Abstract
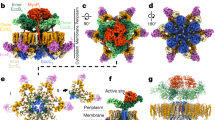
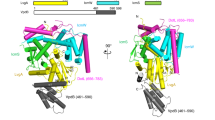
Legionella pneumophila survives and replicates inside host cells by secreting ~300 effectors through the defective in organelle trafficking (Dot)/intracellular multiplication (Icm) type IVB secretion system (T4BSS). Here, we used complementary electron cryotomography and immunofluorescence microscopy to investigate the molecular architecture and biogenesis of the Dot/Icm secretion apparatus. Electron cryotomography mapped the location of the core and accessory components of the Legionella core transmembrane subcomplex, revealing a well-ordered central channel that opens into a large, windowed secretion chamber with an unusual 13-fold symmetry. Immunofluorescence microscopy deciphered an early-stage assembly process that begins with the targeting of Dot/Icm components to the bacterial poles. Polar targeting of this T4BSS is mediated by two Dot/Icm proteins, DotU and IcmF, that, interestingly, are homologues of the T6SS membrane complex components TssL and TssM, suggesting that the Dot/Icm T4BSS is a hybrid system. Together, these results revealed that the Dot/Icm complex assembles in an ‘axial-to-peripheral’ pattern.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The subtomogram average of the Dot/Icm (DotF-sfGFP) complex that supports the findings of this study has been deposited in the Electron Microscopy Data Bank (EMDB) under the accession code: EMD-0566. All other density maps are available from the corresponding authors on request. The authors declare that all data supporting the findings of this study are available within the paper and its Supplementary Information documents.
References
Christie, P. J., Whitaker, N. & Gonzalez-Rivera, C. Mechanism and structure of the bacterial type IV secretion systems. Biochim. Biophys. Acta 1843, 1578–1591 (2014).
Costa, T. R. D. et al. Secretion systems in Gram-negative bacteria: structural and mechanistic insights. Nat. Rev. Microbiol. 13, 343–359 (2015).
Christie, P. J. & Vogel, J. P. Bacterial type IV secretion: conjugation systems adapted to deliver effector molecules to host cells. Trends Microbiol. 8, 354–360 (2000).
Guglielmini, J. et al. Key components of the eight classes of type IV secretion systems involved in bacterial conjugation or protein secretion. Nucleic Acids Res. 42, 5715–5727 (2014).
Segal, G., Feldman, M. & Zusman, T. The Icm/Dot type-IV secretion systems of Legionella pneumophila and Coxiella burnetii. FEMS Microbiol. Rev. 29, 65–81 (2005).
Nagai, H. & Kubori, T. Type IVB secretion systems of Legionella and other Gram-negative bacteria. Front. Microbiol. 2, 136 (2011).
Sutherland, M. C., Nguyen, T. L., Tseng, V. & Vogel, J. P. The Legionella IcmSW complex directly interacts with DotL to mediate translocation of adaptor-dependent substrates. PLoS Pathog. 8, e1002910 (2012).
Vincent, C. D. et al. Identification of the core transmembrane complex of the Legionella Dot/Icm type IV secretion system. Mol. Microbiol. 62, 1278–1291 (2006).
Vincent, C. D., Friedman, J. R., Jeong, K. C., Sutherland, M. C. & Vogel, J. P. Identification of the DotL coupling protein subcomplex of the Legionella Dot/Icm type IV secretion system. Mol. Microbiol. 85, 378–391 (2012).
Kwak, M.-J. et al. Architecture of the type IV coupling protein complex of Legionella pneumophila. Nat. Microbiol. 2, 17114 (2017).
Jeong, K. C., Ghosal, D., Chang, Y.-W., Jensen, G. J. & Vogel, J. P. Polar delivery of Legionella type IV secretion system substrates is essential for virulence. Proc. Natl Acad. Sci. USA 114, 8077–8082 (2017).
Ghosal, D., Chang, Y. W., Jeong, K. C., Vogel, J. P. & Jensen, G. J. In situ structure of the Legionella Dot/Icm type IV secretion system by electron cryotomography. EMBO Rep. 18, 726–732 (2017).
Dinh, T. & Bernhardt, T. G. Using superfolder green fluorescent protein for periplasmic protein localization studies. J. Bacteriol. 193, 4984–4987 (2011).
