Abstract
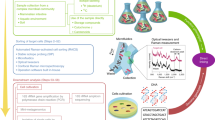
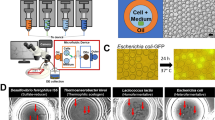
Stable-isotope probing is widely used to study the function of microbial taxa in their natural environment, but sorting of isotopically labelled microbial cells from complex samples for subsequent genomic analysis or cultivation is still in its early infancy. Here, we introduce an optofluidic platform for automated sorting of stable-isotope-probing-labelled microbial cells, combining microfluidics, optical tweezing and Raman microspectroscopy, which yields live cells suitable for subsequent single-cell genomics, mini-metagenomics or cultivation. We describe the design and optimization of this Raman-activated cell-sorting approach, illustrate its operation with four model bacteria (two intestinal, one soil and one marine) and demonstrate its high sorting accuracy (98.3 ± 1.7%), throughput (200–500 cells h−1; 3.3–8.3 cells min−1) and compatibility with cultivation. Application of this sorting approach for the metagenomic characterization of bacteria involved in mucin degradation in the mouse colon revealed a diverse consortium of bacteria, including several members of the underexplored family Muribaculaceae, highlighting both the complexity of this niche and the potential of Raman-activated cell sorting for identifying key players in targeted processes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Code availability
All of the custom codes used in this study can be accessed upon request from the corresponding author. MATLAB GUI (graphical user interface) software for the operation of the RACS platform is provided in Supplementary Files 1 and 2. R code for the calculation in Supplementary Fig. 18 is provided in Supplementary File 3.
Data availability
The data that support the findings of this study are available from the corresponding author upon request. 16S rRNA gene sequence data have been deposited in the NCBI Sequence Read Archive under SRP144990. Metagenomic data have been deposited in the NCBI under SRP144778. MAGs have been deposited as whole-genome shotgun projects at DDBJ/ENA/GenBank under the accessions RYVY00000000–RYWW00000000. All accession numbers with information on the associated samples are provided in Supplementary Table 6.
Change history
12 April 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013).
Marcy, Y. et al. Dissecting biological “dark matter” with single-cell genetic analysis of rare and uncultivated TM7 microbes from the human mouth. Proc. Natl Acad. Sci. USA 104, 11889–11894 (2007).
Blainey, P. C. The future is now: single-cell genomics of bacteria and archaea. FEMS Microbiol. Rev. 37, 407–427 (2013).
Xu, L., Brito, I. L., Alm, E. J. & Blainey, P. C. Virtual microfluidics for digital quantification and single-cell sequencing. Nat. Methods 13, 759–762 (2016).
Ackermann, M. A functional perspective on phenotypic heterogeneity in microorganisms. Nat. Rev. Microbiol. 13, 497–508 (2015).
Ackermann, M. Microbial individuality in the natural environment. ISME J. 7, 465–467 (2013).
Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002).
Stocker, R. Marine microbes see a sea of gradients. Science 338, 628–633 (2012).
Yoon, H. S. et al. Single-cell genomics reveals organismal interactions in uncultivated marine protists. Science 332, 714–717 (2011).
Labonte, J. M. et al. Single-cell genomics-based analysis of virus–host interactions in marine surface bacterioplankton. ISME J. 9, 2386–2399 (2015).
Wagner, M., Nielsen, P. H., Loy, A., Nielsen, J. L. & Daims, H. Linking microbial community structure with function: fluorescence in situ hybridization-microautoradiography and isotope arrays. Curr. Opin. Biotechnol. 17, 83–91 (2006).
Neufeld, J. D., Wagner, M. & Murrell, J. C. Who eats what, where and when? Isotope-labelling experiments are coming of age. ISME J. 1, 103–110 (2007).
Lee, N. et al. Combination of fluorescent in situ hybridization and microautoradiography—a new tool for structure–function analyses in microbial ecology. Appl. Environ. Microbiol. 65, 1289–1297 (1999).
McGlynn, S. E., Chadwick, G. L., Kempes, C. P. & Orphan, V. J. Single cell activity reveals direct electron transfer in methanotrophic consortia. Nature 526, 531–535 (2015).
Krupke, A. et al. The effect of nutrients on carbon and nitrogen fixation by the UCYN-A–haptophyte symbiosis. ISME J. 9, 1635–1647 (2015).
