Abstract
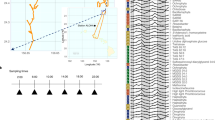
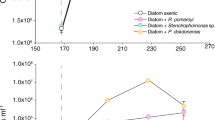
In the surface ocean, phytoplankton transform inorganic substrates into organic matter that fuels the activity of heterotrophic microorganisms, creating intricate metabolic networks that determine the extent of carbon recycling and storage in the ocean. Yet, the diversity of organic molecules and interacting organisms has hindered detection of specific relationships that mediate this large flux of energy and matter. Here, we show that a tightly coupled microbial network based on organic sulfur compounds (sulfonates) exists among key lineages of eukaryotic phytoplankton producers and heterotrophic bacterial consumers in the North Pacific Subtropical Gyre. We find that cultured eukaryotic phytoplankton taxa produce sulfonates, often at millimolar internal concentrations. These same phytoplankton-derived sulfonates support growth requirements of an open-ocean isolate of the SAR11 clade, the most abundant group of marine heterotrophic bacteria. Expression of putative sulfonate biosynthesis genes and sulfonate abundances in natural plankton communities over the diel cycle link sulfonate production to light availability. Contemporaneous expression of sulfonate catabolism genes in heterotrophic bacteria highlights active cycling of sulfonates in situ. Our study provides evidence that sulfonates serve as an ecologically important currency for nutrient and energy exchange between microbial autotrophs and heterotrophs, highlighting the importance of organic sulfur compounds in regulating ecosystem function.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
KM1513 cruise information and associated data for the HOE Legacy II cruise can be found online at http://hahana.soest.hawaii.edu/hoelegacy/hoelegacy.html. Raw sequence data for the diel eukaryotic metatranscriptomes are available in the NCBI Sequence Read Archive under BioProject ID PRJNA492142. Raw sequence data for the prokaryotic metatranscriptomes are available under BioProject ID PRJNA492143. Metabolomics data are available in Metabolomics Workbench under project ID PR000797.
Code availability
The custom code used is available on GitHub at https://github.com/IngallsLabUW and https://github.com/armbrustlab.
References
Field, C. B. Primary production of the biosphere: integrating terrestrial and oceanic components. Science 281, 237–240 (1998).
Azam, F. & Malfatti, F. Microbial structuring of marine ecosystems. Nat. Rev. Microbiol. 5, 782–791 (2007).
Ksionzek, K. B. et al. Dissolved organic sulfur in the ocean: biogeochemistry of a petagram inventory. Science 354, 456–459 (2016).
Levine, N. M. Putting the spotlight on organic sulfur: diverse dissolved organic sulfur compounds play an active role in ocean biogeochemistry. Science 354, 418–419 (2016).
Tripp, H. J. et al. SAR11 marine bacteria require exogenous reduced sulphur for growth. Nature 452, 741–744 (2008).
Dupont, C. L. et al. Genomic insights to SAR86, an abundant and uncultivated marine bacterial lineage. ISME J. 6, 1186–1199 (2012).
Kiene, R. P., Linn, L. J., González, J., Moran, M. A. & Bruton, J. A. Dimethylsulfoniopropionate and methanethiol are important precursors of methionine and protein-sulfur in marine bacterioplankton. Appl. Environ. Microbiol. 65, 4549–4558 (1999).
Autry, A. R. & Fitzgerald, J. W. Sulfonate S: a major form of forest soil organic sulfur. Biol. Fertil. Soils 10, 50–56 (1990).
Cook, A. M., Denger, K. & Smits, T. H. M. Dissimilation of C3-sulfonates. Arch. Microbiol. 185, 83–90 (2006).
Cook, A. M. & Denger, K. Dissimilation of the C2 sulfonates. Arch. Microbiol. 179, 1–6 (2002).
Denger, K. et al. Sulphoglycolysis in Escherichia coli K-12 closes a gap in the biogeochemical sulphur cycle. Nature 507, 114–117 (2014).
Denger, K. & Cook, A. M. Racemase activity effected by two dehydrogenases in sulfolactate degradation by Chromohalobacter salexigens: purification of (S)-sulfolactate dehydrogenase. Microbiology 156, 967–974 (2010).
Mayer, J. et al. 2,3-Dihydroxypropane-1-sulfonate degraded by Cupriavidus pinatubonensis JMP134: purification of dihydroxypropanesulfonate 3-dehydrogenase. Microbiology 156, 1556–1564 (2010).
Warshan, D. et al. Feathermoss and epiphytic Nostoc cooperate differently: expanding the spectrum of plant–cyanobacteria symbiosis. ISME J. 11, 2821–2833 (2017).
