Abstract
Despite its importance, understanding the early phases of human development has been limited by availability of human samples. The recent emergence of stem-cell-derived embryo models, a new field aiming to use stem cells to construct in vitro models to recapitulate snapshots of the development of the mammalian conceptus, opens up exciting opportunities to promote fundamental understanding of human development and advance reproductive and regenerative medicine. This Review provides a summary of the current knowledge of early mammalian development, using mouse and human conceptuses as models, and emphasizes their similarities and critical differences. We then highlight existing embryo models that mimic different aspects of mouse and human development. We further discuss bioengineering tools used for controlling multicellular interactions and self-organization critical for the development of these models. We conclude with a discussion of the important next steps and exciting future opportunities of stem-cell-derived embryo models for fundamental discovery and translation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




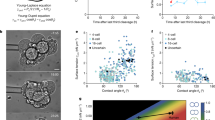
A. Yoney/E. D. Siggia (a); N. Rivron (b); Y. Zheng (c); M. Simunovic (d)
Similar content being viewed by others
References
Solnica-Krezel, L. (ed.) Gastrulation: From Embryonic Pattern to Form (Academic Press, 2020).
Rossant, J. & Tam, P. P. L. New insights into early human development: lessons for stem cell derivation and differentiation. Cell Stem Cell 20, 18–28 (2017).
Fogarty, N. M. E. et al. Genome editing reveals a role for OCT4 in human embryogenesis. Nature 550, 67–73 (2017).
Ma, H. et al. Correction of a pathogenic gene mutation in human embryos. Nature 548, 413–419 (2017).
Deglincerti, A. et al. Self-organization of the in vitro attached human embryo. Nature 533, 251–254 (2016).
Shahbazi, M. N. et al. Self-organization of the human embryo in the absence of maternal tissues. Nat. Cell Biol. 18, 700–708 (2016).
Xiang, L. et al. A developmental landscape of 3D-cultured human pre-gastrulation embryos. Nature 577, 537–542 (2020).
Daley, G. Q. et al. Setting global standards for stem cell research and clinical translation: the 2016 ISSCR guidelines. Stem Cell Rep. 6, 787–797 (2016).
Hyun, I., Wilkerson, A. & Johnston, J. Revisit the 14-day rule. Nature 533, 169–171 (2016).
Ma, H. et al. In vitro culture of cynomolgus monkey embryos beyond early gastrulation. Science 366, eaax7890 (2019).
Niu, Y. et al. Dissecting primate early post-implantation development using long-term in vitro embryo culture. Science 366, eaaw5754 (2019).
Nakamura, T. et al. A developmental coordinate of pluripotency among mice, monkeys and humans. Nature 537, 57–62 (2016).
ten Berge, D. et al. Wnt signaling mediates self-organization and axis formation in embryoid bodies. Cell Stem Cell 3, 508–518 (2008).
Fuchs, C. et al. Self-organization phenomena in embryonic stem cell-derived embryoid bodies: axis formation and breaking of symmetry during cardiomyogenesis. Cells Tissues Organs 195, 377–391 (2012).
Meinhardt, A. et al. 3D reconstitution of the patterned neural tube from embryonic stem cells. Stem Cell Rep. 3, 987–999 (2014).
Poh, Y.-C. et al. Generation of organized germ layers from a single mouse embryonic stem cell. Nat. Commun. 5, 4000 (2014).
van den Brink, S. C. et al. Symmetry breaking, germ layer specification and axial organisation in aggregates of mouse embryonic stem cells. Development 141, 4231–4242 (2014).
Warmflash, A. et al. A method to recapitulate early embryonic spatial patterning in human embryonic stem cells. Nat. Methods 11, 847–854 (2014).
Harrison, S. E. et al. Assembly of embryonic and extra-embryonic stem cells to mimic embryogenesis in vitro. Science 356, eaal1810 (2017).
Shao, Y. et al. A pluripotent stem cell-based model for post-implantation human amniotic sac development. Nat. Commun. 8, 208 (2017).
Beccari, L. et al. Multi-axial self-organization properties of mouse embryonic stem cells into gastruloids. Nature 562, 272–276 (2018).
Rivron, N. C. et al. Blastocyst-like structures generated solely from stem cells. Nature 557, 106–111 (2018).
