Abstract
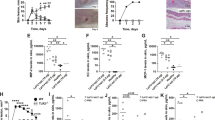
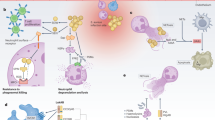
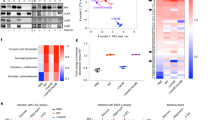
All bacterial infections occur within a polymicrobial environment, from which a pathogen population emerges to establish disease within a host. Emphasis has been placed on prevention of pathogen dominance by competing microflora acting as probiotics1. Here we show that the virulence of the human pathogen Staphylococcus aureus is augmented by native, polymicrobial, commensal skin flora and individual species acting as ‘proinfectious agents’. The outcome is pathogen proliferation, but not commensal. Pathogenesis augmentation can be mediated by particulate cell wall peptidoglycan, reducing the S. aureus infectious dose by over 1,000-fold. This phenomenon occurs using a range of S. aureus strains and infection models and is not mediated by established receptor-mediated pathways including Nod1, Nod2, Myd88 and the NLPR3 inflammasome. During mouse sepsis, augmentation depends on liver-resident macrophages (Kupffer cells) that capture and internalize both the pathogen and the proinfectious agent, leading to reduced production of reactive oxygen species, pathogen survival and subsequent multiple liver abscess formation. The augmented infection model more closely resembles the natural situation and establishes the role of resident environmental microflora in the initiation of disease by an invading pathogen. As the human microflora is ubiquitous2, its role in increasing susceptibility to infection by S. aureus highlights potential strategies for disease prevention.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Isolauri, E., Kirjavainen, P. V. & Salminen, S. Probiotics: a role in the treatment of intestinal infection and inflammation? Gut 50, iii54–iii59 (2002).
Grice, E. A. & Segre, J. A. The skin microbiome. Nat. Rev. Microbiol. 9, 244–253 (2011).
Grice, E. A. et al. Topographical and temporal diversity of the human skin microbiome. Science 324, 1190–1192 (2009).
Naber, C. K. Staphylococcus aureus bacteremia: epidemiology, pathophysiology, and management strategies. Clin. Infect. Dis. 48, S231–S237 (2009).
Naik, S. et al. Commensal–dendritic-cell interaction specifies a unique protective skin immune signature. Nature 520, 104–108 (2015).
Ramsey, M. M., Freire, M. O., Gabrilska, R. A., Rumbaugh, K. P. & Lemon, K. P. Staphylococcus aureus shifts toward commensalism in response to Corynebacterium species. Front. Microbiol. 7, 1230 (2016).
Lo, C.-W., Lai, Y.-K., Liu, Y.-T., Gallo, R. L. & Huang, C.-M. Staphylococcus aureus hijacks a skin commensal to intensify its virulence: immunization targeting β-hemolysin and CAMP factor. J. Invest. Dermatol. 131, 401–409 (2011).
McVicker, G. et al. Clonal expansion during Staphylococcus aureus infection dynamics reveals the effect of antibiotic intervention. PLoS Pathog. 10, e1003959 (2014).
Prajsnar, T. K. et al. A privileged intraphagocyte niche is responsible for disseminated infection of Staphylococcus aureus in a zebrafish model. Cell. Microbiol. 14, 1600–1619 (2012).
Prajsnar, T. K., Cunliffe, V. T., Foster, S. J. & Renshaw, S. A. A novel vertebrate model of Staphylococcus aureus infection reveals phagocyte-dependent resistance of zebrafish to non-host specialized pathogens. Cell. Microbiol. 10, 2312–2325 (2008).
Horsburgh, M. J., Wiltshire, M. D., Crossley, H., Ingham, E. & Foster, S. J. PheP, a putative amino acid permease of Staphylococcus aureus, contributes to survival in vivo and during starvation. Infect. Immun. 72, 3073–3076 (2004).
Bera, A., Biswas, R., Herbert, S. & Götz, F. The presence of peptidoglycan O-acetyltransferase in various staphylococcal species correlates with lysozyme resistance and pathogenicity. Infect. Immun. 74, 4598–4604 (2006).
Cheng, A. G. et al. Contribution of coagulases towards Staphylococcus aureus disease and protective immunity. PLos Pathog. 6, e1001036 (2010).
Grice, E. A. et al. A diversity profile of the human skin microbiota. Genome Res. 18, 1043–1050 (2008).
