Abstract
An enormous diversity of previously unknown bacteria and archaea has been discovered recently, yet their functional capacities and distributions in the terrestrial subsurface remain uncertain. Here, we continually sampled a CO2-driven geyser (Colorado Plateau, Utah, USA) over its 5-day eruption cycle to test the hypothesis that stratified, sandstone-hosted aquifers sampled over three phases of the eruption cycle have microbial communities that differ both in membership and function. Genome-resolved metagenomics, single-cell genomics and geochemical analyses confirmed this hypothesis and linked microorganisms to groundwater compositions from different depths. Autotrophic Candidatus “Altiarchaeum sp.” and phylogenetically deep-branching nanoarchaea dominate the deepest groundwater. A nanoarchaeon with limited metabolic capacity is inferred to be a potential symbiont of the Ca. “Altiarchaeum”. Candidate Phyla Radiation bacteria are also present in the deepest groundwater and they are relatively abundant in water from intermediate depths. During the recovery phase of the geyser, microaerophilic Fe- and S-oxidizers have high in situ genome replication rates. Autotrophic Sulfurimonas sustained by aerobic sulfide oxidation and with the capacity for N2 fixation dominate the shallow aquifer. Overall, 104 different phylum-level lineages are present in water from these subsurface environments, with uncultivated archaea and bacteria partitioned to the deeper subsurface.
Similar content being viewed by others
Main
Much remains to be learned about how microbial communities in the deep terrestrial subsurface vary with depth due to limited access to samples without contamination from drilling fluids or sampling equipment. Studies to date have analysed samples acquired by drilling1,2,3, from deep mines4,5, subsurface research laboratories6,7 and sites of groundwater discharge8,9,10,11. These investigations have shown that the terrestrial subsurface is populated by a vast array of previously undescribed archaea and bacteria. At one site, an aquifer in Colorado (Rifle, USA), the diversity spans much of the tree of life12 and includes organisms of the Candidate Phyla Radiation (CPR)13, which may comprise more than 50% of all bacterial diversity14, and many other previously undescribed bacterial lineages. Also present in the terrestrial subsurface are previously unknown or little known archaea, including members of the DPANN (Diapherotrites, Parvarchaeota, Aenigmarchaeota, Nanoarchaeota, Nanohaloarchaea)11,15, Altiarchaeum10, Lokiarchaeota16 and Aigarchaeota17.
A major question in subsurface microbiology relates to how organisms, and their capacities for carbon, nitrogen and sulfur cycling, vary along depth transects through the terrestrial subsurface. Some evidence pointing to taxonomic variation between 9 m and 52 m below the surface was obtained via a massive 16S ribosomal RNA gene survey at the Hanford Site18. Similar variation and change of two functional genes were also detected for two shallow aquifers that were accessed via drilling in Germany19. However, major groups of archaea and bacteria may have been overlooked due to sampling13 and primer bias13, 20,21 and the spatial variation in metabolic functions over depth transects including the deep subsurface (100 m below the ground) remains unexplored.
Crystal Geyser is a cold-water, CO2-driven geyser located geologically within the Paradox Basin, Colorado Plateau, Utah, USA22. Originally an abandoned oil exploration well, the 800-m deep vertical borehole has served as a geyser conduit whose regular and significant flow rate (since 1936) provides uncontaminated access to organisms present in underlying aquifers. Prior geological studies have defined the region’s hydrostratigraphy, including the transmissive Entrada, Navajo, Wingate and White Rim fractured sandstone aquifers (listed in order of increasing depth), which are separated by low-permeability confining units23,24, through which limited vertical connectivity for CO2, water and microbes is largely restricted to faults and fractures25. A nearby research borehole provided further geologic and aquifer geochemical information to 322 m below ground surface26. Time-series geochemical data collected over the ca. 5-day eruption cycle suggest that Crystal Geyser is primarily sourced from the Navajo Sandstone, with increased contributions from the shallower Entrada Sandstone during major eruptions, and increased fraction of deeper water during minor eruptions26,27.
A survey of ribosomal proteins predicted from metagenome sequences from Crystal Geyser microbial communities revealed the existence of a large phylogenetic diversity of previously unknown bacteria and archaea in this system8, and a genomic resolution study documented a high incidence of carbon-fixation pathways9. A remaining question relates to the source regions and distributions of these organisms. Here, we tracked the microbiology and the associated geyser discharge geochemistry continuously throughout the full 5-day geyser eruption cycle to test the hypothesis that groundwater from stratified aquifers sampled at different stages of the cycle has microbial communities that differ in both membership and function. Our analyses made use of a comprehensive collection of more than 1,000 newly reconstructed genomes, both from metagenomes and single cells, as well as detailed physical and chemical information that enabled linking of fluids to their groundwater source regions.
Results
Continuous in situ (downhole) monitoring of the geyser water pressure throughout the field campaign defined the regular ~5-day period of the eruption cycle (Supplementary Fig. 1). Sampling was conducted over a complete eruption cycle (24–29 May, 2015) during which microbial cells were continuously collected onto 0.1 µm filters. Time series of downhole temperature, electrical conductivity, total dissolved gas pressure and water samples (for major ion, trace metal and dissolved gas analyses) were collected to associate specific microorganisms with water from different geyser eruption intervals and relative aquifer depths (Supplementary Fig. 1).
