Abstract
Rift Valley fever virus (RVFV) is a mosquito-borne pathogen that causes substantial morbidity and mortality in livestock and humans. To date, there are no licensed human vaccines or therapeutics available. Here, we report the isolation of monoclonal antibodies from a convalescent patient, targeting the RVFV envelope proteins Gn and Gc. The Gn-specific monoclonal antibodies exhibited much higher neutralizing activities in vitro and protection efficacies in mice against RVFV infection, compared to the Gc-specific monoclonal antibodies. The Gn monoclonal antibodies were found to interfere with soluble Gn binding to cells and prevent infection by blocking the attachment of virions to host cells. Structural analysis of Gn complexed with four Gn-specific monoclonal antibodies resulted in the definition of three antigenic patches (A, B and C) on Gn domain I. Both patches A and B are major neutralizing epitopes. Our results highlight the potential of antibody-based therapeutics and provide a structure-based rationale for designing vaccines against RVFV.
Similar content being viewed by others
Main
Rift Valley fever virus (RVFV), the causative agent of zoonotic Rift Valley fever (RVF), is one of the most serious arbovirus threats to both human and animal health. Since its identification in 19311, the periodical RVF epizootic has infected millions of ruminants and resulted in major losses in the livestock industry2,3. Meanwhile, from 2000 to June 2018, 4,830 cases of severe RVF in humans were reported to the World Health Organization, including 967 related deaths, with a case fatality rate of approximately 20.0%4. Although formerly restricted to circulation in the African continent, RVFV spread, causing an epidemic in the Kingdom of Saudi Arabia and Yemen in 20005. In 2016, China reported its first imported case, a patient returning from Angola, a non-epidemic country6. Many strategies for RVFV vaccines have been developed since the virus was first characterized7. However, in the 70 years since its characterization, no licensed vaccine preparations nor therapeutics have been approved for use in humans7. Therefore, effective antivirals are urgently needed.
RVFV belongs to the genus Phlebovirus under the family Phenuiviridae in the order Bunyavirales8. The virus, a mosquito-borne arbovirus, contains a tripartite RNA genome, consisting of large, medium (M) and small segments9. Two glycoproteins on the viral envelope, Gn and Gc, which are important components for virus entry and membrane fusion10,11,12, are encoded by the M segment. The structure of RVFV Gn exhibits similar topology as its counterpart of severe fever with thrombocytopenia syndrome virus (SFTSV), another Phlebovirus member13,14. It is presumed that, on the RVFV virion in its pre-fusion state, Gc lines the inner half of the glycoprotein shell and forms an interface with Gn domain II (DII). The Gc fusion loop is shielded from solvents by Gn DI and DII, both of which are externally exposed13,14,15. Considerable data from vaccine studies suggest that neutralizing antibodies elicited by vaccines containing Gn alone or in combination with Gc are sufficient to confer protection against RVFV challenge in mice and livestock16,17,18,19,20,21.
Here, we isolated eight Gn-specific monoclonal antibodies and one Gc-specific monoclonal antibody from a convalescent RVF patient. The results showed that Gn monoclonal antibodies blocked the binding of Gn on the virions to susceptible cells, thus neutralizing viral infection. Additional structural analysis of Gn complexed with monoclonal antibodies revealed two neutralizing hotspots, targeted by most of the isolated Gn monoclonal antibodies. Our results strongly indicate that human infection by RVFV stimulates a robust humoral immune response, and suggest the high potential of antibody-based therapeutics. Moreover, the structural studies presented here shed light on the assembly of virions and provide a structure-based rationale for the design of future RVFV-specific vaccines.
Results
Antibody isolation and binding characterization
We applied both soluble Gn and Gc as baits to isolate specific memory B cells from an RVF convalescent plasma6, which displayed a strong reaction to both Gn and Gc (Supplementary Figs. 1 and 2). Then, the variable regions of IgG antibodies from each cell were determined (Supplementary Table 1). The nine monoclonal antibodies exhibited low somatic hypermutation (SHM) rates compared to their germline genes, with nucleotide identities higher than 97% for the variable regions in both the heavy chain (VH) and the light chain (VL). Additionally, both R13 and R17 incorporated the VL identical to their germline ancestors. R22 was a germline monoclonal antibody without SHM mutations. Interestingly, two pairs of monoclonal antibodies (R12 and R13; R16 and R17) incorporated light chains (L) with the same V and J alleles (Supplementary Tables 1 and 2). Next, each monoclonal antibody was converted to an IgG1 isotype for further experiments.
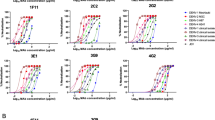
Via fluorescence-activated cell sorting (FACS), nine antibodies bound to cells expressing the M gene (Fig. 1a). However, when Gn and Gc were separated, eight monoclonal antibodies, namely R4, R12, R13, R15, R16, R17, R19 and R22, exclusively interacted with cells containing Gn. Only R5 bound to Gc-expressing cells (Fig. 1a). Consistently, the monoclonal antibodies displayed interactions with either Gn or Gc when assessed by surface plasmon resonance (SPR). The binding affinities between Gn and monoclonal antibodies varied, ranging from tens (R15, R22, R19 and R12) to hundreds of nanomolar (R4, R13, R16 and R17). The Gc monoclonal antibody R5 manifested a Kd of approximately 120 nM (Fig. 1b and Supplementary Fig. 3). No binding was observed between the isolated monoclonal antibodies with the counterparts in SFTSV (Fig. 1b).
a, A flow cytometric assay of the indicated monoclonal antibodies involved in specific antigen binding. HEK293T cells were transfected with plasmids containing coding regions for eGFP-fused M (orange), Gn (red) and Gc (green), respectively, and then tested with the indicated monoclonal antibody. Cells transiently expressing M–eGFP stained with secondary antibodies (anti-hIgG/APC) were used as a negative control (black). b, An SPR assay characterizing the specific binding between the indicated monoclonal antibodies and specific antigens, including RVFV Gn (red), RVFV Gc (green), SFTSV Gn (blue) and SFTSV Gc (cyan). The black lines indicate the fitting curves. The calculated Kd for each monoclonal antibody binding to the specific antigen is listed below the name of the monoclonal antibody. The data are representative of three independent experiments.
Neutralization activities in vitro and protection efficacies in vivo against RVFV infection
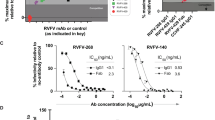
Using a previously reported FACS-based assay with Vero cells22, the Gn monoclonal antibodies exhibited much more intense neutralizing potencies than the Gc monoclonal antibody R5 (Fig. 2 and Supplementary Data Set). The median inhibitory concentration (IC50) of R5 was calculated to be 1,370 ± 530 ng ml−1. In contrast, R15 and R16 displayed high neutralizing activities, with IC50 at 0.53 ± 0.25 ng ml−1 for R15 and 0.29 ± 0.09 ng ml−1 for R16 (Fig. 2c). The IC50 values of both R12 and R17 increased to several nanograms per millilitre (Fig. 2d). R4, R13, R19 and R22 displayed values in the range of tens of nanograms per millilitre (Fig. 2e).
a–e, The neutralization activity of the indicated monoclonal antibody was evaluated by a FACS-based neutralization test using Vero cells. The number in parentheses following the name of the monoclonal antibody represents the mean ± s.d. of the IC50 value (ng ml−1) of n = 3 independent experiments. The IC50 was calculated using the equation of log[inhibitor] versus response in GraphPad Prism 5. The curves are representatives of three independent experiments. Zika monoclonal antibody Z3L1 was used as a negative control (a). RVFV monoclonal antibodies were grouped according to potency: IC50 at micrograms per millilitre (b), sub-nanogram per millilitre (c), several nanograms per millilitre (d) and tens of nanograms per millilitre (e). f–i, For post-exposure treatment, BALB/c mice were challenged with RVFV 1 day before administration of the same volume of the indicated monoclonal antibodies (10 mg kg−1) or PBS (marked as Virus). The PBS group of mice were administered PBS instead of virus or monoclonal antibodies. Mouse survival (f) and body weight (g) were recorded daily. For pre-exposure treatment, mice were injected with the indicated monoclonal antibody (10 mg kg−1) or PBS (Virus), followed by challenge with RVFV 1 day later. Mice without inoculation with monoclonal antibodies or virus were marked as PBS. Mouse survival (h) and body weight (i) were recorded daily. Sample sizes of mice in both post-exposure (left panel) and pre-exposure (right panel) treatments are indicated. The survival curves were generated with a log-rank (Mantel–Cox) test using GraphPad Prism 5. Body weight loss curves were also generated using GraphPad Prism 5.