Berry, D. et al. Host-compound foraging by intestinal microbiota revealed by single-cell stable isotope probing. Proc. Natl Acad. Sci. USA 110, 4720–4725 (2013).
Wang, Y., Huang, W. E., Cui, L. & Wagner, M. Single cell stable isotope probing in microbiology using Raman microspectroscopy. Curr. Opin. Biotechnol. 41, 34–42 (2016).
Jing, X. et al. Raman-activated cell sorting and metagenomic sequencing revealing carbon-fixing bacteria in the ocean. Environ. Microbiol. 20, 2241–2255 (2018).
Li, M. et al. Rapid resonance Raman microspectroscopy to probe carbon dioxide by single cells in microbial communities. ISME J. 6, 875–885 (2012).
Huang, W. E. et al. Raman-FISH: combining stable-isotope Raman spectroscopy and fluorescence in situ hybridization for the single cell analysis of identity and function. Environ. Microbiol. 9, 1878–1889 (2007).
Wagner, M. Single-cell ecophysiology of microbes as revealed by Raman microspectroscopy or secondary ion mass spectrometry imaging. Annu. Rev. Microbiol. 63, 411–429 (2009).
Rinke, C. et al. Obtaining genomes from uncultivated environmental microorganisms using FACS-based single-cell genomics. Nat. Protoc. 9, 1038–1048 (2014).
Dyksma, S. et al. Ubiquitous Gammaproteobacteria dominate dark carbon fixation in coastal sediments. ISME J. 10, 1939–1953 (2016).
Dupont, C. L. et al. Genomic insights to SAR86, an abundant and uncultivated marine bacterial lineage. ISME J. 6, 1186–1199 (2012).
Woyke, T. et al. Assembling the marine metagenome, one cell at a time. PLoS ONE 4, e5299 (2009).
Hatzenpichler, R. et al. Visualizing in situ translational activity for identifying and sorting slow-growing archaeal–bacterial consortia. Proc. Natl Acad. Sci. USA 113, E4069–E4078 (2016).
Berry, D. et al. Tracking heavy water (D2O) incorporation for identifying and sorting active microbial cells. Proc. Natl Acad. Sci. USA 112, E194–E203 (2015).
Huang, W. E., Ward, A. D. & Whiteley, A. S. Raman tweezers sorting of single microbial cells. Environ. Microbiol. Rep. 1, 44–49 (2009).
Zhang, P. et al. Raman-activated cell sorting based on dielectrophoretic single-cell trap and release. Anal. Chem. 87, 2282–2289 (2015).
Mcllvenna, D. et al. Continuous cell sorting in a flow based on single cell resonance Raman spectra. Lab Chip 16, 1420–1429 (2016).
Johansson, M. E. V. et al. The inner of the two Muc2 mucin-dependent mucus layers in colon is devoid of bacteria. Proc. Natl Acad. Sci. USA 105, 15064–15069 (2008).
Schroeder, B. O. et al. Bifidobacteria or fiber protects against diet-induced microbiota-mediated colonic mucus deterioration. Cell Host Microbe 23, 27–40 (2018).
Desai, M. S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339–1353.e21 (2016).
Png, C. W. et al. Mucolytic bacteria with increased prevalence in IBD mucosa augment in vitro utilization of mucin by other bacteria. Am. J. Gastroenterol. 105, 2420–2428 (2010).
Sonnenburg, E. D. & Sonnenburg, J. L. Starving our microbial self: the deleterious consequences of a diet deficient in microbiota-accessible carbohydrates. Cell Metab. 20, 779–786 (2014).
Tailford, L. E., Crost, E. H., Kavanaugh, D. & Juge, N. Mucin glycan foraging in the human gut microbiome. Front. Genet. 6, 81 (2015).
Bae, S. C., Lee, H., Lin, Z. & Granick, S. Chemical imaging in a surface force apparatus: confocal Raman spectroscopy of confined poly(dimethylsiloxane). Langmuir 21, 5685–5688 (2005).
Dochow, S. et al. Quartz microfluidic chip for tumour cell identification by Raman spectroscopy in combination with optical traps. Anal. Bioanal. Chem. 405, 2743–2746 (2013).
Dholakia, K. & Reece, P. Optical micromanipulation takes hold. Nano Today 1, 18–27 (2006).
Lagkouvardos, I. et al. The Mouse Intestinal Bacterial Collection (miBC) provides host-specific insight into cultured diversity and functional potential of the gut microbiota. Nat. Microbiol. 1, 16131 (2016).
