Abstract
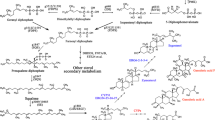
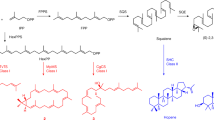
Steroids are essential triterpenoid molecules that are present in all eukaryotes and modulate the fluidity and flexibility of cell membranes. Steroids also serve as signalling molecules that are crucial for growth, development and differentiation of multicellular organisms1,2,3. The steroid biosynthetic pathway is highly conserved and is key in eukaryote evolution4,5,6,7. The flavoprotein squalene epoxidase (SQE) catalyses the first oxygenation reaction in this pathway and is rate limiting. However, despite its conservation in animals, plants and fungi, several phylogenetically widely distributed eukaryote genomes lack an SQE-encoding gene7,8. Here, we discovered and characterized an alternative SQE (AltSQE) belonging to the fatty acid hydroxylase superfamily. AltSQE was identified through screening of a gene library of the diatom Phaeodactylum tricornutum in a SQE-deficient yeast. In accordance with its divergent protein structure and need for cofactors, we found that AltSQE is insensitive to the conventional SQE inhibitor terbinafine. AltSQE is present in many eukaryotic lineages but is mutually exclusive with SQE and shows a patchy distribution within monophyletic clades. Our discovery provides an alternative element for the conserved steroid biosynthesis pathway, raises questions about eukaryote metabolic evolution and opens routes to develop selective SQE inhibitors to control hazardous organisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Benveniste, P. Biosynthesis and accumulation of sterols. Annu. Rev. Plant Biol. 55, 429–457 (2004).
Payne, A. H. & Hales, D. B. Overview of steroidogenic enzymes in the pathway from cholesterol to active steroid hormones. Endocr. Rev. 25, 947–970 (2004).
Weete, J. D., Abril, M. & Blackwell, M. Phylogenetic distribution of fungal sterols. PLoS ONE 5, e10899 (2010).
Gold, D. A., Caron, A., Fournier, G. P. & Summons, R. E. Paleoproterozoic sterol biosynthesis and the rise of oxygen. Nature 543, 420–423 (2017).
Summons, R. E., Bradley, A. S., Jahnke, L. L. & Waldbauer, J. R. Steroids, triterpenoids and molecular oxygen. Phil. Trans. R. Soc. B 361, 951–968 (2006).
Crowe, S. A. et al. Atmospheric oxygenation three billion years ago. Nature 501, 535–538 (2013).
Desmond, E. & Gribaldo, S. Phylogenomics of sterol synthesis: insights into the origin, evolution, and diversity of a key eukaryotic feature. Genome Biol. Evol. 1, 364–381 (2009).
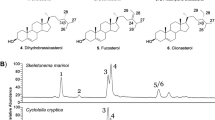
Fabris, M. et al. Tracking the sterol biosynthesis pathway of the diatom Phaeodactylum tricornutum. New Phytol. 204, 521–535 (2014).
Rampen, S. W., Abbas, B. A., Schouten, S. & Damsté, J. S. S. A comprehensive study of sterols in marine diatoms (Bacillariophyta): implications for their use as tracers for diatom productivity. Limnol. Oceanogr. 55, 91–105 (2010).
Gallo, C., d’Ippolito, G., Nuzzo, G., Sardo, A. & Fontana, A. Autoinhibitory sterol sulfates mediate programmed cell death in a bloom-forming marine diatom. Nat. Commun. 8, 1292 (2017).
Goldstein, J. L., DeBose-Boyd, R. A. & Brown, M. S. Protein sensors for membrane sterols. Cell 124, 35–46 (2006).
Gill, S., Stevenson, J., Kristiana, I. & Brown, A. J. Cholesterol-dependent degradation of squalene monooxygenase, a control point in cholesterol synthesis beyond HMG-CoA reductase. Cell Metab. 13, 260–273 (2011).
Pollier, J. et al. The protein quality control system manages plant defence compound synthesis. Nature 504, 148–152 (2013).
Dong, L. et al. Co-expression of squalene epoxidases with triterpene cyclases boosts production of triterpenoids in plants and yeast. Metab. Eng. 49, 1–12 (2018).
Huysman, M. J. J. et al. AUREOCHROME1a-mediated induction of the diatom-specific cyclin dsCYC2 controls the onset of cell division in diatoms (Phaeodactylum tricornutum). Plant Cell 25, 215–228 (2013).
Miettinen, K. et al. The ancient CYP716 family is a major contributor to the diversification of eudicot triterpenoid biosynthesis. Nat. Commun. 8, 14153 (2017).