The protection efficacies of monoclonal antibodies against RVFV infection were evaluated in a mouse model23 (Supplementary Fig. 4). In the post-exposure setting, mice challenged with RVFV died within 5 days post-infection. All mice in the eight Gn monoclonal-antibody-treated groups survived, except for one, injected with R22 (n = 4), which died at 11 days post-infection (Fig. 2f). Compared with the PBS group, Gn monoclonal-antibody-treated mice displayed similar body weight increases over the 14 days of observation (Fig. 2g). In contrast, in the Gc monoclonal antibody R5-treated group, only one survived (n = 5, Fig. 2f). Similarly, Gn monoclonal antibodies prevented RVFV infection in the pre-exposure setting. All Gn monoclonal-antibody-treated mice survived, whereas four mice injected with R5 died (n = 5, Fig. 2h,i).
Neutralizing mechanisms of Gn monoclonal antibodies
Viral attachment to susceptible cells is the initial step of viral infection24,25. We prepared a Gn tetramer to amplify the interaction signals and used them to stain Huh7 cells26. Pre-incubation of the Gn tetramer with the Gn monoclonal antibodies, not the Gc monoclonal antibody R5, before staining Huh7 cells can prevent fluorescence shift (Fig. 3a). Following a reported procedure with some modifications27 (Supplementary Fig. 5), we found that all RVFV monoclonal antibodies associated with both Huh7 and Vero cells, if pre-incubated with the virus (Supplementary Fig. 6), suggesting that all RVFV monoclonal antibodies could bind to the virions. Compared with cells incubated with RVFV and tested with the indicated monoclonal antibody (marked as Virus binding in Fig. 3b), pre-incubation of the virus with each Gn monoclonal antibody before binding to Vero cells (marked as Indicated monoclonal antibody in Fig. 3b) led to reduced fluorescence intensity, whereas the Gc monoclonal antibody R5 displayed no effect on the shift (Fig. 3b and Supplementary Fig. 5). In addition, the blocking effect of Gn monoclonal antibodies was observed in a concentration-dependent manner (Supplementary Fig. 7).
a, The Gn tetramer was prepared and tested by flow cytometry for its interaction with Huh7 cells, with or without the indicated RVFV monoclonal antibody. The MHC tetramer was used as a negative control for the Gn tetramer. Z3L1 was used as a negative control for the RVFV monoclonal antibodies. The data were processed with FlowJo software and are representative of two independent experiments. b, RVFV virions (5 × 104 PFUs) were incubated with Vero cells (2 × 105) for 30 min at 4 °C, and the bound virions were detected using the indicated RVFV monoclonal antibody (200 μg ml−1) by flow cytometry (cyan). For blocking evaluation, the virus was pre-incubated with the indicated monoclonal antibody at 37 °C for 1 hour before its addition to Vero cells. The bound virions were then detected (orange). Cells interacting with the indicated monoclonal antibody and anti-hIgG/APC were used as negative controls to gate the cell (grey). The data are representative of two independent experiments.
Molecular determinants of neutralizing monoclonal antibodies
To delineate the molecular basis of Gn monoclonal antibodies neutralizing RVFV infection, we solved the crystal structures of Gn complexed with the Fab form of the monoclonal antibodies R12, R13 and R15 and the scFv of R17. The resolutions for the complex structures are 2.7 Å for R12, 2.0 Å for R13, 2.4 Å for R15, and 3.2 Å for R17, respectively (Supplementary Table 3). Gn is composed of three domains, DI, DII and DIII13,14. Interestingly, each monoclonal antibody applies a similar perpendicular angle to interact with equal molar Gn (Fig. 4). In addition, all four monoclonal antibodies bind to DI, albeit R12 also interacts with Lys 395 in DIII (Supplementary Table 4). With the exception of complementarity-determining region 1 (CDR1) in the R13 heavy chain (H), all CDRs from both heavy and light chains of R12, R13, R15 and R17 are involved in direct Gn interaction. In the four complexes, the buried surfaces on R12, R13, R15 and R17 are 672.3 Å2, 668.3 Å2, 622.7 Å2 and 793.5 Å2, respectively. Meanwhile, Gn is buried 703.1 Å2, 719.6 Å2, 713 Å2 and 830.3 Å2 by the monoclonal antibodies R12, R13, R15 and R17, respectively.
The overall complex structures of Gn with the indicated monoclonal antibodies are represented in cartoon form in the upper panel. The lower panel delineates the epitope of the indicated monoclonal antibodies on Gn DI. Magenta, blue and green indicate domains I, II and III of RVFV Gn, respectively. For each monoclonal antibody, cyan indicates the heavy chain and light blue represents the light chain. CDRs in the heavy chain are shown in gold. CDRs in the light chain are shown in gold. Accordingly, residues in Gn that form contacts with the heavy or light chains are also highlighted as orange and yellow-orange sticks in the lower panel. The red sticks indicate the residues interacting with both heavy and light chains.
Further scrutiny of the binding interface revealed that the epitopes of R12, R13 and R15 overlap substantially (Figs. 4 and 5a−i). R12H forms a patch of electrostatic interactions with Gn residues in ƞ2α1 (Asp 176) and residues adjacent to ƞ2α1 (Lys 164, His 166, Tyr 168 and Lys 180). R12L interacts with Gn ƞ2α1, with Gln 174 and Lys 294 in the centre (Fig. 5b,c). Consistently, a single mutated residue of either D176A, Q174A, K180A or K294A in Gn decreased or destroyed the association with R12 (Supplementary Fig. 8). Here, we designated the region including ƞ2α1 (173TQEDATCK180) and K294 as antigenic patch A. Deleting antigenic site A (Gn-ΔA) abolished Gn binding by R12 (Fig. 6a,b). Gn residues in the β3/β4 loop (271CPPK274; named antigenic patch B) also confer several van der Waals contacts with R12L (Supplementary Table 4).
a–l, Cartoon representation of the interface between Gn and the indicated monoclonal antibody. Gn DI is marked in magenta. Antigenic site A is highlighted in cyan, antigenic site B in blue, and antigenic site C in green. Orange indicates the heavy chain and yellow-orange represents the light chain. Residues in monoclonal antibodies responsible for the interaction with Gn are displayed as grey sticks. The figures in the four columns, from left to right, indicate the detailed interaction of Gn with R12, R13, R15 and R17, respectively. In each row, the pictures indicate the interface of Gn with the indicated monoclonal antibody (the upper panel), and the heavy (the middle panel), and light chains (the lower panel). The black dashed lines suggest the hydrogen bonds probably formed between the indicated atoms.
a, A flow cytometric assay of the indicated monoclonal antibodies involved in mutated Gn binding. HEK293T cells were transfected with plasmids containing eGFP-fused Gn (red), Gn without epitope A (Gn-ΔA in cyan), Gn without epitope B (Gn-ΔB in blue) and Gc (grey), respectively, and were then tested for binding with the indicated monoclonal antibody. Cells transiently expressing M–eGFP stained with secondary antibodies (anti-hIgG/APC) were used as a negative control and are indicated by black filling. The data are representative of two independent experiments. b, Distribution of epitopes A (cyan), B (blue) and C (green) on Gn as a surface representation. Gn monoclonal antibodies are grouped on the basis of their critical targets on Gn and are highlighted with arrows. c, Antigenic sites docking to the cryogenic electron microscopy structure of RVFV virions. The Gn structure (PDB no. 5Y0W)13 is docked to a model of the hexameric Gn–Gc on the RVFV virions (PDB no. 6F9C)14. Grey indicates the underlined Gc. Gn DI is displayed in magenta, DII in slate and DIII in pale green. Antigenic sites A, B and C are highlighted in cyan, blue and green, respectively. A previously reported neutralizing epitope located on DIII is shown in red35.