Sherertz, R. J. et al. Three-year experience with sonicated vascular catheter cultures in a clinical microbiology laboratory. J. Clin. Microbiol. 28, 76–82 (1990).
Sorbara, M. T. & Philpott, D. J. Peptidoglycan: a critical activator of the mammalian immune system during infection and homeostasis. Immunol. Rev. 243, 40–60 (2011).
Wheeler, R., Chevalier, G., Eberl, G. & Gomperts Boneca, I. The biology of bacterial peptidoglycans and their impact on host immunity and physiology. Cell. Microbiol. 16, 1014–1023 (2014).
Baba, T., Bae, T., Schneewind, O., Takeuchi, F. & Hiramatsu, K. Genome sequence of Staphylococcus aureus strain Newman and comparative analysis of staphylococcal genomes: polymorphism and evolution of two major pathogenicity islands. J. Bacteriol. 190, 300–310 (2008).
Surewaard, B. G. J. et al. Identification and treatment of the Staphylococcus aureus reservoir in vivo. J. Exp. Med. 213, 1141–1151 (2016).
Thammavongsa, V., Missiakas, D. M. & Schneewind, O. Staphylococcus aureus degrades neutrophil extracellular traps to promote immune cell death. Science 342, 863–866 (2013).
Wang, R. et al. Identification of novel cytolytic peptides as key virulence determinants for community-associated MRSA. Nat. Med. 13, 1510–1514 (2007).
Elek, S. D. & Conen, P. E. The virulence of Staphylococcus pyogenes for man; a study of the problems of wound infection. Br. J. Exp. Pathol. 38, 573–586 (1957).
Schleifer, K. H. & Kandler, O. Peptidoglycan types of bacterial cell walls and their taxonomic implications. Bacteriol. Rev. 36, 407–477 (1972).
Hashimoto, M. et al. Lipoprotein is a predominant Toll-like receptor 2 ligand in Staphylococcus aureus cell wall components. Int. Immunol. 18, 355–362 (2006).
Verdrengh, M. & Tarkowski, A. Role of neutrophils in experimental septicemia and septic arthritis induced by Staphylococcus aureus. Infect. Immun. 65, 2517–2521 (1997).
Rigby, K. M. & DeLeo, F. R. Neutrophils in innate host defense against Staphylococcus aureus infections. Semin. Immunopathol. 34, 237–259 (2012).
Heyworth, P. G., Cross, A. R. & Curnutte, J. T. Chronic granulomatous disease. Curr. Opin. Immunol. 15, 578–584 (2003).
Zeng, Z. et al. CRIg functions as a macrophage pattern recognition receptor to directly bind and capture blood-borne Gram-positive bacteria. Cell Host Microbe 20, 99–106 (2016).
Clarke, T. B. et al. Recognition of peptidoglycan from the microbiota by Nod1 enhances systemic innate immunity. Nat. Med. 16, 228–231 (2010).
Gauguet, S. et al. Intestinal microbiota of mice influences resistance to Staphylococcus aureus pneumonia. Infect. Immun. 83, 4003–4014 (2015).
Stoll, H., Dengjel, J., Nerz, C. & Götz, F. Staphylococcus aureus deficient in lipidation of prelipoproteins is attenuated in growth and immune activation. Infect. Immun. 73, 2411–2423 (2005).
Chamaillard, M. et al. An essential role for NOD1 in host recognition of bacterial peptidoglycan containing diaminopimelic acid. Nat. Immunol. 4, 702–707 (2003).
Wolf, A. J. et al. Hexokinase is an innate immune receptor for the detection of bacterial peptidoglycan. Cell 166, 624–636 (2016).
Pollock, J. D. et al. Mouse model of X-linked chronic granulomatous disease, an inherited defect in phagocyte superoxide production. Nat. Genet. 9, 202–209 (1995).
Boyle, J. P., Parkhouse, R. & Monie, T. P. Insights into the molecular basis of the NOD2 signalling pathway. Open Biol. 4, 140178 (2014).
Philpott, D. J., Sorbara, M. T., Robertson, S. J., Croitoru, K. & Girardin, S. E. NOD proteins: regulators of inflammation in health and disease. Nat. Rev. Immunol. 14, 9–23 (2014).
Korgaonkar, A., Trivedi, U., Rumbaugh, K. P. & Whiteley, M. Community surveillance enhances Pseudomonas aeruginosa virulence during polymicrobial infection. Proc. Natl Acad. Sci. USA 110, 1059–1064 (2013).