Time series of water pressure, electrical conductivity and temperature showed three Crystal Geyser eruption phases previously observed26,27: the recovery (relatively low water level, no eruptions, light CO2 bubbling), minor eruptions (short eruptions of ~10 min every hour with elevated CO2 discharge) and major eruptions (constant eruption and heavy CO2 discharge28; Fig. 1). Average chloride concentrations ([Cl]) and baseline water temperature (16.9 °C; also observed in a year-long monitoring period; Supplementary Fig. 1d) indicate that, overall, the geyser water is primarily sourced from ~320 to 480 m depth, which mainly corresponds to the Navajo aquifer; Supplementary Fig. 1a–d). In minor eruptions, elevated electrical conductivity and [Cl] indicate increased contribution from deeper, more saline water (that is, possibly the Wingate aquifer or Paradox brine sourced from even greater depth; Supplementary Fig. 1c). In the major eruption phase, decreased electrical conductivity and [Cl], and elevated Ca, Sr and Fe concentrations, were consistent with an increased contribution from the shallower Entrada aquifer (Supplementary Fig. 1b). During the eruption-free recovery phase, in which the Crystal Geyser borehole slowly refilled after the major eruption phase, electrical conductivity gradually increased with the relative contribution of deeper groundwater up to (and during) the minor eruptions. During this time the water level increased ~3.5 m over 33.5 h, potentially enabling microbes to thrive in microaerophilic borehole-affected conditions. To simplify terminology, we henceforth refer to the source water compositions as relatively ‘deep’ during the minor eruptions, ‘intermediate’ (and borehole-affected) during the recovery phase and ‘shallow’ during the major eruptions. Similar phase variations in the relative depths of water composition were recently observed27.
In each phase, electrical conductivity (EC) and geochemical measurements (6,710 measurements each, no technical replicates; Supplementary Fig. 1) are used to identify relative depths of source water compositions: intermediate for the recovery phase (2,330 measurements), deep for the minor eruptions (2,820 measurements) and shallow for the major eruptions (1,560 measurements). The numbered horizontal grey bars indicate the time periods for each metagenomic sample (25 samples in total) and coloured numbers indicate the grouping of samples from each phase.
Analysis of microbial community composition in the 2015 bulk samples made use of a relatively comprehensive database of genomic information for the Crystal Geyser system (see Supplementary Fig. 2 for sample processing and analysis overview). The genomic dataset included previously reported draft genomes from this system9 and new genomes reconstructed from size-fractionated samples obtained in April and October of 2014 (Supplementary Table 1). Samples included a post-0.2 µm fraction collected onto a 0.1-µm filter to enrich for community members with ultra-small cell sizes. Binning of assembled metagenomes from 27 different samples from five time points in 2014 used seven different algorithms (see Supplementary Methods) and resulted in 30,574 genomic bins (with multiple bins for the same genome generated by different algorithms). Selection of the best quality bin generated for any organism in each sample resulted in 5,795 bins for bacteria and archaea, of which 2,216 were considered to be at least medium quality (>70% completeness based on single-copy genes with less than three multiple single-copy genes). After curation based on guanine-cytosine content, coverage and taxonomy the database contains 1,215 genome sequences for 503 different archaeal and bacterial species (for details on genome numbers for each step, please see Supplementary Fig. 2; genome completeness is provided in Supplementary Table 2).
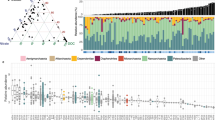
To augment the genome-resolved metagenomics, we acquired 206 single amplified genomes (SAGs) from cells collected at one time point in the minor eruption and one time point during the recovery phase. SAGs were chosen for full sequencing and analysis if PCR-screening for 16S rRNA genes was positive, agnostic to the specific sequence. Only SAGs with assembly size of >100 kbp after multi-step contamination screening were considered further. This set comprised 183 SAGs, seven of which were of sufficient quality to be classified as medium-quality draft genomes ( > 70% complete, less than three multiple single-copy genes). We required alignments ≥98% nucleotide identity over >30% of the SAG to establish a match between SAGs and genomes from metagenomes. This approach was chosen because almost all of the SAGs were less complete than related draft genomes from metagenomes (Supplementary Fig. 3). In general, SAG sequences aligned well to the sequences of genomes from the metagenomes (Supplementary Fig. 3). We found that >70% of the SAGs (145, of which five were draft quality) were represented in the set of 503 draft-quality genomes from the metagenomes. Conversely, 63 of the 503 genomes from metagenomes were also detected by single-cell genomics. Two draft-quality SAG genomes were not binned from the metagenomes and thus were added to the database. One SAG is entirely absent in the metagenomes based on sequences of the ribosomal protein S3 and read mapping, and probably derived from a very rare organism. The 505 genomes in the database (Supplementary Table 2), which were derived via dereplication from a total set of 1,208 genomes (984 genomes from metagenomes, 222 genomes from a previous study9 and two single-cell genomes), represent archaeal and bacterial species that belong to 104 different phylum-level lineages (Fig. 2). Nine lineages were named as they were represented by least two genomes with significant phylogenetic distance to neighbouring phyla and thus may constitute previously unrecognized phylum-level lineages. In addition, six genomes may be from previously unknown phylum-level lineages but the lineages are currently only represented by a single genome. The majority of diversity was attributed to members of the CPR (Fig. 2).
Genomes were reconstructed for organisms from 104 different phylum-level lineages; 503 different lineages are shown (two lineages did not exhibit >50% alignment coverage and are thus not displayed). Phyla in bold were assigned names in this study. The scale corresponds to per cent average amino acid substitution over the alignment. Asterisks mark yet-to-be-cultivated phyla, which thus have a Candidatus status. OD1, Parcubacteria. A full tree with reference sequences can be found in Supplementary File 1.