R13L is the germline sequence of R12L (Supplementary Table 2). It adopts the same interaction mode as R12L to associate with Gn (Fig. 5c,f). Accordingly, the root-mean-square deviation between Gn with R12L and R13L reaches 0.388 (338 Cα atoms). Compared with the germline R13L, R12L acquired Ser 31 through SHMs, which optimized its function (Supplementary Table 4 and Supplementary Fig. 9). Concomitantly, the long CDR3 (17 residues) in R13H interacts hydrophilically with Gn α1 helix (176DAT178), Lys 180 and Lys 294 (Fig. 5e). Introduction of either Q174A, D176A, K180A or K294A decreased the binding to the monoclonal antibody R13 (Supplementary Fig. 8). Gn-ΔA lost interaction with R13 (Fig. 6a,b).
R15, composed of two distinct chains, also targets antigenic patches A and B (Figs. 4 and 5g–i). A single residue mutation in Gn, such as Q174A, D176A, K180A or K294A, led to the varied loss of binding ability to R15 (Supplementary Fig. 8). Gn-ΔA did not support R15 binding (Fig. 6a,b). Additionally, the introduction of both P273A and K274A (Gn-ΔB) to Gn also abolished binding to the monoclonal antibody R15 (Fig. 6a,b).
R17 mainly interacts with antigenic patch B and the β1 strand, which was designated as antigenic patch C (223KADPPSCD230) (Figs. 4 and 5j–l). Through hydrophilic interactions, R17 anchors both terminal regions (Lys 223 and Asp 225 in the amino-terminal region, Asp 230 at the carboxy-terminal region) and the central region of the β1 strand (Ser 228) (Fig. 5k). The Gn mutant with D225A completely lost its ability to bind to R17 (Supplementary Fig. 8). Simultaneously, R17 also interacts with antigenic patch B (Fig. 5l) and the deletion of this region (Gn-ΔB) destroyed binding to R17. R17 maintained association with Gn-ΔA (Fig. 6). HEK293T cells failed to express Gn without the β1 strand, indicating its importance for the structural stability of Gn.
By docking the Gn monoclonal antibodies on the RVFV virions (PDB no. 6F9B, 6F9C and 6F9F)14, we found that all four monoclonal antibodies perpendicularly bind to the virions (Fig. 6c and Supplementary Fig. 10). The shortest distances between monoclonal antibodies associating with adjacent Gns are 51.9 Å for R12, 67.5 Å for R13, and 40.3 Å for R15, respectively. Notably, R17 scFv, on certain positions of the virions, is indicated to sterically hinder the association of others (Supplementary Fig. 10d,e). R17 Fabs were simulated utilizing R12 Fabs and measured to be as short as approximately 20 Å for the ones without clash on the virions (Supplementary Fig. 10).
Both antigenic patches A and B are hotspot neutralizing targets
We further tested the ability of the monoclonal antibodies without structural information to stain cells expressing different Gn mutants. R4 failed to bind to cells expressing Gn-ΔB. Although R4 maintained its interaction with Gn-ΔA, D176A or K180A decreased this association. Residues in patch C did not exert an effect on binding to R4. R16 could not bind to Gn-ΔA or Gn-ΔB, but normally interacted with Gn mutants in patch C. R19 associated with Gn-ΔB, but not Gn-ΔA. Moreover, either D225A or D230A weakened the interaction. Although R22 could bind to both Gn-ΔA and Gn-ΔB, a single mutation of either D176A, K180A, D225A or D230A could reduce the level of interaction with R22 (Fig. 6 and Supplementary Fig. 8).
Thus, according to the critical patches that affect the Gn–monoclonal antibody interaction, the eight Gn monoclonal antibodies can be divided into four groups. Group 1 is that for which antigenic patch A, not B or C, is pivotal for the interaction, and contains R12, R13 and R19. For group 2 monoclonal antibodies, containing R15 and R16, both antigenic patches A and B, but not C, are critical for binding. For monoclonal antibodies in group 3, containing R4 and R17, antigenic patch B, not A, plays a key role. In addition, antigenic patch C also determines the association with R17 (Fig. 6). The essential targets of R22, the only monoclonal antibody of group 4, require further study. Moreover, none of the Gn monoclonal antibodies or the Gc monoclonal antibody R5 could recognize the linearized epitopes, as indicated by western blot experiments (Supplementary Fig. 11).
Notably, residues that are equivalent to RVFV patches A, B and C among Phlebovirus are highly diverse (Supplementary Fig. 12). Further sequence analyses of 108 strains (see Methods) indicate that residues constituting patch B are completely conserved. Three strains carry T173L28, and one strain carries E175G29. Another strain carries K294E in patch A as well as D230N in patch C (Supplementary Fig. 13). Using flow cytometry analysis, Gn with T173L exerted no effect on binding to R17, but decreased or destroyed the association with other Gn monoclonal antibodies (Supplementary Fig. 14). Both E175G and K294E-D230N led to decreased binding to Gn monoclonal antibodies to various degrees (Supplementary Fig. 14).
Discussion and conclusion
In this study, we characterized human memory B cells encoding monoclonal antibodies targeting RVFV Gn and Gc. It seems that Gn is the immune-dominant antigen in human RVFV infections. Since the same primers were used to sequence genes in the isolated cells, there is little possibility that this discrepancy resulted from sequencing biases. However, we cannot exclude the possibility that the soluble Gc, which was used for screening, might exist in different conformations from the one that appears on the surface of RVFV14, and was thus missed by relevant Gc-targeting antibodies. Further studies are needed using labelled virus or virus-like particles to investigate the composition of Gn-specific and Gc-specific monoclonal antibodies in additional human patients. Among the nine monoclonal antibodies, seven incorporated λ light chains. This might result from the different efficiencies of the primers used in PCR. Notably, similar λ light chain-biased antibody responses were also reported in ferrets and humans infected with influenza virus30,31. Further studies are needed to determine whether this phenomenon is primer-dependent or case-dependent.
The isolated Gn monoclonal antibodies manifested relatively low binding affinities to Gn. Consequently, complete blocking of the interaction between the Gn tetramer and Huh7 by these Gn monoclonal antibodies required that they be used in high concentrations. The relatively low SHM rates in the isolated monoclonal antibodies might explain their low binding affinities and imply that this infection was probably the primary infection in this patient. In fact, the patient was a temporary worker in Angola, but originated from China, where RVFV has not been recorded before. Despite having low maturation, Gn monoclonal antibodies demonstrate high neutralizing activities, preventing RVFV infection with an IC50 in the sub-nanogram per millilitre range, based on a previously reported FACS-based assay with Vero cells. This method is widely used for evaluation of the neutralizing activities of flavivirus monoclonal antibodies22,32 and respiratory syncytial virus monoclonal antibodies33. However, further studies are needed to investigate the comparability of this method with the standard plaque reduction neutralization test (PRNT80 or PRNT50) with overlay, in terms of RVFV infection. Although R5, which targets Gc, showed relatively low biological activity, the antibody still prevents RVFV infection at high concentrations, helping to clear the virus. Accordingly, vaccines containing both Gn and Gc stimulate sera with higher neutralizing activities than vaccines containing Gn alone34.
