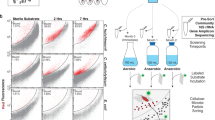
Abstract
Cultivation of microbial consortia provides low-complexity communities that can serve as tractable models to understand community dynamics. Time-resolved metagenomics demonstrated that an aerobic cellulolytic consortium cultivated from compost exhibited community dynamics consistent with the definition of an endogenous heterotrophic succession. The genome of the proposed pioneer population, ‘Candidatus Reconcilibacillus cellulovorans’, possessed a gene cluster containing multidomain glycoside hydrolases (GHs). Purification of the soluble cellulase activity from a 300litre cultivation of this consortium revealed that ~70% of the activity arose from the ‘Ca. Reconcilibacillus cellulovorans’ multidomain GHs assembled into cellulase complexes through glycosylation. These remarkably stable complexes have supramolecular structures for enzymatic cellulose hydrolysis that are distinct from cellulosomes. The persistence of these complexes during cultivation indicates that they may be active through multiple cultivations of this consortium and act as public goods that sustain the community. The provision of extracellular GHs as public goods may influence microbial community dynamics in native biomass-deconstructing communities relevant to agriculture, human health and biotechnology.
Similar content being viewed by others
Plant polysaccharide hydrolysis is a critical process in the human microbiome1, soil microbiomes2 and microbiomes related to bioenergy production3,4,5. Identifying glycoside hydrolases (GHs) responsible for polysaccharide hydrolysis in these ecosystems has important implications for improving human health, managing agriculture and implementing biotechnological advances. Characterizing GHs from uncultivated organisms may also expand the diversity of protein structures that hydrolyse polysaccharides6. However, these communities harbour substantial diversity, complicating the assignment of specific enzymatic roles to individual community members.
Model microbial consortia with simplified community compositions relative to native consortia have been identified as important systems to develop a mechanistic understanding of community function7. Methods to cultivate these model consortia include combining isolates from native consortia and adapting native communities through selective pressure8. These model consortia have enabled the assignment of function to specific microbial community members, clarified successional structures in communities9 and identified previously unknown protein functions10. Cultivation of model consortia that hydrolyse cellulose, the most abundant plant polysaccharide11, has produced low-complexity communities where cellulose hydrolysis can be assigned to specific populations and linked to community structure and dynamics12,13.
Here, we report that a model cellulolytic consortium derived from compost was reproducibly cultivated aerobically at 15 l and 300 l. The proposed pioneer population7 in this community, present at ~1% abundance at the time of culture harvest, produced multidomain cellulases that persisted through microbial succession and were the most active cellulases in the culture. These cellulases were organized in protein complexes that are distinct from cellulosomes8 isolated from anaerobic bacteria. The persistence of these unusually stable complexes indicates that they may act as public goods14 that sustain the community.
Results
A thermophilic cellulolytic bacterial consortium cultivated aerobically at 60 °C from compost obtained from Vacaville, CA, USA produced extracellular cellulases that released glucose from pretreated plant biomass at temperature up to 80 °C15,16. This consortium was maintained for >3 years by passaging every 2 weeks in 50 ml shake flasks. Cultivation of the consortium was scaled to 15 l, and the culture from this 15 l bioreactor was used to inoculate a 300 l bioreactor to provide enzymes for bioprocess studies of biomass deconstruction17 and to purify individual soluble cellulases. The dynamics of the consortium were analysed by time-resolved metagenomics and population genomes recovered by automated binning. At the end of the cultivation, the most abundant populations in the 15 l cultivation were closely related to Rhodothermus marinus (45%), Thermus thermophilus (29%) and Thermobispora bispora (14%) (Fig. 1a and Supplementary Tables 1 and 2). The 300 l cultivation had a similar community composition, with a lower proportion of T. thermophilus (Supplementary Fig. 1a and Supplementary Tables 3 and 4). CMCase activity, a proxy for cellulase activity, increased between days 4 and 5 (from ~0.03–0.05 µmol ml−1 min−1 to ~0.2 µmol ml−1 min−1) and levelled off in both cultivations, while the xylanase activity showed divergent behaviour (Fig. 1b and Supplementary Fig. 1b). Surprisingly, in the 15 l cultivation, CMCase activity correlated with the population dynamics of a single population, affiliated with the Paenibacillaceae (Paenibacillaceae 1; Bin 008). This population rapidly increased in relative abundance from day 2 (4%) to day 4 (40%), but diminished to below 5% by day 8 (Fig. 1c). Similar dynamics were observed in a more restricted sampling of the 300 l cultivation (Supplementary Fig. 1c). The dynamic behaviour of the Paenibacillaceae 1 population is consistent with a pioneer population that commences a community succession7. This succession resembles the model for endogenous heterotrophic succession described by Fierer et al.18, in which the Paenibacillaceae 1 population responded rapidly but was outcompeted by other populations, particularly R. marinus, which becomes the most abundant population in both bioreactor experiments as well as previous cultivations of this consortium in shake flasks12,16. These compositional profiles demonstrated the cellulolytic consortium displayed reproducible behaviour at volumes from 50 ml to 300 l.
a, Relative abundance of dominant populations (≥1%) at the end of the cultivation. Detailed genome information and average coverage are provided in Supplementary Tables 1 and 2. Single DNA samples were isolated from the 15 l culture on each indicated day for metagenomic sequencing. b, CMCase (red) and xylanase (green) activity measurements obtained by daily sampling of the 15 l cultivation. Enzymatic assays are reported as the mean of technical replicates (n = 3) and error bars represent standard error of the mean. c, Daily relative abundances (calculated using the average number of reads of binned scaffolds from time-series metagenomic data for the 15 l cultivation) of the Paenibacillaceae 1 population (red) during a 14-day cultivation of the consortium grown with microcrystalline cellulose.
Phylogenetic trees constructed from conserved proteins and rRNA genes recovered from the population genome of the Paenibacillaceae 1 bin demonstrated that this population is distinct from isolates in the Paenibacillaceae and represents a distinct genus (Fig. 2a,b, Supplementary Fig. 2 and Supplementary Tables 5 and 6). The species represented by this population was named ‘Candidatus Reconcilibacillus cellulovorans’ (from reconciliare in Latin, present active infinitive, to recover) because of evidence that this population is closely related to an incompletely characterized cellulolytic isolate, ‘Caldibacillus cellulovorans’, based on the similarity of its DNA polymerase (Supplementary Fig. 3)19,20. The genome of ‘Ca. R. cellulovorans’ is distantly related to sequenced Paenibacillaceae isolates (<70% amino acid identity). However, a nearly identical population genome was recovered from the metagenome of a similar adaptation of compost microbiota from Milipitas, CA, USA to grow on crystalline cellulose at 60 °C (Supplementary Table 7)13. This ‘Ca. R. cellulovorans NIC (Newby Island Compost)’ population was at <1% relative abundance after a two-week cultivation. Time-series analysis of this consortium using 16S rRNA marker genes provided evidence for a succession transitioning from Paenibacillaceae populations, including operational taxonomic units (OTUs) > 99% identical to the 16S rRNA gene sequence of ‘Ca. R. cellulovorans’, to a thermophilic population in the Chitinophagaceae. The importance of succession in this consortium was underscored by the inability of an isolated representative of the Chitinophagaceae population, strain NYFB, to grow with insoluble cellulose as the sole carbon source.
a, Population genome of ‘Ca. R. cellulovorans’ recovered from metagenomics data from the 15 l cultivation. The genome was dispersed on 114 scaffolds (blue), with 2,814 predicted CDS (coding DNA sequences) in forward (red) and reverse (green) and average (orange) coverage. N50 is the shortest sequence length that includes 50% of the assembled genome, summing from the largest contig. b, Maximum-likelihood phylogenetic tree based on 86 concatenated amino acid sequences that are conserved in the Paenibacillaceae (Supplementary Table 6). c, Molecular organization of multidomain GH genes from the population genome of ‘Ca. R. cellulovorans’ arranged in a 17-kb gene cluster.
