicon sequencing, metagenomic data and whole-genome sequences generated in this study are publicly available from the NCBI Sequence Read Archive (BioProject: PRJNA522699).
References
Curson, A. R. J., Todd, J. D., Sullivan, M. J. & Johnston, A. W. B. Catabolism of dimethylsulphoniopropionate: microorganisms, enzymes and genes. Nat. Rev. Microbiol. 9, 849–859 (2011).
Sievert, S. M., Kiene, R. P. & Schulz-Vogt, H. N. The sulfur cycle. Oceanography 20, 117–123 (2007).
Andreae, M. O. Ocean-atmosphere interactions in the global biogeochemical sulfur cycle. Mar. Chem. 30, 1–29 (1990).
Seymour, J. R., Simó, R., Ahmed, T. & Stocker, R. Chemoattraction to dimethylsulfoniopropionate throughout the marine microbial food web. Science 329, 342–345 (2010).
Nevitt, G. A. The neuroecology of dimethyl sulfide: a global-climate regulator turned marine infochemical. Integr. Comp. Biol. 51, 819–825 (2011).
Stefels, J., Steinke, M., Turner, S., Malin, G. & Belviso, S. Environmental constraints on the production and removal of the climatically active gas dimethylsulphide (DMS) and implications for ecosystem modelling. Biogeochemistry 83, 245–275 (2007).
Vallina, S. M. & Simó, R. Strong relationship between DMS and the solar radiation dose over the global surface ocean. Science 315, 506–508 (2007).
Curson, A. R. J. et al. Dimethylsulfoniopropionate biosynthesis in marine bacteria and identification of the key gene in this process. Nat. Microbiol. 2, 17009 (2017).
Galí, M., Devred, E., Levasseur, M., Royer, S. J. & Babin, M. A remote sensing algorithm for planktonic dimethylsulfoniopropionate (DMSP) and an analysis of global patterns. Remote Sens. Environ. 171, 171–184 (2015).
van Bergeijk, S. A., Schönefeldt, K., Stal, L. J. & Huisman, J. Production and consumption of dimethylsulfide (DMS) and dimethylsulfoniopropionate (DMSP) in a diatom-dominated intertidal sediment. Mar. Ecol. Prog. Ser. 231, 37–46 (2002).
Zhuang, G. C. et al. Distribution and isotopic composition of trimethylamine, dimethylsulfide and dimethylsulfoniopropionate in marine sediments. Mar. Chem. 196, 35–46 (2017).
Dacey, J. W. H., King, G. M. & Wakeham, S. G. Factors controlling emission of dimethylsulfide from salt marshes. Nature 330, 643–645 (1987).
Steudler, P. A. & Peterson, B. J. Contribution of gaseous sulphur from salt marshes to the global sulphur cycle. Nature 311, 455–457 (1984).
Mincer, T. J., Jensen, P. R., Kauffman, C. A. & Fenical, W. Widespread and persistent populations of a major new marine actinomycete taxon in ocean sediments. Appl. Environ. Microbiol. 68, 5005–5011 (2002).
Speeckaert, G., Borges, A. V., Champenois, W., Royer, C. & Gypens, N. Annual cycle of dimethylsulfoniopropionate (DMSP) and dimethylsulfoxide (DMSO) related to phytoplankton succession in the Southern North Sea. Sci. Total Environ. 622–623, 362–372 (2018).
Curson, A. et al. DSYB catalyses the key step of dimethylsulfoniopropionate biosynthesis in many phytoplankton. Nat. Microbiol. 3, 430–439 (2018).
Dickschat, J. S., Rabe, P. & Citron, C. A. The chemical biology of dimethylsulfoniopropionate. Org. Biomol. Chem. 13, 1954–1968 (2015).
Hanson, A. D., Rivoal, J., Paquet, L. & Cage, D. A. Biosynthesis of 3-dimethylsulfoniopropionate in Wollastonia biflora (L.) DC. Evidence that S-methylmethionine is an intermediate. Plant Physiol. 105, 103–110 (1994).
