Abstract
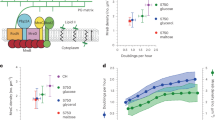
Rod-shaped bacteria grow by adding material into their cell wall via the action of two spatially distinct enzymatic systems: the Rod complex moves around the cell circumference, whereas class A penicillin-binding proteins (aPBPs) do not. To understand how the combined action of these two systems defines bacterial dimensions, we examined how each affects the growth and width of Bacillus subtilis as well as the mechanical anisotropy and orientation of material within their sacculi. Rod width is not determined by MreB, rather it depends on the balance between the systems: the Rod complex reduces diameter, whereas aPBPs increase it. Increased Rod-complex activity correlates with an increased density of directional MreB filaments and a greater fraction of directional PBP2a enzymes. This increased circumferential synthesis increases the relative quantity of oriented material within the sacculi, making them more resistant to stretching across their width, thereby reinforcing rod shape. Together, these experiments explain how the combined action of the two main cell wall synthetic systems builds and maintains rods of different widths. Escherichia coli Rod mutants also show the same correlation between width and directional MreB filament density, suggesting this model may be generalizable to bacteria that elongate via the Rod complex.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All datasets and raw data generated during and/or analysed during the current study are available from the corresponding author on reasonable request. Raw and proteomic data are available at https://garnerlab.fas.harvard.edu/Dion2019/Raw-MS-data-Dion2019.zip.
Code availability
All particle tracking was done with the Trackmate plugin within FIJI, then analysed using code available at https://bitbucket.org/garnerlab/hussain-2017-elife. Filament density calculations and filament simulations were performed with custom code available at https://bitbucket.org/garnerlab/dion-2018/src/.
Change history
06 June 2019
In the version of this Article originally published, Supplementary Video 1 was incorrectly linked to Supplementary Video 3, Supplementary Video 2 was incorrectly linked to Supplementary Video 1 and Supplementary Video 3 was incorrectly linked to Supplementary Video 2. The files have now been replaced to rectify this.
References
Vadia, S. & Levin, P. A. Growth rate and cell size: a re-examination of the growth law. Curr. Opin. Microbiol. 24, 96–103 (2015).
Sharpe, M. E., Hauser, P. M., Sharpe, R. G. & Errington, J. Bacillus subtilis cell cycle as studied by fluorescence microscopy: constancy of cell length at initiation of DNA replication and evidence for active nucleoid partitioning. J. Bacteriol. 180, 547–555 (1998).
Vollmer, W., Blanot, D. & de Pedro, M. A. Peptidoglycan structure and architecture. FEMS Microbiol. Rev. 32, 149–167 (2008).
Lebar, M. D. et al. Reconstitution of peptidoglycan cross-linking leads to improved fluorescent probes of cell wall synthesis. J. Am. Chem. Soc. 136, 10874–10877 (2014).
Banzhaf, M. et al. Cooperativity of peptidoglycan synthases active in bacterial cell elongation. Mol. Microbiol. 85, 179–194 (2012).
Meeske, A. J. et al. SEDS proteins are a widespread family of bacterial cell wall polymerases. Nature 537, 634–638 (2016).
Jones, L. J., Carballido-López, R. & Errington, J. Control of cell shape in bacteria: helical, actin-like filaments in Bacillus subtilis. Cell 104, 913–922 (2001).
van den Ent, F., Amos, L. & Löwe, J. Bacterial ancestry of actin and tubulin. Curr. Opin. Microbiol. 4, 634–638 (2001).
van den Ent, F., Izoré, T., Bharat, T. A., Johnson, C. M. & Lowe, J. Bacterial actin MreB forms antiparallel double filaments. eLife 3, e02634 (2014).
Salje, J., van den Ent, F., de Boer, P. & Lowe, J. Direct membrane binding by bacterial actin MreB. Mol. Cell 43, 478–487 (2011).
Hussain, S. et al. MreB filaments align along greatest principal membrane curvature to orient cell wall synthesis. eLife 7, e32471 (2018).
Garner, E. C. et al. Coupled, circumferential motions of the cell wall synthesis machinery and MreB filaments in B. subtilis. Science 333, 222–225 (2011).
van Teeffelen, S. et al. The bacterial actin MreB rotates, and rotation depends on cell-wall assembly. Proc. Natl Acad. Sci. USA 108, 15822–15827 (2011).
Domínguez-Escobar, J. et al. Processive movement of MreB-associated cell wall biosynthetic complexes in bacteria. Science 333, 225–228 (2011).