Analysis of the genome of ‘Ca. R. cellulovorans’ focused on its glycoside hydrolases (GHs) to assess its role in cellulose depolymerization. The initial population genome contained six partial genes on small discontinuous contigs encoding for multidomain GHs containing catalytic subunits that were linked to family 3 CBMs (CBM3b and CBM3c). PCR-based reconstruction demonstrated that these genes formed a 17-kb gene cluster containing multidomain GHs and a lytic polysaccharide monooxygenase (LPMO) (Fig. 2c, Supplementary Fig. 4 and Supplementary Table 8). The multidomain structure of the encoded proteins in this gene cluster resembled multidomain cellulases identified in the genomes of Caldicellulosiruptor isolates, a genus of extremely thermophilic, anaerobic bacteria that also contain multidomain GHs (Supplementary Fig. 5)21. However, the ‘Ca. R. cellulovorans’ GHs were capped at the C terminus by a CBM3 domain, while many of the Caldicellulosiruptor GHs are multidomain proteins that contain multiple catalytic domains linked by CBM3s22,23. The ‘Ca. R. cellulovorans’ CelA has a GH9 catalytic domain directly linked to a CBM3c and two CBM3b linked to the GH9-CBM3c sequence through poly-proline linkers. The N-terminal portion of Caldicellulosiruptor bescii CelA has an identical domain structure to ‘Ca. R. cellulovorans’ CelA, with 58% sequence identity, but has an additional GH48 domain appended to the C terminus. Despite the similarity in domain structure and sequence between ‘Ca. R. cellulovorans’ and C. bescii CelA, a phylogenetic tree of the catalytic domains demonstrated that the GH9 catalytic domain of these proteins clustered with members of their own phylogenetic group, which was not consistent with horizontal gene transfer (Supplementary Fig. 6a). Multidomain GHs with GH9 catalytic domains have been identified in members of the Paenibacillaceae, exemplified by the GH9 from Paenibacillus barcinonensis, which has a GH9 domain directly linked to a CBM3c, identical to ‘Ca. R. cellulovorans’ CelA, but has a fibronectin-like III domain and a CBM3b domain at the C terminus, rather than two CBM3b domains as in CelA24. In ‘Ca. R. cellulovorans’, the downstream gene CelB contains a GH48 domain that is 68 amino acid residues longer than the Caldicellulosiruptor GH48 domains (Supplementary Fig. 6b). XynA contains two CBM3b modules and an N-terminal GH10 catalytic domain that is 99% identical to a GH10 domain of a xylanase from ‘C. cellulovorans’ (Supplementary Fig. 6c)25. CelC contains an N-terminal GH6 domain and a C-terminal GH5 domain, with both domains clustering with members of the Paenibacillaceae (Supplementary Fig. 6d,e). The N-terminal AA10 domain of PmoA clustered with the N-terminal domain of ManA of ‘C. cellulovorans’ (>99% amino acid identity) (Supplementary Fig. 6f).
Purification of the active GH components from the 300 l cultivation was performed to determine whether the ‘Ca. R. cellulovorans’ proteins were present in the supernatant. The majority (66.3% CMCase and 54.1% xylanase) of the GH activity eluted from an anion-exchange column at 260 mM NaCl (Supplementary Fig. 7a). Visualization and activity staining demonstrated that the GH activities arose from two broad protein bands with molecular weights centred at ~600 kDa (CMCase) and ~300 kDa (xylanase) (Fig. 3a). Separation of these broad bands by SDS–PAGE and subsequent proteomics analysis (Fig. 3b, Supplementary Figs. 8 and 9a–d and Supplementary Tables 9 and 10a–d) indicated that these bands represented protein complexes containing CelABC and XynA (bound to CelB) from ‘Ca. R. cellulovorans’. In an alternative method, the supernatant was treated by affinity digestion to remove proteins that lacked the capacity to bind and hydrolyse insoluble cellulose26. The affinity digestion procedure provided a protein preparation in which the CelABC complexes were enriched but the CelB–XynA complex was not recovered (Fig. 3c). Anion exchange chromatography of the affinity-digested preparation confirmed the retention of CMCase activity and loss of most of the xylanase activity (Supplementary Fig. 7b). A second dimension SDS–PAGE of the heat-denatured samples demonstrated that the CelABC formed three distinct complexes: a high-molecular-weight complex primarily containing CelC and CelA, a lower-molecular-weight complex containing CelABC, and a homocomplex containing only CelB (Fig. 3d). These three complexes were also visualized by denaturing the samples at room temperature and separating them by SDS–PAGE, which demonstrated that the complexes remained intact in the presence of 2% SDS (Fig. 3e). Zymography established that the CelC- and CelA-containing complexes accounted for the CMCase activity of the preparation, while no CelB activity was evident in the gel. The affinity-digested preparation had higher hydrolytic activity on carboxymethylcellulose (CMC) (~25×), phosphoric acid-swollen cellulose (PASC) (~3×) and Avicel (~3×) compared to the original supernatant (Fig. 4a,b).
a, Separation of cellulase and xylanase complexes eluted from an anion-exchange chromatography column at 260 mM NaCl and visualized by 2D BN–PAGE58. Complexes containing S-Layer proteins (S), CMCase (C) and xylanase (X) activity were separated in the first dimension according to their indicated masses by BN–PAGE. Protein staining was accompanied by zymography with gels embedded with CMC and xylan. b, Subunits of the native complexes were separated in a second dimension by SDS–PAGE (8%) and identified as CelABC and XynA by proteomics. The XynA molecular weight (~80 kDa) is indicative of a truncation of the full-length protein (100 kDa). Zymography with CMC revealed the activity of bands corresponding to CelC and CelA. c, GH complexes enriched by affinity digestion were separated by BN–PAGE and protein stains were accompanied by zymography with gels embedded with CMC and xylan. d, Native complexes were separated by SDS–PAGE into subunits CelABC and visualized by protein and CMCase activity staining. e, SDS–PAGE was also performed without initial heat denaturation, and three abundant individual complexes with different compositions of CelABC were identified by proteomics. Detailed proteomics data are provided in Supplementary Figs. 8 and 9. Images were cropped for clarity. Each gel is representative of five gels performed on multiple protein preparations. Gels stained with Coomassie and analysed by zymography were run in parallel in the same electrophoretic cell to ensure comparability. The gels in this figure are from one individual protein preparation for each of the two purification techniques described in the text. The gel images were cropped for clarity and the original gel images are provided in Supplementary Fig. 14.
