Abstract
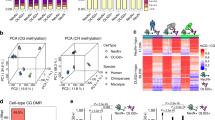
Mammalian brains feature exceptionally high levels of non-CpG DNA methylation alongside the canonical form of CpG methylation. Non-CpG methylation plays a critical regulatory role in cognitive function, which is mediated by the binding of MeCP2, the transcriptional regulator that when mutated causes Rett syndrome. However, it is unclear whether the non-CpG neural methylation system is restricted to mammalian species with complex cognitive abilities or has deeper evolutionary origins. To test this, we investigated brain DNA methylation across 12 distantly related animal lineages, revealing that non-CpG methylation is restricted to vertebrates. We discovered that in vertebrates, non-CpG methylation is enriched within a highly conserved set of developmental genes transcriptionally repressed in adult brains, indicating that it demarcates a deeply conserved regulatory program. We also found that the writer of non-CpG methylation, DNMT3A, and the reader, MeCP2, originated at the onset of vertebrates as a result of the ancestral vertebrate whole-genome duplication. Together, we demonstrate how this novel layer of epigenetic information assembled at the root of vertebrates and gained new regulatory roles independent of the ancestral form of the canonical CpG methylation. This suggests that the emergence of non-CpG methylation may have fostered the evolution of sophisticated cognitive abilities found in the vertebrate lineage.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The sequencing data have been deposited in the Gene Expression Omnibus under the accession number GSE141609.
Code availability
The analysis code is available at https://github.com/AlexdeMendoza/BrainZoo.
References
Schübeler, D. Function and information content of DNA methylation. Nature 517, 321–326 (2015).
Luo, C., Hajkova, P. & Ecker, J. R. Dynamic DNA methylation: in the right place at the right time. Science 361, 1336–1340 (2018).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Suzuki, M. M. & Bird, A. DNA methylation landscapes: provocative insights from epigenomics. Nat. Rev. Genet. 9, 465–476 (2008).
de Mendoza, A., Lister, R. & Bogdanovic, O. Evolution of DNA methylome diversity in eukaryotes. J. Mol. Biol. 432, 1687–1705 (2019).
He, Y. & Ecker, J. R. Non-CG methylation in the human genome. Annu. Rev. Genomics Hum. Genet. 16, 55–77 (2015).
Lister, R. et al. Global epigenomic reconfiguration during mammalian brain development. Science 341, 1237905 (2013).
Mo, A. et al. Epigenomic signatures of neuronal diversity in the mammalian brain. Neuron 86, 1369–1384 (2015).
Lister, R. et al. Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature 471, 68–73 (2011).
Guo, J. U. et al. Distribution, recognition and regulation of non-CpG methylation in the adult mammalian brain. Nat. Neurosci. 17, 215–222 (2014).
Ziller, M. J. et al. Genomic distribution and inter-sample variation of non-CpG methylation across human cell types. PLoS Genet. 7, e1002389 (2011).
Gabel, H. W. et al. Disruption of DNA-methylation-dependent long gene repression in Rett syndrome. Nature 522, 89–93 (2015).
Stroud, H. et al. Early-life gene expression in neurons modulates lasting epigenetic states. Cell 171, 1151–1164 (2017).
Luo, C. et al. Single-cell methylomes identify neuronal subtypes and regulatory elements in mammalian cortex. Science 357, 600–604 (2017).
Amir, R. E. et al. Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nat. Genet. 23, 185–188 (1999).
Lyst, M. J. & Bird, A. Rett syndrome: a complex disorder with simple roots. Nat. Rev. Genet. 16, 261–275 (2015).
Tatton-Brown, K. et al. Mutations in the DNA methyltransferase gene DNMT3A cause an overgrowth syndrome with intellectual disability. Nat. Genet. 46, 385–388 (2014).
Laine, V. N. et al. Evolutionary signals of selection on cognition from the great tit genome and methylome. Nat. Commun. 7, 10474 (2016).
Derks, M. F. L. et al. Gene and transposable element methylation in great tit (Parus major) brain and blood. BMC Genomics 17, 332 (2016).
Sugahara, F. et al. Evidence from cyclostomes for complex regionalization of the ancestral vertebrate brain. Nature 531, 97–100 (2016).
Roth, G. Convergent evolution of complex brains and high intelligence. Phil. Trans. R. Soc. B 370, 20150049 (2015).
Holland, L. Z. et al. Evolution of bilaterian central nervous systems: a single origin? EvoDevo 4, 27 (2013).
Arendt, D., Tosches, M. A. & Marlow, H. From nerve net to nerve ring, nerve cord and brain—evolution of the nervous system. Nat. Rev. Neurosci. 17, 61–72 (2016).