Sozen, B. et al. Self-assembly of embryonic and two extra-embryonic stem cell types into gastrulating embryo-like structures. Nat. Cell Biol. 20, 979–989 (2018).
Xue, X. et al. Mechanics-guided embryonic patterning of neuroectoderm tissue from human pluripotent stem cells. Nat. Mater. 17, 633–641 (2018).
Haremaki, T. et al. Self-organizing neuruloids model developmental aspects of Huntington’s disease in the ectodermal compartment. Nat. Biotechnol. 37, 1198–1208 (2019).
Simunovic, M. et al. A 3D model of a human epiblast reveals BMP4-driven symmetry breaking. Nat. Cell Biol. 21, 900–910 (2019).
Zheng, Y. et al. Controlled modelling of human epiblast and amnion development using stem cells. Nature 573, 421–425 (2019).
van den Brink, S. C. et al. Single-cell and spatial transcriptomics reveal somitogenesis in gastruloids. Nature 582, 405–409 (2020).
Zheng, Y. et al. Dorsal-ventral patterned neural cyst from human pluripotent stem cells in a neurogenic niche. Sci. Adv. 5, eaax5933 (2019).
Li, R. et al. Generation of blastocyst-like structures from mouse embryonic and adult cell cultures. Cell 179, 687–702 (2019).
Rossant, J. & Tam, P. P. L. Blastocyst lineage formation, early embryonic asymmetries and axis patterning in the mouse. Development 136, 701–713 (2009).
Wennekamp, S., Mesecke, S., Nédélec, F. & Hiiragi, T. A self-organization framework for symmetry breaking in the mammalian embryo. Nat. Rev. Mol. Cell Biol. 14, 452–459 (2013).
Boroviak, T. & Nichols, J. Primate embryogenesis predicts the hallmarks of human naïve pluripotency. Development 144, 175–186 (2017).
O’Rahilly, R. & Müller, F. Developmental Stages in Human Embryos: Revised and New Measurements. Cells Tissues Organs 192, 73–84 (2010).
Tam, P. P. L. & Loebel, D. A. F. Gene function in mouse embryogenesis: get set for gastrulation. Nat. Rev. Genet. 8, 368–381 (2007).
Arnold, S. J. & Robertson, E. J. Making a commitment: cell lineage allocation and axis patterning in the early mouse embryo. Nat. Rev. Mol. Cell Biol. 10, 91–103 (2009).
Rivera-Pérez, J. A., Mager, J. & Magnuson, T. Dynamic morphogenetic events characterize the mouse visceral endoderm. Dev. Biol. 261, 470–487 (2003).
Thomas, P. & Beddington, R. Anterior primitive endoderm may be responsible for patterning the anterior neural plate in the mouse embryo. Curr. Biol. 6, 1487–1496 (1996).
Yamamoto, M. et al. Nodal antagonists regulate formation of the anteroposterior axis of the mouse embryo. Nature 428, 387–392 (2004).
Yoon, Y. et al. Extra-embryonic Wnt3 regulates the establishment of the primitive streak in mice. Dev. Biol. 403, 80–88 (2015).
Tortelote, G. G. et al. Wnt3 function in the epiblast is required for the maintenance but not the initiation of gastrulation in mice. Dev. Biol. 374, 164–173 (2013).
Ohinata, Y. et al. Blimp1 is a critical determinant of the germ cell lineage in mice. Nature 436, 207–213 (2005).
Saitou, M., Barton, S. C. & Surani, M. A. A molecular programme for the specification of germ cell fate in mice. Nature 418, 293–300 (2002).
Chen, D. et al. Human primordial germ cells are specified from lineage-primed progenitors. Cell Rep. 29, 4568–4582e4565 (2019).
Sasaki, K. et al. The germ cell fate of cynomolgus monkeys is specified in the nascent amnion. Dev. Cell 39, 169–185 (2016).
Colas, J.-F. & Schoenwolf, G. C. Towards a cellular and molecular understanding of neurulation. Dev. Dynam. 221, 117–145 (2001).
Vijayraghavan, D. S. & Davidson, L. A. Mechanics of neurulation: from classical to current perspectives on the physical mechanics that shape, fold, and form the neural tube. Birth Defects Res. 109, 153–168 (2017).