The Gn monoclonal antibodies inhibit the attachment of RVFV virions to the cells in a monoclonal antibody concentration-dependent manner. Interestingly, R15 and R16 seem to have the strongest blocking effects. The blocking effects of R12 and R17 are probably weaker than those of R15 and R16, but stronger than those of R4, R13, R19 and R22, which are consistent with their neutralizing activities.
Through structural analysis of Gn–monoclonal antibody complexes, the monoclonal antibodies R12, R13 and R15 are demonstrated to bind to highly overlapping epitopes. Interestingly, their binding affinity to Gn seems to be correlated with their neutralizing activities. R15, which displays the highest binding affinity to Gn, confers the best neutralization. R13 exhibits the lowest binding affinity with the lowest inhibitory effect. R12 is in the middle. Additionally, the decreased interaction between R12H–13L also results in a reduced preventive effect, as compared with the parental R12. Thus, for monoclonal antibodies targeting similar epitopes, higher binding affinities are indicated to be linked to better neutralizing activities. Notably, R17 displays the weakest interaction with Gn but inhibits RVFV infection at the nanogram per millilitre level. Further docking of this monoclonal antibody to the RVFV virion indicates approximately 20 Å as the shortest distance between R17 associating with adjacent Gns without steric hindrance. Based on the reported structures (PDB nos. 1IGY, 1IGT and 1HZH), the distance between two Fabs in one IgG is approximately 40–52 Å, suggesting a high possibility that one R17 molecule interacts with more than one Gn molecule on the virion, thereby resulting in high neutralizing activities. Additionally, one R12 or R15 may also bind two Gns. Although the distance between the two most adjacent R13 Fab regions exceeds 60 Å, we cannot exclude the possibility that one R13 binds to two Gns, due to the flexibility of IgG.
Further alanine screening indicates that patch A is critical for five of the monoclonal antibody interactions (R12, R13, R19, R15 and R16), while patch B is essential for four (R15, R16, R4 and R17). R17 also depends on patch C. Additionally, other residues, including Ser 291, Leu 292 and Lys 293, also contribute to the interaction between Gn and the monoclonal antibodies. Further studies are still needed to validate the roles of either patches A and B, or other regions, for the neutralization process, by using live RVFV virus possessing the mutations. The isolation of multiple monoclonal antibodies targeting similar regions suggests that these regions are truly hotspot neutralizing targets. There are several strains containing substitutions in individual residues within patches A and C. These naturally occurring variations could reduce the interaction with all Gn monoclonal antibodies to an extent, highlighting the necessity to identify monoclonal antibodies targeting other epitopes.
A previous report has indicated that DIII also contains neutralizing epitopes, including those targeted by murine monoclonal antibody 4-39-CC35. Recently, monoclonal antibody 4-5, an SFTSV-specific neutralizing antibody, was also determined to bind to α6 on SFTSV DIII13. Due to the diverse residues within the epitope, monoclonal antibody 4-5 does not interact with RVFV Gn. The RVFV Gn monoclonal antibodies in this study do not bind to SFTSV Gn either. Usually, accessible epitopes on the virions stimulate neutralizing monoclonal antibodies more easily than do cryptic ones. The distribution pattern of neutralizing epitopes on both Phlebovirus Gns supports the model recently proposed for Gn organization13,14. The fact that antibodies binding to Gn DI can block the interaction between soluble Gn protein and host cells implies that Gn DI is probably responsible for receptor binding.
In summary, we isolated eight Gn-specific monoclonal antibodies and one Gc-specific monoclonal antibody from a patient infected with RVFV. The Gn monoclonal antibodies exert high neutralizing activities in vitro and complete protection against RVFV challenge in vivo, and are worthy of future development as biological therapeutics and prophylaxis.
Methods
Isolation of antigen-specific single memory B cells
A patient returning from Angola to China was diagnosed with RVF, representing the first imported case of RVFV infection in China6. On day 46 of the disease, blood was taken after written consent using guidelines approved by the ethics committee of the Beijing Ditan Hospital. Peripheral blood mononuclear cells were isolated following the manufacturer’s instructions (GE Healthcare) before being subjected to cell sorting as previously reported22,36. Briefly, peripheral blood mononuclear cells were incubated with both His-tagged Gn and His-tagged Gc at 100 nM before staining with anti-CD3, anti-CD16, anti-CD235a, anti-CD19, anti-CD38, anti-CD27 and anti-His. Antigen-specific memory B cells, which were CD3−, CD16−, CD235a−, CD38−, CD19+, CD27+, hIgG+ and His+ were sorted into 96-well PCR plates with one cell per well. Flow cytometric analysis and cell sorting were performed on a BD FACSAria ΙΙΙ flow cytometer (BD Biosciences) and the data were analysed using FlowJo.
Determination of Ig variable region sequences from sorted single B cells by reverse transcription-PCR
The genes encoding Ig VH and VL chains were amplified by reverse transcription and nested PCR using an optimized version of a previously reported method, and the primers used were similar to those reported earlier37. Briefly, synthesis of Ig VH and VL was performed in 96-well PCR plates containing sorted single B cells. The reverse transcription reaction was carried out following the manufacturer’s instructions (Invitrogen). The primers used for reverse transcription were a mixture, including primers specific for IgA, IgD, IgE, IgG, IgM, Igκ and Igλ constant regions. After complementary DNA synthesis, VH, Vκ and Vλ genes were amplified separately by two rounds of PCR in 96-well PCR plates. As a template, 2 μl of reverse transcription reaction product was used in the first-round of PCR, 1 μl product of which was used to further amplify variable genes in the nested second round of PCR. Samples of VH, Vκ and Vλ chain PCR products were analysed on 1.2% agarose gels. Bands at about 400 bp were cut, purified by the Gel Extraction Micro Kit (CWBIO) and sequenced. Sequences were analysed using the IMGT information system38 to identify variable region gene segments and somatic mutations.
Gene construction
The PCR-amplified and sequence-confirmed VH, Vκ and Vλ were separately linked to the indicated constant region by overlapping PCR. These constant regions included the IgG1 constant region (GeneBank Accession no. BC041037) for the heavy chain, the Ig kappa constant region (GeneBank accession no. BC073791) for the κ chain and the Ig lambda constant region (GeneBank Accession no. BC073769) for the L fragment. The Ig leader sequence (METDTLLLWVLLLWVPGSTGD) coding region was introduced to the 5′ end of the variable region using primers. The overlapping product containing the leader sequence, variable region and constant region was ligated into the pCAGGS plasmid (Addgene) for monoclonal antibody expression, using EcoRI and XhoI restriction sites for IgG and Igλ cloning, and SacI and XhoI restriction sites for Igκ cloning. The coding regions of the Gn monoclonal antibodies were also constructed in VL-(GGGGS)4-VH form and cloned into the pET21a expression vector (Invitrogen) through NheI and XhoI restriction sites.
The constructs used for the expression and purification of RVFV and SFTSV glycoproteins were the same as those previously reported12,13. Specifically, coding regions for RVFV Gn (M residues 154–469, GenBank accession number: JQ068143.1), RVFV Gc (M residues 691–1119, GenBank accession number: JQ068143.1), SFTSV Gn (M residues 20–452, GenBank accession number: JF906057.1) and SFTSV Gc (M residues 563–996, GenBank accession number: JF906057.1) were individually cloned into the EcoRI and XhoI restriction sites of the pFastBac1 vector. For each protein, an N-terminal gp67 signal peptide39 and a C-terminal hexa-His were added to facilitate protein secretion and purification.