Bogdanović, O. et al. Active DNA demethylation at enhancers during the vertebrate phylotypic period. Nat. Genet. 48, 417–426 (2016).
Marlétaz, F. et al. Amphioxus functional genomics and the origins of vertebrate gene regulation. Nature 564, 64–70 (2018).
Albuixech-Crespo, B. et al. Molecular regionalization of the developing amphioxus neural tube challenges major partitions of the vertebrate brain. PLoS Biol. 15, e2001573 (2017).
Lyko, F. The DNA methyltransferase family: a versatile toolkit for epigenetic regulation. Nat. Rev. Genet. 19, 81–92 (2018).
Mugal, C. F., Arndt, P. F., Holm, L. & Ellegren, H. Evolutionary consequences of DNA methylation on the GC content in vertebrate genomes. G3 5, 441–447 (2015).
Albertin, C. B. et al. The octopus genome and the evolution of cephalopod neural and morphological novelties. Nature 524, 220–224 (2015).
Zhang, Z. et al. Genome-wide and single-base resolution DNA methylomes of the sea lamprey (Petromyzon marinus) reveal gradual transition of the genomic methylation pattern in early vertebrates. Preprint at https://doi.org/10.1101/033233 (2015).
Shen, J. C., Rideout, W. M. 3rd & Jones, P. A. The rate of hydrolytic deamination of 5-methylcytosine in double-stranded DNA. Nucleic Acids Res. 22, 972–976 (1994).
Pfeifer, G. P. Mutagenesis at methylated CpG sequences. Curr. Top. Microbiol. Immunol. 301, 259–281 (2006).
Smith, J. J. et al. The sea lamprey germline genome provides insights into programmed genome rearrangement and vertebrate evolution. Nat. Genet. 50, 270–277 (2018).
Kinde, B., Gabel, H. W., Gilbert, C. S., Griffith, E. C. & Greenberg, M. E. Reading the unique DNA methylation landscape of the brain: non-CpG methylation, hydroxymethylation, and MeCP2. Proc. Natl Acad. Sci. USA 112, 6800–6806 (2015).
Xie, W. et al. Base-resolution analyses of sequence and parent-of-origin dependent DNA methylation in the mouse genome. Cell 148, 816–831 (2012).
Wienholz, B. L. et al. DNMT3L modulates significant and distinct flanking sequence preference for DNA methylation by DNMT3A and DNMT3B in vivo. PLoS Genet. 6, e1001106 (2010).
Herculano-Houzel, S. The glia/neuron ratio: how it varies uniformly across brain structures and species and what that means for brain physiology and evolution. Glia 62, 1377–1391 (2014).
Olkowicz, S. et al. Birds have primate-like numbers of neurons in the forebrain. Proc. Natl Acad. Sci. USA 113, 7255–7260 (2016).
Clemens, A. W. et al. MeCP2 represses enhancers through chromosome topology-associated DNA methylation. Mol. Cell 77, 279–293 (2020).
Brawand, D. et al. The evolution of gene expression levels in mammalian organs. Nature 478, 343–348 (2011).
Villar, D. et al. Enhancer evolution across 20 mammalian species. Cell 160, 554–566 (2015).
Jeong, M. et al. Large conserved domains of low DNA methylation maintained by DNMT3A. Nat. Genet. 46, 17–23 (2014).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Sendžikaitė, G., Hanna, C. W., Stewart-Morgan, K. R., Ivanova, E. & Kelsey, G. A DNMT3A PWWP mutation leads to methylation of bivalent chromatin and growth retardation in mice. Nat. Commun. 10, 1884 (2019).
Heyn, P. et al. Gain-of-function DNMT3A mutations cause microcephalic dwarfism and hypermethylation of Polycomb-regulated regions. Nat. Genet. 51, 96–105 (2019).
Lee, J.-H., Park, S.-J. & Nakai, K. Differential landscape of non-CpG methylation in embryonic stem cells and neurons caused by DNMT3s. Sci. Rep. 7, 11295 (2017).
Albalat, R., Martí-Solans, J. & Cañestro, C. DNA methylation in amphioxus: from ancestral functions to new roles in vertebrates. Brief. Funct. Genomics 11, 142–155 (2012).
Liu, J., Hu, H., Panserat, S. & Marandel, L. Evolutionary history of DNA methylation related genes in chordates: new insights from multiple whole genome duplications. Sci. Rep. 10, 970 (2020).
Greenberg, M. V. C. & Bourc’his, D. The diverse roles of DNA methylation in mammalian development and disease. Nat. Rev. Mol. Cell Biol. 20, 590–607 (2019).