Mapping of metagenome reads to the set of 505 genomes showed that the genomes account for ~50% of the sequence data collected through the 2015 eruption cycle (Supplementary Table 4) and thus is representative of the community found in the Crystal Geyser ecosystem (Supplementary Fig. 4; morphological diversity of organisms is provided in Supplementary Fig. 5). Analysis of the community structure of the 25 different metagenomes using this approach revealed strong shifts in microbial composition over the cycle (Fig. 3b; Supplementary Table 5). The accuracy of relative abundance measures of individual genomes was confirmed for three species using quantitative digital droplet PCR (Supplementary Fig. 6). No physical or geochemical factor besides time, which corresponds to the source regions of the sampled water, could explain the observed changes in community composition (based on multivariate statistics; Supplementary Table 6). The community was dominated by species of the taxa Candidatus “Altiarchaeum”, Sulfurimonas, Piscirickettsiaceae, Gallionellaceae and Betaproteobacteria (in order of decreasing abundance, Fig. 3c). The set of CPR and archaea from the DPANN superphylum showed several peaks in relative abundance at several time points during the eruption cycle (Fig. 3d,e). Both groups had the highest cumulative abundance during the minor eruptions, when groundwater from the deepest source was sampled. The most abundant CPR (Moranbacteria13) was, however, prominent during the recovery phase (Fig. 3d). Overall, the cumulative abundances of DPANN and other archaea were significantly higher in the deep groundwater compared to shallow or intermediate (Supplementary Fig. 7).
a, The Crystal Geyser site lies within one of the several natural CO2 reservoirs within the Paradox Basin. The CO2 was probably generated from thermal decomposition of Pennsylvanian-aged carbonate rocks26, 51,52. CO2 gas and brine formed by groundwater dissolution of Paradox evaporites migrate via faults and fractures53,54. b, The community profile of 505 organisms strongly followed the succession of the geyser eruptions (blue lines, NMDS). One data point corresponds to one metagenomic sample. The samples show a clear pattern following the succession of the geyser cycle. c, Entire community profile of 505 organisms tracked across the 5-day cycle of the geyser. Each colour corresponds to one genome. d,e, Profiles of the CPR and DPANN community, respectively, show an increase in the overall abundance during the minor eruptions when groundwater has the deepest source composition. f, Downhole electrical conductivity time series during the sampling of the cycle illustrating the individual phases of the geyser (6,710 samples were measured, see Fig. 1 and Supplementary Fig. 1). Number of biological replicates in panels b–e was 24. EC, electrical conductivity; GW, groundwater.
When analysed one at a time, the majority of organisms (289, ~57%) were significantly enriched (false discovery rate-corrected P value <0.05) in one specific phase of the geyser and could thus be sourced to one of the groundwater depths (Fig. 4a–c). The shallowest groundwater was mainly populated by one Sulfurimonas sp. along with a few other bacteria and some archaea. Based on the genome sequence of Sulfurimonas sp., this organism was inferred to be a chemolithoautotroph, capable of nitrogen and carbon fixation as well as sulfide oxidation through oxygen respiration (Supplementary Table 7). The capacity for carbon fixation via the low-cost reductive TCA cycle (two ATP per pyruvate29) coupled to oxygen respiration may provide an ecological advantage for this species and is also indicative of microaerophilic conditions in the relatively shallow aquifer. In contrast to the shallow source, groundwater from intermediate depths had a great diversity of different organisms, the majority of which belonged to the CPR. The most abundant organism was a member of the Gallionellaceae, a family of bacteria well known for microaerophilic iron and sulfur oxidation at Crystal Geyser8,9. This organism also exhibited the highest genome replication rates of all bacteria in the study (average in situ replication rate (iRep) value of 2.5, maximum iRep value of 4.2; Supplementary Table 8), suggesting that it was also proliferating in the geyser conduit over the 33.5 h of the recovery phase. Its growth was probably favoured by microaerophilic conditions as well as sulfide and reduced iron in the geyser fluids. Potentially, other microorganisms enriched in this fraction may also have favoured the conditions in the borehole over the 33.5-hour recovery phase, during which the geyser had no water discharge. Consequently, the community sampled from the recovery phase represents the community from intermediate depths with distortions from microbial growth in the borehole. When deeper groundwater was discharged, the abundances of different DPANN archaea and Ca. “Altiarchaeum” were significantly increased. Diverse members of the CPR were still present in deep groundwater, although at low relative abundance.
a–c, The abundances of organisms that are significantly enriched in groundwater from the different depths (for details on organisms please see Supplementary Table 5, number of biological replicates are given in parenthesis of panels a–c). The pie charts indicate the diversity of CPR, DPANN, other bacteria, and other archaea associated with each relative depth. d, Different carbon fixation pathways predominate in groundwater from the three different depths. Nitrogen fixation and the reverse TCA cycle occur in one organism, Sulfurimonas (a). e, Metabolic pathway analysis shows distinct metabolic profiles associated with the groundwater from the different depths (individual metabolic capacities of each organism are listed in Supplementary Table 7). Each circle displays the cumulative relative abundance of genomes contributing to this single metabolic process. Arrows display if an increase or decrease is significant (P < 0.05). CBB, Calvin-Benson-Bassham; disprop., disproportionation.
The shallowest groundwater had a substantially higher capacity for microbial sulfide oxidation, nitrogen fixation and oxygen respiration, probably due to the presence of atmospheric gases. In contrast, the intermediate source and borehole community had the highest microbial capacity for reduction of various nitrogen compounds as well as thiosulfate disproportionation, metal reduction and oxidation. The deepest groundwater was enriched in several bacteria with the capacity for sulfite reduction, with carbon fixation mediated by the Ca. “Altiarchaeum”. The capacity for oxygen respiration decreased with increasing depth to the sourced groundwater.
Previously, we reported the operation of three carbon fixation pathways in bacteria and archaea from the Crystal Geyser communities, each of which requires substantially different amounts of energy9. While the Wood–Ljungdahl pathway requires approximately 1 mol of ATP for the generation of 1 mol pyruvate, the Calvin–Benson–Bassham cycle and the reverse TCA cycle require 7 mol and 2 mol, respectively29. Here, we show that the three carbon fixation pathways used by Crystal Geyser microorganisms were most abundant in different eruption phases, which reflect varying depths of source water composition (and borehole; Fig. 4e). The reductive TCA cycle was associated with a Sulfurimonas sp. that dominates the shallowest groundwater and also has the capacity for N2 fixation. The Calvin–Benson–Bassham cycle was enriched in bacteria associated with the intermediate groundwater as well as the borehole, and the Wood–Ljungdahl pathway was encoded in Ca. “Altiarchaeum” and Deltaproteobacteria genomes, and thus most highly represented in the deepest groundwater (Fig. 4d).
