Abstract
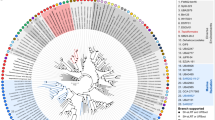
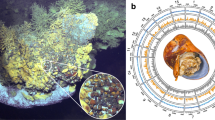
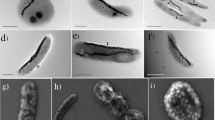
Mutualistic symbioses are often a source of evolutionary innovation and drivers of biological diversification1. Widely distributed in the microbial world, particularly in anoxic settings2,3, they often rely on metabolic exchanges and syntrophy2,4. Here, we report a mutualistic symbiosis observed in marine anoxic sediments between excavate protists (Symbiontida, Euglenozoa)5 and ectosymbiotic Deltaproteobacteria biomineralizing ferrimagnetic nanoparticles. Light and electron microscopy observations as well as genomic data support a multi-layered mutualism based on collective magnetotactic motility with division of labour and interspecies hydrogen-transfer-based syntrophy6. The guided motility of the consortia along the geomagnetic field is allowed by the magnetic moment of the non-motile ectosymbiotic bacteria combined with the protist motor activity, which is a unique example of eukaryotic magnetoreception7 acquired by symbiosis. The nearly complete deltaproteobacterial genome assembled from a single consortium contains a full magnetosome gene set8, but shows signs of reduction, with the probable loss of flagellar genes. Based on the metabolic gene content, the ectosymbiotic bacteria are anaerobic sulfate-reducing chemolithoautotrophs that likely reduce sulfate with hydrogen produced by hydrogenosome-like organelles6 underlying the plasma membrane of the protist. In addition to being necessary hydrogen sinks, ectosymbionts may provide organics to the protist by diffusion and predation, as shown by magnetosome-containing digestive vacuoles. Phylogenetic analyses of 16S and 18S ribosomal RNA genes from magnetotactic consortia in marine sediments across the Northern and Southern hemispheres indicate a host–ectosymbiont specificity and co-evolution. This suggests a historical acquisition of magnetoreception by a euglenozoan ancestor from Deltaproteobacteria followed by subsequent diversification. It also supports the cosmopolitan nature of this type of symbiosis in marine anoxic sediments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Gene sequences of 18S and 16S rRNA amplified from magnetically purified populations or sorted single holobionts have been assigned to GenBank accession numbers MK131721–MK131747 and MK153697–MK153721, respectively. Sequencing reads and the annotated genome of strain CR-1 were deposited to the European Nucleotide Archive database under the BioProject numbers PRJEB29359 and PRJEB30760, respectively.
References
Margulis, L. & Fester, R. Symbiosis as a Source of Evolutionary Innovation: Speciation and Morphogenesis (MIT Press, 1991).
López-García, P., Eme, L. & Moreira, D. Symbiosis in eukaryotic evolution. J. Theor. Biol. 434, 20–33 (2017).
Bernhard, J. M., Buck, K. R., Farmer, M. A. & Bowser, S. S. The Santa Barbara Basin is a symbiosis oasis. Nature 403, 77–80 (2000).
Morris, B. E. L., Henneberger, R., Huber, H. & Moissl-Eichinger, C. Microbial syntrophy: interaction for the common good. FEMS Microbiol. Rev. 37, 384–406 (2013).
Yubuki, N. & Leander, B. S. Diversity and evolutionary history of the symbiontida (Euglenozoa). Front. Ecol. Evol. 6, 100 (2018).
Müller, M. et al. Biochemistry and evolution of anaerobic energy metabolism in eukaryotes. Microbiol. Mol. Biol. Rev. 76, 444–495 (2012).
Mouritsen, H. Long-distance navigation and magnetoreception in migratory animals. Nature 558, 50–59 (2018).
Uebe, R. & Schüler, D. Magnetosome biogenesis in magnetotactic bacteria. Nat. Rev. Microbiol. 14, 621–637 (2016).
Walker, M. M. et al. Structure and function of the vertebrate magnetic sense. Nature 390, 371–376 (1997).
Mora, C. V., Davison, M., Wild, J. M. & Walker, M. M. Magnetoreception and its trigeminal mediation in the homing pigeon. Nature 432, 508–511 (2004).
Nordmann, G. C., Hochstoeger, T. & Keays, D. A. Magnetoreception—a sense without a receptor. PLoS Biol. 15, e2003234 (2017).
Bazylinski, D. A. & Frankel, R. B. Magnetosome formation in prokaryotes. Nat. Rev. Microbiol. 2, 217–230 (2004).
Popp, F., Armitage, J. P. & Schüler, D. Polarity of bacterial magnetotaxis is controlled by aerotaxis through a common sensory pathway. Nat. Commun. 5, 5398 (2014).