Smith, T. H. L., Collins, T. M. & McGowan, R. A. Expression of the dnmt3 genes in zebrafish development: similarity to Dnmt3a and Dnmt3b. Dev. Genes Evol. 220, 347–353 (2011).
Lagger, S. et al. MeCP2 recognizes cytosine methylated tri-nucleotide and di-nucleotide sequences to tune transcription in the mammalian brain. PLoS Genet. 13, e1006793 (2017).
Tillotson, R. et al. Neuronal non-CG methylation is an essential target for MeCP2 function. Preprint at https://doi.org/10.1101/2020.07.02.184614 (2020).
Albalat, R. Evolution of DNA-methylation machinery: DNA methyltransferases and methyl-DNA binding proteins in the amphioxus Branchiostoma floridae. Dev. Genes Evol. 218, 691–701 (2008).
Millar, C. B. et al. Enhanced CpG mutability and tumorigenesis in MBD4-deficient mice. Science 297, 403–405 (2002).
Lyst, M. J. et al. Rett syndrome mutations abolish the interaction of MeCP2 with the NCoR/SMRT co-repressor. Nat. Neurosci. 16, 898–902 (2013).
Jones, P. L. et al. Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nat. Genet. 19, 187–191 (1998).
Nan, X. et al. Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex. Nature 393, 386–389 (1998).
Skene, P. J. et al. Neuronal MeCP2 is expressed at near histone-octamer levels and globally alters the chromatin state. Mol. Cell 37, 457–468 (2010).
Klose, R. J. et al. DNA binding selectivity of MeCP2 due to a requirement for A/T sequences adjacent to methyl-CpG. Mol. Cell 19, 667–678 (2005).
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220 (2010).
Bewick, A. J. et al. On the origin and evolutionary consequences of gene body DNA methylation. Proc. Natl Acad. Sci. USA 113, 9111–9116 (2016).
Yaari, R. et al. RdDM-independent de novo and heterochromatin DNA methylation by plant CMT and DNMT3 orthologs. Nat. Commun. 10, 1613 (2019).
Bonasio, R. et al. Genome-wide and caste-specific DNA methylomes of the ants Camponotus floridanus and Harpegnathos saltator. Curr. Biol. 22, 1755–1764 (2012).
Harris, K. D., Lloyd, J. P. B., Domb, K., Zilberman, D. & Zemach, A. DNA methylation is maintained with high fidelity in the honey bee germline and exhibits global non-functional fluctuations during somatic development. Epigenetics Chromatin 12, 62 (2019).
de Mendoza, A. et al. Convergent evolution of a vertebrate-like methylome in a marine sponge. Nat. Ecol. Evol. 3, 1464–1473 (2019).
Ross, S. E., Angeloni, A., Geng, F. S., de Mendoza, A. & Bogdanovic, O. Developmental remodelling of non-CG methylation at satellite DNA repeats. Nucleic Acids Res. 48, 12675–12688 (2020).
Torres-Méndez, A. et al. A novel protein domain in an ancestral splicing factor drove the evolution of neural microexons. Nat. Ecol. Evol. 3, 691–701 (2019).
Urich, M. A., Nery, J. R., Lister, R., Schmitz, R. J. & Ecker, J. R. MethylC-seq library preparation for base-resolution whole-genome bisulfite sequencing. Nat. Protoc. 10, 475–483 (2015).
Peat, J. R., Ortega-Recalde, O., Kardailsky, O. & Hore, T. A. The elephant shark methylome reveals conservation of epigenetic regulation across jawed vertebrates. F1000Research 6, 526 (2017).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Guo, W. et al. BS-Seeker2: a versatile aligning pipeline for bisulfite sequencing data. BMC Genomics 14, 774 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Guo, W. et al. CGmapTools improves the precision of heterozygous SNV calls and supports allele-specific methylation detection and visualization in bisulfite-sequencing data. Bioinformatics 34, 381–387 (2018).
Zemach, A., McDaniel, I. E., Silva, P. & Zilberman, D. Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science 328, 916–919 (2010).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Landau, D. A. et al. Locally disordered methylation forms the basis of intratumor methylome variation in chronic lymphocytic leukemia. Cancer Cell 26, 813–825 (2014).
Hansen, K. D., Langmead, B. & Irizarry, R. A. BSmooth: from whole genome bisulfite sequencing reads to differentially methylated regions. Genome Biol. 13, R83 (2012).
Wagih, O. ggseqlogo: a versatile R package for drawing sequence logos. Bioinformatics 33, 3645–3647 (2017).