a, Saccharification of phosphoric acid swollen cellulose (PASC) and Avicel for the culture supernatant, affinity digested preparation (AD) and recombinant CelABC individually expressed in E. coli (see Methods for details). Enzymatic assays are reported as the mean of technical replicates (n = 3) and error bars represent standard error of the mean. b, CMCase-specific activities of the supernatant, AD fraction and in E. coli-expressed CelABC. Enzymatic assays are reported as the mean of technical replicates (n = 3) and error bars represent standard error of the mean. c, SDS–PAGE (8–16% gradient; stained with Coomassie Brilliant Blue) of the AD fraction and E. coli-expressed CelABC. The gel depicted is representative of three gels that displayed very similar results. The gel images were cropped for clarity; the original gel images are provided in Supplementary Fig. 15.
Characterization of recombinant CelABC was performed to determine their cellulase activities. E. coli-expressed CelB and CelC were ~10–50 kDa smaller than their native source (Fig. 4c), indicating extensive post-translational modifications. Glycosylation of CelB and CelC was demonstrated using a periodic acid–Schiff base stain (Supplementary Fig. 10)27. CelA was not glycosylated. Interestingly, C. bescii CelA has been shown to have extensive glycosylation, further distinguishing the two proteins28. CelB is the most abundant protein in the complexes, but had low activities on CMC, PASC and Avicel (68.7 ± 27.9, 26.3 ± 0.1 and 0.9 ± 0.0 mU mg–1), consistent with the results obtained for the complex-bound CelB by zymography. CelC had the highest activities of the three proteins on CMC (2.06 ± 0.11 U mg–1) and PASC (0.19 ± 0.01 U mg–1), but had slightly lower activity on Avicel (32.0 ± 0.1 mU mg–1) compared to CelA (40.9 ± 0.8 mU mg–1). CelA had lower activities on PASC (67.1 ± 1.6 mU mg–1) and CMC (1.1 ± 0.1 U mg–1).
The CelABC complexes are distinct from cellulosomes isolated from anaerobic Firmicutes, because the cellulases in the CelABC complex are multidomain proteins containing carbohydrate-binding modules rather than dockerin domains8. Also absent in CelABC complexes is a non-catalytic scaffoldin protein containing the cohesin domains that bind the individual GHs. Glycosylation analysis of the complex revealed that it mainly contains O-linked galactooligosaccharides units (2–6 monomers) that are arranged in predominantly 1,2-glycosyl linkages (Supplementary Table 11 and Supplementary Fig. 11). The identification of O-glycosylation in CelB and CelC is consistent with the presence of Ser-Pro-Thr-rich linkers in both proteins (Supplementary Fig. 12). These linkers are often glycosylated in multidomain cellulases and in the scaffoldin protein CipA29,30. Although CelA also contains these linkers in its sequence, it was not glycosylated. The contribution of these O-linked galactooligosaccharides to the formation and stability of the CelABC complex was demonstrated by comparison of the proteolytic stability of the complex compared to E. coli-produced CelA and CelC, which form the complex with the highest molecular weight. The CelABC complexes were stable to Proteinase K treatment for 60 min, while both CelC and CelA were proteolysed (Supplementary Fig. 13).
Discussion
This work has demonstrated that an aerobic cellulolytic microbial consortium derived from compost can be reproducibly cultivated and can be scaled to grow at 300 l. Therefore, this community provides an excellent model with which to understand the community dynamics of cellulolytic consortia and uncover enzymes missed by analysis focused solely on culturable isolates. Detailed study of the dynamics of the community using metagenomic methods established a succession in the community that is consistent with the definition of endogenous heterotrophic succession18. These metagenomic studies provided the basis for biochemical studies that identified a class of cellulase complexes in which O-linked glycosylation was critical for complex formation and stability.
The observation of O-linked galactooligosaccharides in CelB and CelC provides a hypothetical mechanism for complex formation. Although O-linked glycosylation is widespread in proteins from bacterial pathogens, it has not been shown to be involved in complex formation for GHs31. We propose that the galactooligosaccharides are bound to the threonine and serine linkages in the protein, and these glycosylated linkers form interprotein interactions with the CBM3 domains present in each of the CelABC component proteins. CBM3s have two predicted carbohydrate-binding sites on opposite faces of the domain32. The first binding site is a planar array of aromatic and polar residues that are predicted to bind crystalline cellulose. The second binding site on the opposite face, which also has polar and aromatic residues, has a shallow groove whose function is unknown. A previous study demonstrated weak binding between the shallow groove of a CBM3 and a peptide representing a linker consensus sequence from CipA33. Glycosylation of these linkers may strengthen the interaction with the shallow groove of the CBM3, providing a mechanism for the formation of the CelABC complexes. Glycosylation-dependent formation of a protein complex has been observed for insulin growth factors (IGFs), in which N-linked glycans are required for an 85 kDa glycoprotein to form a ternary complex with the IGFs in human serum34.
The unusual stability of the CelABC complexes, as demonstrated by their presence at the end of the cultivation and their resistance to proteolysis, complicates a simple mechanistic description linking the succession observed in the microbial community with activity of the celluases. Initial CMCase activity measurements for the 15 l and 300 l cultures indicated that some residual activity remains at the beginning of the culture (Fig. 1b and Supplementary Fig. 1b). This residual activity may be responsible for initial hydrolysis of the crystalline cellulose independent of the enzyme produced during subsequent cultivation. During the 2 week cultivations, the CMCase activity increased ~4–7-fold in the cultures at approximately the same time as the relative abundances in the community shifted from the dominance of ‘Ca. R. cellulovorans’, suggesting that the majority of this cellulase activity is produced during cultivation. Therefore, the cellulase activities that may influence the composition of the consortium may arise from residual enzymes carried over from previous cultivations, as well as enzymes produced during each 2 week cultivation by ‘Ca. R. cellulovorans’. This interplay may be critical for maintaining the remarkable stability of the consortium, which has retained similar community membership and dynamics through multiyear serial cultivations and scaling by 6,000-fold. A second complication in mechanistic interpretation is that the nature of the cellulose substrate may change during the cultivation, so that the CelABC complexes may be more effective over time, independent of the influence of the ‘Ca. R. cellulovorans’ population that initially produced them. Evidence for this phenomenon has been observed in the cellulolytic consortium adapted from Milipitas, CA, compost microbiota described above13, in which the residual cellulose becomes decrystallized during the 2 week cultivation. The difficulty in distinguishing the effects of microbial and enzymatic deconstruction and accounting for the dynamic nature of the cellulose substrate during cultivation emphasizes that the mechanisms underlying the proposed endogenous heterotrophic succession may be more complicated than a simple microbial succession.