Ribes, V. & Briscoe, J. Establishing and interpreting graded sonic hedgehog signaling during vertebrate neural tube patterning: The role of negative feedback. Cold Spring Harbor Persp. Biol. 1, a002014 (2009).
Briscoe, J. & Small, S. Morphogen rules: Design principles of gradient-mediated embryo patterning. Development 142, 3996–4009 (2015).
Nichols, J. & Smith, A. Naive and primed pluripotent states. Cell Stem Cell 4, 487–492 (2009).
Smith, A. Formative pluripotency: the executive phase in a developmental continuum. Development 144, 365–373 (2017).
Boroviak, T. et al. The ability of inner-cell-mass cells to self-renew as embryonic stem cells is acquired following epiblast specification. Nat. Cell Biol. 16, 513–525 (2014).
Ying, Q.-L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008).
Brons, I. G. M. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007).
Tesar, P. J. et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 448, 196–199 (2007).
Kojima, Y. et al. The transcriptional and functional properties of mouse epiblast stem cells resemble the anterior primitive streak. Cell Stem Cell 14, 107–120 (2014).
Hayashi, K. et al. Reconstitution of the mouse germ cell specification pathway in culture by pluripotent stem cells. Cell 146, 519–532 (2011).
Tanaka, S. et al. Promotion of trophoblast stem cell proliferation by FGF4. Science 282, 2072–2075 (1998).
Niakan, K. K., Schrode, N., Cho, L. T. Y. & Hadjantonakis, A.-K. Derivation of extraembryonic endoderm stem (XEN) cells from mouse embryos and embryonic stem cells. Nat. Protocols 8, 1028–1041 (2013).
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Guo, H. et al. The DNA methylation landscape of human early embryos. Nature 511, 606–610 (2014).
Yan, L. et al. Single-cell RNA-Seq profiling of human preimplantation embryos and embryonic stem cells. Nature Struct. Mol. Biol. 20, 1131–1139 (2013).
Chia, N.-Y. et al. A genome-wide RNAi screen reveals determinants of human embryonic stem cell identity. Nature 468, 316–320 (2010).
Irie, N. et al. SOX17 is a critical specifier of human primordial germ cell fate. Cell 160, 253–268 (2015).
Sasaki, K. et al. Robust in vitro induction of human germ cell fate from pluripotent stem cells. Cell Stem Cell 17, 178–194 (2015).
Theunissen, T. W. et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 15, 471–487 (2014).
Guo, G. et al. Naive pluripotent stem cells derived directly from isolated cells of the human inner cell mass. Stem Cell Rep. 6, 437–446 (2016).
Theunissen, T. W. et al. Molecular criteria for defining the naive human pluripotent state. Cell Stem Cell 19, 502–515 (2016).
Takashima, Y. et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell 158, 1254–1269 (2014).
Yamanaka, S. & Blau, H. M. Nuclear reprogramming to a pluripotent state by three approaches. Nature 465, 704–712 (2010).
Choi, J. et al. A comparison of genetically matched cell lines reveals the equivalence of human iPSCs and ESCs. Nat. Biotechnol. 33, 1173–1181 (2015).
Quadrato, G. et al. Cell diversity and network dynamics in photosensitive human brain organoids. Nature 545, 48–53 (2017).
Yang, Y. et al. Derivation of pluripotent stem cells with in vivo embryonic and extraembryonic potency. Cell 169, 243–257 (2017).
Gao, X. et al. Establishment of porcine and human expanded potential stem cells. Nat. Cell Biol. 21, 687–699 (2019).
Okae, H. et al. Derivation of human trophoblast stem cells. Cell Stem Cell 22, 50–63e56 (2018).
Linneberg-Agerholm, M. et al. Naïve human pluripotent stem cells respond to Wnt, Nodal and LIF signalling to produce expandable naïve extra-embryonic endoderm. Development 146, dev180620 (2019).
Dong, C. et al. Derivation of trophoblast stem cells from naïve human pluripotent stem cells. eLife 9, e52504 (2019).
Guo, G. et al. Trophectoderm potency is retained exclusively in human naïve cells. Preprint at https://doi.org/10.1101/2020.02.04.933812 (2020).