Nieto, C. & Espinosa, M. Construction of the mobilizable plasmid pMV158GFP, a derivative of pMV158 that carries the gene encoding the green fluorescent protein. Plasmid 49, 281–285 (2003).
Cheng, A. G. et al. Genetic requirements for Staphylococcus aureus abscess formation and persistence in host tissues. FASEB J. 23, 3393–3404 (2009).
Turner, R. D. et al. Peptidoglycan architecture can specify division planes in Staphylococcus aureus. Nat. Commun. 1, 26 (2010).
Sabroe, I., Williams, T. J., Hébert, C. A. & Collins, P. D. Chemoattractant cross-desensitization of the human neutrophil IL-8 receptor involves receptor internalization and differential receptor subtype regulation. J. Immunol. 158, 1361–1369 (1997).
Dockrell, D. H., Lee, M., Lynch, D. H. & Read, R. C. Immune-mediated phagocytosis and killing of Streptococcus pneumoniae are associated with direct and bystander macrophage apoptosis. J. Infect. Dis. 184, 713–722 (2001).
Bremell, T. et al. Outbreak of spontaneous staphylococcal arthritis and osteitis in mice. Arthritis Rheum. 33, 1739–1744 (1990).
Ali, A. et al. CTLA4 immunoglobulin but not anti-tumor necrosis factor therapy promotes staphylococcal septic arthritis in mice. J. Infect. Dis. 212, 1308–1316 (2015).
Kwiecinski, J., Jin, T. & Josefsson, E. Surface proteins of Staphylococcus aureus play an important role in experimental skin infection. APMIS 122, 1240–1250 (2014).
Wong, C. H. Y., Jenne, C. N., Lee, W.-Y., Léger, C. & Kubes, P. Functional innervation of hepatic iNKT cells is immunosuppressive following stroke. Science 334, 101–105 (2011).
Wilson, R. et al. Protection against Streptococcus pneumoniae lung infection after nasopharyngeal colonization requires both humoral and cellular immune responses. Mucosal Immunol. 8, 627–639 (2015).
Nusslein-Volhard, C. & Dahm, R. Zebrafish. A Practical Approach (Oxford Univ. Press, New York, 2002).
Horsburgh, M. J. et al. σB modulates virulence determinant expression and stress resistance: characterization of a functional rsbU strain derived from Staphylococcus aureus 8325-4. J. Bacteriol. 184, 5457–5467 (2002).
Mainiero, M. et al. Differential target gene activation by the Staphylococcus aureus two-component system saeRS. J. Bacteriol. 192, 613–623 (2010).
Duthie, E. S. & Lorenz, L. L. Staphylococcal coagulase: mode of action and antigenicity. J. Gen. Microbiol. 6, 95–107 (1952).
Pang, Y. Y. et al. agr-dependent interactions of Staphylococcus aureus USA300 with human polymorphonuclear neutrophils. J. Innate Immun. 2, 546–559 (2010).
Surewaard, B. G. J. et al. Inactivation of staphylococcal phenol soluble modulins by serum lipoprotein particles. PLoS Pathog. 8, e1002606 (2012).
Fey, P. D. et al. A genetic resource for rapid and comprehensive phenotype screening of nonessential Staphylococcus aureus genes. mBio 4, e00537-12 (2013).
Modun, B. J., Cockayne, A., Finch, R. & Williams, P. The Staphylococcus aureus and Staphylococcus epidermidis transferrin-binding proteins are expressed in vivo during infection. Microbiology 144, 1005–1012 (1998).
Hasenberg, A. et al. Catchup: a mouse model for imaging-based tracking and modulation of neutrophil granulocytes. Nat. Methods 12, 445–452 (2015).
Gomez de Agüero, M. et al. The maternal microbiota drives early postnatal innate immune development. Science 351, 1296–1302 (2016).