The activity of the CelABC complexes on crystalline cellulose is sufficient (0.38 mmol h−1 l−1) to hydrolyse ~15% of the cellulose in the 300 l cultivation. Therefore, the complexes may serve as public goods14, releasing soluble glucans that support the other community members over the 2 week cultivation. A similar provision of GHs as public goods has been observed in defined co-cultures of human gut Bacteroidales isolates, in which species that generate extracellular GHs (producers) support the growth of species that lack these genes (recipients), and the recipients outcompete the producers when the growth of the producer is limited35. Interestingly, the inferred recipients in this cellulolytic consortium, T. bispora and R. marinus, generate substantial amounts of extracellular cellulases under other cultivation conditions36,37, but are not the dominant cellulase producers in the community cultivations described here, which suggests that glucans released by the CelABC complexes may repress subsequent synthesis of cellulases from other community members.
This work demonstrates that adapted consortia can provide microbial community models that can be used to begin to understand the interplay of enzymes and microbes in the deconstruction of plant biomass by microbial consortia. Metagenomic studies of complex microbial communities that hydrolyse plant polysaccharides contain highly abundant populations with genomes that have large numbers of GH genes38,39. These abundant populations are often assigned as the primary GH producers in the environments they inhabit. The results described here suggest that transient community members in these complex consortia may produce highly active and stable extracellular GHs that are active independently of the populations that produce them and can act as public goods for the community.
Methods
Sample collection and enrichment of thermophilic consortia
The sample collection and enrichment procedures have been described previously15. Briefly, compost samples were collected from Jepson Prairie (JP) Organics, located in Vacaville, CA, in 2008. The compost-derived microbial consortium was initially grown aerobically with unpretreated switchgrass and then switched to grow on microcrystalline cellulose (1% wt/vol; Sigma) as the sole carbon source in liquid M9 medium augmented with vitamins. The enrichments were grown at 60 °C and 200 r.p.m. under aerobic conditions in an aerial rotary shaker and serially passed every 14 days with 4% vol/vol inoculum, referred to as passages. Cultivation of the 50 ml culture after passage 80 was scaled at the Advanced Biofuels Process Demonstration Unit, Lawrence Berkeley National Laboratory. A 500 ml culture was inoculated with a 2.5% vol/vol sample from the 50 ml culture and incubated at 60 °C for 14 days at 150 r.p.m. on a rotary shaker. This culture was inoculated into a 19 l bioreactor (Bioengineering USA) to a total volume of 15 l. The 15 l culture was grown at 60 °C, 150 r.p.m. and 0.26 volume gas (sterile air) per volume liquid per minute (VVM) for 14 days. A 400 l bioreactor (ABEC) was inoculated with 7.5 l of culture from the 19 l bioreactor to a volume of 300 l and incubated for 14 days at 60 °C, 150 r.p.m. agitation and 0.25 VVM air sparging. After 14 days, the final fermentation broth was centrifuged using an Alfa Laval Disc Stack centrifuge at 9,000g at 100 l h–1 with 30 min cell discharge interval and 125 kPa back pressure. The clarified broth was collected in the 300 l holding tank bioreactor and the pelleted biomass collected and stored at –80 °C. The supernatant was concentrated through a tangential flow filtration system with a 10 kDa Biomax filter membrane (EMD Millipore). The transmembrane pressure was set at 13 p.s.i. with feed pressure of 30 p.s.i. The concentrated supernatant was freeze-dried under vacuum for 24 h using a lyophilizer (Labconco) and the resulting powder was stored at –20 °C. CMCase and xylanase activities were measured daily for the 15 l and 300 l bioreactor cultivations by removal of samples each day from the bioreactors and the supernatant assayed as described below.
Sequencing, assembly and binning of metagenomics reads
DNA purification from samples extracted from the 15 l and 300 l bioreactors was performed as previously described13. Illumina sequencing (250 bp × 2) of the metagenomic samples was carried out by the Joint Genome Institute (JGI) and performed as previously described40. The sequencing reads of the DNA samples recovered from the 15 l bioreactor (days 1–14 were trimmed using Trimmomatics (with parameter ILLUMINACLIP:TruSeq3-PE.fa:2:30:10 LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:36)41 and co-assembled using IDBA-UD42 with the --pre-correction parameter. The 300 l samples (days 4, 5, 7, 10, 12 and 14) were also co-assembled, using the same settings as the 15 l samples. The co-assembled samples for the 15 l and 300 l bioreactor experiments were then binned using MaxBin 2.043 with default parameters, yielding population genomes. The completeness and contamination ratios of these population genomes were assessed using CheckM44. Genomes with >10% of contamination rates were re-binned using MaxBin 2.0 by setting the input contig to the genome file and the input abundance to the extracted abundance files during the whole metagenome binning. The output bins were re-examined using CheckM and the produced bins with higher completeness were chosen to replace the original genome, while the other bins with higher levels of contaminants were discarded. The most likely taxonomic ranks of the recovered genomes were predicted by searching the predicted proteins against the NCBI non-redundant (NR) database, collecting and processing the hits using the least common ancestor (LCA) algorithm proposed by MEGAN445 and assigning the most probable taxonomic rank to the recovered genomes according to the LCA results. GH genes present in the recovered population genomes were identified by using Prodigal46 and then annotated using dbCAN47.
DNA was isolated from a 50 ml aerobic shake flask culture with a bacterial consortium that was adapted from green waste compost obtained from Newby Island Sanitary Landfill in Milpitas, CA, by growth on crystalline cellulose. The enzymatic activities and microbial community membership of this consortium have been described previously13. Metagenomic sequencing was performed by the JGI as described above, and the sequenced reads were assembled as previously described40. Population genomes were recovered by automated binning with Maxbin43 and checked for completeness and contamination with CheckM44.
16S and 23S rRNA gene analysis
A partial 16S rRNA gene (706 bp) was recovered from the ‘Ca. R. cellulovorans’ metagenomic bin. This fragment was used to identify a nearly full-length 16S rRNA gene (1,597 bp; 99.7% identical JGI gene ID Ga0074251_1085371) that was recovered from the initial assembled metagenome (JGI taxon ID 3300005442) obtained for this consortium when it had been adapted to switchgrass40. This observation indicated that ‘Ca. R. cellulovorans’ was present in the consortium when it was adapted to switchgrass, before transferring the consortium to grow on microcrystalline cellulose. Sequences from three clones (GenBank accessions KC978751, KC978760 and KC978763) were >99% identical to portions of the full-length rRNA ‘Ca. R. cellulovorans’ sequence. These clones were recovered from DNA samples isolated from a time series (14 days) of an adaptation of compost from Newby Island Landfill (Milipitas, CA, USA) to grow with microcrystalline cellulose as the sole carbon source at 60 °C13. As described above, a partial genome was recovered from this cultivation that was >99% identical at the amino acid level to the ‘Ca. R. cellulovorans’. A partial 23S rRNA gene (1,444 bp) was recovered from the ‘Ca. R. cellulovorans’ metagenomic bin. The 16S and 23S rRNA gene sequences were aligned using MUSCLE48 trimmed using Gblocks49, and the phylogenetic tree was constructed using MEGA550 with the Tamura–Nei model. Bootstrap values were calculated with 1,000 replicates.