Chetrit, D., Hu, B., Christie, P. J., Roy, C. R. & Liu, J. A unique cytoplasmic ATPase complex defines the Legionella pneumophila type IV secretion channel. Nat. Microbiol. 3, 678–686 (2018).
Kubori, T. et al. Native structure of a type IV secretion system core complex essential for Legionella pathogenesis. Proc. Natl Acad. Sci. USA 111, 11804–11809 (2014).
Chandran, V. et al. Structure of the outer membrane complex of a type IV secretion system. Nature 462, 1011–1015 (2009).
Rivera-Calzada, A. et al. Structure of a bacterial type IV secretion core complex at subnanometre resolution. EMBO J. 32, 1195–1204 (2013).
Yerushalmi, G., Zusman, T. & Segal, G. Additive effect on intracellular growth by Legionella pneumophila Icm/Dot proteins containing a lipobox motif. Infect. Immun. 73, 7578–7587 (2005).
Nakano, N., Kubori, T., Kinoshita, M., Imada, K. & Nagai, H. Crystal structure of Legionella DotD: insights into the relationship between type IVB and type II/III secretion systems. PLoS Pathog. 6, e1001129 (2010).
Souza, D. P. et al. A component of the Xanthomonadaceae type IV secretion system combines a VirB7 motif with a N0 domain found in outer membrane transport proteins. PLoS Pathog. 7, e1002031 (2011).
Matthews, M. & Roy, C. R. Identification and subcellular localization of the Legionella pneumophila IcmX protein: a factor essential for establishment of a replicative organelle in eukaryotic host cells. Infect. Immun. 68, 3971–3982 (2000).
Nagai, H. & Roy, C. R. The DotA protein from Legionella pneumophila is secreted by a novel process that requires the Dot/Icm transporter. EMBO J. 20, 5962–5970 (2001).
Durand, E. et al. Structural characterization and oligomerization of the TssL protein, a component shared by bacterial type VI and type IVb secretion systems. J. Biol. Chem. 287, 14157–14168 (2012).
Nguyen, V. S. et al. Towards a complete structural deciphering of type VI secretion system. Curr. Opin. Struct. Biol. 49, 77–84 (2018).
Sexton, J. A., Miller, J. L., Yoneda, A., Kehl-Fie, T. E. & Vogel, J. P. Legionella pneumophila DotU and IcmF are required for stability of the Dot/Icm complex. Infect. Immun. 72, 5983–5992 (2004).
VanRheenen, S. M., Dumenil, G. & Isberg, R. R. IcmF and DotU are required for optimal effector translocation and trafficking of the Legionella pneumophila vacuole. Infect. Immun. 72, 5972–5982 (2004).
Zusman, T., Feldman, M., Halperin, E. & Segal, G. Characterization of the icmH and icmF genes required for Legionella pneumophila intracellular growth, genes that are present in many bacteria associated with eukaryotic cells. Infect. Immun. 72, 3398–3409 (2004).
Durand, E. et al. Biogenesis and structure of a type VI secretion membrane core complex. Nature 523, 555–560 (2015).
Chang, Y.-W., Shaffer, C. L., Rettberg, L. A., Ghosal, D. & Jensen, G. J. In vivo structures of the Helicobacter pylori cag type IV secretion system. Cell Rep. 23, 673–681 (2018).
Frick-Cheng, A. E. et al. Molecular and structural analysis of the Helicobacter pylori cag type IV secretion system core complex. mBio 7, e02001-15 (2016).
Low, H. H. et al. Structure of a type IV secretion system. Nature 508, 550–553 (2014).
Basler, M. Type VI secretion system: secretion by a contractile nanomachine. Phil. Trans. R. Soc. B 370, 20150021 (2015).
Kubori, T. & Nagai, H. The type IVB secretion system: an enigmatic chimera. Curr. Opin. Microbiol. 29, 22–29 (2016).
Bardill, J. P., Miller, J. L. & Vogel, J. P. IcmS-dependent translocation of SdeA into macrophages by the Legionella pneumophila type IV secretion system. Mol. Microbiol. 56, 90–103 (2005).
