Abstract
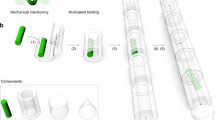
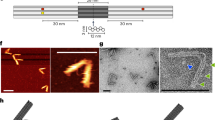
Molecular devices with information-processing capabilities hold great promise for developing intelligent nanorobotics. Here we demonstrate a DNA navigator system that can perform single-molecule parallel depth-first search on a ten-vertex rooted tree defined on a two-dimensional DNA origami platform. Pathfinding by the DNA navigators exploits a localized strand exchange cascade, which is initiated at a unique trigger site on the origami with subsequent automatic progression along paths defined by DNA hairpins containing a universal traversal sequence. Each single-molecule navigator autonomously explores one of the possible paths through the tree. A specific solution path connecting a given pair of start and end vertices can then be easily extracted from the set of all paths taken by the navigators collectively. The solution path laid out on origami is illustrated with single-molecule imaging. Our approach points towards the realization of molecular materials with embedded computational functions operating at the single-molecule level.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All the data that support the findings of this study are available within the paper and its Supplementary Information files, and from the corresponding authors upon reasonable request.
References
Kudernac, T. et al. Electrically driven directional motion of a four-wheeled molecule on a metal surface. Nature 479, 208–211 (2011).
Hagiya, M., Konagaya, A., Kobayashi, S., Saito, H. & Murata, S. Molecular robots with sensors and intelligence. Acc. Chem. Res. 47, 1681–1690 (2014).
Kassem, S. et al. Artificial molecular motors. Chem. Soc. Rev. 46, 2592–2621 (2017).
Yin, P., Yan, H., Daniell, X. G., Turberfield, A. J. & Reif, J. H. A unidirectional DNA walker that moves autonomously along a track. Angew. Chem. Int. Ed. 43, 4906–4911 (2004).
Shin, J.-S. & Pierce, N. A. A synthetic DNA walker for molecular transport. J. Am. Chem. Soc. 126, 10834–10835 (2004).
Tian, Y., He, Y., Chen, Y., Yin, P. & Mao, C. A DNAzyme that walks processively and autonomously along a one-dimensional track. Angew. Chem. Int. Ed. 44, 4355–4358 (2005).
Gu, H., Chao, J., Xiao, S.-J. & Seeman, N. C. A proximity-based programmable DNA nanoscale assembly line. Nature 465, 202–205 (2010).
Lund, K. et al. Molecular robots guided by prescriptive landscapes. Nature 465, 206–210 (2010).
Wickham, S. F. J. et al. A DNA-based molecular motor that can navigate a network of tracks. Nat. Nanotech. 7, 169–173 (2012).
Tomov, T. E. et al. Rational design of DNA motors: Fuel optimization through single-molecule fluorescence. J. Am. Chem. Soc. 135, 11935–11941 (2013).
Cha, T.-G. et al. A synthetic DNA motor that transports nanoparticles along carbon nanotubes. Nat. Nanotech. 9, 39–43 (2014).
Kopperger, E., Pirzer, T. & Simmel, F. C. Diffusive transport of molecular cargo tethered to a DNA origami platform. Nano. Lett. 15, 2693–2699 (2015).
Yang, Y. et al. Direct visualization of walking motions of photocontrolled nanomachine on the DNA nanostructure. Nano. Lett. 15, 6672–6676 (2015).
Thubagere, A. J. et al. A cargo-sorting DNA robot. Science 357, eaan6558 (2017).
Fan, C. & Pei, H. DNA nanotechnology. Chin. J. Chem. 34, 251 (2016).
Zhou, C., Duan, X. & Liu, N. A plasmonic nanorod that walks on DNA origami. Nat. Commun. 6, 8102 (2015).
Zhu, D. et al. A surface-confined proton-driven DNA pump using a dynamic 3D DNA scaffold. Adv. Mater. 28, 6860–6865 (2016).
Adleman, L. M. Molecular computation of solutions to combinatorial problems. Science 266, 1021–1024 (1994).
Elbaz, J. et al. DNA computing circuits using libraries of DNAzyme subunits. Nat. Nanotech. 5, 417–422 (2010).
Pei, R., Matamoros, E., Liu, M., Stefanovic, D. & Stojanovic, M. N. Training a molecular automaton to play a game. Nat. Nanotech. 5, 773–777 (2010).
Macdonald, J. et al. Medium scale integration of molecular logic gates in an automaton. Nano. Lett. 6, 2598–2603 (2006).
Qian, L., Winfree, E. & Bruck, J. Neural network computation with DNA strand displacement cascades. Nature 475, 368–372 (2011).
Douglas, S. M., Bachelet, I. & Church, G. M. A logic-gated nanorobot for targeted transport of molecular payloads. Science 335, 831–834 (2012).
Han, D. et al. A cascade reaction network mimicking the basic functional steps of adaptive immune response. Nat. Chem. 7, 835–841 (2015).
Li, J., Green, A. A., Yan, H. & Fan, C. H. Engineering nucleic acid structures for programmable molecular circuitry and intracellular biocomputation. Nat. Chem. 9, 1056–1067 (2017).