Reconstruction of ‘Ca. R. cellulovorans’ GH gene cluster
Six small contigs were identified in the ‘Ca. R. cellulovorans’ population genome, which contained partial genes containing a catalytic domain (GH9, GH48, GH6/5, 2×GH10, AA10) linked to at least one CBM3. The clustering of these genes was confirmed by PCR amplification of DNA isolated from the cellulolytic consortium. PCR primers (Supplementary Table 8) were designed using the CLC Main Workbench (Qiagen) and PCR products were cloned into pJET1.2/blunt Cloning Vector (Fermentas) and sequenced with an ABI system according to the manufacturer’s instructions. Assembly of gene sequences into a gene cluster and annotation of genes was performed with the CLC Main Workbench and checked for chimerae using the Bellerophon algorithm51.
Phylogenetic analysis
Alignments of protein sequences were performed using the CLUSTALW multiple alignment accessory application in the CLC Main Workbench (Qiagen). In brief, phylogenetic trees were constructed using the CLC Main Workbench applying the maximum likelihood method based on the Whelan and Goldman protein substitution model52. Bootstrap values were calculated with 1,000 replicates.
To build the concatenated protein tree, genes were first searched against the PFAM profiles53 using HMMER354. Genes with PFAM annotations that appear once and only once across all involved genomes were aligned separately using MUSCLE48. After the alignments were concatenated and trimmed using Gblocks49, the concatenated maximum-likelihood protein tree was constructed using MEGA550 with the JTT (Jones, Taylor, Thorton) model. Bootstrap values were estimated with 1,000 replicates.
Protein purification
Lyophilized supernatant (170 mg) obtained from the 300 l cultivation was dissolved in 5 ml H2O and passed through a 0.2 µm filter. The supernatant was desalted by dialysis against the buffer (20 mM Tris, pH 8.0) for 24 h with three buffer changes, followed by a 30 ml NaCl gradient fractionation (0–2 M NaCl) using a 5 ml HiTrap Q HP column on an ÄKTA Protein Purification System (GE Healthcare).
Cellulases in the supernatant from the 300 l cultivation were also enriched by binding to phosphoric acid swollen cellulose (PASC), an adaptation of a procedure previously described for cellulosome purification from Clostridium thermocellum 26. Briefly, 250 mg of lyophilized PASC produced from Avicel PH-105 was added to 500 mg of supernatant dissolved in 10 ml H2O and mixed at room temperature with a magnetic stir bar for 30 min. After a binding step at 4 °C for 2 h, the amorphous cellulose was centrifuged for 10 min at 3,000g and rigorously washed for 6 cycles with 25 ml reaction buffer (25 mM 2-(N-morpholino)ethanesulfonic acid (MES), pH 6.0). Washed PASC was resuspended in 10 ml reaction buffer and transferred into dialysis membranes (SnakeSkin and Slide-A-Lyzer; Fisher Scientific) with a 3.5–10 kDa cutoff and dialysed at 60 °C against 4 l reaction buffer at 55–60 °C for up to 48 h with three buffer exchanges per day to prevent possible product inhibition. Dialysis membranes used in this study consisted of regenerated cellulose and were destabilized by cellulases of the substrate, and thus needed to be exchanged every 24 h to prevent membrane rupture. The reaction was considered complete after no visible changes to the substrate were observable. By centrifugation for 20 min at 3,000g the enrichment was split into residual biomass (in the pellet) and the affinity digestion protein fraction (in the supernatant, AD).
Measurement of protein concentration and GH activity
Protein concentrations were determined using the bicinchoninic (BCA) assay (Pierce BCA Protein Assay Kit, Thermo Scientific) method using a 96-well plate (200 μl reaction volume) with bovine serum albumin as the standard. CMCase and xylanase activity assays were conducted as described previously55. Enzyme activity units (U) were defined as µmol of sugar liberated per min. Enzyme activity units for supernatant preparations were calculated as U per ml of supernatant volume. CMC activity units of purified heterologously expressed proteins were reported as U per mg, representing specific activity measurements.
Soluble substrates (p-nitrophenyl (pNP)-labelled) with cellobiohydrolase (pNPC), β-d-dglucosidase (pNPG), β-D-xylosidase (pNPX) and α-l-arabinofuranosidase (pNPA) activities were used to determine enzyme activities on their respective substituents56. The p-nitrophenyl substrate (90 μl) was incubated with 10 μl of diluted enzyme, incubated for 30 min, and quenched with 50 μl of 2% cold sodium bicarbonate. The absorbance of released p-nitrophenyl was measured at 410 nm. Activities using p-nitrophenyl substrates were calculated as U ml−1.
Saccharification of cellulose substrates
Saccharifications were performed in the presence of 2% (wt/vol) Avicel (Sigma) and PASC. Each mixture was prepared in 50 mM MES, pH 6.0 with 10 mg protein per g glucan in biomass to a final volume of 625 µl in a 2 ml screw-cap vial. Saccharifications were carried out at 70 °C in a shaker for 72 h, with 50 µl samples taken every 24 h. All hydrolysates were collected via centrifugation at 21,000g for 5 min and 0.45 μm filtered to remove large biomass particles prior to sugar analysis. After filtration, samples were kept frozen at −20 °C and thawed before analysis. Glucose concentrations were measured on an Agilent 1200 Series HPLC system equipped with an Aminex HPX-87H column (Bio-Rad) and refractive index detector. Samples were run with an isocratic 4 mM sulfuric acid mobile phase. Sugar concentrations were determined using standards containing cellotriose, cellobiose, glucose, xylose and arabinose.
PAGE and zymograms
SDS–PAGE was performed with 8–16% Protean TGX protein gradient gels (Bio-Rad) with the Tris-glycine-SDS buffer57. Blue Native (BN)–PAGE58 was performed with 3–12% NativePAGE Bis-Tris protein gradient gels (Thermo Scientific) in presence of 0.02% Coomassie Blue G-250. For subunit analysis of native complexes, individual lanes from the BN–PAGE were excised, incubated in 2% SDS and 160 mM dithiothreitol (DTT), and denatured at 95 °C for 10 min, unless otherwise indictated (Fig. 3e). Proteins were separated with 8% polyacrylamide gels, which were hand cast. Protein bands were stained with SimplyBlue SafeStain Coomassie Blue dye (Thermo Scientific) according to the manufacturer’s instructions.
Protein bands with activity on CMC and xylan were visualized using modification of the zymogram technique, as described previously15. Gels were incubated in 2% wt/vol CMC or 2% wt/vol birchwood xylan solutions followed by incubation at 60 °C for up to 2 h in reaction buffer (25 mM MES, pH 6.0). In-gel enzymatic activities were visualized by incubating gels with a 0.5% Congo Red solution for 15 min and subsequent multiple washing steps with 20% NaCl.