One previously undescribed archaeal phylum-level lineage within the DPANN branched next to Parvarchaeota (ARMAN-5) and Nanoarchaeum equitans. The 16S rRNA gene sequence of this species shared less than 67% identity with any 16S rRNA gene available in the SILVA database30 (and <78% with 16S rRNA gene fragments from environmental samples in the National Center for Biotechnology Information). We reconstructed 11 genomes for this species (including one from a single isolated cell) and estimated the genome size to be about 0.5 Mbp, which is similar to those of Nanoarchaeum equitans and some other DPANN15,31. We propose the name Candidatus “Huberarchaeum crystalense” (phylum Ca. “Huberarchaea”) for this archaeal lineage in honour of Prof. Robert Huber, pioneer in research on psychrophilic archaea and discoverer of Ca. “Altiarchaeum”.
Although enzymes for modification of purine and pyrimidine bases were encoded in the genome of Ca. “H. crystalense” (for example, via methylated folate), it is predicted to be incapable of de novo nucleotide synthesis (Supplementary File 2). The genome encodes a near-complete set of aminoacyl transfer RNA synthetases, proteins for replication and repair of DNA and translation and transcription machinery. Amino acids, whose biosynthesis pathways were lacking, are probably acquired via five different proteases. It has enzymes for glycosylating proteins and lipids and a near-complete pathway for lipid biosynthesis, observations that support the claim that this is a cellular organism. Protein export was probably accomplished via an encoded sec-pathway. Based on the limited metabolism of Ca. “H. crystalense”, we infer a symbiotic lifestyle. Interaction of the symbiont and a host may be mediated via large surface proteins, some of which are Cys-rich32. One of the extracellular, membrane-anchored Cys-rich proteins is predicted to bind calcium (Supplementary Fig. 8)33, a function also commonly found in hemolysin proteins. Hemolysin proteins destroy cell membranes, an activity that might be pivotal for Ca. “H. crystalense” to access metabolites from its host.
Within the whole geyser community, Ca. “H. crystalense” is the seventh most abundant organism. Notably, its abundance correlated significantly with that of the dominant organism, Ca. “Altiarchaeum” (linear correlation, P value < 2.8 × 10E-12; Fig. 5a). Based on the correlation of abundance patterns, we suggest that Ca. “H. crystalense” is a symbiont of the Ca. “Altiarchaeum”. Some support for this may be provided by scanning electron microscope images, which showed small rounded structures of approximately 0.15-µm diameter attached to larger cells (Fig. 5b, Supplementary Fig. 9). We infer that the larger cells are Ca. “Altiarchaeum”, based on the distinct hami-like appendages10, and that Ca. “H. crystalense” are episymbionts. Interestingly, both genomes exhibited very high levels of fragmentation, an indication of high levels of strain heterogeneity within both populations. The diversification of the Ca. “Altiarchaeum” host in its deep subsurface habitat might drive coevolution of Ca. “H. crystalense”. The shared characteristic of strain heterogeneity may also support the inference of their interaction.
a, Linear correlation analysis of relative abundance of the two archaea across 25 metagenome samples (full cycle of the geyser). b, Scanning electron micrograph of what are inferred to be Ca. “Altiarchaeum” cells (“SM1”) taken during the minor eruptions of the geyser. Tiny cell-like structures appear to be attached to the surface (“?N”). This structure was observed in two out of five samples taken for scanning electron microscopy analysis from the geyser fluids. cor., correlation coefficient. More images are available under Supplementary Fig. 9.
Discussion
Our microbiological investigation clearly demonstrated a strong stratification of microbial community composition and microbial function with relative groundwater source depths. Groundwater sampled from all three relative depths was dominated by autotrophs. The main pathway used for carbon fixation in the deeper subsurface is the one with the lowest energy cost, the Wood–Ljungdahl pathway, possibly because the deep biosphere is the most energy limited. Use of this pathway for provision of organic carbon was reported recently for other deep biosphere communities3,6. This pathway is also central to metabolism of methanogens, archaeal autotrophs found in the deep subsurface34. Reliance on the Wood–Ljungdahl pathway for CO2 fixation may be a widespread phenomenon in such environments. Our results indicate that the carbon provided by primary producers operating different carbon fixation pathways sustains a wide variety of bacteria and archaea in the subsurface.
This study adds to a growing body of literature that suggests that terrestrial subsurface regions are biodiversity hotspots6,12. Notably, we find that deeper regions can be particularly enriched in candidate phyla bacteria (especially CPR), DPANN archaea and other deep-branching archaea. The CPR were the most diverse organisms in the system. Intriguingly, many of these enigmatic CPR and DPANN are inferred to be symbionts13,15, probably episymbionts of other bacteria or archaea35. One highly abundant DPANN is a putative episymbiont of the most abundant archaeon, Ca. “Altiarchaeum”; however, further investigations are necessary to confirm this association. The putative symbiotic relationship between Ca. “Altiarchaeum” and Ca. “H. crystalense” could be analogous to that described between Ignicoccus hospitalis and Nanoarchaeum equitans36. Although Ca. “Altiarchaeum” is found elsewhere in the subsurface10, Ca. “H. crystalense” has not been detected in other metagenomic studies. The frequent detection of CPR and DPANN in groundwater, as found in this and other studies6, 8, 9, 12, 13, 15, may reflect the advantage of existence as ultra-small cells that can be readily distributed through sediment pore spaces, allowing periodic encounters with potential host organisms (Fig. 6). Highly interdependent lifestyles and intimate metabolic connections among community members may be an adaptation to constant low-nutrient conditions at depth.