For Gn tetramer preparation, coding regions for RVFV Gn were cloned into the KpnI and XhoI restriction sites of pCAGGS. The sequential biotinylation tag and His tag were linked to the 3′ end of Gn by a GS linker.
The constructs of pEGFP-M (M residues 1–1197, GenBank accession number: JQ068143.1), pEGFP-Gn (M residues 154–616, GenBank accession number: JQ068143.1) and pEGFP-Gc (M residues 635–1197, GenBank accession number: JQ068143.1) that were used for FACS were cloned by insertion of the indicated coding regions into pEGFP-N1 at the restriction sites of XhoI and BamHI. Thus, each protein was expressed as a fusion protein with GFP. For pEGFP-Gn and pEGFP-Gc, the MMKVIWFSSLICFVIQCSG sequence was used as a signal peptide and linked to the 5′ end of both genes. Gns incorporating mutations were designed on the basis of the pEGFP-Gn vector. Residues in antigenic patch A (173TQEDATCK180 and Lys 294) were mutated to alanine, except for Cys 179, to maintain the disulfide bond between Cys 179–Cys 188 (Gn-ΔA). P273A and K274A were introduced to Gn-ΔB to delete antigenic patch B. Residues in antigenic patch C (223KADPPSCD230) were also mutated to 223AAAAAACA230 (Gn-ΔC). We constructed eGFP-fused Gn mutants that contained single-residue mutations, including Q174A, D176A, K180A, K294A, D225A, S228A or D230A, to investigate the minimum epitopes in patches A and C. T173L, E175G and K294E-D230N were also designed on the basis of the pEGFP-Gn vector.
Protein expression and purification
The proteins used for crystallization and SPR experiments were prepared using the Bac-to-Bac baculovirus expression system (Invitrogen). Briefly, recombinant bacmids were prepared and then transfected into sf9 cells to obtain a baculovirus stock, which was then used to infect High5 cells for protein expression. Target proteins in the supernatant were sequentially collected, affinity-purified by a HisTrap HP column (GE Healthcare) and then subjected to gel filtration (Superdex 200 column, GE Healthcare).
The monoclonal antibodies were expressed in HEK293T cells by transient transfection. Supernatant containing monoclonal antibodies was collected and passed through a HiTrap Protein A FF (1 ml, GE Healthcare) column. The bound protein was detached from the column using 0.1 M sodium citrate, pH 4.5, and further purified by gel filtration (Superdex 200 column, GE Healthcare). For the crystallization study, purified human monoclonal antibodies were digested with the immobilized papain (Thermo Scientific) according to the manufacturer’s instructions. The monoclonal antibodies were first buffered in 20 mM Na3PO4, 10 mM EDTA (pH 7.0), and then digested in buffer containing 20 mM Na3PO4, 10 mM EDTA, 10 μM cysteine-HCl (pH 7.0) at 37 °C overnight. The fragments of Fab were purified by HiTrap Protein A FF (5 ml, GE Healthcare) and gel filtration (Superdex 200 column, GE Healthcare).
The Gn monoclonal antibody in the scFv form was overexpressed in Escherichia coli as inclusion bodies, and then sequentially dissolved in 8 M urea and diluted against a refolding buffer (100 mM Tris, pH 8.0; 400 mM l-arginine; 5 mM EDTA-Na; 5 mM glutathione; 0.5 mM glutathione disulfide). After 12 h of slow stirring at 4 °C, the refolded scFv was then concentrated and changed to a 20 mM Tris-HCl (pH 8.0) and 150 mM NaCl buffer, and further analysed by gel filtration (Superdex 200 column, GE Healthcare).
Gn with both the biotinylation tag (GLNDIFEAQKIEWHE) and 6 × His tag at the C terminus was expressed in HEK293T cells. After purification by nickel affinity purification (HisTrap HP column, GE Healthcare) and gel filtration (Superdex 200 column, GE Healthcare), Gn was biotinylated using the Bir A enzyme (Avidity). Biotin is a small protein that forms a strong bond with streptavidin. PE-tagged streptavidin (BioSource International) was added at a molar ratio of 4:1 to the bioengineered Gn monomers, with the biotin–streptavidin interaction causing four Gn monomers to bind to streptavidin and create a tetramer. The Gn tetramers were then used to analyse the interaction between Gn and cells. MHC tetramers were also prepared as previously reported40 and used as a negative control.
Enzyme-linked immunosorbent assay
The binding activities of patient plasma to RVFV Gn and Gc were determined by an enzyme-linked immunosorbent assay. The purified Gn, Gc and Ebola Gp (negative control) were immobilized at 200 ng per well in PBS on 96-well plates at 4 °C overnight. After blocking with 1% BSA at room temperature for 1 h, diluted patient plasma was added and incubated at room temperature for 1 h. The wells were then washed with PBST (0.5% Tween 20) for 5 × 5 min. Goat anti-human IgG/HRP (Santa Cruz Biotechnology) diluted with PBST at 1:3,000 was added and incubated at room temperature for 1 h, followed by washing with PBST for 5 × 5 min. TMB (3,3′,5′,5′-tetramethylbenzidine) substrate (CWBio) was added and 2 M HCl was applied to stop the reaction. Absorbance at 450 nm was measured and recorded. The bar chart was generated using GraphPad Prism 5.
Flow cytometry
HEK293T cells transiently transfected with plasmids of pEGFP-M, pEGFP-Gn, pEGFP-Gc or pEGFP-Gn mutants (T173L, E175G, K294E-D230N, Q174A, D176A, K180A, D225A, S228A, D230A and K294A) were used for the binding test. At 48 h post-transfection, cells were collected and stained with the indicated purified RVFV monoclonal antibodies at a concentration of 20 μg ml−1, except for R12, R13 and Z3L1, which were used at 50 μg ml−1. After washing, anti-hIgG/APC was added. The cells were then subjected to analysis using a BD FACSCalibur.
Huh7 cells were incubated with the Gn tetramer at a concentration of 10 μg ml−1. For the binding-blocking assay, Gn tetramers were pre-incubated with the indicated RVFV monoclonal antibody (0.6 mg ml−1) before their addition to Huh7 cells. The MHC tetramer was used as a negative control for the Gn tetramer, and Z3L1 was used as a negative control for the RVFV monoclonal antibody. After washing, streptavidin-PE (1 μg ml−1) was added. Then, the cells were subjected to analysis using a BD FACSCalibur.
Evaluation of the blocking effect of RVFV monoclonal antibody on viral attachment
We performed these experiments using a previously reported method with some modifications27. To test the association of antibodies with Huh7 and Vero cells (virus binding setting), both cell lines (2 × 105) were incubated with RVFV virions at 5 × 104 plaque-forming unit (PFUs) at 4 °C for 30 min. In addition, 2.5 × 105 PFU RVFV was also used to test the interaction with Huh7 (marked as 5*virus in Supplementary Fig. 6). After washing with PBS 3 times, the cells were fixed with 0.5% paraformaldehyde (PFA) on ice for 30 min, and then washed with PBS twice. Final concentrations of 200 μg ml−1 of either Z3L1 (negative control) or the relative RVFV monoclonal antibody were used to stain cells for 30 min on ice. After washing with PBS three times, anti-hIgG/APC (final concentration of 5 μg ml−1, BioLegend) was incubated with cells for 30 min on ice. Afterwards, the cells were washed with PBS three times and subjected to analysis by flow cytometry.