Posfai, E. et al. Defining totipotency using criteria of increasing stringency. Preprint at https://doi.org/10.1101/2020.03.02.972893 (2020).
Frias-Aldeguer, J. et al. Embryonic signals perpetuate polar-like trophoblast stem cells and pattern the blastocyst axis. Preprint at https://doi.org/10.1101/510362 (2019).
Vrij, E. J. et al. Chemically-defined induction of a primitive endoderm and epiblast-like niche supports post-implantation progression from blastoids. Preprint at https://doi.org/10.1101/510396 (2019).
Sozen, B. et al. Self-organization of mouse stem cells into an extended potential blastoid. Dev. Cell 51, 698–712 (2019).
Taniguchi, K. et al. Lumen formation is an intrinsic property of isolated human pluripotent stem cells. Stem Cell Rep. 5, 954–962 (2015).
Shahbazi, M. N. et al. Pluripotent state transitions coordinate morphogenesis in mouse and human embryos. Nature 552, 239–243 (2017).
Minn, K. T. et al. High-resolution transcriptional and morphogenetic profiling of cells from micropatterned human embryonic stem cell gastruloid cultures. Preprint at https://doi.org/10.1101/2020.01.22.915777 (2020).
Tewary, M. et al. A stepwise model of reaction-diffusion and positional-information governs self-organized human peri-gastrulation-like patterning. Development 144, 4298–4312 (2017).
Etoc, F. et al. A balance between secreted inhibitors and edge sensing controls gastruloid self-organization. Dev. Cell 39, 302–315 (2016).
Heemskerk, I. et al. Rapid changes in morphogen concentration control self-organized patterning in human embryonic stem cells. eLife 8, e40526 (2019).
Nemashkalo, A., Ruzo, A., Heemskerk, I. & Warmflash, A. Morphogen and community effects determine cell fates in response to BMP4 signaling in human embryonic stem cells. Development 144, 3042–3053 (2017).
Chhabra, S. et al. Dissecting the dynamics of signaling events in the BMP, WNT, and NODAL cascade during self-organized fate patterning in human gastruloids. PLoS Biol. 17, e3000498 (2019).
Martyn, I., Brivanlou, A. H. & Siggia, E. D. A wave of WNT signaling balanced by secreted inhibitors controls primitive streak formation in micropattern colonies of human embryonic stem cells. Development 146, dev172791 (2019).
Morgani, S. M. et al. Micropattern differentiation of mouse pluripotent stem cells recapitulates embryo regionalized cell fate patterning. eLife 7, e32839 (2018).
Manfrin, A. et al. Engineered signaling centers for the spatially controlled patterning of human pluripotent stem cells. Nat. Methods 16, 640–648 (2019).
Veenvliet, J. V. et al. Mouse embryonic stem cells self-organize into trunk-like structures with neural tube and somites. Preprint at https://doi.org/10.1101/2020.03.04.974949 (2020).
Rossi, G. et al. Embryonic organoids recapitulate early heart organogenesis. Preprint at https://doi.org/10.1101/802181 (2019).
Ranga, A. et al. Neural tube morphogenesis in synthetic 3D microenvironments. Proc. Natl Acad. Sci. USA 113, E6831–E6839 (2016).
Britton, G. et al. A novel self-organizing embryonic stem cell system reveals signaling logic underlying the patterning of human ectoderm. Development 146, dev179093 (2019).
McBeath, R. et al. Cell shape, cytoskeletal tension, and RhoA regulate stem cell lineage commitment. Dev. Cell 6, 483–495 (2004).
Lam, R. H. W., Sun, Y., Chen, W. & Fu, J. Elastomeric microposts integrated into microfluidics for flow-mediated endothelial mechanotransduction analysis. Lab Chip 12, 1865–1873 (2012).
Nelson, C. M. et al. Tissue geometry determines sites of mammary branching morphogenesis in organotypic cultures. Science 314, 298–300 (2006).
Habib, S. J. et al. A localized Wnt signal orients asymmetric stem cell division in vitro. Science 339, 1445–1448 (2013).
Repina, N. A. et al. Optogenetic control of Wnt signaling for modeling early embryogenic patterning with human pluripotent stem cells. Preprint at https://doi.org/10.1101/665695 (2019).