Glycosylation analysis
Protein glycosylations were visualized in-gel by the periodic acidic Schiff stain27 using a Pierce Glycoprotein Staining Kit (Thermo Scientific) according to the manufacturer’s instructions.
N-glycan analysis was performed as described previously59. However, no N-linked glycans were detected. Total glycosyl compositional analysis was performed by combined gas chromatography/mass spectrometry (GC/MS) of the per-O-trimethylsilyl (TMS) derivatives of the monosaccharide methyl glycosides produced from the sample by acidic methanolysis60. O-linked glycans were released by β-elimination and permethylated59. The permethylated O-linked glycans were analysed by matrix assisted laser desorption/ionization-time of flight (MALDI–TOF) and electrospray ionization tandem mass spectrometry (ESI MS/MS)61 and gas chromatography/mass spectrometry (GC/MS) for linkage analysis62.
Proteinase K digestion
The E. coli-expressed CelA and CelC and the AD fraction were digested at 50 °C for 60 min in reaction buffer (20 mM Tris-HCl, 400 mM NaCl and 0.3% SDS, 5 mM EDTA containing 75 µg of respective enzyme and 3.75 µg proteinase K). After heat inactivation of proteinase K at 95 °C, the reaction mixture was analysed by SDS–PAGE (8–16% gradient).
Proteomic analysis
Proteins were digested from SDS–PAGE gels as previously described63. Samples were analysed on an Agilent 6550 iFunnel QTOF mass spectrometer coupled to an Agilent 1290 UHPLC system, as described in ref. 64. Briefly, peptides were loaded onto an Ascentis Express Peptide ES-C18 column (10 cm length × 2.1 mm internal diameter, 2.7 µm particle size; Sigma Aldrich) operating at 60 °C and at a flow rate of 400 µl min–1. A 13.5 min chromatography method with the following gradient was used: the initial starting condition (95% Buffer A (0.1% formic acid) and 5% Buffer B (99.9% acetonitrile, 0.1% formic acid)) was held for 1 min. Buffer B was then increased to 35% in 5.5 min, followed by an increase to 80% B in 1 min, where it was held at 600 µl min–1 for 3.5 min. Buffer B was decreased to 5% over 0.5 min, where it was held for 2 min at 400 µl min–1 to re-equilibrate the column with the starting conditions. Peptides were introduced into the mass spectrometer from the UHPLC by using a Dual Agilent Jet Stream Electrospray Ionization source operating in positive-ion mode. The source parameters used include a gas temperature of 250 °C, drying gas at 14 l min–1, nebulizer at 35 p.s.i.g, sheath gas temp of 250 °C, sheath gas flow of 11 l min–1), V Cap of 5,000 V, fragmentor V of 180 V and OCT(octopole) 1 RF (radio frequency) V pp of 750 V. The data were acquired with Agilent MassHunter Workstation Software, LC/MS Data Acquisition B.06.01 (Build 6.01.6157). The resultant data files were searched against a data set containing reconstructed population genomes from the 300 l bioreactor, with common contaminants appended, with Mascot version 2.3.02 (Matrix Science), then filtered and refined using Scaffold version 4.6.1 (Proteome Software).
Heterologous protein expression
Constructs for the CelABC genes were obtained both by PCR amplification from metagenomic DNA with specific primers (Supplementary Table 8) and synthesis of codon-optimized versions for expression in E. coli (Gen9). Genes were cloned into the modified bacterial expression vector pET39b(+) vector with a T7/lac promoter and a TEV-cleavable C-terminal 6xHis tag but lacking the DsbA secretion sequence (Novagen) using Gibson assembly65. All reagents were purchased from New England Biolabs. The desired genes without their signal sequences and the expression vector were PCR-amplified, DpnI-digested and incubated with 1× Gibson assembly Master Mix for 15 min at 50 °C. The product was then transformed into chemically competent E. coli DH10α cells for storage and for heterologous protein expression into chemically competent E. coli BL21 (DE3). Starter cultures (50 ml) of E. coli BL21 (DE3) harbouring plasmids were grown overnight in LB medium containing 25 μg ml–1 kanamycin at 37 °C and shaken at 200 r.p.m. in rotary shakers. Expression was performed in Terrific broth with 2% glycerol, 25 μg ml–1 kanamycin and 2 mM MgSO4. Starter cultures were used to inoculate 1 l of expression medium in a 2 l baffled Erlenmeyer flask and incubated at 18 °C while shaking (200 r.p.m.), and induced with 500 µM isopropyl β-d-thiogalactopyranoside (IPTG). Following induction, cultures were again incubated at 18 °C. At 22 h, cultures were centrifuged at 15,500g for 30 min. Cell pellets were resuspended in 25 ml lysis buffer (50 mM NaPO4, 300 mM NaCl, 5 mM imidazole; pH 7.4) and homogenized with an EmulsiFlex-C3 instrument (Avestein). After incubation at 60 °C for 30 min, lysates were collected via centrifugation at 75,000g for 30 min and 0.45 μm filtered to remove large particles before purification. Polyhistidine-tagged proteins were purified on Cobalt-NTA resin (Thermo Scientific). To cleave the 6xHis Tag, 1 g purified protein was incubated with 50 mg His-tagged TEV-protease and simultaneously dialysed against 4 l reaction buffer (50 mM NaPO4, 300 mM NaCl; pH 7.4) for 24 h and three reaction buffer exchanges. After a second purification step via Cobalt-NTA resin, the flow-through fractions contained the purified and untagged proteins. Proteins were stored at 4 °C until ready for use. The proteins were >90% pure as visualized by SDS–PAGE (Fig. 4c).
Life Sciences Reporting Summary
Further information on experimental design and reagents is available in the Life Sciences Reporting Summary.
Data availability
Metagenomic sequencing data can be accessed at the JGI IMG website (http://img.jgi.doe.gov/) or the JGI Genome Portal (http://genome.jgi.doe.gov/), and the specific IMG genome IDs are listed in Supplementary Table 12. The draft genome sequence for ‘Candidatus Reconcilbacillus cellulovorans’ has been deposited at GenBank (MOXJ00000000). The gene sequences and plasmid constructs for the ‘Ca. Reconcilbacillus’ cellulases CelA (JPUB_007824), CelB (JPUB_007826) and CelC (JPUB_007828) are available from the public version of the JBEI Registry (https://public-registry.jbei.org) and are physically available from the authors and/or Addgene (http://www.addgene.org) upon request.
References
Cantarel, B. L., Lombard, V. & Henrissat, B. Complex carbohydrate utilization by the healthy human microbiome. PLoS ONE 7, e28742 (2012).
Pankratov, T. A., Ivanova, A. O., Dedysh, S. N. & Liesack, W. Bacterial populations and environmental factors controlling cellulose degradation in an acidic sphagnum peat. Environ. Microbiol. 13, 1800–1814 (2011).