We used Vero cells to evaluate the blocking effects of RVFV monoclonal antibodies against viral attachment (blocking setting) using a previously published method with some modifications27. Virus (5 × 104 PFUs) was pre-incubated with 200 μg ml−1 of RVFV monoclonal antibody at 37 °C for 1 h, and then placed on ice for 30 min. The virus and the indicated monoclonal antibody mixture were added to Vero cells (2 × 105) and placed on ice for another 30 min. After washing with PBS 3 times, the cells were fixed with 0.5% PFA for 30 min on ice. Subsequently, the cells were washed with PBS twice and incubated with anti-hIgG/APC (final concentration of 5 μg ml−1). Finally, the cells were washed with PBS three times and subjected to analysis by flow cytometry (marked as Indicated monoclonal antibody in Fig. 3b). In parallel, Vero cells (2 × 105) were incubated with cold RVFV (5 × 104 PFUs) for 30 min on ice. After washing with PBS three times, the cells were subsequently fixed with PFA and stained with the indicated monoclonal antibody at a final concentration of 200 μg ml−1 and anti-hIgG/APC (final concentration of 5 μg ml−1), as depicted in the above paragraph (virus binding setting, marked as Virus binding in Fig. 3b).
The blocking effects of the Gn monoclonal antibodies were further evaluated using serially diluted monoclonal antibodies. In this experiment, the fluorescence of cells was evaluated in both virus binding and blocking settings as depicted above, except that 2.5 × 104 PFUs RVFV were used. Each monoclonal antibody was tested at a final concentration of 100, 20, 4 and 0.8 μg ml−1. Compared with the sample in the virus binding test, the decreased proportion of hIgG/APC+ cells in the blocking setting is the inhibitory effect of the indicated monoclonal antibody at the tested concentration.
SPR analysis
The interaction between purified antibodies with RVFV Gn or Gc was monitored by SPR using a BIAcore T100 system (GE Healthcare) carried out at 25 °C in single-cycle mode. The CM5 biosensor chip (GE Healthcare) was first immobilized with anti-human antibody for flow cells (Fc) 1 and 2, according to the manufacturer’s amine-coupling chemistry protocol (GE Healthcare). The indicated monoclonal antibody was then injected and captured on Fc 2. Fc 1 was used as the negative control. Gn or Gc protein was serially diluted and injected. Specifically, both SFTSV Gn and Gc were loaded at concentrations of 50, 100, 200, 400 and 800 nM. RVFV Gc was loaded at concentration of 50, 100, 200, 400, 800 and 1,600 nM for all monoclonal antibodies, except for R5, which was tested at 100, 200, 400, 800 and 1,600 nM. In addition, RVFV Gn at concentrations of 50, 100, 200, 400 and 800 nM was used for R5. The serial dilutions of RVFV Gn used for R13, R16, R17, R12H-13L and R13H-12L were 100, 200, 400, 800 and 1,600 nM; for R4 and R12 were 25, 50, 100, 200 and 400 nM; for R19 were 12.5, 25, 50, 100 and 200 nM; and for R15 and R22 were 6.25, 12.5, 25, 50 and 100 nM. The binding responses were recorded. All proteins used in the experiment were exchanged into the HBS-EP buffer consisting of 10 mM HEPES, pH 7.4, 150 mM NaCl and 0.005% v/v Tween 20. The equilibrium dissociation constants (binding affinity, Kd) for each antigen–monoclonal antibody interaction were calculated using Biacore T100 Evaluation Software, version 1.0 (GE Healthcare). The Kd values for R5 to RVFV Gc and R22 to RVFV Gn were calculated using the model of 1:1 (Langmuir) binding mode, while those for R4, R12, R13, R15, R16, R17, R19, R12H–13L and R13H–12L to Gn were calculated using the model of steady-state affinity.
Neutralization assay by flow cytometry
RVFV was propagated in Vero cells. For the antibody-neutralizing activity of each RVFV monoclonal antibody, a flow cytometry-based neutralization assay with Vero cells was conducted using a similar protocol to that previously reported22 with some modifications. Briefly, 2 × 105 cells per well were plated in a 24-well plate 24 h before conducting the experiment. The isolated monoclonal antibodies were serially diluted and incubated with RVFV (8 × 103 PFUs). Then, the mixtures were used to infect Vero cells. After 48 h, cells were collected and stained by R4 monoclonal antibody at 2 μg ml−1 and anti-hIgG/FITC. After washing, the percentages of Gn-positive cells were measured using a BD FACSCanto II. Sigmoidal neutralization curves were generated using GraphPad Prism 5.
Animal protection
All animal experiments were carried out according to the procedures approved by the Chinese Academy of Sciences and comply with all relevant ethical regulations regarding animal research. To determine the median lethal dose (LD50) of RVFV infection to BALB/c mice, groups of 5 randomly assigned female mice at 6–8 weeks were challenged intraperitoneally with the indicated PFUs of RVFV. Mice were monitored daily for survival for 14 days post-infection.
In the post-exposure setting, groups of 4 (for the monoclonal antibodies R4, R12, R13, R16, R17, R19 and R22, respectively) or 5 (for the monoclonal antibodies R5 and R15, respectively) randomly assigned BALB/c mice were challenged intraperitoneally with 1 × 103 PFU RVFV. On the first day post-infection, a single dose of monoclonal antibody (10 mg kg−1) or PBS (marked as Virus, n = 5) was intraperitoneally administered. The PBS group (n = 4), in which mice were administered PBS instead of virus, was also included. In the pre-exposure setting, groups of 4 (for the monoclonal antibodies R4, R12, R13, R16, R17 and R19, respectively) or 5 randomly assigned mice (for the monoclonal antibodies R5, R15 and R22, respectively) were injected with either monoclonal antibodies (10 mg kg−1) or PBS (n = 4), followed by administration of 1 × 103 PFU RVFV, 1 day later. Mice were monitored daily for survival for 14 days post-infection. Mice exhibiting weight loss of over 20% of their initial weight on day 0 were euthanized.
Crystallization, data collection and structure determination
The indicated Fabs or scFvs were mixed with Gn at a stoichiometry of 1:1 and incubated on ice for 2 h. The mixture was then subjected to gel filtration. The eluted peak containing the complex of the indicated Fab and Gn were pooled, buffered in 20 mM Tris and 50 mM NaCl (pH 8.0), and concentrated to 10 mg ml−1. Crystallization trials were performed by sitting-drop (300 nl + 300 nl) vapour diffusion at 18 °C. Diffractable crystals of the complex were finally obtained in a solution consisting of 0.2 M magnesium chloride hexahydrate, 0.1 M HEPES sodium, pH 7.5, and 30% v/v 2-propanol, with a protein concentration of 10 mg ml−1 for Gn–R12 Fab; a solution containing 0.1 M bicine, pH 9.0, and 20% w/v PEG 6000, with a protein concentration of 10 mg ml−1 for Gn–R13 Fab; a solution containing 0.1 M amino acids, 0.1 M imidazole, MES, pH 6.5, and 30% w/v P550MME_P20K, with a protein concentration of 10 mg ml−1 for Gn–R15 Fab; a solution containing 0.2 M ammonium sulfate, 0.1 M Tris, pH 7.5, and 20% w/v PEG 5000 MME, with a protein concentration of 15 mg ml−1 for Gn–R17 scFv. Diffraction data were collected at 100 K at Shanghai Synchrotron Radiation Facility (SSRF) BL19U1. For data collection, the crystals were cryo-protected by briefly soaking in reservoir solution supplemented with 20% (v/v) glycerol before flash-cooling in liquid nitrogen. The wavelengths used to collect the data for Gn–R12 Fab, Gn–R13 Fab, Gn–R15 Fab and Gn–R17 scFv were 0.97776 Å, 0.97853 Å, 0.97915 Å and 0.97853 Å, respectively. The datasets were processed with HKL2000 software41.