Liu, H., Wang, J., Song, S., Fan, C. & Gothelf, K. V. A DNA-based system for selecting and displaying the combined result of two input variables. Nat. Commun. 6, 10089 (2015).
Boemo, M. A., Lucas, A. E., Turberfield, A. J. & Cardelli, L. The formal language and design principles of autonomous DNA walker circuits. ACS Synth. Biol. 5, 878–884 (2016).
Chatterjee, G., Dalchau, N., Muscat, R. A., Phillips, A. & Seelig, G. A spatially localized architecture for fast and modular DNA computing. Nat. Nanotech. 12, 920–927 (2017).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Voigt, N. V. et al. Single-molecule chemical reactions on DNA origami. Nat. Nanotech. 5, 200–203 (2010).
Lin, C. et al. Submicrometre geometrically encoded fluorescent barcodes self-assembled fromDNA. Nat. Chem. 4, 832–839 (2012).
Kuzyk, A. et al. DNA-based self-assembly of chiral plasmonic nanostructures with tailored optical response. Nature 483, 311–314 (2012).
Knudsen, J. B. et al. Routing of individual polymers in designed patterns. Nat. Nanotech. 10, 892–898 (2015).
Timm, C. & Niemeyer, C. M. Assembly and purification of enzyme-functionalized DNA origami structures. Angew. Chem. Int. Ed. 54, 6745–6750 (2015).
Edwardson, T. G. W., Lau, K. L., Bousmail, D., Serpell, C. J. & Sleiman, H. F. Transfer of molecular recognition information from DNA nanostructures to gold nanoparticles. Nat. Chem. 8, 162–170 (2016).
Zhang, Y. et al. Transfer of two-dimensional oligonucleotide patterns onto stereocontrolled plasmonic nanostructures through DNA-origami-based nanoimprinting lithography. Angew. Chem. Int. Ed. 55, 8036–8040 (2016).
Zhang, Z., Yang, Y., Pincet, F., C. Llaguno, M. & Lin, C. Placing and shaping liposomes with reconfigurable DNA nanocages. Nat. Chem. 9, 653–659 (2017).
Rao, V. N. & Kumar, V. Parallel depth first search. Part I. implementation. Int. J. Parallel Prog. 16, 479–499 (1987).
Dirks, R. M. & Pierce, N. A. Triggered amplification by hybridization chain reaction. Proc. Natl Acad. Sci. USA 101, 15275–15278 (2004).
Lakin, M. R., Youssef, S., Polo, F., Emmott, S. & Phillips, A. Visual DSD: a design and analysis tool for DNA strand displacement systems. Bioinformatics 27, 3211–3213 (2011).
Moore, C. & Mertens, S. The Nature of Computation (Oxford Univ. Press, Oxford, 2011).
Scheible, M. B., Pardatscher, G., Kuzyk, A. & Simmel, F. C. Single molecule characterization of DNA binding and strand displacement reactions on lithographic DNA origami microarrays. Nano. Lett. 14, 1627–1633 (2014).
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 11, 313–318 (2014).
Ovesný, M., Křížek, P., Borkovec, J., Švindrych, Z. & Hagen, G. M. ThunderSTORM: a comprehensive ImageJ plug-in for PALM and STORM data analysis and super-resolution imaging. Bioinformatics 30, 2389–2390 (2014).
Acknowledgements
We greatly appreciate financial support from the Ministry of Science and Technology of China (2016YFA0201200), National Science Foundation of China (21390414, 21473236, 21675167, 21505148, 21722310, 61771253), and the Chinese Academy of Sciences (QYZDJ-SSW-SLH031). We further acknowledge support by the Deutsche Forschungsgemeinschaft through SFB1032 Project A2 and by the Technical University Munich International Graduate School of Science and Engineering.
Author information
Authors and Affiliations
Contributions
C.F. directed the research. C.F., F.C.S., H.L. and J.C. conceived the project and designed the experiments. J.C., J.W., F.W., X.O. and E.K. designed the DNA sequences, constructed the navigator system and performed the single-molecule studies. F.W., Q.L. and J.S. carried out the theoretical simulation. J.C., J.W., F.W., E.K., H.L., Lihua W., J.H., Lianhui W., W.H. and F.C.S. analysed the data. All authors discussed the results and commented on the manuscript. C.F., F.C.S. and H.L. co-wrote the paper.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes 1–4, Supplementary Schemes 1–3, Supplementary Software, Supplementary Figures 1–26, Supplementary References 1–3
Rights and permissions
About this article
Cite this article
Chao, J., Wang, J., Wang, F. et al. Solving mazes with single-molecule DNA navigators. Nature Mater 18, 273–279 (2019). https://doi.org/10.1038/s41563-018-0205-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41563-018-0205-3
This article is cited by
-
DNA as a universal chemical substrate for computing and data storage
Nature Reviews Chemistry (2024)
-
Advances of medical nanorobots for future cancer treatments
Journal of Hematology & Oncology (2023)
-
DNA-based programmable gate arrays for general-purpose DNA computing
Nature (2023)
-
High-entropy alloy nanopatterns by prescribed metallization of DNA origami templates
Nature Communications (2023)
-
Throwing and manipulating and cheating with a DNA nano-dice
Nature Communications (2023)