Abstract
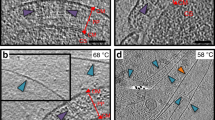
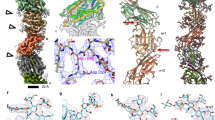
Pili on the surface of Sulfolobus islandicus are used for many functions, and serve as receptors for certain archaeal viruses. The cells grow optimally at pH 3 and ~80 °C, exposing these extracellular appendages to a very harsh environment. The pili, when removed from cells, resist digestion by trypsin or pepsin, and survive boiling in sodium dodecyl sulfate or 5 M guanidine hydrochloride. We used electron cryo-microscopy to determine the structure of these filaments at 4.1 Å resolution. An atomic model was built by combining the electron density map with bioinformatics without previous knowledge of the pilin sequence—an approach that should prove useful for assemblies where all of the components are not known. The atomic structure of the pilus was unusual, with almost one-third of the residues being either threonine or serine, and with many hydrophobic surface residues. While the map showed extra density consistent with glycosylation for only three residues, mass measurements suggested extensive glycosylation. We propose that this extensive glycosylation renders these filaments soluble and provides the remarkable structural stability. We also show that the overall fold of the archaeal pilin is remarkably similar to that of archaeal flagellin, establishing common evolutionary origins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
References
Prangishvili, D. et al. The enigmatic archaeal virosphere. Nat. Rev. Microbiol. 15, 724–739 (2017).
Chaudhury, P., Quax, T. E. F. & Albers, S. V. Versatile cell surface structures of archaea. Mol. Microbiol. 107, 298–311 (2018).
DiMaio, F. et al. A virus that infects a hyperthermophile encapsidates A-form DNA. Science 348, 914–917 (2015).
Liu, Y. et al. Structural conservation in a membrane-enveloped filamentous virus infecting a hyperthermophilic acidophile. Nat. Commun. 9, 3360 (2018).
Ptchelkine, D. et al. Unique architecture of thermophilic archaeal virus APBV1 and its genome packaging. Nat. Commun. 8, 1436 (2017).
Kasson, P. et al. Model for a novel membrane envelope in a filamentous hyperthermophilic virus. eLife 6, e26268 (2017).
Veesler, D. et al. Atomic structure of the 75 MDa extremophile Sulfolobus turreted icosahedral virus determined by CryoEM and X-ray crystallography. Proc. Natl Acad. Sci. USA 110, 5504–5509 (2013).
Berry, J. L. & Pelicic, V. Exceptionally widespread nanomachines composed of type IV pilins: the prokaryotic Swiss Army knives. FEMS Microbiol. Rev. 39, 134–154 (2015).
Makarova, K. S., Koonin, E. V. & Albers, S. V. Diversity and evolution of type IV pili systems in archaea. Front. Microbiol. 7, 667 (2016).
Pohlschroder, M., Ghosh, A., Tripepi, M. & Albers, S. V. Archaeal type IV pilus-like structures—evolutionarily conserved prokaryotic surface organelles. Curr. Opin. Microbiol. 14, 357–363 (2011).
Pohlschroder, M. & Esquivel, R. N. Archaeal type IV pili and their involvement in biofilm formation. Front. Microbiol. 6, 190 (2015).
Albers, S. V. & Jarrell, K. F. The archaellum: an update on the unique archaeal motility structure. Trends Microbiol. 26, 351–362 (2018).
Wang, F. et al. Cryoelectron microscopy reconstructions of the Pseudomonas aeruginosa and Neisseria gonorrhoeae type IV pili at sub-nanometer resolution. Structure 25, 1423–1435 (2017).
Kolappan, S. et al. Structure of the Neisseria meningitidis type IV pilus. Nat. Commun. 7, 13015 (2016).
Lopez-Castilla, A. et al. Structure of the calcium-dependent type 2 secretion pseudopilus. Nat. Microbiol. 2, 1686–1695 (2017).
Craig, L. et al. Type IV pilus structure by cryo-electron microscopy and crystallography: implications for pilus assembly and functions. Mol. Cell 23, 651–662 (2006).
Craig, L. et al. Type IV pilin structure and assembly: X-ray and EM analyses of Vibrio cholerae toxin-coregulated pilus and Pseudomonas aeruginosa PAK pilin. Mol. Cell 11, 1139–1150 (2003).
Hartung, S. et al. Ultrahigh resolution and full-length pilin structures with insights for filament assembly, pathogenic functions, and vaccine potential. J. Biol. Chem. 286, 44254–44265 (2011).