The complex structures were determined by the molecular replacement method using Phaser42 with previously reported structures of RVFV Gn (PDB code: 5Y0W) and Fab (PDB code: 1HZH). The atomic models were completed with Coot43 and refined with phenix.refine in Phenix44, and the stereochemical qualities of the final models were assessed with MolProbity45. Of all the residues, a total of 96.59, 97.32, 95.96 and 94.94% were located in the favoured regions of the Ramachandran plot for Gn–R12 Fab, Gn–R13 Fab, Gn–R15 Fab and Gn–R17 scFv, respectively. The outlier regions were determined to be 0.16, 0.14, 0.00 and 0.19% for Gn–R12 Fab, Gn–R13 Fab, Gn–R15 Fab and Gn–R17 scFv, respectively. The data collection and structure refinement statistics are summarized in Supplementary Table 3. The structural figures were generated using Pymol.
Western blotting
To test whether the isolated monoclonal antibodies could recognize the linearized epitope, each monoclonal antibody was subjected to western blotting. We loaded 0.1 μg of both purified RVFV Gn and Gc for SDS–polyacrylamide gel electrophoresis, then transferred the proteins to nitrocellulose membranes and incubated them with the identified monoclonal antibodies at a final concentration of 1 μg ml−1. A secondary antibody of anti-hIgG/HRP was used at a final concentration of 0.1 μg ml−1 (Zhongshanjinqiao Corp.). Directly HRP-conjugated anti-His antibodies were used at a final concentration of 0.1 μg ml−1 (CWBio). Then, the TMB substrate (CWBio) was added.
Sequence alignments
The GenBank accession number of the sequences used for analysing the conservation of epitopes among Phlebovirus are as follows: Heartland virus, AIF75092.1; Candiru virus, YP_004347992.1; Punta Toro virus, AAA47110.1; Chagres virus, AEL29641.1; sandfly fever Naples virus, AIS25027.1; Toscana virus, ABZ85665.1; Uukuniemi virus, NP_941979; Bhanja virus, YP_009141014.1; and Palma virus, AGC60100.1. The 108 sequences of the RVFV strains, with their complete genomes stored in the NCBI Viral Genomes Resource, are as follows: P03518.2, ABD38806.1, AEB20462.1, ABD38803.1, ABD38804.1, ABD38805.1, ABD38800.1, ABD38807.1, AEF79993.1, ABD38802.1, AJA32703.1, ARW71759.1, ABD38801.1, ABD38808.1, ABD38835.1, AMD08950.1, ABD38798.1, ABD38799.1, ABD38797.1, ABD38796.1, ABD38823.1, ABD38821.1, ABD38820.1, YP_003848705.1, ABD38819.1, ABD38812.1, ABD38815.1, ABD38813.1, ABD38814.1, ABD38817.1, ABD38822.1, ABD38818.1, ABD38816.1, ABD38809.1, ABD38810.1, ABD38811.1, AEC14287.1, AEC14288.1, AGB13833.1, ACE78370.1, ACE78388.1, ACE78379.1, AGB13831.1, AGB13830.1, AGB13832.1, AGB13834.1, ACE78394.1, ACE78383.1, AJQ19496.1, ABD38830.1, ABD38829.1, ABD38828.1, ABD38825.1, ABD38824.1, ABD38826.1, ABD38834.1, ABD38827.1, AJQ19492.1, ABD38831.1, ABD38833.1, ABD38832.1, AEB20468.1, AEB20467.1, AEB20463.1, AEB20472.1, AEB20466.1, AEB20465.1, AEB20464.1, AEC14283.1, ACE78377.1, ACE78376.1, AEC14290.1, ACE78378.1, ACE78382.1, ACE78384.1, ACE78375.1, ACE78386.1, ACE78387.1, ACE78385.1, ACE78381.1, AEB20469.1, AEZ53084.1, AEZ53085.1, CCG28447.1, CCG28444.1, AEZ53088.1, AEZ53087.1, AEZ53086.1, AEL31325.1, AEE10383.1, AEC14291.1, AEC14289.1, AEC14286.1, AEC14285.1, AEC14284.1, AEB20471.1, AEB20470.1, ACE78395.1, ACE78393.1, ACE78392.1, ACE78391.1, ACE78390.1, ACE78389.1, ACE78380.1, ACE78374.1, ACE78373.1, ACE78372.1 and ACE78371.1. The sequence logos were generated using the web server of Weblogo (http://weblogo.berkeley.edu/logo.cgi).
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The crystal structures of Gn complexed with the Fab form of the monoclonal antibodies R12, R13 and R15 and the scFv of R17 have been deposited in the Protein Data Bank under accession codes 6IEK, 6IEA, 6IEB and 6IEC, respectively. The data that support the findings of this study are available from the corresponding author upon request.
References
Daubney, R., Hudson, J. R. & Garnham, P. C. Enzootic hepatitis or Rift Valley fever. An undescribed virus disease of sheep cattle and man from East Africa. J. Pathol. Bacteriol. 34, 545–579 (1931).
Nanyingi, M. O. et al. A systematic review of Rift Valley Fever epidemiology 1931–2014. Infect. Ecol. Epidemiol. 5, 28024 (2015).
Rift Valley Fever (RVF). World Organisation for Animal Health http://www.oie.int/en/animal-health-in-the-world/animal-diseases/rift-valley-fever (2019).
Rift Valley Fever. World Health Organization http://www.who.int/csr/don/archive/disease/rift_valley_fever/en/ (2018).
Shoemaker, T. et al. Genetic analysis of viruses associated with emergence of Rift Valley fever in Saudi Arabia and Yemen, 2000–01. Emerg. Infect. Dis. 8, 1415–1420 (2002).
Liu, J. et al. The first imported case of Rift Valley fever in China reveals a genetic reassortment of different viral lineages. Emerg. Microbes Infect. 6, e4 (2017).
Mansfield, K. L. et al. Rift Valley fever virus: A review of diagnosis and vaccination, and implications for emergence in Europe. Vaccine 33, 5520–5531 (2015).
Adams, M. J. et al. Changes to taxonomy and the International Code of Virus Classification and Nomenclature ratified by the International Committee on Taxonomy of Viruses (2017). Arch. Virol. 162, 2505–2538 (2017).
Knipe, D. M. & Howley, P. M. Fields Virology 6th edn (Wolters Kluwer/Lippincott Williams & Wilkins Health, 2013).
de Boer, S. M. et al. Acid-activated structural reorganization of the Rift Valley fever virus Gc fusion protein. J. Virol. 86, 13642–13652 (2012).
Dessau, M. & Modis, Y. Crystal structure of glycoprotein C from Rift Valley fever virus. Proc. Natl Acad. Sci. USA 110, 1696–1701 (2013).
Zhu, Y. et al. The post-fusion structure of the Heartland virus Gc glycoprotein supports taxonomic separation of the bunyaviral families Phenuiviridae and Hantaviridae. J. Virol. 92, e01558–01517 (2017).
Wu, Y. et al. Structures of phlebovirus glycoprotein Gn and identification of a neutralizing antibody epitope. Proc. Natl Acad. Sci. USA 114, E7564–E7573 (2017).
Halldorsson, S. et al. Shielding and activation of a viral membrane fusion protein. Nat. Commun. 9, 349 (2018).
Huiskonen, J. T., Overby, A. K., Weber, F. & Grunewald, K. Electron cryo-microscopy and single-particle averaging of Rift Valley fever virus: evidence for GN–GC glycoprotein heterodimers. J. Virol. 83, 3762–3769 (2009).
Lagerqvist, N. et al. Characterisation of immune responses and protective efficacy in mice after immunisation with Rift Valley Fever virus cDNA constructs. Virol. J. 6, 6 (2009).
de Boer, S. M. et al. Rift Valley fever virus subunit vaccines confer complete protection against a lethal virus challenge. Vaccine 28, 2330–2339 (2010).
Wallace, D. B. et al. Protective immune responses induced by different recombinant vaccine regimes to Rift Valley fever. Vaccine 24, 7181–7189 (2006).
Faburay, B. et al. A recombinant Rift Valley fever virus glycoprotein subunit vaccine confers full protection against Rift Valley fever challenge in sheep. Sci. Rep. 6, 27719 (2016).