Ranocha, P. et al. Characterization and functional expression of cDNAs encoding methionine-sensitive and -insensitive homocysteine S-methyltransferases from Arabidopsis. J. Biol. Chem. 275, 15962–15968 (2000).
Kocsis, M. G. et al. Dimethylsulfoniopropionate biosynthesis in Spartina alterniflora. Plant Physiol. 117, 273–281 (1998).
Shapiro, S. K. Biosynthesis of methionine from homocysteine and S-methylmethionine in bacteria. J. Bacteriol. 72, 730–735 (1956).
Albers, E. Metabolic characteristics and importance of the universal methionine salvage pathway recycling methionine from 5′‐methylthioadenosine. IUBMB Life 61, 1132–1142 (2009).
Liao, C. & Seebeck, F. P. In vitro reconstitution of bacterial DMSP biosynthesis. Angew. Chem. Int. Ed. 58, 3591–3594 (2019).
Bourgis, F. et al. S-methylmethionine plays a major role in phloem sulfur transport and is synthesized by a novel type of methyltransferase. Plant Cell 11, 1485–1497 (1999).
Sunagawa, S. et al. Structure and function of the global ocean microbiome. Science 348, 1261359 (2015).
Hobbie, J. E., Daley, R. J. & Jasper, S. Use of nuclepore filters for counting bacteria by fluorescence microscopy. Appl. Environ. Microbiol. 33, 1225–1228 (1977).
Todd, J. D. et al. Molecular dissection of bacterial acrylate catabolism—unexpected links with dimethylsulfoniopropionate catabolism and dimethyl sulfide production. Environ. Microbiol. 12, 327–343 (2010).
Caddick, S. E., Harrison, C. J., Stavridou, I., Johnson, S. & Brearley, C. A. A lysine accumulation phenotype of ScIpk2Δ mutant yeast is rescued by Solanum tuberosum inositol phosphate multikinase. Biochem. J. 403, 381–389 (2007).
Franklin, D. J., Steinke, M., Young, J., Probert, I. & Malin, G. Dimethylsulphoniopropionate (DMSP), DMSP-lyase activity (DLA) and dimethylsulphide (DMS) in 10 species of coccolithophore. Mar. Ecol. Prog. Ser. 410, 13–23 (2010).
Zhang, S.-H., Yang, G.-P., Zhang, H.-H. & Yang, J. Spatial variation of biogenic sulfur in the south Yellow Sea and the East China Sea during summer and its contribution to atmospheric sulfate aerosol. Sci. Total Environ. 488, 157–167 (2014).
Murphy, J. & Riley, J. P. A modified single solution method for the determination of phosphate in natural waters. Anal. Chim. Acta 27, 31–36 (1962).
Jones, R. D. An improved fluorescence method for the determination of nanomolar concentrations of ammonium in natural waters. Limnol. Oceanogr. Methods 36, 814–819 (1991).
Armstrong, F. A. J., Stearns, C. R. & Strickland, J. D. H. The measurement of upwelling and subsequent biological process by means of the technicon autoanalyzer and associated equipment. Deep Sea Res. 14, 381–389 (1967).
Slezak, D., Kiene, R. P., Toole, D. A., Simó, R. & Kieber, D. J. Effects of solar radiation on the fate of dissolved DMSP and conversion to DMS in seawater. Aquat. Sci. 69, 377–393 (2007).
Boyd, T. J. et al. in Microbial Ecology Research Trends (ed. Van Dijk, T.) Chap. 1 (Nova Science, 2008).
Morris, J. T. et al. Contributions of organic and inorganic matter to sediment volume and accretion in tidal wetlands at steady state. Earth’s Future 4, 110–121 (2016).
Guillard, R. R. L. & Ryther, J. H. Studies of marine planktonic diatoms: I. Cyclotella nana Hustedt and Detonula confervacea (Cleve) Gran. Can. J. Microbiol. 8, 229–239 (1962).