Weinitschke, S., Sharma, P. I., Stingl, U., Cook, A. M. & Smits, T. H. M. Gene clusters involved in isethionate degradation by terrestrial and marine bacteria. Appl. Environ. Microbiol. 76, 618–621 (2010).
Denger, K. et al. Bifurcated degradative pathway of 3-sulfolactate in Roseovarius nubinhibens ISM via sulfoacetaldehyde acetyltransferase and (S)-cysteate sulfolyase. J. Bacteriol. 191, 5648–5656 (2009).
Denger, K., Smits, T. H. M. & Cook, A. M. l-Cysteate sulpho-lyase, a widespread pyridoxal 5′-phosphate-coupled desulphonative enzyme purified from Silicibacter pomeroyi DSS-3T. Biochem. J. 394, 657–664 (2006).
Landa, M., Burns, A. S., Roth, S. J. & Moran, M. A. Bacterial transcriptome remodeling during sequential co-culture with a marine dinoflagellate and diatom. ISME J. 11, 2677–2690 (2017).
Durham, B. P. et al. Cryptic carbon and sulfur cycling between surface ocean plankton. Proc. Natl Acad. Sci. USA 112, 453–457 (2015).
Durham, B. P. et al. Recognition cascade and metabolite transfer in a marine bacteria–phytoplankton model system. Environ. Microbiol. 19, 3500–3513 (2017).
Amin, S. A. et al. Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature 522, 98–101 (2015).
Celik, E. et al. Metabolism of 2,3-dihydroxypropane-1-sulfonate by marine bacteria. Org. Biomol. Chem. 15, 2919–2922 (2017).
Jackson, A. E., Ayer, S. W. & Laycock, M. V. The effect of salinity on growth and amino acid composition in the marine diatom Nitzschia pungens. Can. J. Bot. 70, 2198–2201 (1992).
Boroujerdi, A. F. B. et al. Identification of isethionic acid and other small molecule metabolites of Fragilariopsis cylindrus with nuclear magnetic resonance. Anal. Bioanal. Chem. 404, 777–784 (2012).
Graham, D. E., Taylor, S. M., Wolf, R. Z. & Namboori, S. C. Convergent evolution of coenzyme M biosynthesis in the Methanosarcinales: cysteate synthase evolved from an ancestral threonine synthase. Biochem. J. 424, 467–478 (2009).
Götz, F. et al. Targeted metabolomics reveals proline as a major osmolyte in the chemolithoautotroph Sulfurimonas denitrificans. MicrobiologyOpen 7, e00586 (2018).
Ho, T. et al. The elemental composition of some marine phytoplankton. J. Phycol. 39, 1145–1159 (2003).
Hunter, J. E. et al. Lipidomics of Thalassiosira pseudonana under phosphorus stress reveal underlying phospholipid substitution dynamics and novel diglycosylceramide substitutes. Appl. Environ. Microbiol. 84, e02034-17 (2018).
Helgadóttir, S., Rosas-Sandoval, G., Söll, D. & Graham, D. E. Biosynthesis of phosphoserine in the Methanococcales. J. Bacteriol. 189, 575–582 (2007).
Tevatia, R. et al. The taurine biosynthetic pathway of microalgae. Algal Res. 9, 21–26 (2015).
Agnello, G., Chang, L. L., Lamb, C. M., Georgiou, G. & Stone, E. M. Discovery of a substrate selectivity motif in amino acid decarboxylases unveils a taurine biosynthesis pathway in prokaryotes. ACS Chem. Biol. 8, 2264–2271 (2013).
Krejcik, Z., Hollemeyer, K., Smits, T. H. M. & Cook, A. M. Isethionate formation from taurine in Chromohalobacter salexigens: purification of sulfoacetaldehyde reductase. Microbiology 156, 1547–1555 (2010).
Felux, A.-K., Spiteller, D., Klebensberger, J. & Schleheck, D. Entner–Doudoroff pathway for sulfoquinovose degradation in Pseudomonas putida SQ1. Proc. Natl Acad. Sci. USA 112, E4298–E4305 (2015).
Busby, W. F. & Benson, A. A. Sulfonic acid metabolism in the diatom Navicula pelliculosa. Plant Cell Physiol. 14, 1123–1132 (1973).
Busby, W. Studies on the Identification and Metabolism of the Sulfonic Acids, Cysteinolic Acid and Sulfopropanediol, in the Diatom Navicula pelliculosa, and their Distribution in the Major Algal Groups. PhD dissertation, University of California San Diego (1966).
Graupner, M., Xu, H. & White, R. H. Identification of an archaeal 2-hydroxy acid dehydrogenase catalyzing reactions involved in coenzyme biosynthesis in methanoarchaea. J. Bacteriol. 182, 3688–3692 (2000).