Stepanauskas, R. & Sieracki, M. E. Matching phylogeny and metabolism in the uncultured marine bacteria, one cell at a time. Proc. Natl Acad. Sci. USA 104, 9052–9057 (2007).
Qin, D., Xia, Y. & Whitesides, G. M. Soft lithography for micro- and nanoscale patterning. Nat. Protoc. 5, 491–502 (2010).
Schuster, K. C., Reese, I., Urlaub, E., Gapes, J. R. & Lendl, B. Multidimensional information on the chemical composition of single bacteria cells by confocal Raman microspectroscopy. Anal. Chem. 72, 5529–5534 (2000).
Carlo, D. D., Irimia, D., Tompkins, R. G. & Toner, M. Continuous inertial focusing, ordering, and separation of particles in microchannels. Proc. Natl Acad. Sci. USA 104, 18892–18897 (2007).
Ha, B. H., Lee, K. S., Jung, J. H. & Sung, H. J. Three-dimensional hydrodynamic flow and particle focusing using four vortices Dean flow. Microfluid. Nanofluid. 17, 647–655 (2014).
Chu, H., Doh, I. & Cho, Y. A three-dimensional (3D) particle focusing channel using the positive dielectrophoresis (pDEP) guided by a dielectric structure between two planar electrodes. Lab Chip 9, 686–691 (2009).
Shi, J. et al. Three-dimensional continuous particle focusing in a microfluidic channel via standing surface acoustic waves (SSAW). Lab Chip 11, 2319–2324 (2011).
Rhee, M. et al. Synthesis of size-tunable polymeric nanoparticles enabled by 3D hydrodynamic flow focusing in single-layer microchannels. Adv. Mater. 23, H79–H83 (2011).
Ozkumur, E. et al. Inertial focusing for tumor antigen-dependent and -independent sorting of rare circulating tumor cells. Sci. Transl. Med. 5, 179ra47 (2013).
Svoboda, K. & Block, S. M. Biological applications of optical forces. Annu. Rev. Biophys. Biomol. Struct. 23, 247–285 (1994).
Dienerowitz, M. et al. Optically trapped bacteria pairs reveal discrete motile response to control aggregation upon cell–cell approach. Curr. Microbiol. 69, 669–674 (2014).
Chang, C. B., Huang, W.-X., Lee, K. H. & Sung, H. J. Optical levitation of a non-spherical particle in a loosely focused Gaussian beam. Opt. Express 20, 24068–24084 (2012).
Lee, K. S. et al. Radiation forces on a microsphere in an arbitrary refractive index profile. J. Opt. Soc. Am. B 29, 407–414 (2012).
Koppel, D. E., Axelrod, D., Schlessinger, J., Elson, E. L. & Web, W. W. Dynamics of fluorescence marker concentration as a probe of mobility. Biophys. J. 16, 1315–1329 (1976).
Kiefer, J., Ebel, N., Schlucker, E. & Leipertz, A. Characterization of Escherichia coli suspensions using UV/Vis/NIR absorption spectroscopy. Anal. Methods 2, 123–128 (2010).
Lucker, S., Nowka, B., Rattei, T., Spieck, E. & Daims, H. The genome of Nitrospina gracilis illuminates the metabolism and evolution of the major marine nitrite oxidizer. Front. Microbiol. 4, 27 (2013).
Patzold, R. et al. In situ mapping of nitrifiers and anammox bacteria in microbial aggregates by means of confocal resonance Raman microscopy. J. Microbiol. Methods 72, 241–248 (2008).
Milucka, J. et al. Zero-valent sulphur is a key intermediate in marine methane oxidation. Nature 491, 541–546 (2012).
Song, Y. et al. Single-cell genomics based on Raman sorting reveals novel carotenoid-containing bacteria in the Red Sea. Microb. Biotechnol. 10, 125–137 (2017).
Griffiths, R. I., Whiteley, A. S., O’Donnell, A. G. & Bailey, M. J. Rapid method for coextraction of DNA and RNA from natural environments for analysis of ribosomal DNA- and rRNA-based microbial community composition. Appl. Environ. Microbiol. 66, 5488–5491 (2000).
Herbold, C. W. et al. A flexible and economical barcoding approach for highly multiplexed amplicon sequencing of diverse target genes. Front. Microbiol. 6, 731 (2015).