Shanklin, J. & Cahoon, E. B. Desaturation and related modifications of fatty acids. Annu. Rev. Plant Physiol. Plant Mol. Biol. 49, 611–641 (1998).
Zhu, G., Koszelak-Rosenblum, M., Connelly, S. M., Dumont, M. E. & Malkowski, M. G. The crystal structure of an integral membrane fatty acid α-hydroxylase. J. Biol. Chem. 290, 29820–29833 (2015).
Liu, X. et al. Addressing various compartments of the diatom model organism Phaeodactylum tricornutum via sub-cellular marker proteins. Algal Res. 20, 249–257 (2016).
Leber, R. et al. Dual localization of squalene epoxidase, Erg1p, in yeast reflects a relationship between the endoplasmic reticulum and lipid particles. Mol. Biol. Cell 9, 375–386 (1998).
Laranjeira, S. et al. Arabidopsis squalene epoxidase 3 (SQE3) complements SQE1 and is important for embryo development and bulk squalene epoxidase activity. Mol. Plant 8, 1090–1102 (2015).
Bai, Y. et al. X-ray structure of a mammalian stearoyl-CoA desaturase. Nature 524, 252–256 (2015).
Johnsson, N. & Varshavsky, A. Split ubiquitin as a sensor of protein interactions in vivo. Proc. Natl Acad. Sci. USA 91, 10340–10344 (1994).
Monier, A. et al. Horizontal gene transfer of an entire metabolic pathway between a eukaryotic alga and its DNA virus. Genome Res. 19, 1441–1449 (2009).
Keeling, P. J. & Inagaki, Y. A class of eukaryotic GTPase with a punctate distribution suggesting multiple functional replacements of translation elongation factor 1α. Proc. Natl Acad. Sci. USA 101, 15380–15385 (2004).
Szabová, J., Růžička, P., Verner, Z., Hampl, V. & Lukeš, J. Experimental examination of EFL and MATX eukaryotic horizontal gene transfers: coexistence of mutually exclusive transcripts predates functional rescue. Mol. Biol. Evol. 28, 2371–2378 (2011).
Wang, D. Z. Neurotoxins from marine dinoflagellates: a brief review. Mar. Drugs 6, 349–371 (2008).
Vasconcelos, V., Azevedo, J., Silva, M. & Ramos, V. Effects of marine toxins on the reproduction and early stages development of aquatic organisms. Mar. Drugs 8, 59–79 (2010).
Trainic, M. et al. Infection dynamics of a bloom-forming alga and its virus determine airborne coccolith emission from seawater. iScience 6, 327–335 (2018).
Moses, T. et al. Combinatorial biosynthesis of sapogenins and saponins in Saccharomyces cerevisiae using a C-16α hydroxylase from Bupleurum falcatum. Proc. Natl Acad. Sci. USA 111, 1634–1639 (2014).
Nelson, B. K., Cai, X. & Nebenführ, A. A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J. 51, 1126–1136 (2007).
Deslandes, L. et al. Physical interaction between RRS1-R, a protein conferring resistance to bacterial wilt, and PopP2, a type III effector targeted to the plant nucleus. Proc. Natl Acad. Sci. USA 100, 8024–8029 (2003).
De Riso, V. et al. Gene silencing in the marine diatom Phaeodactylum tricornutum. Nucleic Acids Res. 37, e96 (2009).
Berges, J. A., Franklin, D. J. & Harrison, P. J. Evolution of an artificial seawater medium: improvements in enriched seawater, artificial water over the last two decades. J. Phycol. 37, 1138–1145 (2001).
Lin, H.-Y. et al. Alkaline phosphatase promoter as an efficient driving element for exogenic recombinant in the marine diatom Phaeodactylum tricornutum. Algal Res. 23, 58–65 (2017).
Nagai, T. et al. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat. Biotechnol. 20, 87–90 (2002).
Strand, T. A., Lale, R., Degnes, K. F., Lando, M. & Valla, S. A new and improved host-independent plasmid system for RK2-based conjugal transfer. PLoS ONE 9, e90372 (2014).
Diner, R. E., Bielinski, V. A., Dupont, C. L., Allen, A. E. & Weyman, P. D. Refinement of the diatom episome maintenance sequence and improvement of conjugation-based DNA delivery methods. Front. Bioeng. Biotechnol. 4, 65 (2016).
Siaut, M. et al. Molecular toolbox for studying diatom biology in Phaeodactylum tricornutum. Gene 406, 23–25 (2007).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Johnson, L. K., Alexander, H. B. & Brown, C. T. Re-assembly, quality evaluation, and annotation of 678 microbial eukaryotic reference transcriptomes. Preprint at https://www.biorxiv.org/content/early/2018/09/18/323576 (2018).