The microbiome is dominated by Ca. “Altiarchaeum” (SM1) and their putative DPANN symbionts and populated by many CPR and other bacteria, some of which are probably symbiotic partners for CPR. We envision facile distribution of the very small CPR and DPANN cells through the sandstone pore spaces, providing periodic opportunities for establishment of the symbiont–host interactions that are probably required for CPR and DPANN cell replication. This figure provides a conceptual diagram of generalized microbial habitats in the aquifer based on an approximate pore size of sandstone. However, we note the subsurface is a heterogeneous three-dimensional system and physical properties will vary substantially55. The Carmel and Kayenta formations are expected to act as aquitards (confining barriers) that separate the high permeability sandstone aquifers (Fig. 3a), with each aquifer largely confined, both hydrologically and microbiologically, from other aquifers by these low-permeability shale/mudstone units56. This physical separation by low-permeability units probably contributes to the distinctive microbial communities associated with the three relative groundwater source depths as documented in the study.
Methods
Water chemistry and isotopes
Downhole electrical conductivity, water pressure and temperature were monitored using a Solinst LTC logger located in Crystal Geyser borehole about 8.5 m below ground surface. Water samples for major ions and trace metals were collected hourly at a pumping rate of 0.2 l min-1 over the eruption cycle from the borehole from a sampling tube inserted to 8.5 m below ground surface, and from the Green River. During two individual minor eruptions, samples were collected approximately every 10 min. Samples were field filtered to 0.2 μm by hand using Acrodisc syringe filters and a plastic syringe before collection into prerinsed 60 ml scintillation vials and then acidified to pH 2 with high-purity nitric acid for sample preservation37. The bottles were frozen for transport to the University of Calgary. Alkalinity was measured in the laboratory using an Orion Autochemistry 960 Autotitrator with 0.2 N sulfuric acid within 1 month of collection and expressed as HCO3 concentration. Major element and trace metal concentrations were determined using Inductively Coupled Plasma Emission Spectrometry and Inductively Coupled Plasma Mass Spectrometry, respectively38, at the Geologic Survey of Canada in Calgary.
Dissolved gas collection and analysis
Water samples for dissolved gas composition were taken simultaneously with water chemistry and isotope samples, collected using the inverted bottle method39 in 12 ml glass bottles capped with precision seal silicone septa caps. All samples were refrigerated until analysis at the University of Calgary. Dissolved gas compositions were determined by gas chromatography using headspace extraction. Due to our primary interest in CO2 gas in this system, we used a headspace to sample water ratio of 3:1 and shaking time of 12 min at 400 rpm40. Headspace samples were injected onto an HP 5890 (Hewlett-Packard) gas chromatograph with a Hamilton gas-tight syringe via a six-port, two-position sampling valve. The gas chromatograph was outfitted with parallel Rt-Msieve 5 A (Restek, 30 m × 0.32 mm) and Rt-Q-PLOT (Restek, 30 m × 0.53 mm), and data were collected using an HP 3396 Series II Integrator (Hewlett-Packard).
Genome-resolved metagenomics and single-cell genomics
Methods for genomic analysis of the 2014 datasets (including estimation of genome completeness) can be found in the Supplementary Methods.
Crystal geyser genome database
The genome database was constructed from genomes, from metagenomes and from single-cell genomes (SAGs) collected in 2014. First, all curated, newly binned genomes from metagenomes (985 in total) were combined with 222 previously published genomes9 and clustered based on 98% nucleotide identity. One representative of each genome cluster was chosen based on the highest completeness (single-copy genes) and lowest amount of contamination (multiple single-copy genes) following the formula: score = single-copy genes—2x multiple single-copy genes9,12. In cases of ties, the genome with the highest N50 was chosen. The resulting 503 archaea and bacteria were then compared against draft-quality SAGs (at least 70% complete) using 98% nucleotide identity. Two draft-quality SAGs were not covered by the genomes from metagenomes and were thus added to the Crystal Geyser database that consists of 505 archaeal and bacterial species used for downstream analyses. A schematic overview of the procedure is presented in Supplementary Fig. 2.
Comparison of genomes from metagenomes to SAGs
Whole-genome alignment of genomes from metagenomes9 was performed at 98% nucleotide identity. If a SAG shared more than 30% of its genomic content with a genome from a metagenome (which were at least 70% complete), the SAG was considered to be represented by the genome from the metagenome (Supplementary Fig. 4).
Phylogeny of bacteria and archaea
Phylogenetic placements of the 505 archaea and bacteria in the Crystal Geyser database were determined from a tree computed from 16 ribosomal proteins14 and included 3,609 sequences (including reference sets from previous studies12,14). Bacterial ribosomal proteins were extracted using usearch41 against a public database9 (https://github.com/AJProbst/sngl_cp_gn), while archaeal ribosomal proteins were first selected by searching against Hidden Markov Models (HMMs)42 built from a previous dataset14 (to exclude A/E type) and then annotated against UniRef43. Individually aligned protein sequences44 were end trimmed and gaps (<5% coverage) were removed before concatenation of protein sequences. Only sequences of genomes that spanned at least 50% of the alignment were included in the phylogenetic analysis; others were classified using ribosomal protein S3 or 16S rRNA gene sequences. Trees were computed as described earlier14. Taxa from Probst et al., 2016 that were redefined in their taxonomy are listed in Supplementary Table 2.