Hamady, M., Walker, J., Harris, J., Gold, N. & Knight, R. Error-correcting barcoded primers allow hundreds of samples to be pyrosequenced in multiplex. Nat. Methods 5, 235–237 (2008).
Schubert, M., Lindgreen, S. & Orlando, L. AdapterRemoval v2: rapid adapter trimming, identification, and read merging. BMC Res. Notes 9, 88 (2016).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Kang, D. D. et al. MetaBAT, an efficient tool for accurately reconstructing single genomes from complex microbial communities. PeerJ 3, e1165 (2015).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genome de-replication that enables tracking of microbial genotypes and improved genome recovery from metagenomes. ISME J. 11, 2864–2868 (2017).
Aziz, R. K. et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9, 75 (2008).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Richter, M. & Rosselló-Móra, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl Acad. Sci. USA 106, 19126–19131 (2009).
Ravcheev, D. A. & Thiele, I. Comparative genomic analysis of the human gut microbiome reveals a broad distribution of metabolic pathways for the degradation of host-synthetized mucin glycans and utilization of mucin-derived monosaccharides. Front. Genet. 8, 111 (2017).
Abu-Ali, G. S. et al. Metatranscriptome of human faecal microbial communities in a cohort of adult men. Nat. Microbiol. 3, 356–366 (2018).
Arike, L., Holmén-Larsson, J. & Hansson, G. C. Intestinal Muc2 mucin O-glycosylation is affected by microbiota and regulated by differential expression of glycosyltranferases. Glycobiology 27, 318–328 (2017).
Evans, C. C. et al. Exercise prevents weight gain and alters the gut microbiota in a mouse model of high fat diet-induced obesity. PLoS ONE 9, e92193 (2014).
Rooks, M. G. et al. Gut microbiome composition and function in experimental colitis during active disease and treatment-induced remission. ISME J. 8, 1403–1417 (2014).
Ormerod, K. L. et al. Genomic characterization of the uncultured Bacteroidales family S24-7 inhabiting the guts of homeothermic animals. Microbiome 4, 36 (2016).
Eichorst, S. A. et al. Advancements in the application of NanoSIMS and Raman microspectroscopy to investigate the activity of microbial cells in soils. FEMS Microbiol. Ecol. 91, fiv106 (2015).
Ashkin, A. Forces of a single-beam gradient laser trap on a dielectric sphere in the ray optics regime. Biophys. J. 61, 569–582 (1992).
Novotny, L., Bian, R. X. & Xie, X. S. Theory of nanometric optical tweezers. Phys. Rev. Lett. 79, 645–648 (1997).
Samek, O. et al. Quantitative Raman spectroscopy analysis of polyhydroxyalkanoates produced by Cupriavidus necator H16. Sensors 16, 1808 (2016).
Gruber-Vodicka, H. R. et al. Paracatenula, an ancient symbiosis between thiotrophic Alphaproteobacteria and catenulid flatworms. Proc. Natl Acad. Sci. USA 108, 12078–12083 (2011).
Majed, N., Matthaus, C., Diem, M. & Gu, A. Z. Evaluation of intracellular polyphosphate dynamics in enhanced biological phosphorus removal process using Raman microscopy. Environ. Sci. Technol. 43, 5436–5442 (2009).
Kumar, B. N. V., Kampe, B., Rosch, P. & Popp, J. Characterization of carotenoids in soil bacteria and investigation of their photodegradation by UVA radiation via resonance Raman spectroscopy. Analyst 140, 4584–4593 (2015).
Lutz, M. Resonance Raman spectra of chlorophyll in solution. J. Raman Spectrosc. 2, 497–516 (1974).
Salama, S. & Spiro, T. G. Visible and near-ultraviolet resonance Raman spectra of photolabile vitamin B12 derivatives with a rapid-flow technique. J. Raman Spectrosc. 6, 57–60 (1977).
Spiro, T. G. & Strekas, T. C. Resonance Raman spectra of heme proteins. Effects of oxidation and spin state. J. Am. Chem. Soc. 96, 338–345 (1974).
Palings, I. et al. Assignment of fingerprint vibrations in the resonance Raman spectra of rhodopsin, isorhodopsin, and bathorhodopsin: implications for chromophore structure and environment. Biochemistry 26, 2544–2556 (1987).