Burki, F. et al. Untangling the early diversification of eukaryotes: a phylogenomic study of the evolutionary origins of Centrohelida, Haptophyta and Cryptista. Proc. R. Soc. B 283, 20152802 (2016).
Yoshida, Y. et al. De novo assembly and comparative transcriptome analysis of Euglena gracilis in response to anaerobic conditions. BMC Genomics 17, 182 (2016).
Van Bel, M. et al. TRAPID: an efficient online tool for the functional and comparative analysis of de novo RNA-seq transcriptomes. Genome. Biol. 14, R134 (2013).
Keller, O., Kollmar, M., Stanke, M. & Waack, S. A novel hybrid gene prediction method employing protein multiple sequence alignments. Bioinformatics 27, 757–763 (2011).
Villar, E. et al. The Ocean Gene Atlas: exploring the biogeography of plankton genes online. Nucleic Acids Res. 46, W289–W295 (2018).
Sunagawa, S. et al. Structure and function of the global ocean microbiome. Science 348, 1261359 (2015).
Wilson, W. H. et al. Complete genome sequence and lytic phase transcription profile of a Coccolithovirus. Science 309, 1090–1092 (2005).
Hurwitz, B. L. & Sullivan, M. B. The Pacific Ocean Virome (POV): a marine viral metagenomic dataset and associated protein clusters for quantitative viral ecology. PLoS ONE 8, e57355 (2013).
Goodacre, N., Aljanahi, A., Nandakumar, S., Mikailov, M. & Khan, A. S. A Reference Viral Database (RVDB) to enhance bioinformatics analysis of high-throughput sequencing for novel virus detection. mSphere 3, e00069-18 (2018).
Nishimura, Y. et al. Environmental viral genomes shed new light on virus–host interactions in the ocean. mSphere 2, e00359-16 (2017).
Mihara, T. et al. Linking virus genomes with host taxonomy. Viruses 8, 66 (2016).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Acknowledgements
We thank R. De Clercq and T. Lapshina for technical assistance, Y. Bai, M. Johnson and R. Abbriano-Burke for support with confocal microscopy and M. Huysman and L. De Veylder for providing the P. tricornutum cDNA library. J.P. is a postdoctoral fellow of the Research Foundation-Flanders. E.V. is funded by the BOF project GOA01G01715. M.F. is supported by a CSIRO Synthetic Biology Future Science Fellowship, co-funded by CSIRO and the University of Technology Sydney.
Author information
Authors and Affiliations
Contributions
J.P., E.V., U.K. and M.F. carried out the experiments. J.P., K.V., A.G. and M.F. designed the experiments. All authors contributed to writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–8, Supplementary Table 1.
Supplementary Dataset 1
Overview of all detected alternative and conventional SQE protein sequences in all queried organisms.
Supplementary Dataset 2
Maximum-likelihood phylogeny of eukaryotic and viral AltSQE proteins constructed from an alignment of homologues from 202 organisms and 403 informative aligned sites, rooted with ERG3. Proteins are coloured based on their phylogenetic affiliations and bootstrap values are mentioned at the right of every node.
Supplementary Dataset 3
Maximum-likelihood phylogeny of eukaryotic conventional SQE proteins constructed from an alignment of homologues from 235 organisms and 624 informative aligned sites, rooted with UbiH. Proteins are coloured based on their phylogenetic affiliations and bootstrap values are mentioned at the right of every node.
Rights and permissions
About this article
Cite this article
Pollier, J., Vancaester, E., Kuzhiumparambil, U. et al. A widespread alternative squalene epoxidase participates in eukaryote steroid biosynthesis. Nat Microbiol 4, 226–233 (2019). https://doi.org/10.1038/s41564-018-0305-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0305-5
This article is cited by
-
Identification of squalene epoxidase in triterpenes biosynthesis in Poria cocos by molecular docking and CRISPR-Cas9 gene editing
Microbial Cell Factories (2024)
-
Diverse uses of valuable seafood processing industry waste for sustainability: a review
Environmental Science and Pollution Research (2023)
-
Research progress on biological regulation and biosynthesis of isosteroid alkaloids in Fritillaria
Plant Growth Regulation (2023)
-
Lipid biomarkers: molecular tools for illuminating the history of microbial life
Nature Reviews Microbiology (2022)
-
The Physalis floridana genome provides insights into the biochemical and morphological evolution of Physalis fruits
Horticulture Research (2021)