Tracking community members across time
In 2015, time resolution of the Crystal Geyser community was achieved by near-continuous filtration of groundwater onto 0.1-µm filters (ZTECG, Graver Technologies, Glasgow, USA) that we recovered at 25 different time points over a time course of nearly 5 days (114 h). Filtration was performed for an average of 4.6 h per sample and sampling spanned one entire eruption cycle of the geyser (Fig. 1). Metagenomic DNA was extracted from the samples8 and paired-end sequenced (Illumina HiSeq 2500, Supplementary Table 1). Reads of 150 bp were quality filtered (see Supplementary Methods) and mapped onto the de-replicated genome set of 505 organisms using bowtie245 (default settings), allowing three mismatches per 150-bp read (98% identity)46. Read coverage was normalized by genome size and relative abundances of each genome in each sample were normalized by number of reads per sample using the equation Ar = Nm/Ns * (r * l) g-1, where Ar is the relative abundance of the genome in a particular sample, Nm is the maximum number of reads of all metagenome samples, Ns is the total number of reads of that particular sample, r is the number of reads of that particular sample that mapped to the genome, l is the average read length and g is the length of the genome.
iRep values were calculated with one mismatch per read as described earlier46. Relative abundance measure using metagenomics was confirmed using quantitative digital droplet PCR (ddPCR), for which the method can be found in the Supplementary Methods.
Microbial metabolism from genomics
Microbial metabolism from genomics was predicted as described earlier9,12. In brief, genes for each genome were predicted using prodigal47 with the respective genetic code and key metabolic genes for various sulfur, nitrogen, hydrogen and metal redox processes were predicted using HMMs12,42. In addition, functional predictions against KEGG were performed on the basis of HMMs including hits with e-values < E-109. As such, the genetic potential of organisms for carbon fixation and oxygen respiration was based on the presence of all key enzymes and a pathway coverage of at least 60% in the KEGG module.
Microbial community statistics
Ordination analyses of microbial community structure was performed using a Bray–Curtis distance measure and non-metric multidimensional scaling (NMDS) in the R programming environment48. Influence of environmental factors as provided in Supplementary Table 1 were determined by BioENV (Bray–Curtis dissimilarity, Spearman correlation) and plotted onto the NMDS49. Microbial source tracking of organisms and changes in microbial metabolism between different groundwater source depths based on cumulative abundance of organisms was performed using analysis of variance coupled to a Tukey honest significant difference post hoc test. Sample designations of the different depths correspond to those provided in Fig. 1. All P values that were affected by multiple testing were corrected for false discovery rate using the Benjamini–Hochberg procedure50.
Scanning electron microscopy
Methods can be found in the supplementary documents.
Life Sciences Reporting Summary
Further information on experimental design is available in the Life Sciences Reporting Summary.
Data availability
SRA accession numbers for metagenomes of each sample are provided in Supplementary Table 1. All genomes from metagenomes included in this study were deposited at NCBI under Bioprojects PRJNA362739, PRJNA349044 and PRJNA297582. Genomes from metagenomes and single-cell genomes are also available under: http://ggkbase.berkeley.edu/CG_2014_505_non-redundant_genomes/organisms, http://ggkbase.berkeley.edu/CG_2014_genomes_from_metagenomes/organisms and http://ggkbase.berkeley.edu/CG_2014_SAGs/organisms.
References
Wrighton, K. C. et al. Fermentation, hydrogen, and sulfur metabolism in multiple uncultivated bacterial phyla. Science 337, 1661–1665 (2012).
Pedersen, K., Arlinger, J., Ekendahl, S. & Hallbeck, L. 16S rRNA gene diversity of attached and unattached bacteria in boreholes along the access tunnel to the Äspö hard rock laboratory, Sweden. Fems. Microbiol. Ecol. 19, 249–262 (1996).
Magnabosco, C. et al. A metagenomic window into carbon metabolism at 3 km depth in Precambrian continental crust. ISME J. 10, 730–741 (2016).
Chivian, D. et al. Environmental genomics reveals a single-species ecosystem deep within Earth. Science 322, 275–278 (2008).
Bagnoud, A. et al. Reconstructing a hydrogen-driven microbial metabolic network in Opalinus Clay rock. Nat. Commun. 7, 12770 (2016).
Hernsdorf, A. W. et al. Potential for microbial H2 and metal transformations associated with novel bacteria and archaea in deep terrestrial subsurface sediments. ISME J. 11, 1915–1929 (2017).
McKelvie, J. R., Korber, D. R. & Wolfaardt, G. M. in Their World: A Diversity of Microbial Environments (ed. Hurst, C. J.) 251–300 (Springer International Publishing, Switzerland, 2016).
Emerson, J. B., Thomas, B. C., Alvarez, W. & Banfield, J. F. Metagenomic analysis of a high carbon dioxide subsurface microbial community populated by chemolithoautotrophs and bacteria and archaea from candidate phyla. Environ. Microbiol. 18, 1686–1703 (2016).
Probst, A. J. et al. Genomic resolution of a cold subsurface aquifer community provides metabolic insights for novel microbes adapted to high CO2 concentrations. Environ. Microbiol. 19, 459–474 (2017).
Probst, A. J. et al. Biology of a widespread uncultivated archaeon that contributes to carbon fixation in the subsurface. Nat. Commun. 5, 5497 (2014).
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013).
Anantharaman, K. et al. Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nat. Commun. 7, 13219 (2016).
Brown, C. T. et al. Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523, 208–211 (2015).
Hug, L. A. et al. A new view of the tree of life. Nat. Microbiol. 1, 16048 (2016).
Castelle, C. J. et al. Genomic expansion of domain archaea highlights roles for organisms from new phyla in anaerobic carbon cycling. Curr. Biol. 25, 690–701 (2015).
Zaremba-Niedzwiedzka, K. et al. Asgard archaea illuminate the origin of eukaryotic cellular complexity. Nature 541, 353–358 (2017).
Nunoura, T. et al. Insights into the evolution of Archaea and eukaryotic protein modifier systems revealed by the genome of a novel archaeal group. Nucleic Acids Res. 39, 3204–3223 (2011).