Kopriva, S. et al. Light regulation of assimilatory sulphate reduction in Arabidopsis thaliana. Plant J. 20, 37–44 (1999).
Wilson, S. T. et al. Coordinated regulation of growth, activity and transcription in natural populations of the unicellular nitrogen-fixing cyanobacterium Crocosphaera. Nat. Microbiol. 2, 17118 (2017).
Bates, T. S. et al. The cycling of sulfur in surface seawater of the northeast Pacific. J. Geophys. Res. 99, 7835–7843 (1994).
Johnson, W. M., Kido Soule, M. C. & Kujawinski, E. B. Extraction efficiency and quantification of dissolved metabolites in targeted marine metabolomics. Limnol. Oceanogr. Methods 15, 417–428 (2017).
McCarren, J. et al. Microbial community transcriptomes reveal microbes and metabolic pathways associated with dissolved organic matter turnover in the sea. Proc. Natl Acad. Sci. USA 107, 16420–16427 (2010).
Fuhrman, J. A. & Hagstrom, A. in Microbial Ecology of the Oceans (ed. Kirchman, D. L.) 45–90 (John Wiley & Sons, 2008).
Carini, P., Steindler, L., Beszteri, S. & Giovannoni, S. J. Nutrient requirements for growth of the extreme oligotroph ‘Candidatus Pelagibacter ubique’ HTCC1062 on a defined medium. ISME J. 7, 592–602 (2013).
Schwalbach, M. S., Tripp, H. J., Steindler, L., Smith, D. P. & Giovannoni, S. J. The presence of the glycolysis operon in SAR11 genomes is positively correlated with ocean productivity. Environ. Microbiol. 12, 490–500 (2010).
Tripp, H. J. et al. Unique glycine-activated riboswitch linked to glycine-serine auxotrophy in SAR11. Environ. Microbiol. 11, 230–238 (2009).
Reisch, C. R., Moran, M. A. & Whitman, W. B. Bacterial catabolism of dimethylsulfoniopropionate (DMSP). Front. Microbiol. 2, 172 (2011).
Sun, J. et al. The abundant marine bacterium Pelagibacter simultaneously catabolizes dimethylsulfoniopropionate to the gases dimethyl sulfide and methanethiol. Nat. Microbiol. 1, 16065 (2016).
Seymour, J. R., Amin, S. A., Raina, J.-B. & Stocker, R. Zooming in on the phycosphere: the ecological interface for phytoplankton–bacteria relationships. Nat. Microbiol. 2, 17065 (2017).
Luo, H., Csuros, M., Hughes, A. L. & Moran, M. A. Evolution of divergent life history strategies in marine alphaproteobacteria. mBio 4, e00373-13 (2013).
Giovannoni, S. J. SAR11 bacteria: the most abundant plankton in the oceans. Ann. Rev. Mar. Sci. 9, 231–255 (2017).
Boysen, A. K., Heal, K. R., Carlson, L. T. & Ingalls, A. E. Best-matched internal standard normalization in liquid chromatography–mass spectrometry metabolomics applied to environmental samples. Anal. Chem. 90, 1363–1369 (2018).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Smith, C. A., Want, E. J., O’Maille, G., Abagyan, R. & Siuzdak, G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Boiteau, R. M. & Repeta, D. J. An extended siderophore suite from Synechococcus sp. PCC 7002 revealed by LC-ICPMS-ESIMS. Metallomics 7, 877–884 (2015).
Tsugawa, H. et al. MS-DIAL: data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat. Methods 12, 523–526 (2015).
Patiny, L. & Borel, A. ChemCalc: a building block for tomorrow’s chemical infrastructure. J. Chem. Inf. Model. 53, 1223–1228 (2013).
Satinsky, B. M., Gifford, S. M., Crump, B. C. & Moran, M. A. Use of internal standards for quantitative metatranscriptome and metagenome analysis. Methods Enzymol. 531, 237–250 (2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Magoč, T., Magoč, M. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet. 16, 276–277 (2000).
Keeling, P. J. et al. The Marine Microbial Eukaryote Transcriptome Sequencing Project (MMETSP): illuminating the functional diversity of eukaryotic life in the oceans through transcriptome sequencing. PLoS Biol. 12, e1001889 (2014).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–1165 (2011).
Stamatakis, A., Hoover, P. & Rougemont, J. A rapid bootstrap algorithm for the RAxML web servers. Syst. Biol. 57, 758–771 (2008).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Matsen, F. A., Kodner, R. B. & Armbrust, V. pplacer: linear time maximum-likelihood and Bayesian phylogenetic placement of sequences onto a fixed reference tree. BMC Bioinformatics 11, 538 (2010).