Tivol, W. F., Briegel, A. & Jensen, G. J. An improved cryogen for plunge freezing. Microsc. Microanal. 14, 375–379 (2008).
Zheng, S. Q. et al. UCSF tomography: an integrated software suite for real-time electron microscopic tomographic data collection, alignment, and reconstruction. J. Struct. Biol. 157, 138–147 (2007).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Agulleiro, J.-I. & Fernandez, J.-J. Tomo3D 2.0—exploitation of advanced vector extensions (AVX) for 3D reconstruction. J. Struct. Biol. 189, 147–152 (2015).
Nicastro, D. et al. The molecular architecture of axonemes revealed by cryoelectron tomography. Science 313, 944–948 (2006).
The UniProt Consortium, T. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 46, 2699 (2018).
Emanuelsson, O., Brunak, S., von Heijne, G. & Nielsen, H. Locating proteins in the cell using TargetP, SignalP and related tools. Nat. Protoc. 2, 953–971 (2007).
Käll, L., Krogh, A. & Sonnhammer, E. L. L. A combined transmembrane topology and signal peptide prediction method. J. Mol. Biol. 338, 1027–1036 (2004).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Yang, J. et al. The I-TASSER Suite: protein structure and function prediction. Nat. Methods 12, 7–8 (2015).
Xu, D. & Zhang, Y. Ab initio protein structure assembly using continuous structure fragments and optimized knowledge-based force field. Proteins 80, 1715–1735 (2012).
Schneidman-Duhovny, D., Inbar, Y., Nussinov, R. & Wolfson, H. J. PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res. 33, W363–W367 (2005).
Acknowledgements
We thank R. Isberg (Tufts University, Medford, MA, USA) for antibodies that recognize DotF and DotH, E. Buford for technical assistance and P. Levin (Washington University, St Louis, MO, USA) for assistance with deconvolution microscopy. ECT data were recorded at the Beckman Institute Resource Center for Transmission Electron Microcopy at Caltech and the cryo-EM facility at Janelia Research Campus. We thank C. Oikonomou for the creation of the domain maps and for help structuring and revising the text. We also recognize E. Darwin for key suggestions and critical appraisal of this manuscript. This work was funded by the NIH grant R01AI127401 to G.J.J. and the NIH grant R01AI48052 to J.P.V.
Author information
Authors and Affiliations
Contributions
D.G., K.C.C.J., J.P.V. and G.J.J. conceived the project. K.C.C.J., J.P.V. and J.G. constructed and characterized the L. pneumophila expression plasmids and strains. K.C.J. and J.P.V. collected the immunofluorescence data. D.G. collected the tomography data. D.G., K.C.C.J., J.P.V., G.J.J., Y.-W.C. and L.T. analysed the data. A.G. made Supplementary Video 1. D.G., J.P.V., K.C.C.J. and G.J.J. wrote the manuscript with input from other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–32, Supplementary Video Legend, Supplementary Tables 1 and 2, and Supplementary References.
Supplementary Video 1
3D representation of the Dot/Icm complex showing a windowed secretion chamber (salmon, DotH; grey, DotD; green, DotK; and cyan, DotC), wings (yellow, DotF), a secretion channel (red, DotG) and the top-view of the complex. Cytoplasmic components are not shown. In this 3D representation, IcmF, IcmX and DotA are not visible. The blue structure below the secretion channel represents the UF seed that initiates polar Dot/Icm complex assembly.
Rights and permissions
About this article
Cite this article
Ghosal, D., Jeong, K.C., Chang, YW. et al. Molecular architecture, polar targeting and biogenesis of the Legionella Dot/Icm T4SS. Nat Microbiol 4, 1173–1182 (2019). https://doi.org/10.1038/s41564-019-0427-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0427-4
This article is cited by
-
Contributions of lipopolysaccharide and the type IVB secretion system to Coxiella burnetii vaccine efficacy and reactogenicity
npj Vaccines (2021)
-
Architecture of the outer-membrane core complex from a conjugative type IV secretion system
Nature Communications (2021)
-
Structural basis for effector protein recognition by the Dot/Icm Type IVB coupling protein complex
Nature Communications (2020)
-
In vivo structure of the Legionella type II secretion system by electron cryotomography
Nature Microbiology (2019)