Lin, X., Kennedy, D., Fredrickson, J., Bjornstad, B. & Konopka, A. Vertical stratification of subsurface microbial community composition across geological formations at the Hanford Site. Environ. Microbiol. 14, 414–425 (2012).
Herrmann, M. et al. Large fractions of CO2-fixing microorganisms in pristine limestone aquifers appear to be involved in the oxidation of reduced sulfur and nitrogen compounds. Appl. Environ. Microbiol. 81, 2384–2394 (2015).
Baker, B. J. et al. Lineages of acidophilic archaea revealed by community genomic analysis. Science 314, 1933–1935 (2006).
Eloe-Fadrosh, E. A., Ivanova, N. N., Woyke, T. & Kyrpides, N. C. Metagenomics uncovers gaps in amplicon-based detection of microbial diversity. Nat. Microbiol. 1, 15032 (2016).
Nuccio, V. F. & Condon, S. M. Burial and thermal history of the Paradox Basin, Utah and Colorado, and petroleum potential of the Middle Pennsylvanian Paradox Formation. US Geol. Surv. Bull. 2000–O (1996).
Trimble, L. M. & Doelling, H. H. The geology and uranium vanadium deposits of the San Rafael River Mining Area, Emery County, Utah. Utah Geol. Surv. Bull. 1046-D, 113 (1978).
Hood, J. W. & Patterson, D. J. Bedrock Aquifers in the Northern San Rafael Swell Area, Utah, with Special Emphasis on the Navajo Sandstone 139 (Utah Department of Natural Resources, Division of Water Rights, 1984).
Freethey, G. W. & Cordy, G. E. Geohydrology of Mesozoic Rocks in the Upper Colorado River Basin in Arizona, Colorado, New Mexico, Utah, and Wyoming, excluding the San Juan Basin. US Geol. Surv. Prof. Pap. 1411-C (1991).
Kampman, N. et al. Drilling and sampling a natural CO2 reservoir: implications for fluid flow and CO2-fluid–rock reactions during CO2 migration through the overburden. Chem. Geol. 369, 51–82 (2014).
Han, W. S. et al. Periodic changes in effluent chemistry at cold-water geyser: Crystal geyser in Utah. J. Hydrol. 550, 54–64 (2017).
Watson, Z. T., Han, W. S., Keating, E. H., Jung, N.-H. & Lu, M. Eruption dynamics of CO2-driven cold-water geysers: Crystal, Tenmile geysers in Utah and Chimayó geyser in New Mexico. Earth. Planet. Sci. Lett. 408, 272–284 (2014).
Berg, I. A. et al. Autotrophic carbon fixation in archaea. Nat. Rev. Microbiol. 8, 447 (2010).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Waters, E. et al. The genome of Nanoarchaeum equitans: insights into early archaeal evolution and derived parasitism. Proc. Natl Acad. Sci. USA 100, 12984–12988 (2003).
Anantharaman, K. et al. Analysis of five complete genome sequences for members of the class Peribacteria in the recently recognized Peregrinibacteria bacterial phylum. Peer J. 4, e1607 (2016).
Sharon, I. et al. Time series community genomics analysis reveals rapid shifts in bacterial species, strains, and phage during infant gut colonization. Genome Res. 23, 111–120 (2013).
Lau, M. C. Y. et al. An oligotrophic deep-subsurface community dependent on syntrophy is dominated by sulfur-driven autotrophic denitrifiers. Proc. Natl Acad. Sci. USA 113, E7927–E7936 (2016).
He, X. et al. Cultivation of a human-associated TM7 phylotype reveals a reduced genome and epibiotic parasitic lifestyle. Proc. Natl Acad. Sci. USA 112, 244–249 (2015).
Huber, H. et al. A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont. Nature 417, 63–67 (2002).
Standard Guide for Sampling Ground-Water Monitoring Wells ASTM D4448-01 (ASTM International, West Conshohocken, 2013).
Rice, E. W., Baird, E. B., Eaton, A. D. & Clesceri, L. S. Standard Methods for the Examination of Water and Wastewater (American Public Health Association, American Water Works Association, Water Environment Federation, 2012).
Clark, I. D. & Fritz, P. Environmental Isotopes in Hydrogeology (CRC Press, New York, 1997).
Jahangir, M. M. et al. Evaluation of headspace equilibration methods for quantifying greenhouse gases in groundwater. J. Environ. Manag. 111, 208–212 (2012).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Johnson, L.S., Eddy, S. R. & Portugaly, E. Hidden Markov model speed heuristic and iterative HMM search procedure. BMC Bioinformatics 11, 431 (2010).
Suzek, B. E., Huang, H., McGarvey, P., Mazumder, R. & Wu, C. H. UniRef: comprehensive and non-redundant UniProt reference clusters. Bioinformatics 23, 1282–1288 (2007).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Brown, C. T., Olm, M. R., Thomas, B. C. & Banfield, J. F. Measurement of bacterial replication rates in microbial communities. Nat. Biotechnol. 34, 1256–1263 (2016).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. Bmc. Bioinforma. 11, 119 (2010).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2016); http://www.r-proj.org
Weinmaier, T. et al. A viability-linked metagenomic analysis of cleanroom environments: eukarya, prokaryotes, and viruses. Microbiome 3, 62 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol. 57, 289–300 (1995).
Mayo, A. L., Shrum, D. B. & Chidsey Jr., T. C. Factors contributing to exsolving carbon dioxide in ground water systems in the Colorado Plateau, Utah. Utah Geol. Assoc. Publ. 19, 335–342 (1991).
Shipton, Z. K. et al. Analysis of CO2 leakage through ‘low-permeability’ faults from natural reservoirs in the Colorado Plateau, east-central Utah. Geol. Soc. Lond. Spec. Publ. 233, 43–58 (2004).