Turner, R. D., Mesnage, S., Hobbs, J. K. & Foster, S. J. Molecular imaging of glycan chains couples cell-wall polysaccharide architecture to bacterial cell morphology. Nat. Commun. 9, 1263 (2018).
McPherson, D. C. & Popham, D. L. Peptidoglycan synthesis in the absence of class A penicillin-binding proteins in Bacillus subtilis. J. Bacteriol. 185, 1423–1431 (2003).
Cho, H. et al. Bacterial cell wall biogenesis is mediated by SEDS and PBP polymerase families functioning semi-autonomously. Nat. Microbiol. 1, 16172 (2016).
Ursell, T. S. et al. Rod-like bacterial shape is maintained by feedback between cell curvature and cytoskeletal localization. Proc. Natl Acad. Sci. USA 111, E1025–E1034 (2014).
Ouzounov, N. et al. MreB orientation correlates with cell diameter in Escherichia coli. Biophys. J. 111, 1035–1043 (2016).
Wang, S. & Wingreen, N. S. Cell shape can mediate the spatial organization of the bacterial cytoskeleton. Biophys. J. 104, 541–552 (2013).
Tropini, C. et al. Principles of bacterial cell-size determination revealed by cell-wall synthesis perturbations. Cell Rep. 9, 1520–1527 (2014).
Shi, H. et al. Deep phenotypic mapping of bacterial cytoskeletal mutants reveals physiological robustness to cell size. Curr. Biol. 27, 3419–3429 (2017).
Colavin, A., Shi, H. & Huang, K. C. RodZ modulates geometric localization of the bacterial actin MreB to regulate cell shape. Nat. Commun. 9, 1280 (2018).
Olshausen, P. V. et al. Superresolution imaging of dynamic MreB filaments in B. subtilis—a multiple-motor-driven transport? Biophys. J. 105, 1171–1181 (2013).
Shi, H., Bratton, B. P., Gitai, Z. & Huang, K. C. How to build a bacterial cell: MreB as the foreman of E. coli construction. Cell 172, 1294–1305 (2018).
Schirner, K. & Errington, J. Influence of heterologous MreB proteins on cell morphology of Bacillus subtilis. Microbiology 155, 3611–3621 (2009).
Harris, L. K., Dye, N. A. & Theriot, J. A. A Caulobacter MreB mutant with irregular cell shape exhibits compensatory widening to maintain a preferred surface area to volume ratio. Mol. Microbiol. 5, 988–1005 (2014).
Bisson-Filho, A. W. et al. FtsZ filament capping by MciZ, a developmental regulator of bacterial division. Proc. Natl Acad. Sci. USA 112, E2130–E2138 (2015).
Formstone, A. & Errington, J. A magnesium-dependent mreB null mutant: implications for the role of mreB in Bacillus subtilis. Mol. Microbiol. 55, 1646–1657 (2005).
Vigouroux, A., Oldewurtel, E., Cui, L., Bikard, D. & van Teeffelen, S. Tuning dCas9’s ability to block transcription enables robust, noiseless knockdown of bacterial genes. Mol. Sys. Biol. 14, e7899 (2018).
Henriques, A. O., Glaser, P., Piggot, P. J. & Moran, C. P. Jr Control of cell shape and elongation by the rodA gene in Bacillus subtilis. Mol. Microbiol. 28, 235–247 (1998).
Fraipont, C. et al. The integral membrane FtsW protein and peptidoglycan synthase PBP3 form a subcomplex in Escherichia coli. Microbiology 157, 251–259 (2010).
Taheri-Araghi, S. et al. Cell-size control and homeostasis in bacteria. Curr. Biol. 25, 385–391 (2015).
Zheng, H. et al. Interrogating the Escherichia coli cell cycle by cell dimension perturbations. Proc. Natl Acad. Sci. USA 113, 15000–15005 (2016).
Murray, T., Popham, D. L. & Setlow, P. Bacillus subtilis cells lacking penicillin-binding protein 1 require increased levels of divalent cations for growth. J. Bacteriol. 180, 4555–4563 (1998).
Popham, D. L. & Setlow, P. Phenotypes of Bacillus subtilis mutants lacking multiple class a high-molecular-weight penicillin-binding proteins. J. Bacteriol. 178, 2079–2085 (1996).
Emami, K. et al. RodA as the missing glycosyltransferase in Bacillus subtilis and antibiotic discovery for the peptidoglycan polymerase pathway. Nat. Microbiol. 2, 16253 (2017).