Acknowledgements
This work was funded by the Wellcome Trust (099957/Z/12/Z, 089981), Innovate UK (27486-188210), the Swedish Research Council (2013-09302), an MRC programme grant to S.A.R. (MR/M004864/1), an MRC grant to S.J.F. (MR/R001111/1) and the University of Sheffield 2022 Futures programme via the Florey Institute. Imaging used the Wolfson Light Microscopy Facility (supported by MRC grant MR/K015753/1). P.K. is supported by Alberta Innovates Health Solutions (AIHS), the Canadian Institutes of Health Research (CIHR) and the Canada Research Chairs Program. B.G.J.S. is partially funded by a postdoctoral fellowship from CIHR. The authors acknowledge use of the Bateson Centre aquarium, Biological Services Unit, Core Histology Service and the Flow Cytometry Facility at the University of Sheffield. The authors thank the International Microbiome Centre from the University of Calgary for assistance, the Bateson Centre aquarium staff for assistance with zebrafish husbandry, L. Prince, D. Yang, J. Hooker, F. Wright and A. Hendriks for advice and assistance, F. Götz for providing SA113lgt::ermB and M. Gunzer and A. Hasenberg for providing Ly6G-tdTomato reporter mice.
Author information
Authors and Affiliations
Contributions
E.B., B.G.J.S., D.S., M.N., Y.F., A.A., A.W., E.J.G.P., P.S., P.M. and T.K.P. performed and analysed the experiments. K.D.M., T.J., D.H.D., J.A.G.S., P.K., S.A.R. and S.J.F. contributed to study design and data analysis. E.B. and S.J.F. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Information
Supplementary Figures 1–4
Supplementary Video 1
Capture of S. aureus and PGN by Kupffer cells. Kupffer cells (F4/80, purple) capture most intravenous injected S. aureus (5 × 107 c.f.u., BSG1) from the circulation in female C57BL/6J mice. Neutrophils (Ly6g; red). Bacteria were injected 1 min after initiation of 20 min time interval visualized by SD-IVM. Videos represent one out of three independent experiments. Scale bar, 50 µm.
Supplementary Video 2
Kupffer cells (F4/80, purple) capture most intravenous injected S. aureus + PGN from the circulation in female C57BL/6J mice. Neutrophils (Ly6g; red). Bacteria were injected 1 min after initiation of 20 min time interval visualized by SD-IVM. Videos represent one out of three independent experiments. Scale bar, 50 µm.
Supplementary Video 3
Oxidative burst associated with S. aureus and PGN within Kupffer cells. SD-IVM video of mouse livers with labelled Kupffer cells (F4/80, purple), injected with pHrodo S. aureus bioparticles (red) additionally labelled with AF647 (blue) as a reference fluorophore, and OxyBURST (green) without co-injection of 500 µg PGN. Panels show individual label channels and a merge. S. aureus bioparticles were injected 1 min after initiation of 50 min time interval visualized by SD-IVM. Videos represent one out of three independent experiments. Scale bar, 25 µm.
Supplementary Video 4
Oxidative burst associated with S. aureus and PGN within Kupffer cells. SD-IVM video of mouse livers with labelled Kupffer cells (F4/80, purple), injected with pHrodo S. aureus bioparticles (red) additionally labelled with AF647 (blue) as a reference fluorophore, and OxyBURST (green) with co-injection of 500 µg PGN. Panels show individual label channels and a merge. S. aureus bioparticles were injected 1 min after initiation of 50 min time interval visualized by SD-IVM. Videos represent one out of three independent experiments. Scale bar, 25 µm.
Supplementary Video 5
Co-phagocytosis of S. aureus and latex beads in the zebrafish embryo model of infection. In vivo imaging of 9,000 latex beads (green) and S. aureus SH1000-mCherry (red, 1,500 c.f.u.) 2 hpi. Imaging for 5 minutes. Video represents one out of two independent experiments. Scale bar, 10 μm.
Rights and permissions
About this article
Cite this article
Boldock, E., Surewaard, B.G.J., Shamarina, D. et al. Human skin commensals augment Staphylococcus aureus pathogenesis. Nat Microbiol 3, 881–890 (2018). https://doi.org/10.1038/s41564-018-0198-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0198-3
This article is cited by
-
Formation of a biofilm matrix network shapes polymicrobial interactions
The ISME Journal (2023)
-
Clonal population expansion of Staphylococcus aureus occurs due to escape from a finite number of intraphagocyte niches
Scientific Reports (2023)
-
CuS nanoenzyme against bacterial infection by in situ hydroxyl radical generation on bacteria surface
Rare Metals (2023)
-
Characterization of Brazilian green propolis as a photosensitizer for LED light-induced antimicrobial photodynamic therapy (aPDT) against methicillin-resistant Staphylococcus aureus (MRSA) and Vancomycin-intermediate Staphylococcus aureus (VISA)
Photochemical & Photobiological Sciences (2023)
-
Skeletal infections: microbial pathogenesis, immunity and clinical management
Nature Reviews Microbiology (2022)