Andersen, R. A. & Kawachi, M. in Algal Culturing Techniques (ed. Anderson, R. A.) 83–101 (Elsevier Acad. Press, 2005).
Naz, T., Burhan, Z., Munir, S. & Siddiqui, P. J. A. Biovolume and biomass of common diatom species from the coastal waters of Karachi, Pakistan. Pak. J. Bot. 45, 325–328 (2013).
Yin, Q. et al. Spatial variations in microbial community composition in surface seawater from the ultra-oligotrophic center to rim of the South Pacific Gyre. PLoS ONE 8, e55148 (2013).
Carrión, O. et al. Methanethiol-dependent dimethylsulfide production in soil environments. ISME J. 11, 2379 (2017).
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004).
Rose, T. M., Henikoff, J. G. & Henikoff, S. CODEHOP (COnsensus-DEgenerate hybrid oligonucleotide primer) PCR primer design. Nucleic Acids Res. 31, 3763–3766 (2003).
Lane, D. J. et al. Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc. Natl Acad. Sci. USA 82, 6955–6959 (1985).
Farhan Ul Haque, M. et al. Correlated production and consumption of chloromethane in the Arabidopsis thaliana phyllosphere. Sci. Rep. 7, 17589 (2017).
Sun, D. L., Jiang, X., Wu, Q. L. & Zhou, N. Y. Intragenomic heterogeneity of 16S rRNA genes causes overestimation of prokaryotic diversity. Appl. Environ. Microbiol. 79, 5962–5969 (2013).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Trifinopoulos, J., Nguyen, L. T., von Haeseler, A. & Minh, B. Q. W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 44, 232–235 (2016).
Minh, B. Q., Nguyen, M. A. T. & von Haeseler, A. Ultrafast approximation for phylogenetic bootstrap. Mol. Biol. Evol. 30, 1188–1195 (2013).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T. Y. Ggtree: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2008).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Wickham H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2010).
Kassambara A. ggpubR: ‘ggplot2’ Based Publication Ready Plots. R package version 2.2.1 (2018); https://github.com/kassambara/ggpubr
Wilke C. O. cowplot: Streamlined Plot Theme and Plot Annotations for ‘ggplot2’. R package version 0.9.2 (2017); https://CRAN.R-project.org/package=cowplot
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
González, J. M., Whitman, W. B., Hodson, R. E. & Moran, M. A. Identifying numerically abundant culturable bacteria from complex communities: an example from a lignin enrichment culture. Appl. Environ. Microbiol. 62, 4433–4440 (1996).
Baumann, P. & Baumann, L. in The Prokaryotes: A Handbook on Habitats, Isolation and Identification of Bacteria (eds Starr, M. P. et al.) 1302–1331 (Springer, 1981).
Sambrook, J., Fritsch, E., Maniatis, T. & Nolan, C. Molecular Cloning: a Laboratory Manual (Cold Spring Harbor Laboratory Press, 1989).
Beringer, J. E. R. Factor transfer in Rhizobium leguminosarum. J. Gen. Microbiol. 84, 188–198 (1974).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Li H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Aziz, R. K. et al. The RAST server: rapid annotations using subsystems technology. BMC Genom. 9, 75 (2008).
Figurski, D. H. & Helinski, D. R. Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc. Natl Acad. Sci. USA 76, 1648–1652 (1979).
Carrión, O. et al. A novel pathway producing dimethylsulphide in bacteria is widespread in soil environments. Nat. Commun. 6, 6579 (2015).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Todd, J. D. et al. DddQ, a novel, cupin-containing, dimethylsulfoniopropionate lyase in marine roseobacters and in uncultured marine bacteria. Environ. Microbiol. 13, 427–438 (2011).
Schäfer, A. et al. Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 145, 69–73 (1994).