Grimm, J. B. et al. A general method to improve fluorophores for live-cell and single-molecule microscopy. Nat. Methods 12, 244–250 (2015).
Oldenbourg, R. Polarized light microscopy: principles and practice. Cold Spring Harb. Protoc. 2013, pdb.top078600 (2013).
Probine, M. C. & Preston, R. D. Cell growth and the structure and mechanical properties of the wall in internodal cells of Nitella opacai. Wall structure and growth. J. Exp. Bot. 12, 261–282 (1961).
Inoué, S. Polarization Microscopy. Curr. Protoc. Cell Biol. 13, 4.9.1–4.9.27 (2002).
Verwer, R. W., Beachey, E. H., Keck, W., Stoub, A. M. & Poldermans, J. E. Oriented fragmentation of Escherichia coli sacculi by sonication. J. Bacteriol. 141, 327–332 (1980).
Yao, X., Jericho, M., Pink, D. & Beveridge, T. Thickness and elasticity of gram-negative murein sacculi measured by atomic force microscopy. J. Bacteriol. 181, 6865–6875 (1999).
Bratton, B. P., Shaevitz, J. W., Gitai, Z. & Morgenstein, R. M. MreB polymers and curvature localization are enhanced by RodZ and predict E. coli’s cylindrical uniformity. Nat. Commun. 9, 2797 (2018).
Kurita, K., Shin, R., Tabei, T. & Shiomi, D. Relation between rotation of MreB actin and cell width of Escherichia coli. Genes Cells 24, 259–265 (2018).
Rojas, E. R., Huang, K. C. & Theriot, J. A. Homeostatic cell growth is accomplished mechanically through membrane tension inhibition of cell-wall synthesis. Cell Syst. 5, 578–590 (2017).
Leaver, M. & Errington, J. Roles for MreC and MreD proteins in helical growth of the cylindrical cell wall in Bacillus subtilis. Mol. Microbiol. 57, 1196–1209 (2005).
Kawai, Y., Daniel, R. A. & Errington, J. Regulation of cell wall morphogenesis in Bacillus subtilis by recruitment of PBP1 to the MreB helix. Mol. Microbiol. 71, 1131–1144 (2009).
Lai, G. C., Cho, H. & Bernhardt, T. G. The mecillinam resistome reveals a role for peptidoglycan endopeptidases in stimulating cell wall synthesis in Escherichia coli. PLoS Genet. 13, e1006934 (2017).
Lee, T. K., Meng, K., Shi, H. & Huang, K. C. Single-molecule imaging reveals modulation of cell wall synthesis dynamics in live bacterial cells. Nat. Commun. 7, 13170 (2016).
Agresti, A. & Coull, B. A. Approximate is better than ‘exact’ for interval estimation of binomial proportions. Am. Stat. 52, 119–126 (1998).
Ursell, T. et al. Rapid, precise quantification of bacterial cellular dimensions across a genomic-scale knockout library. BMC Biol. 15, 17 (2017).
Grimm, J. B. et al. A general method to improve fluorophores for live-cell and single-molecule microscopy. Nat. Methods 12, 244–250 (2015).
Tinevez, J.-Y. et al. TrackMate: an open and extensible platform for single-particle tracking. Methods 115, 80–90 (2017).
Billaudeau, C. et al. Contrasting mechanisms of growth in two model rod-shaped bacteria. Nat. Commun. 8, 15370 (2017).
Kner, P., Chhun, B. B., Griffis, E. R., Winoto, L. & Gustafsson, M. G. L. Super-resolution video microscopy of live cells by structured illumination. Nat. Methods 6, 339–342 (2009).
Kall, L., Storey, J. D. & Noble, W. S. Non-parametric estimation of posterior error probabilities associated with peptides identified by tandem mass spectrometry. Bioinformatics 24, i42–i48 (2008).
Morgenstein, R. M. et al. RodZ links MreB to cell wall synthesis to mediate MreB rotation and robust morphogenesis. Proc. Natl Acad. Sci. USA 112, 12510–12515 (2015).
Acknowledgements
We thank C. Wivagg, Y. J. Eun and L. Harris for their helpful discussions, G. Squyres for TIRF-SIM, L. Lavis for JF dyes and Z. Gitai and K. C. Huang for bacterial strains. TIRF-SIM was performed at the Advanced Imaging Center at the Janelia Research Campus, a facility jointly supported by the Gordon and Betty Moore Foundation and Howard Hughes Medical Institute. This work was funded by the National Institute of Health (grant nos R01GM114274 to R.O. and DP2AI117923 to E.C.G) and support from the Volkswagen Foundation to E.C.G. and S.v.T. Some work was performed at the Center for Nanoscale Systems at Harvard University, supported by NSF ECS-0335765.