Brulc, J. M. et al. Gene-centric metagenomics of the fiber-adherent bovine rumen microbiome reveals forage specific glycoside hydrolases. Proc. Natl Acad. Sci. USA 106, 1948–1953 (2009).
Pope, P. B. et al. Metagenomics of the Svalbard reindeer rumen microbiome reveals abundance of polysaccharide utilization loci. PLoS ONE 7, e38571 (2012).
Hanreich, A. et al. Metagenome and metaproteome analyses of microbial communities in mesophilic biogas-producing anaerobic batch fermentations indicate concerted plant carbohydrate degradation. Syst. Appl. Microbiol. 36, 330–338 (2013).
Hess, M. et al. Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331, 463–467 (2011).
Wolfe, B. E., Button, J. E., Santarelli, M. & Dutton, R. J. Cheese rind communities provide tractable systems for in situ and in vitro studies of microbial diversity. Cell 158, 422–433 (2014).
Cole, J. K. et al. Phototrophic biofilm assembly in microbial-mat-derived unicyanobacterial consortia: model systems for the study of autotroph–heterotroph interactions. Front. Microbiol. 5, 109 (2014).
Konopka, A., Lindemann, S. & Fredrickson, J. Dynamics in microbial communities: unraveling mechanisms to identify principles. ISME J. 9, 1488–1495 (2015).
Romine, M. F. et al. Elucidation of roles for vitamin B12 in regulation of folate, ubiquinone, and methionine metabolism. Proc. Natl Acad. Sci. USA 114, E1205–E1214 (2017).
Lynd, L. R., Weimer, P. J., van Zyl, W. H. & Pretorius, I. S. Microbial cellulose utilization: fundamentals and biotechnology. Microbiol. Mol. Biol. Rev. 66, 506–580 (2002).
Gladden, J. M., Eichorst, S. A., Hazen, T. C., Simmons, B. A. & Singer, S. W. Substrate perturbation alters the glycoside hydrolase activities and community composition of switchgrass-adapted bacterial consortia. Biotechnol. Bioeng. 109, 1140–1145 (2012).
Eichorst, S. A. et al. Community dynamics of cellulose-adapted thermophilic bacterial consortia. Environ. Microbiol. 15, 2573–2587 (2013).
Folse, H. J. & Allison, S. D. Cooperation, competition, and coalitions in enzyme-producing microbes: social evolution and nutrient depolymerization rates. Front. Microbiol. 3, 338 (2012).
Gladden, J. M. et al. Glycoside hydrolase activities of thermophilic bacterial consortia adapted to switchgrass. Appl. Environ. Microbiol. 77, 5804–5812 (2011).
Park, J. I. et al. A thermophilic ionic liquid-tolerant cellulase cocktail for the production of cellulosic biofuels. PLoS ONE 7, e37010 (2012).
Shi, J. et al. One-pot ionic liquid pretreatment and saccharification of switchgrass. Green Chem. 15, 2579–2589 (2013).
Fierer, N., Nemergut, D., Knight, R. & Craine, J. M. Changes through time: integrating microorganisms into the study of succession. Res. Microbiol. 161, 635–642 (2010).
Huang, X. P. & Monk, C. Purification and characterization of a cellulase (CMCase) from a newly isolated thermophilic aerobic bacterium Caldibacillus cellulovorans gen. nov., sp. nov. World J. Microbiol. Biotechnol. 20, 85–92 (2004).
Shandilya, H. et al. Thermophilic bacterial DNA polymerases with reverse-transcriptase activity. Extremophiles 8, 243–251 (2004).
Blumer-Schuette, S. E. et al. Caldicellulosiruptor core and pangenomes reveal determinants for noncellulosomal thermophilic deconstruction of plant biomass. J. Bacteriol. 194, 4015–4028 (2012).
Yi, Z., Su, X., Revindran, V., Mackie, R. I. & Cann, I. Molecular and biochemical analyses of CbCel9A/Cel48A, a highly secreted multi-modular cellulase by Caldicellulosiruptor bescii during growth on crystalline cellulose. PLoS ONE 8, e84172 (2013).
Brunecky, R. et al. Revealing nature’s cellulase diversity: the digestion mechanism of Caldicellulosiruptor bescii CelA. Science 342, 1513–1516 (2013).
Chiriac, A. I. et al. Engineering a family 9 processive endoglucanase from Paenibacillus barcinonensis displaying a novel architecture. Appl. Microbiol. Biotechnol. 86, 1125–1134 (2010).
Sunna, A., Gibbs, M. D. & Bergquist, P. L. A novel thermostable multidomain 1,4-β-xylanase from ‘Caldibacillus cellulovorans’ and effect of its xylan-binding domain on enzyme activity. Microbiology 146, 2947–2955 (2000).
Morag, E., Bayer, E. A. & Lamed, R. Affinity digestion for the near-total recovery of purified cellulosome from Clostridium thermocellum. Enzyme Microbial Technol. 14, 289–292 (1992).
Kligman, A. M. & Mescon, H. The periodic-acid-Schiff stain for the demonstration of fungi in animal tissue. J. Bacteriol. 60, 415–421 (1950).
Chung, D. et al. Homologous expression of the Caldicellulosiruptor bescii CelA reveals that the extracellular protein is glycosylated. PLoS ONE 10, e0119508 (2015).
Beckham, G. T. et al. Harnessing glycosylation to improve cellulase activity. Curr. Opin. Biotechnol. 23, 338–345 (2012).
Gerwig, G. J. et al. The nature of the carbohydrate-peptide linkage region in glycoproteins from the cellulosomes of Clostridium thermocellum and Bacteroides cellulosolvens. J. Biol. Chem. 268, 26956–26960 (1993).
Nothaft, H. & Szymanski, C. M. Protein glycosylation in bacteria: sweeter than ever. Nat. Rev. Microbiol. 8, 765–778 (2010).
Tormo, J. et al. Crystal structure of a bacterial family-III cellulose-binding domain: a general mechanism for attachment to cellulose. EMBO J. 15, 5739–5751 (1996).
Yaniv, O., Frolow, F., Levy-Assraf, M., Lamed, R. & Bayer, E. A. Interactions between family 3 carbohydrate binding modules (CBMs) and cellulosomal linker peptides. Methods Enzymol. 510, 247–259 (2012).
Janosi, J. B., Firth, S. M., Bond, J. J., Baxter, R. C. & Delhanty, P. J. N-linked glycosylation and sialylation of the acid-labile subunit. J. Biol. Chem. 274, 5292–5298 (1999).
Rakoff-Nahoum, S., Coyne, M. J. & Comstock, L. E. An ecological network of polysaccharide utilization among human intestinal symbionts. Curr. Biol. 24, 40–49 (2014).
Hreggvidsson, G. O. et al. An extremely thermostable cellulase from the thermophilic eubacterium Rhodothermus marinus. Appl. Environ. Microbiol. 62, 3047–3049 (1996).