Acknowledgements
Funding from the Natural Environmental Research Council (NE/N002385, NE/P012671 and NE/S001352) supported work in J.D.T.’s laboratory. Funding from the National Natural Science Foundation of China (grant numbers 91751202 and 41730530) supported the research in X.-H.Z.’s laboratory. B.T.W. was supported by a NERC EnvEast grant (NE/L002582/1). A.B.M. and K.C. were supported by the BBSRC Norwich Research Park Biosciences Doctoral Training Partnership grant number BB/M011216/1. J.P. was supported by a NERC Independent Research Fellowship (NE/L010771/2). We thank P. Wells for general technical support, and P. Nelson and R. Whiting at the CEFAS, Lowestoft for sediment nutrient analysis. We acknowledge the Tara Oceans Consortium for providing metagenomic sequence data. We also thank the late R. Kiene, whose work on DMSP was an inspiration for this study.
Author information
Authors and Affiliations
Contributions
J.D.T. wrote the paper, designed all of the experiments and performed experiments. B.T.W. wrote the paper, designed all of the experiments and performed or contributed to all of the experiments and prepared figures and tables. K.C. performed experiments (genomic library screening, mutant complementation and characterization of MMT+ bacteria). A.B.M. performed experiments (LC–MS work). A.R.J.C. performed experiments (genomic library construction, MMT assays, mutant construction and rate experiments). Y.Z. performed experiments (qPCR, degenerate primer design, sampling and DMSP quantification in the Mariana Trench). Jingli Liu and Ji Liu performed experiments (seawater incubations, qPCR, sediment sampling, purge-trap analysis, DNA/RNA purification from water). S.N.-P., M.P. and C.-Y.L. designed and performed experiments (MmtN protein characterization). P.P.L.R. performed experiments (DMSP quantification in sediment, isolation and characterization of eukaryotic species). L.G.S. wrote the paper and performed experiments (evolutionary analysis of MmtN sequences and phylogenetic tree construction). C.A.B. devised experiments for measuring DMSP pathway intermediates in sediment and cell lysate by HPLC, carried out LC–MS experiments and discussed results. B.W.M. performed experiments (16S rRNA amplicon sequencing analysis) and prepared figures. B.J.P. performed experiments (cell lysate assays); J.P. performed experiments (degenerate primer design, sediment sampling and bioinformatics analysis of metagenomic sequencing). O.C., X.-H.Z., Y.-Z.Z. and J.C.M. designed experiments and discussed results.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–18, Supplementary Tables 1–3, Supplementary Tables 5 and 6, Supplementary Tables 9–11, Supplementary Tables 13–17, Supplementary references.
Supplementary Table 4
Species shown to produce DMSP, including those containing dsyB, mmtN or unknown DMSP-synthesis genes, as of June 2018.
Supplementary Table 7
Metagenome information and results of mmtN/dsyB metagenomic searches in OM-RGC and Stiffkey metagenomes.
Supplementary Table 8
DMSP production in selected strains of bacteria and activity of the corresponding cloned mmtN genes.
Supplementary Table 12
Tara Oceans metatranscriptome mmtN transcript abundance, alongside dsyB, DSYB and DMSP lyases calculated in ref. 1.
Rights and permissions
About this article
Cite this article
Williams, B.T., Cowles, K., Bermejo Martínez, A. et al. Bacteria are important dimethylsulfoniopropionate producers in coastal sediments. Nat Microbiol 4, 1815–1825 (2019). https://doi.org/10.1038/s41564-019-0527-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0527-1
This article is cited by
-
Methanolobus use unspecific methyltransferases to produce methane from dimethylsulphide in Baltic Sea sediments
Microbiome (2024)
-
Optimized methyl donor and reduced precursor degradation pathway for seleno-methylselenocysteine production in Bacillus subtilis
Microbial Cell Factories (2023)
-
Ubiquitous occurrence of a dimethylsulfoniopropionate ABC transporter in abundant marine bacteria
The ISME Journal (2023)
-
Microbial drivers of DMSO reduction and DMS-dependent methanogenesis in saltmarsh sediments
The ISME Journal (2023)
-
The biogeochemistry of marine dimethylsulfide
Nature Reviews Earth & Environment (2023)