Abstract
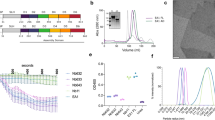
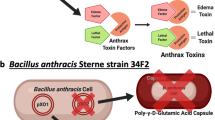
Anthrax is an ancient and deadly disease caused by the spore-forming bacterial pathogen Bacillus anthracis. At present, anthrax mostly affects wildlife and livestock, although it remains a concern for human public health—primarily for people who handle contaminated animal products and as a bioterrorism threat due to the high resilience of spores, a high fatality rate of cases and the lack of a civilian vaccination programme1,2. The cell surface of B. anthracis is covered by a protective paracrystalline monolayer—known as surface layer or S-layer—that is composed of the S-layer proteins Sap or EA1. Here, we generate nanobodies to inhibit the self-assembly of Sap, determine the structure of the Sap S-layer assembly domain (SapAD) and show that the disintegration of the S-layer attenuates the growth of B. anthracis and the pathology of anthrax in vivo. SapAD comprises six β-sandwich domains that fold and support the formation of S-layers independently of calcium. Sap-inhibitory nanobodies prevented the assembly of Sap and depolymerized existing Sap S-layers in vitro. In vivo, nanobody-mediated disruption of the Sap S-layer resulted in severe morphological defects and attenuated bacterial growth. Subcutaneous delivery of Sap inhibitory nanobodies cleared B. anthracis infection and prevented lethality in a mouse model of anthrax disease. These findings highlight disruption of S-layer integrity as a mechanism that has therapeutic potential in S-layer-carrying pathogens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Coordinates and structure factors of the SapAD–NbsAF684–NbsAF694 and SapAD–NbsAF683–NbsAF694 complexes have been deposited in PDB under accession codes 6HHU and 6QX4, respectively. All other data are available in the manuscript or the Supplementary Information.
References
Jernigan, D. B. et al. Investigation of bioterrorism-related anthrax, United States, 2001: epidemiologic findings. Emerg. Infect. Dis. 8, 1019–1028 (2002).
Sweeney, D. A., Hicks, C. W., Cui, X., Li, Y. & Eichacker, P. Q. Anthrax infection. Am. J. Respir. Crit. Care Med. 184, 1333–1341 (2011).
Couture-Tosi, E. et al. Structural analysis and evidence for dynamic emergence of Bacillus anthracis S-layer networks. J. Bacteriol. 184, 6448–6456 (2002).
Collier, R. J. & Young, J. A. Anthrax toxin. Annu. Rev. Cell Dev. Biol. 19, 45–70 (2003).
Zwartouw, H. T. & Smith, H. Polyglutamic acid from Bacillus anthracis grown in vivo; structure and aggressin activity. Biochem. J. 63, 437–442 (1956).
Makino, S., Uchida, I., Terakado, N., Sasakawa, C. & Yoshikawa, M. Molecular characterization and protein analysis of the cap region, which is essential for encapsulation in Bacillus anthracis. J. Bacteriol. 171, 722–730 (1989).
Weiner, Z. P. & Glomski, I. J. Updating perspectives on the initiation of Bacillus anthracis growth and dissemination through its host. Infect. Immun. 80, 1626–1633 (2012).
Candela, T. & Fouet, A. Bacillus anthracis CapD, belonging to the gamma-glutamyltranspeptidase family, is required for the covalent anchoring of capsule to peptidoglycan. Mol. Microbiol. 57, 717–726 (2005).
Sara, M. & Sleytr, U. B. S-layer proteins. J. Bacteriol. 182, 859–868 (2000).
Gerbino, E., Carasi, P., Mobili, P., Serradell, M. A. & Gomez-Zavaglia, A. Role of S-layer proteins in bacteria. World J. Microbiol. Biotechnol. 31, 1877–1887 (2015).
Mignot, T., Mesnage, S., Couture-Tosi, E., Mock, M. & Fouet, A. Developmental switch of S-layer protein synthesis in Bacillus anthracis. Mol. Microbiol. 43, 1615–1627 (2002).
Mesnage, S., Tosi-Couture, E., Mock, M., Gounon, P. & Fouet, A. Molecular characterization of the Bacillus anthracis main S-layer component: evidence that it is the major cell-associated antigen. Mol. Microbiol. 23, 1147–1155 (1997).
Kern, V. J., Kern, J. W., Theriot, J. A., Schneewind, O. & Missiakas, D. Surface-layer (S-layer) proteins Sap and EA1 govern the binding of the S-layer-associated protein BslO at the cell septa of Bacillus anthracis. J. Bacteriol. 194, 3833–3840 (2012).
Baranova, E. et al. SbsB structure and lattice reconstruction unveil Ca2+ triggered S-layer assembly. Nature 487, 119–122 (2012).
Kern, J. et al. Structure of surface layer homology (SLH) domains from Bacillus anthracis surface array protein. J. Biol. Chem. 286, 26042–26049 (2011).
Sychantha, D. et al. Molecular basis for the attachment of S-layer proteins to the cell wall of Bacillus anthracis. Biochemistry 57, 1949–1953 (2018).
Candela, T., Mignot, T., Hagnerelle, X., Haustant, M. & Fouet, A. Genetic analysis of Bacillus anthracis Sap S-layer protein crystallization domain. Microbiology 151, 1485–1490 (2005).
Pavkov, T. et al. The structure and binding behavior of the bacterial cell surface layer protein SbsC. Structure 16, 1226–1237 (2008).