Hiras, J. et al. Comparative community proteomics demonstrates the unexpected importance of actinobacterial glycoside hydrolase family 12 protein for crystalline cellulose hydrolysis. mBio 7, e01106-16 (2016).
Naas, A. E. et al. Do rumen Bacteroidetes utilize an alternative mechanism for cellulose degradation? mBio 5, e01401-14 (2014).
Güllert, S. et al. Deep metagenome and metatranscriptome analyses of microbial communities affiliated with an industrial biogas fermenter, a cow rumen, and elephant feces reveal major differences in carbohydrate hydrolysis strategies. Biotechnol. Biofuels 9, 121 (2016).
D’haeseleer, P. et al. Proteogenomic analysis of a thermophilic bacterial consortium adapted to deconstruct switchgrass. PLoS ONE 8, e68465 (2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Peng, Y., Leung, H. C., Yiu, S. M. & Chin, F. Y. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28, 1420–1428 (2012).
Wu, Y.-W., Simmons, B. A. & Singer, S. W. MaxBin 2.0: an automated binning algorithm to recover genomes from multiple metagenomic datasets. Bioinformatics 32, 605–607 (2015).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Huson, D. H., Auch, A. F., Qi, J. & Schuster, S. C. MEGAN analysis of metagenomic data. Genome Res. 17, 377–386 (2007).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Yin, Y. et al. dbCAN: a web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 40, W445–W451 (2012).
Edgar, R. C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5, 1–19 (2004).
Talavera, G. & Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst. Biol. 56, 564–577 (2007).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Huber, T., Faulkner, G. & Hugenholtz, P. Bellerophon: a program to detect chimeric sequences in multiple sequence alignments. Bioinformatics 20, 2317–2319 (2004).
Whelan, S. & Goldman, N. A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol. Biol. Evol. 18, 691–699 (2001).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Xiao, Z. Z., Storms, R. & Tsang, A. Microplate-based carboxymethylcellulose assay for endoglucanase activity. Anal. Biochem. 342, 176–178 (2005).
Bowers, G. N., McComb, R. B., Christensen, R. & Schaffer, R. High-purity 4-nitrophenol: purification, characterization, and specifications for use as a spectrophotometric reference material. Clin. Chem. 26, 724–729 (1980).
Laemmli, U. K. Cleavage of structural proteins during assembly of head of bacteriophage-T4. Nature 227, 680–685 (1970).
Wittig, I., Braun, H.-P. & Schängger, H. Blue native PAGE. Nat. Protoc. 1, 418–428 (2006).
Wada, Y. et al. Comparison of the methods for profiling glycoprotein glycans–HUPO Human Disease Glycomics/Proteome Initiative multi-institutional study. Glycobiology 17, 411–422 (2007).
Santander, J. et al. Mechanisms of intrinsic resistance to antimicrobial peptides of Edwardsiella ictaluri and its influence on fish gut inflammation and virulence. Microbiology 159, 1471–1486 (2013).
North, S. J. et al. Mass spectrometric analysis of mutant mice. Methods Enzymol. 478, 27–77 (2010).
Heiss, C., Klutts, J. S., Wang, Z., Doering, T. L. & Azadi, P. The structure of Cryptococcus neoformans galactoxylomannan contains β-d-glucuronic acid. Carbohydrate Res. 344, 915–920 (2009).
Shevchenko, A., Tomas, H., Havli, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
González Fernández-Niño, S. M. et al. Standard flow liquid chromatography for shotgun proteomics in bioenergy research. Front. Bioeng. Biotechnol. 3, 44 (2015).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Acknowledgements
This work was performed as part of the DOE Joint BioEnergy Institute (http://www.jbei.org), supported by the US DOE, Office of Science, Office of Biological and Environmental Research, through contract DE-AC02-05CH11231 between Lawrence Berkeley National Laboratory and the US DOE. Metagenomic sequencing was conducted by the Joint Genome Institute, which is supported by the Office of Science of the US DOE under contract no. DE-AC02-05CH11231. Work was performed in the Advanced Biofuels Process Development Unit, which is supported by the US DOE, Office of Energy Efficiency and Renewable Energy, Bioenergy Technologies Office through contract DE-AC02-05CH11231 between Lawrence Berkeley National Laboratory and the US DOE. S.K. was supported by a research fellowship from the German Research Foundation (Deutsche Forschungsgemeinschaft KO 5295/1-1). Glycosylation analysis was supported by the Chemical Sciences, Geosciences and Biosciences Division, Office of Basic Energy Sciences, US DOE grant (DE-SC0015662) and in part by the NIH-funded grant no. 1S10OD018530 and grant no. P41GM10349010 to P.A. The authors thank P. Coffman and S. Hubbard (Lawrence Berkeley National Laboratory) for technical assistance and S. Baker (Pacific Northwest National Laboratory) for discussions.
Author information
Authors and Affiliations
Contributions
S.W.S, J.M.G., B.A.S. and P.D.A. designed the project. F.T., D.T. and T.R.P. designed and performed scale-up of the cellulolytic consortium. J.H. and N.B. isolated DNA from samples from the consortium and prepared the DNA for metagenomic sequencing. S.A.E. cultivated the cellulolytic community from Newby Island compost samples and isolated DNA. Y.-W.W. performed bioinformatic analysis on all metagenomic data. S.K. designed primers and performed PCR amplification to recover gene cluster. S.K. and E.D. developed the purification strategy for cellulase complexes and S.K. characterized the complexes. C.J.P. and L.J.C. performed proteomics analysis to identify the components of the complexes. S.K. designed constructs that express individual cellulases from the complexes. S.K. and D.F. performed heterologous expression and purification of these individual cellulases. R.G., Q.C. and P.A. performed glycosylation analysis. S.K., Y.-W.W. and S.W.S. wrote the manuscript, and all authors approved the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
S.K., E.D., J.H., J.M.G. and S.W.S. are inventors on a patent application related to this work (PCT/US2016/063198). The remaining authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Information
Supplementary Tables 1–12 and Supplementary Figures 1–17.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kolinko, S., Wu, YW., Tachea, F. et al. A bacterial pioneer produces cellulase complexes that persist through community succession. Nat Microbiol 3, 99–107 (2018). https://doi.org/10.1038/s41564-017-0052-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-017-0052-z
This article is cited by
-
Structural and functional analysis of the active cow rumen’s microbial community provides a catalogue of genes and microbes participating in the deconstruction of cardoon biomass
Biotechnology for Biofuels and Bioproducts (2024)
-
Origin, Selection, and Succession of Coastal Intertidal Zone-Derived Bacterial Communities Associated with the Degradation of Various Lignocellulose Substrates
Microbial Ecology (2023)
-
Low-abundance populations distinguish microbiome performance in plant cell wall deconstruction
Microbiome (2022)
-
Genome-resolved metagenome and metatranscriptome analyses of thermophilic composting reveal key bacterial players and their metabolic interactions
BMC Genomics (2021)
-
CAZymes from the thermophilic fungus Thermoascus aurantiacus are induced by C5 and C6 sugars
Biotechnology for Biofuels (2021)