Papin, J. F. et al. Recombinant Rift Valley fever vaccines induce protective levels of antibody in baboons and resistance to lethal challenge in mice. Proc. Natl Acad. Sci. USA 108, 14926–14931 (2011).
Heise, M. T. et al. An alphavirus replicon-derived candidate vaccine against Rift Valley fever virus. Epidemiol. Infect. 137, 1309–1318 (2009).
Wang, Q. et al. Molecular determinants of human neutralizing antibodies isolated from a patient infected with Zika virus. Sci. Transl. Med. 8, 369ra179 (2016).
Smith, D. R. et al. The pathogenesis of Rift Valley fever virus in the mouse model. Virology 407, 256–267 (2010).
Gao, G. F. From "A"IV to "Z"IKV: Attacks from emerging and re-emerging pathogens. Cell 172, 1157–1159 (2018).
Shi, Y., Wu, Y., Zhang, W., Qi, J. & Gao, G. F. Enabling the ‘host jump’: structural determinants of receptor-binding specificity in influenza A viruses. Nat. Rev. Microbiol. 12, 822–831 (2014).
Hofmann, H. et al. Severe fever with thrombocytopenia virus glycoproteins are targeted by neutralizing antibodies and can use DC-SIGN as a receptor for pH-dependent entry into human and animal cell lines. J. Virol. 87, 4384–4394 (2013).
Yamashita, T. et al. Biological and immunological characteristics of hepatitis E virus-like particles based on the crystal structure. Proc. Natl Acad. Sci. USA 106, 12986–12991 (2009).
Nderitu, L. et al. Sequential Rift Valley fever outbreaks in eastern Africa caused by multiple lineages of the virus. J. Infect. Dis. 203, 655–665 (2011).
Bird, B. H., Khristova, M. L., Rollin, P. E., Ksiazek, T. G. & Nichol, S. T. Complete genome analysis of 33 ecologically and biologically diverse Rift Valley fever virus strains reveals widespread virus movement and low genetic diversity due to recent common ancestry. J. Virol. 81, 2805–2816 (2007).
Frank, A. L., Six, H. R. & Marchini, A. Human monoclonal antibodies to influenza virus: IgG subclass and light chain distribution. Viral Immunol. 2, 31–36 (1989).
Kirchenbaum, G. A., Allen, J. D., Layman, T. S., Sautto, G. A. & Ross, T. M. Infection of ferrets with influenza virus elicits a light chain-biased antibody response against hemagglutinin. J. Immunol. 199, 3798–3807 (2017).
Kraus, A. A., Messer, W., Haymore, L. B. & De Silva, A. M. Comparison of plaque- and flow cytometry-based methods for measuring dengue virus neutralization. J. Clin. Microbiol. 45, 3777–3780 (2007).
Chen, M. et al. A flow cytometry-based assay to assess RSV-specific neutralizing antibody is reproducible, efficient and accurate. J. Immunol. Methods 362, 180–184 (2010).
Schmaljohn, C. S. et al. Baculovirus expression of the M genome segment of Rift Valley fever virus and examination of antigenic and immunogenic properties of the expressed proteins. Virology 170, 184–192 (1989).
Keegan, K. & Collett, M. S. Use of bacterial expression cloning to define the amino-acid-sequences of antigenic determinants on the G2-glycoprotein of Rift-Valley fever virus. J. Virol. 58, 263–270 (1986).
Zhang, Q. et al. Potent neutralizing monoclonal antibodies against Ebola virus infection. Sci. Rep. 6, 25856 (2016).
Liao, H. X. et al. High-throughput isolation of immunoglobulin genes from single human B cells and expression as monoclonal antibodies. J. Virol. Methods 158, 171–179 (2009).
Ehrenmann, F., Kaas, Q. & Lefranc, M. P. IMGT/3Dstructure-DB and IMGT/DomainGapAlign: a database and a tool for immunoglobulins or antibodies, T cell receptors, MHC, IgSF and MhcSF. Nucleic Acids Res. 38, D301–D307 (2010).
Zhang, W. et al. Crystal structure of the swine-origin A (H1N1)-2009 influenza A virus hemagglutinin (HA) reveals similar antigenicity to that of the 1918 pandemic virus. Protein Cell 1, 459–467 (2010).
Zhou, M. et al. Screening and identification of severe acute respiratory syndrome-associated coronavirus-specific CTL epitopes. J. Immunol. 177, 2138–2145 (2006).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Read, R. J. Pushing the boundaries of molecular replacement with maximum likelihood. Acta Crystallogr. D 57, 1373–1382 (2001).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Acknowledgements
We thank G. Salazar (University of Texas Health Science Center at Houston) for her critical editing of the manuscript. We acknowledge L. Zhang and Q. Zhang (Comprehensive AIDS Research Center, and Collaborative Innovation Center for Diagnosis and Treatment of Infectious Diseases, School of Medicine, Tsinghua University) for their instruction and help with isolating single B cells. We thank the staff of the BL19U1 beamline at Shanghai Synchrotron Radiation Facility (Shanghai, People’s Republic of China) for assistance during data collection. We are grateful to J. Jia (Institute of Biophysics, Chinese Academy of Sciences) for technical support of BDAria II manipulation and Y. Chen and Z. Yang (Institute of Biophysics, Chinese Academy of Sciences) for technical help with BIAcore experiments. This work was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (grant no. XDB29040201), the National Science and Technology Major Projects for ‘Major New Drugs Innovation and Development’ (grant no. 2018ZX09711003-002-001), the National Science and Technology Major Project (grant nos. 2016ZX10004222-008 and 2018ZX10101004-001), the External Cooperation Program of CAS (153211KYSB20160001) and the National Natural Science Foundation of China (NSFC, grant nos. 31872745, 31502078 and 81502972). Y.S. is supported by the National Science and Technology Major Project (grant no. 2018ZX10101004-001). G.F.G. is also supported by the External Cooperation Program of CAS (grant no. 153211KYSB20160001). Q.W. is supported by Young Elite Scientist Sponsorship Program by China Association for Science and Technology (grant no. 2015QNRC001), the Youth Innovation Promotion Association CAS (grant no. 2018119) and the grant from China Scholarship Council (grant no. 201704910327). Y.W. is supported by the Youth Innovation Promotion Association CAS (grant no. 2016086). J.Y. and G.F.G. are supported by the foundation of the NSFC Innovative Research Group (grant no. 81621091).
Author information
Authors and Affiliations
Contributions
Q.W., Z.T., G.F.G. and J.Y. initiated and coordinated the project. Q.W., Y.W., Z.T., G.F.G. and J.Y. designed the experiments. Q.W., T.M., Y.W., Z.T. and H.Y. conducted the experiments with the assistance of Z.Z., L.W., R.S. and M.Y. F.G., J.Q. and Y.C. collected the crystallographic data and solved the structures. Z.C. and H.Z. coordinated the isolation of peripheral blood mononuclear cells from the RVFV patient. M.L., C.Q., G.F.G. and J.Y. coordinated the viral experiments. T.M. and G.W. manipulated the viruses with the help of Y.B., J.S. and H.J. Q.W., Y.W., Z.T., Z.A., Y.S., G.F.G. and J.Y. analysed the data. Q.W. wrote the manuscript. Y.W., G.W., Z.A., J.W., T.D.Y., Y.S., W.J.L., G.F.G. and J.Y. helped revise the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–4, Supplementary Figures 1–14, Supplementary References and the legend for the Supplementary Dataset.
Rights and permissions
About this article
Cite this article
Wang, Q., Ma, T., Wu, Y. et al. Neutralization mechanism of human monoclonal antibodies against Rift Valley fever virus. Nat Microbiol 4, 1231–1241 (2019). https://doi.org/10.1038/s41564-019-0411-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0411-z