Reardon, P. N. & Mueller, K. T. Structure of the type IVa major pilin from the electrically conductive bacterial nanowires of Geobacter sulfurreducens. J. Biol. Chem. 288, 29260–29266 (2013).
Szabo, Z. et al. Identification of diverse archaeal proteins with class III signal peptides cleaved by distinct archaeal prepilin peptidases. J. Bacteriol. 189, 772–778 (2007).
Albers, S. V., Szabo, Z. & Driessen, A. J. Archaeal homolog of bacterial type IV prepilin signal peptidases with broad substrate specificity. J. Bacteriol. 185, 3918–3925 (2003).
Strom, M. S., Nunn, D. N. & Lory, S. A single bifunctional enzyme, PilD, catalyzes cleavage and N-methylation of proteins belonging to the type IV pilin family. Proc. Natl Acad. Sci. USA 90, 2404–2408 (1993).
Jaubert, C. et al. Genomics and genetics of Sulfolobus islandicus LAL14/1, a model hyperthermophilic archaeon. Open Biol. 3, 130010 (2013).
Quemin, E. R. et al. First insights into the entry process of hyperthermophilic archaeal viruses. J. Virol. 87, 13379–13385 (2013).
DiMaio, F. et al. Atomic-accuracy models from 4.5-A cryo-electron microscopy data with density-guided iterative local refinement. Nat. Methods 12, 361–365 (2015).
Braun, T. et al. Archaeal flagellin combines a bacterial type IV pilin domain with an Ig-like domain. Proc. Natl Acad. Sci. USA 113, 10352–10357 (2016).
Yu, X. et al. Filaments from Ignicoccus hospitalis show diversity of packing in proteins containing N-terminal type IV pilin helices. J. Mol. Biol. 422, 274–281 (2012).
Poweleit, N. et al. CryoEM structure of the Methanospirillum hungatei archaellum reveals structural features distinct from the bacterial flagellum and type IV pili. Nat. Microbiol. 2, 16222 (2016).
Daum, B. et al. Structure and in situ organisation of the Pyrococcus furiosus archaellum machinery. eLife 6, e27470 (2017).
Bayley, D. P. & Jarrell, K. F. Further evidence to suggest that archaeal flagella are related to bacterial type IV pili. J. Mol. Evol. 46, 370–373 (1998).
Ng, S. Y., Chaban, B. & Jarrell, K. F. Archaeal flagella, bacterial flagella and type IV pili: a comparison of genes and posttranslational modifications. J. Mol. Microbiol. Biotechnol. 11, 167–191 (2006).
Faguy, D. M., Jarrell, K. F., Kuzio, J. & Kalmokoff, M. L. Molecular analysis of archael flagellins: similarity to the type IV pilin-transport superfamily widespread in bacteria. Can. J. Microbiol. 40, 67–71 (1994).
Song, Y. et al. High-resolution comparative modeling with RosettaCM. Structure 21, 1735–1742 (2013).
Henche, A. L. et al. Structure and function of the adhesive type IV pilus of Sulfolobus acidocaldarius. Environ. Microbiol. 14, 3188–3202 (2012).
Kryndushkin, D., Pripuzova, N., Burnett, B. & Shewmaker, F. Non-targeted identification of prions and amyloid-forming proteins from yeast and mammalian cells. J. Biol. Chem. 288, 27100–27111 (2013).
Nelson, R. et al. Structure of the cross-β spine of amyloid-like fibrils. Nature 435, 773–778 (2005).
Gromiha, M. M. & Selvaraj, S. Comparison between long-range interactions and contact order in determining the folding rate of two-state proteins: application of long-range order to folding rate prediction. J. Mol. Biol. 310, 27–32 (2001).
Broom, A. et al. Designed protein reveals structural determinants of extreme kinetic stability. Proc. Natl Acad. Sci. USA 112, 14605–14610 (2015).
Stranges, P. B. & Kuhlman, B. A comparison of successful and failed protein interface designs highlights the challenges of designing buried hydrogen bonds. Protein Sci. 22, 74–82 (2013).
Pohlschroder, M., Pfeiffer, F., Schulze, S. & Halim, M. F. A. Archaeal cell surface biogenesis. FEMS Microbiol. Rev. 42, 694–717 (2018).
Mann, M. & Jensen, O. N. Proteomic analysis of post-translational modifications. Nat. Biotechnol. 21, 255–261 (2003).
Wall, J. S. & Hainfeld, J. F. Mass mapping with the scanning transmission electron microscope. Annu. Rev. Biophys. Biophys. Chem. 15, 355–376 (1986).