Author information
Authors and Affiliations
Contributions
B. subtilis strains were cloned by M.F.D., Y.S. and M.K. All width, length and bulk growth measurements of B. subtilis were performed by M.F.D. and E. coli widths by Y.S. E. coli CRISPRi strains were cloned by A.V., supervised by S.v.T. Single-cell growth rates were done by Y.S. and M.K. TIRFM and tracking of PBP2a was conducted by M.K. All TIRFM of MreB was done by Y.S. TIRF-SIM of MreB was performed by Y.S., E.C.G. and J.R. Proteomic preparations and analyses, and sacculi purifications were done by M.F.D. Y.S. wrote the code for the analysis of single-cell growth rates, filament density and simulations of data. Polarization microscopy and analysis was conducted by J.R., R.O. and E.C.G. S.W. did the FDAA synthesis, osmotic shocks and transmission electron microscopy. The paper was written by E.C.G., M.K., M.F.D. and Y.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–7 and Supplementary Video legends.
Supplementary Data 1
Mass spectrometry data.
Supplementary Video 1
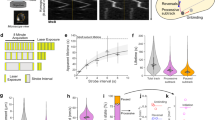
Example single molecule videos (taken with TIRF microscopy) of Halo–PBP2a (labelled with JF549) at different mreBCD induction levels in bMK385 (amyE::erm Pxyl-mreBCD, ΔmreBCD::spc, pbpA::cat HaloTag-11aa-pbpA). Xylose concentrations are indicated in each panel. Frames are 300 ms apart.
Supplementary Video 2
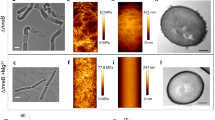
Z-stack of LC-PolScope images of sacculi purified from WT strain PY79. Left, Z-stack of the retardance of each plane. Right, colour shows slow axis orientation, intensity corresponds to retardance in that direction (reference, upper left colour wheel). Z-steps were taken in 100 nm increments.
Supplementary Video 3
Z-stack colour map of birefringence of sacculi purified from strain bMD620 (amyE::erm Pxyl-mreBCD, ΔmreBCD::spc, yhdG::cat Pspank-ponA, ΔponA::kan), where cells were induced to “Wide” (0.5 mM xylose, 0.1 mM IPTG) and “Normal” widths (5 mM xylose, 0.025 mM IPTG). Colour shows slow axis orientation, intensity corresponds to retardance in that direction (reference, upper left colour wheel). Z-steps were taken in 100 nm increments.
Supplementary Video 4
TIRF-SIM videos of different MreB-msfGFPsw mutants. Cells were grown in LB at 37 °C, then placed under an LB agarose pad and imaged at 37 °C. Frames are 1 s apart. Scale bars are 1 μm.
Supplementary Video 5
TIRF-SIM videos of strain KC717 (mreB::msfGFP-mreB, ProdZ < > (frt araC PBAD)). Cells were grown for 5 h in LB with the indicated arabinose concentrations (to induce different amounts of RodZ expression) then placed under an LB agarose pad and imaged at 37 °C. Frames are 1 s apart. Scale bars are 1 μm.
Rights and permissions
About this article
Cite this article
Dion, M.F., Kapoor, M., Sun, Y. et al. Bacillus subtilis cell diameter is determined by the opposing actions of two distinct cell wall synthetic systems. Nat Microbiol 4, 1294–1305 (2019). https://doi.org/10.1038/s41564-019-0439-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0439-0
This article is cited by
-
Molecular motor tug-of-war regulates elongasome cell wall synthesis dynamics in Bacillus subtilis
Nature Communications (2024)
-
Identification and optimization of genes potentially related to protein expression for enhancing α-amylase production in Bacillus subtilis
Systems Microbiology and Biomanufacturing (2024)
-
Barriers to simultaneous multilocus integration in Bacillus subtilis tumble down: development of a straightforward screening method for the colorimetric detection of one-step multiple gene insertion using the CRISPR-Cas9 system
Microbial Cell Factories (2023)
-
Coordinated peptidoglycan synthases and hydrolases stabilize the bacterial cell wall
Nature Communications (2023)
-
Evidence of two differentially regulated elongasomes in Salmonella
Communications Biology (2023)