Torres de Araujo, F. F., Pires, M. A., Frankel, R. B. & Bicudo, C. E. M. Magnetite and magnetotaxis in algae. Biophys. J. 50, 375–378 (1986).
Simpson, A. G. B., VandenHoff, J., Bernard, C., Burton, H. R. & Patterson, D. J. The ultrastructure and systematic position of the euglenozoon Postgaardi mariagerensis, Fenchel et al. Arch. Protistenkd. 147, 213–225 (1997).
Blakemore, R. P., Frankel, R. B. & Kalmijn, A. J. South-seeking magnetotactic bacteria in the Southern Hemisphere. Nature 286, 384–385 (1980).
Edgcomb, V. P. et al. Identity of epibiotic bacteria on symbiontid euglenozoans in O2-depleted marine sediments: evidence for symbiont and host co-evolution. ISME J. 5, 231–243 (2011).
Breglia, S. A., Yubuki, N., Hoppenrath, M. & Leander, B. S. Ultrastructure and molecular phylogenetic position of a novel euglenozoan with extrusive episymbiotic bacteria: Bihospites bacati n. gen. et sp. (Symbiontida). BMC Microbiol. 10, 145 (2010).
Yubuki, N., Edgcomb, V. P., Bernhard, J. M. & Leander, B. S. Ultrastructure and molecular phylogeny of Calkinsia aureus: cellular identity of a novel clade of deep-sea euglenozoans with epibiotic bacteria. BMC Microbiol. 9, 16 (2009).
Abreu, F. et al. Deciphering unusual uncultured magnetotactic multicellular prokaryotes through genomics. ISME J. 8, 1055–1068 (2014).
Lefèvre, C. T. et al. A cultured greigite-producing magnetotactic bacterium in a novel group of sulfate-reducing bacteria. Science 334, 1720–1723 (2011).
Pósfai, M., Lefèvre, C. T., Trubitsyn, D., Bazylinski, D. A. & Frankel, R. B. Phylogenetic significance of composition and crystal morphology of magnetosome minerals. Front. Microbiol. 4, 344 (2013).
Moran, N. A., McLaughlin, H. J. & Sorek, R. The dynamics and time scale of ongoing genomic erosion in symbiotic bacteria. Science 323, 379–382 (2009).
McCutcheon, J. P. & Moran, N. A. Extreme genome reduction in symbiotic bacteria. Nat. Rev. Microbiol. 10, 13–26 (2011).
Leão, P. et al. North-seeking magnetotactic Gammaproteobacteria in the Southern Hemisphere. Appl. Environ. Microbiol. 82, 5595–5602 (2016).
Simmons, S. L., Bazylinski, D. A. & Edwards, K. J. South-seeking magnetotactic bacteria in the Northern Hemisphere. Science 311, 371–374 (2006).
Bray, D. (ed.) Cell Movements: From Molecules to Motility 2nd edn, 3–60 (Garland Science, 2001)..
Leander, B. S. & Keeling, P. J. Symbiotic innovation in the oxymonad Streblomastix strix. J. Eukaryot. Microbiol. 51, 291–300 (2004).
Rabus, R., Hansen, T. A. & Widdel, F. in The Prokaryotes (eds Rosenberg, E., DeLong, E. F., Lory, S., Stackebrandt, E. & Thompson, F.) 309–404 (Springer, 2013).
Schauder, R., Eikmanns, B., Thauer, R. K., Widdel, F. & Fuchs, G. Acetate oxidation to CO2 in anaerobic bacteria via a novel pathway not involving reactions of the citric acid cycle. Arch. Microbiol. 145, 162–172 (1986).
Hamann, E. et al. Environmental Breviatea harbour mutualistic Arcobacter epibionts. Nature 534, 254–258 (2016).
Monteil, C. L. et al. Accumulation and dissolution of magnetite crystals in a magnetically responsive ciliate. Appl. Environ. Microbiol. 84, e02865-17 (2018).
Schüler, D. The biomineralization of magnetosomes in Magnetospirillum gryphiswaldense. Int. Microbiol. 5, 209–214 (2002).
Jogler, C. et al. Cultivation-independent characterization of ‘Candidatus Magnetobacterium bavaricum’ via ultrastructural, geochemical, ecological and metagenomic methods. Environ. Microbiol. 12, 2466–2478 (2010).
Medlin, L., Elwood, H. J., Stickel, S. & Sogin, M. L. The characterization of enzymatically amplified eukaryotic 16S-like rRNA-coding regions. Gene 71, 491–499 (1988).