Cederquist, G. Y. et al. Specification of positional identity in forebrain organoids. Nat. Biotechnol. 37, 436–444 (2019).
Sun, Y., Chen, C. S. & Fu, J. Forcing stem cells to behave: A biophysical perspective of the cellular microenvironment. Annu. Rev. Biophys. 41, 519–542 (2012).
Bedzhov, I. & Zernicka-Goetz, M. Self-organizing properties of mouse pluripotent cells initiate morphogenesis upon implantation. Cell 156, 1032–1044 (2014).
Tam, P. P. Postimplantation mouse development: whole embryo culture and micro-manipulation. Int. J. Dev. Biol. 42, 895–902 (1998).
Wang, H. et al. A novel model of human implantation: 3D endometrium-like culture system to study attachment of human trophoblast (Jar) cell spheroids. Mol. Human Reprod. 18, 33–43 (2011).
Tyser, R. C. V. et al. A spatially resolved single cell atlas of human gastrulation. Preprint at https://doi.org/10.1101/2020.07.21.213512 (2020).
Rivron, N. et al. Debate ethics of embryo models from stem cells. Nature 564, 183–185 (2018).
Hyun, I. et al. Toward guidelines for research on human embryo models formed from stem cells. Stem Cell Rep. 14, 169–174 (2020).
Petropoulos, S. et al. Single-Cell RNA-Seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016).
Dobreva, M. P. et al. Amniotic ectoderm expansion in mouse occurs via distinct modes and requires SMAD5-mediated signalling. Development 145, dev157222 (2018).
Pereira, P. N. G. et al. Amnion formation in the mouse embryo: the single amniochorionic fold model. BMC Dev. Biol. 11, 48 (2011).
Lawson, K. A. et al. Bmp4 is required for the generation of primordial germ cells in the mouse embryo. Genes Dev. 13, 424–436 (1999).
Munoz-Sanjuan, I. & Brivanlou, A. H. Neural induction, the default model and embryonic stem cells. Nat. Rev. Neurosci. 3, 271–280 (2002).
Stern, C. D. Neural induction: old problem, new findings, yet more questions. Development 132, 2007–2021 (2005).
Barnat, M. et al. Huntington’s disease alters human neurodevelopment. Science 369, 787–793 (2020).
Acknowledgements
The authors thank S. Vianello for invaluable work on illustrations and N. Rivron, A. Yoney, E.D. Siggia, M. Simunovic and Y. Zheng for microscopic images shown in Fig. 4. J.F.’s research is supported by the University of Michigan Mechanical Engineering Department, the Michigan–Cambridge Research Initiative, the University of Michigan Mcubed Fund, the National Institutes of Health (R21 NS113518 and R21 HD100931), and the National Science Foundation (CMMI 1917304 and CBET 1901718). A.W.’s work is supported by the Rice University, the Welch Foundation (C-2021), the Simons Foundation (511709), the National Institutes of Health (R01 GM126122), and the National Science Foundation (MCB-1553228). M.P.L.’s work is supported by the École Polytechnique Fédérale de Lausanne, the National Center of Competence in Research (NCCR) ‘Bio-Inspired Materials’, and the Swiss National Science Foundation Sinergia Grant (no. 3189956). The authors apologize to colleagues whose work they could not cite owing to space restrictions.
Author information
Authors and Affiliations
Contributions
J.F., A.W. and M.P.L. wrote the manuscript. All authors edited and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Fu, J., Warmflash, A. & Lutolf, M.P. Stem-cell-based embryo models for fundamental research and translation. Nat. Mater. 20, 132–144 (2021). https://doi.org/10.1038/s41563-020-00829-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41563-020-00829-9
This article is cited by
-
Precise and scalable self-organization in mammalian pseudo-embryos
Nature Structural & Molecular Biology (2024)
-
Time-integrated BMP signaling determines fate in a stem cell model for early human development
Nature Communications (2024)
-
Reprogramming fibroblast into human iBlastoids
Nature Protocols (2024)
-
Exploring the promising potential of induced pluripotent stem cells in cancer research and therapy
Molecular Cancer (2023)
-
Generating human blastoids modeling blastocyst-stage embryos and implantation
Nature Protocols (2023)