Sojar, H. T. & Bahl, O. P. Chemical deglycosylation of glycoproteins. Methods Enzymol. 138, 341–350 (1987).
Ellen, A. F. et al. Proteomic analysis of secreted membrane vesicles of archaeal Sulfolobus species reveals the presence of endosome sorting complex components. Extremophiles 13, 67–79 (2009).
Altschul, S. F. & Lipman, D. J. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Holm, L. & Rosenstrom, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–W549 (2010).
Esquivel, R. N., Schulze, S., Xu, R., Hippler, M. & Pohlschroder, M. Identification of Haloferax volcanii pilin N-glycans with diverse roles in pilus biosynthesis, adhesion, and microcolony formation. J. Biol. Chem. 291, 10602–10614 (2016).
Gault, J. et al. Neisseria meningitidis type IV pili composed of sequence invariable pilins are masked by multisite glycosylation. PLoS Pathog. 11, e1005162 (2015).
Mer, G., Hietter, H. & Lefevre, J. F. Stabilization of proteins by glycosylation examined by NMR analysis of a fucosylated proteinase inhibitor. Nat. Struct. Biol. 3, 45–53 (1996).
Shental-Bechor, D. & Levy, Y. Effect of glycosylation on protein folding: a close look at thermodynamic stabilization. Proc. Natl Acad. Sci. USA 105, 8256–8261 (2008).
Sola, R. J. & Griebenow, K. Effects of glycosylation on the stability of protein pharmaceuticals. J. Pharm. Sci. 98, 1223–1245 (2009).
Zillig, W. et al. Screening for Sulfolobales, their plasmids and their viruses in Icelandic solfataras. Syst. Appl. Microbiol. 16, 609–628 (1993).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Afonine, P. V. et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr. D Struct. Biol. 74, 814–840 (2018).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43, W389–W394 (2015).
Wang, R. Y. et al. De novo protein structure determination from near-atomic-resolution cryo-EM maps. Nat. Methods 12, 335–338 (2015).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D Biol. Crystallogr. 66, 213–221 (2010).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013).
Wall, J. S. & Simon, M. N. Scanning transmission electron microscopy of DNA–protein complexes. Methods Mol. Biol. 148, 589–601 (2001).
Acknowledgements
This work was supported by NIH grants GM122510 (to E.H.E.) and GM123089 (to F.D.), as well as l’Agence Nationale de la Recherche project ENVIRA (ANR-17-CE15-0005-01 to M.K.). M.A.B.K. was supported by NIH grant T32 GM080186. The cryo-EM imaging conducted at the Molecular Electron Microscopy Core facility at the University of Virginia was supported by the School of Medicine and built with NIH grant G20-RR31199. The Titan Krios and Falcon II direct electron detectors were obtained with NIH grants S10-RR025067 and S10-OD018149, respectively. We thank V. Conticello for the suggestion of TFMS. We are also grateful to the Ultrastructural BioImaging (UTechS UBI) unit of Institut Pasteur for access to electron microscopes.
Author information
Authors and Affiliations
Contributions
V.C.-K. isolated and purified the pili. J.S.W. performed the STEM analysis. N.S. performed the mass spectrometry. G.A.P.d.O. performed the amyloid assays. F.D. performed the interfacial analysis. F.W. performed the cryo-EM, three-dimensional reconstruction and model building, with assistance from T.O. and E.H.E. M.K. and Z.S. performed the bioinformatics analysis. M.A.B.K. performed the TFMS deglycosylation. D.P., M.K. and E.H.E. designed the project. F.W., D.P., M.K. and E.H.E. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Table 1 and Supplementary Figures 1–10.
Rights and permissions
About this article
Cite this article
Wang, F., Cvirkaite-Krupovic, V., Kreutzberger, M.A.B. et al. An extensively glycosylated archaeal pilus survives extreme conditions. Nat Microbiol 4, 1401–1410 (2019). https://doi.org/10.1038/s41564-019-0458-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0458-x
This article is cited by
-
Archaeal DNA-import apparatus is homologous to bacterial conjugation machinery
Nature Communications (2023)
-
A general approach to explore prokaryotic protein glycosylation reveals the unique surface layer modulation of an anammox bacterium
The ISME Journal (2022)
-
An archaellum filament composed of two alternating subunits
Nature Communications (2022)
-
Flagellin outer domain dimerization modulates motility in pathogenic and soil bacteria from viscous environments
Nature Communications (2022)
-
Cryo-EM structure of an extracellular Geobacter OmcE cytochrome filament reveals tetrahaem packing
Nature Microbiology (2022)