Teixeira, L. M. et al. Entropically driven self-assembly of Lysinibacillus sphaericus S-layer proteins analyzed under various environmental conditions. Macromol. Biosci. 10, 147–155 (2010).
Rad, B. et al. Ion-specific control of the self-assembly dynamics of a nanostructured protein lattice. ACS Nano 9, 180–190 (2015).
Kirk, J. A. et al. New class of precision antimicrobials redefines role of Clostridium difficile S-layer in virulence and viability. Sci. Transl. Med. 9, eaah6813 (2017).
Wang, X. Y. et al. A S-layer protein of Bacillus anthracis as a building block for functional protein arrays by in vitro self-assembly. Small 11, 5826–5832 (2015).
Domanska, K. et al. Atomic structure of a nanobody-trapped domain-swapped dimer of an amyloidogenic β2-microglobulin variant. Proc. Natl Acad. Sci. USA 108, 1314–1319 (2011).
Conrath, K. E. et al. β-lactamase inhibitors derived from single-domain antibody fragments elicited in the Camelidae. Antimicrob. Agents Chemother. 45, 2807–2812 (2001).
Pardon, E. et al. A general protocol for the generation of nanobodies for structural biology. Nat. Protoc. 9, 674–693 (2014).
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010).
Vonrhein, C. et al. Data processing and analysis with the autoPROC toolbox. Acta Crystallogr. D 67, 293–302 (2011).
Sheldrick, G. M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D 66, 479–485 (2010).
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D 59, 2023–2030 (2003).
Abrahams, J. P. & Leslie, A. G. W. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D 52, 30–42 (1996).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Cowtan, K. D. & Main, P. Phase combination and cross validation in iterated density-modification calculations. Acta Crystallogr. D 52, 43–48 (1996).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
BUSTER v.2.10.3 (Global Phasing Ltd, 2017).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007).
Holm, L. & Sander, C. Dali: a network tool for protein structure comparison. Trends Biochem. Sci. 20, 478–480 (1995).
Jones, D. T., Taylor, W. R. & Thornton, J. M. The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. 8, 275–282 (1992).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Petoukhov, M. V. et al. New developments in the ATSAS program package for small-angle scattering data analysis. J. Appl. Cryst. 45, 342–350 (2012).
Svergun, D. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Cryst. 25, 495–503 (1992).
Svergun, D., Barberato, C. & Koch, M. H. J. CRYSOL—a program to evaluate X-ray solution scattering of biological macromolecules from atomic coordinates. J. Appl. Cryst. 28, 768–773 (1995).
Kozin, M. B. & Svergun, D. I. Automated matching of high- and low-resolution structural models. J. Appl. Cryst. 34, 33–41 (2001).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Gipson, B., Zeng, X., Zhang, Z. Y. & Stahlberg, H. 2dx—user-friendly image processing for 2D crystals. J. Struct. Biol. 157, 64–72 (2007).
Leppla, S. H. Production and purification of anthrax toxin. Methods Enzymol. 165, 103–116 (1988).
Zwietering, M. H., de Koos, J. T., Hasenack, B. E., de Witt, J. C. & van’t Riet, K. Modeling of bacterial growth as a function of temperature. Appl. Environ. Microbiol. 57, 1094–1101 (1991).
Acknowledgements
We thank P. Wattiau (CODA–CERVA Brussels) for providing the B. anthracis 34F2 strain as well as P. Goossens (Institut Pasteur, Paris) for providing the S-layer knockout mutants RBA91 and SM91; P. Borghgraef for assistance with SEM acquisition; BEI Resources, NIAID, NIH for providing us with anti-PA monoclonal antibodies; A. E. Pirro Lundqvist for assistance with selection and identification of Nbs; R. K. Singh for assistance with SAXS data analysis; and the beamline staff at I03, Diamond Light Source, UK, for support with the data collection under proposal MX12718-10. This research was supported by VIB and FWO Flanders through project grant number G028113N.
Author information
Authors and Affiliations
Contributions
A.F. and H.R. conceived the project and wrote the manuscript; H.D.G. performed gene assembly; A.F. performed cloning, protein production, functional and biophysical analysis, bacterial work as well as identification of Nbs; E.P. and J.S. supervised Ilama immunization and identification of Nbs; W.J. assisted in protein production; A.F. and H.R performed structural studies; S.E.V.d.V. performed and analysed TEM experiments, supervised by A.F. and H.R.; A.F. performed all microscopy experiments, with assistance of A.G. for fluorescent microscopy; F.V.H. performed mouse experiments with the assistance of A.F. and supervised by M.L. and H.R.
Corresponding authors
Ethics declarations
Competing interests
A priority application on compounds used to inhibit bacterial S-layer protein assembly has been filed by VIB and Vrije Universiteit Brussel at the European Patent Office listing A.F. and H.R. as inventors. The other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–3, legend for Supplementary Video 1, Supplementary Figs. 1–9, Supplementary Table 1 and full length blots.
Supplementary Video 1
Timelapse of B. anthracis growth in the presence or absence of NbsSAI.
Rights and permissions
About this article
Cite this article
Fioravanti, A., Van Hauwermeiren, F., Van der Verren, S.E. et al. Structure of S-layer protein Sap reveals a mechanism for therapeutic intervention in anthrax. Nat Microbiol 4, 1805–1814 (2019). https://doi.org/10.1038/s41564-019-0499-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0499-1
This article is cited by
-
Structure and function of the EA1 surface layer of Bacillus anthracis
Nature Communications (2023)
-
Structure and assembly of the S-layer in C. difficile
Nature Communications (2022)