Thaben, P. F. & Westermark, P. O. Detecting rhythms in time series with RAIN. J. Biol. Rhythms 29, 391–400 (2014).
Marshall, K. T. & Morris, R. M. Isolation of an aerobic sulfur oxidizer from the SUP05/Arctic96BD-19 clade. ISME J. 7, 452–455 (2013).
Acknowledgements
The data sets presented here resulted from the contributions of many scientists. We acknowledge laboratory assistance from M. Schatz, R. Boccamazzo, G. Workman, N. Kellogg, A. Weid and R. Lionheart. Cultures were kindly provided by R. Cattolico, S. Chisholm, A. Coe, C. Deodato, M. Moran and M. Saito. Assistance with untargeted metabolomics data analysis was generously provided by R. Boiteau. We also thank J. Becker, J. Collins, F. Ribalet, J. Saunders, M. Moran and S. Amin for helpful discussion and feedback. We thank the crew members of the R/V Kilo Moana and S. Wilson for cruise leadership (KM1513), the crew members of the R/V Ka’imikai-O-Kanaloa (KOK1606) and the operational staff of the Simons Collaboration on Ocean Processes and Ecology (SCOPE). This work was supported by grants from the National Science Foundation (Award ID OCE PRF-1521564 to B.P.D. and Award ID OCE-1558483 to R.M.M. and A.E.I.), the Simons Foundation (SCOPE Award ID 329108 to E.V.A. and A.E.I., SF Award ID 385428 to A.E.I. and SF Award ID 426570 to E.V.A.) and the Gordon and Betty Moore Foundation (GBMF3776 to E.V.A.).
Author information
Authors and Affiliations
Contributions
B.P.D., R.M.M., A.E.I. and E.V.A. designed the study. B.P.D., A.K.B., L.T.C. and K.R.H. generated and analysed the metabolomics data. B.P.D., R.D.G., R.L.M. and S.N.C. generated and analysed the metatranscriptomics data. K.R.C. and R.M.M. performed the SAR11 culture experiments. B.P.D., R.M.M., A.E.I. and E.V.A. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5, Supplementary Table 1, Supplementary Table 6, Supplementary Table 9, Supplementary Table 11 and Supplementary References.
Supplementary Table 2
Measurements of cell count, biomass, and sulfonate concentration for cultures used in metabolomics measurements presented in Table 1.
Supplementary Table 3
Relative abundance of DHPS and DMSP in Thalassiosira pseudonana cells during exponential growth at 10 ppt and 35 ppt in artificial seawater medium.
Supplementary Table 4
Homologs for sulfonate metabolism genes in publicly available phytoplankton genomes at JGI and NCBI.
Supplementary Table 5
Transcript abundances in phytoplankton groups over the course of the diel study in the North Pacific Subtropical Gyre and FDR values from RAIN statistical analysis.
Supplementary Table 7
Sulfonate abundances in seawater plankton over the course of the diel study in the North Pacific Subtropical Gyre.
Supplementary Table 8
Sulfur-containing mass features detected in untargeted liquid chromatography mass spectrometry-based metabolomics data (processed using XCMS) collected near Station ALOHA in the North Pacific Subtropical Gyre.
Supplementary Table 10
Gene homologues for sulfonate catabolism genes present in publically available marine bacterial genomes available through Roseobase (http://www.roseobase.org) and IMG-ER (https://img.jgi.doe.gov/).
Supplementary Table 12
Statistics for ANOVA and Tukey’s honest significance testsc for the SAR11 growth experiments.
Rights and permissions
About this article
Cite this article
Durham, B.P., Boysen, A.K., Carlson, L.T. et al. Sulfonate-based networks between eukaryotic phytoplankton and heterotrophic bacteria in the surface ocean. Nat Microbiol 4, 1706–1715 (2019). https://doi.org/10.1038/s41564-019-0507-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0507-5
This article is cited by
-
Algal blooms in the ocean: hot spots for chemically mediated microbial interactions
Nature Reviews Microbiology (2024)
-
Microbial metabolomic responses to changes in temperature and salinity along the western Antarctic Peninsula
The ISME Journal (2023)
-
Taurine as a key intermediate for host-symbiont interaction in the tropical sponge Ianthella basta
The ISME Journal (2023)
-
A comparative study reveals the relative importance of prokaryotic and eukaryotic proton pump rhodopsins in a subtropical marginal sea
ISME Communications (2023)
-
Functional annotation and importance of marine bacterial transporters of plankton exometabolites
ISME Communications (2023)