Lane, D. J. in Nucleic Acid Techniques in Bacterial Systematics (eds Stackebrandt, E. & Goodfellow, M.) 115–175 (John Wiley & Sons, 1991).
Pernthaler, J., Glockner, F. O., Schonhuber, W. & Amann, R. Fluorescence in situ hybridization (FISH) with rRNA-targeted oligonucleotide probes. Methods Microbiol. 30, 207–226 (2001).
Cole, J. R. et al. The Ribosomal Database Project (RDP-II): previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomy. Nucleic Acids Res. 31, 442–443 (2003).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Criscuolo, A. & Gribaldo, S. BMGE (Block Mapping and Gathering with Entropy): a new software for selection of phylogenetic informative regions from multiple sequence alignments. BMC Evol. Biol. 10, 210 (2010).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Antipov, D., Korobeynikov, A., McLean, J. S. & Pevzner, P. A. hybridSPAdes: an algorithm for hybrid assembly of short and long reads. Bioinformatics 32, 1009–1015 (2016).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 13, e1005595 (2017).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Vallenet, D. et al. MicroScope in 2017: an expanding and evolving integrated resource for community expertise of microbial genomes. Nucleic Acids Res. 45, D517–D528 (2017).
Huerta-Cepas, J. et al. Fast genome-wide functional annotation through orthology assignment by eggNOG-mapper. Mol. Biol. Evol. 34, 2115–2122 (2017).
Acknowledgements
This work was supported by a project from the French National Research Agency (ANR Tremplin-ERC, ANR-16-TERC-0025-01). N.M., D.F., C.L.M., D.P. and C.T.L. acknowledge support within the framework of a DFG-ANR project (ANR-14-CE35-0018). R.J.W. and C.T.L. received support from the Dumont d’Urville Science and Technology Programme (grant no. DDU-LVL1501) and New Zealand Ministry for Business, Innovation and Employment (grant no. LVLX1703). D.F. was supported by the Max Planck Society. C.T.L. also received support from the France-Berkeley Fund. P.L.-G. received support from the European Research Council Advanced Grant ‘ProtistWorld’ (grant no. 322669). The SEM facility at IMPMC was supported by funding from Région Ile de France (grant no. SESAME 2006 I-07-593/R); the transmission electron microscopy facility at IMPMC was supported by funding from Region Ile de France (grant no. SESAME 2000 E 1435). We thank A.-L. Monteil and N. Monteil for their help in collecting samples and S. Preveral for her help in confocal observation. Support for the confocal microscope was provided by the Région Provence Alpes Côte d’Azur, Conseil General of Bouches du Rhône, French Ministry of Research, CNRS and Commissariat à l’Energie Atomique et aux Energies Alternatives.
Author information
Authors and Affiliations
Contributions
C.L.M. and C.T.L. designed, performed and analysed most of the experiments. D.V., V.B., S.F. and C.C. designed and performed the genome sequencing and annotation. C.T.L., N.M., K.B. and M.F. performed the electron microscope analyses. E.V. performed the oxygen measurements. G.A. contributed to the identification of microorganisms. N.L. and D.F. contributed to the single-cell sorting. R.J.W. contributed to the sampling. C.L.M. and C.T.L. supervised the project with input from D.P. C.L.M. and C.T.L. wrote the first draft of the manuscript with input from P.L.-G. All authors contributed to data interpretation and editing of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes, Supplementary References, legend for Supplementary Video, Supplementary Figures 1–9 and Supplementary Tables 1 and 3.
Supplementary Table 2
List of proteins involved in metabolic pathways found in the draft genome of Ca. Desulfarcum epimagneticum strain CR-1.
Supplementary Videos
Supplementary Video 1.
Rights and permissions
About this article
Cite this article
Monteil, C.L., Vallenet, D., Menguy, N. et al. Ectosymbiotic bacteria at the origin of magnetoreception in a marine protist. Nat Microbiol 4, 1088–1095 (2019). https://doi.org/10.1038/s41564-019-0432-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0432-7
This article is cited by
-
Impact of direct physical association and motility on fitness of a synthetic interkingdom microbial community
The ISME Journal (2023)
-
Magnetotactic bacteria and magnetofossils: ecology, evolution and environmental implications
npj Biofilms and Microbiomes (2022)
-
Antibiotics affect migratory restlessness orientation
Journal of Ethology (2022)
-
Observations on a magnetotactic bacteria-grazing ciliate in sediment from the intertidal zone of Huiquan Bay, China
Journal of Oceanology and Limnology (2021)
-
Magnetotactic bacteria and magnetoreception
Journal of Oceanology and Limnology (2021)