Allis, R. et al. Implications of results from CO flux surveys over known CO systems for long-term monitoring. In Fourth Annual Conference on Carbon Capture and Sequestration (DOE/NETL, Alexandria, VA, 2005).
Jung, N.-H., Han, W. S., Han, K. & Park, E. Regional-scale advective, diffusive, and eruptive dynamics of CO2 and brine leakage through faults and wellbores. J. Geophys. Res. Solid Earth 120, 2014JB011722 (2015).
Stegen, J. et al. Coupling among microbial communities, biogeochemistry, and mineralogy across biogeochemical facies. Sci. Rep. 6, 30553 (2016).
Krumholz, L. R., McKinley, J. P., Ulrich, G. A. & Suflita, J. M. Confined subsurface microbial communities in Cretaceous rock. Nature 386, 64 (1997).
Acknowledgements
We thank J. Hinshaw for his contribution to fieldwork logistics and year-long temperature monitoring. We also thank C. Brown, S. Spaulding, S. Clingenpeel, D. Barton, B. Rocha and D. Bethune for logistic support during fieldwork. C. Niemann provided technical assistance regarding scanning electron microscopy. D. Goudeau is acknowledged for help with single-cell lab work. We thank T. G. del Rio for handling of metagenomic samples at JGI. We are grateful to S. Gribaldo for discussion of the phylogenetic placement of bacterial and archaeal phyla. A.J.P. was supported by the German Science Foundation under DFG PR 1603/1-1 and by Lawrence Berkeley National Laboratory’s Sustainable Systems Scientific Focus Area funded by the US Department of Energy, Office of Science, Office of Biological and Environmental Research under contract DE-AC02-05CH11231. Work at UCB was funded by the Sloan Foundation (“Deep Life”, grant no. G-2016-20166041). Funding for hydrogeological and geochemical analyses was provided by a Natural Sciences and Engineering Research Council of Canada Discovery Grant to M.C.R. Development of ggKbase was supported by the Office of Science, Office of Biological and Environmental Research, of the US Department of Energy Grant DOE-SC10010566. Work conducted by the US Department of Energy Joint Genome Institute, a DOE Office of Science User Facility, is supported under Contract No. DE-AC02-05CH11231. We thank DOE’s Emerging Technologies Opportunity Program “Development of a pipeline for high-throughput recovery of near-complete and complete microbial genomes from complex metagenomic datasets” for sequencing.
Author information
Authors and Affiliations
Contributions
A.J.P., B.L., J.B.E., K.A., M.C.R. and JFB sampled the ecosystem. B.L. and M.C.R. conducted hydrogeological and geochemical analyses. A.J.P. performed genome-resolved metagenomics, phylogenetic, metabolic and community analyses. D.E.G.M. performed digital droplet PCR. C.M.K.S. and B.C.T. provided software. J.J. and T.W. performed single-cell genomics. A.J.P., J.J. and R.R.M. performed analyses of single-cell genomes. M.S. and A.K. carried out scanning electron microscopy. A.J.P. and J.F.B. designed the study. A.J.P. and J.F.B. wrote the paper with input from B.L. and M.C.R. All authors revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary Figures 1–9, Supplementary Table 4 and Supplementary References.
Supplementary File 1
Phylogenetic tree based on 16 concatenated ribosomal proteins.
Supplementary File 2
HMM profile of all 11 genomes of Candidatus ’Huberarchaeum crystalense’. The completeness of each pathway is displayed and the individual enzymes for each KEGG module are displayed by clicking onto the respective numbers (counts) of each module. The displayed predictions were retrieved via HMM search against each single KEGG enzyme with e-values < E-10 as described in the methods. For details on HMM generation please see Probst et al., 2016.
Supplementary Table 1
Sample overview. All samples had a read length, or were trimmed to a read length, of 150 bp.
Supplementary Table 2
Genome completeness of 983 genomes from metagenomes and 183 single-cell genomes based on 51 bacterial single-copy genes and 38 archaeal single-copy genes.
Supplementary Table 3
Overview of the taxonomy of the 505 genomes and novel phylum names that have Candidatus status.
Supplementary Table 5
Normalized relative abundance of organisms from Crystal Geyser across the 25 metagenome samples (see Supplementary Table 3). Relative abundance values are based on bowtie229 mapping allowing a maximum of three mismatches per read (98% identity; see methods). Column B, C and D indicate where the respective organism was enriched.
Supplementary Table 6
Overview of the environmental variables collected for each metagenomic sample. Continuously collected data for variables like temperature were averaged over the sampling time, in which the metagenomic sample was acquired. BioENV was performed on all continuous variables listed, except sampling date.
Supplementary Table 7
Metabolic profile of 505 species detected at Crystal Geyser based on recovered genomic content.
Supplementary Table 8
iRep33 values of organisms across the different metagenomes of the geyser cycle. iRep analysis is based on mapping reads using bowtie229 allowing one mismatch per read.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Probst, A.J., Ladd, B., Jarett, J.K. et al. Differential depth distribution of microbial function and putative symbionts through sediment-hosted aquifers in the deep terrestrial subsurface. Nat Microbiol 3, 328–336 (2018). https://doi.org/10.1038/s41564-017-0098-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-017-0098-y
This article is cited by
-
Selective lipid recruitment by an archaeal DPANN symbiont from its host
Nature Communications (2024)
-
May microbial ecological baseline exist in continental groundwater?
Microbiome (2023)
-
Genome-centric metagenomics reveals the host-driven dynamics and ecological role of CPR bacteria in an activated sludge system
Microbiome (2023)
-
Chemolithoautotroph distributions across the subsurface of a convergent margin
The ISME Journal (2023)
-
Ultra-small bacteria and archaea exhibit genetic flexibility towards groundwater oxygen content, and adaptations for attached or planktonic lifestyles
ISME Communications (2023)