Bjerg, J. T. et al. Long-distance electron transport in individual, living cable bacteria. Proc. Natl Acad. Sci. USA 115, 5786–5791 (2018).
Cheng, J. X. & Xie, X. S. Vibrational spectroscopic imaging of living systems: an emerging platform for biology and medicine. Science 350, aaa8870 (2015).
Evans, C. L. & Xie, X. S. Coherent anti-Stokes Raman scattering microscopy: chemical imaging for biology and medicine. Annu. Rev. Anal. Chem. 1, 883–909 (2008).
Acknowledgements
We acknowledge support by a US Department of Energy Joint Genome Institute Emerging Technologies Opportunity grant (DE-AC02-05CH11231 to R.S. and M.W.). M.W. and M.P. were also supported by the European Research Council via the Advanced Grant project NITRICARE 294343. R.S. acknowledges support from a Gordon and Betty Moore Marine Microbial Initiative Investigator Award (no. 3783). F.M. was supported by the Engineering and Physical Sciences Research Council (EP/R035350/1 and EP/S001921/1). F.C.P. and D.B. were supported by the Austrian Science Fund (FWF; P26127-B20) and the European Research Council (Starting Grant: FunKeyGut 741623). F.C.P. was also supported by the European Union’s Horizon 2020 Framework Programme for Research and Innovation (no. 658718). A.J.M. was supported by the Austrian Science Fund (FWF) project Microbial Nitrogen Cycling–From Single Cells to Ecosystems (W1257). We thank R. Weiß and N. Ivleva for collaboration on the surface-enhanced Raman spectroscopy, C. Herbold for bioinformatics assistance and input, and T. Woyke and R. Malmstrom for many helpful discussions.
Author information
Authors and Affiliations
Contributions
K.S.L. and R.S. created the software for the RACS. K.S.L., D.B. and R.S. performed the numerical calculations. K.S.L., M.P., J.N., V.I.F. and F.M. developed the RACS system and designed and performed the pure culture RACS experiments. K.S.L., F.C.P. and D.B. designed and performed the mouse colon sample experiments. K.S.L., A.J.M. and H.D. designed and performed the marine enrichment sample experiments. M.W. and R.S. supervised the project. All authors wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–19, Supplementary Tables 1–6, legends for Supplementary Movies and legends for Supplementary Files.
Supplementary File 1
MATLAB GUI code for the operation of RACS.
Supplementary File 2
MATLAB GUI code for the operation of RACS.
Supplementary File 3
R code for the calculation in Supplementary Figure 18.
Supplementary File 4
CAD diagrams for the microfluidic sorter.
Supplementary Video 1
A numerically simulated (COMSOL Multiphysics) depiction of the vertical flow focusing.
Supplementary Video 2
An experimental validation of the vertical flow focusing effect.
Supplementary Video 3
Flow within the device.
Supplementary Video 4
An overview of the fully automated RACS process.
Supplementary Video 5
Flow carries cells passing through the evaluation region into the collection outlet.
Supplementary Video 6
Continuous single cell optical tweezing and release.
Supplementary Video 7
Continuous single cell optical tweezing and release.
Supplementary Video 8
Magnetic stirrer designed to prevent cell sedimentation and allow for continuous cell injection during the experiment.
Supplementary Video 9
Laser-induced (photophoretic) cell damage by a high-power Raman laser.
Rights and permissions
About this article
Cite this article
Lee, K.S., Palatinszky, M., Pereira, F.C. et al. An automated Raman-based platform for the sorting of live cells by functional properties. Nat Microbiol 4, 1035–1048 (2019). https://doi.org/10.1038/s41564-019-0394-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0394-9
This article is cited by
-
The gut microbiota and its biogeography
Nature Reviews Microbiology (2024)
-
Functional gene-guided enrichment plus in situ microsphere cultivation enables isolation of new crucial ureolytic bacteria from the rumen of cattle
Microbiome (2023)
-
Characteristics of the intestinal bacterial microbiota profiles in Bifidobacterium pseudocatenulatum LI09 pre-treated rats with D-galactosamine-induced liver injury
World Journal of Microbiology and Biotechnology (2023)
-
Identification of inulin-responsive bacteria in the gut microbiota via multi-modal activity-based sorting
Nature Communications (2023)
-
Opportunities and challenges of using metagenomic data to bring uncultured microbes into cultivation
Microbiome (2022)