Reimand, J., Kull, M., Peterson, H., Hansen, J. & Vilo, J. g:Profiler—a web-based toolset for functional profiling of gene lists from large-scale experiments. Nucleic Acids Res. 35, W193–W200 (2007).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Venkatesh, B. et al. Elephant shark genome provides unique insights into gnathostome evolution. Nature 505, 174–179 (2014).
Barbosa-Morais, N. L. et al. The evolutionary landscape of alternative splicing in vertebrate species. Science 338, 1587–1593 (2012).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Cardoso-Moreira, M. et al. Gene expression across mammalian organ development. Nature 571, 505–509 (2019).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Nguyen, L. T., Schmidt, H. A., Von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Acknowledgements
We dedicate this paper to the memory of Jose Luis Gomez-Skarmeta, a dear friend and colleague who was instrumental in igniting this project and contributing to this work, but who sadly passed away during the revision process. We also thank N. Maeso for critical reading of this manuscript and suggestions, and M. Irimia for advice on isoform quantification. We thank N. Saunders (University of Melbourne) for sharing the opossum material. We thank J. Pascual-Anaya for granting access to the hagfish genome assembly. We thank Semilleria las Ganchozas for providing advice about the material required for this project. This work was supported by the Australian Research Council (ARC) Centre of Excellence programme in Plant Energy Biology (grant no. CE140100008). R.L. was supported by a Sylvia and Charles Viertel Senior Medical Research Fellowship, ARC Future Fellowship (no. FT120100862) and Howard Hughes Medical Institute International Research Scholarship. A.d.M. was funded by an EMBO long-term fellowship (no. ALTF 144-2014). J.L.G.-S. was supported by the Spanish government (grant no. BFU2016- 74961-P) and the institutional grant Unidad de Excelencia María de Maeztu (no. MDM-2016-0687). B.V. was supported by the Biomedical Research Council of the Agency for Science, Technology and Research of Singapore. F.G. was supported by an ARC Future Fellowship (no. FT160100267). C.W.R. was supported by an NSF grant (no. IOS-1354898). J.R.E. is an investigator of the Howard Hughes Medical Institute. Genomic data was generated at the Australian Cancer Research Foundation Centre for Advanced Cancer Genomics.
Author information
Authors and Affiliations
Contributions
O.B., A.d.M. and R.L. designed the study. A.d.M., O.B., D.P. and R.L. prepared the MethylC-seq libraries, which were sequenced by J.P., J.R.N. and D.P. A.d.M. analysed the data with help from S. Buckberry. J.L.G.-S., E.d.l.C.-M., C.B.A., C.W.R., F.G., T.D., B.V., J.R.E., B.B., S. Bertrand and H.E. provided the biological samples. A.d.M., O.B. and R.L. wrote the manuscript. All authors commented on the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Ecology and Evolution thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Locally disordered methylation characterises the lamprey epigenome.
Proportion of Discordant Reads (PDR) values for a subset of CpGs (100,000) of each species (See Methods). Boxplot centre lines are medians, box limits are quartiles 1 (Q1) and 3 (Q3), whiskers are 1.5 × interquartile range (IQR).
Extended Data Fig. 2 CpG hypermutability is widespread in vertebrates except the lamprey.
Percentage of Single Nucleotide Variants identified from the WGBS libraries from the total number of dinucleotides in the reference genome. In pale blue are those proportions that are equal or lower than the expected (total number of SNVs / total number of dinucleotides), and in dark blue are those that are overrepresented. Note that the mouse has very few SNVs as it is a laboratory isogenic line, however it still shows a slightly higher enrichment for SNVs in CpG dinucleotides, whereas birds have very high SNV rates on CpG dinucleotides despite having intermediate levels of CpG methylation.
Extended Data Fig. 3 CpH methylation is specific to brain tissues across vertebrates.
Sequence motifs found surrounding the highest methylated CpH positions in each sample. CpH positions were required to have a coverage ≥ 10x. hpf = embryo hours post fertilization. Sox10+ cells correspond to developmental neural crest cells in zebrafish. The motif found in the zebrafish brain sample corresponds to the satellite repeat MOSAT, a unique type of non-CpG methylation in zebrafish66. (b) Gene Ontology enrichments for genes showing the highest and lowest gene body methylation levels in the CpA context, as defined by belonging to the top and bottom deciles in each species and tissue. (c) Gene Ontology enrichments for genes showing the highest and lowest methylated levels in the CpG context.
Extended Data Fig. 4 Anticorrelation between CpG and CpA methylation and transcription is restricted to a subset of vertebrate samples.
Distribution of gene body methylation levels on genes separated by expression level on brain tissue. ‘No expression’ category includes all genes with TPM < 1, whereas the rest of genes were classified in 10 deciles of expression (lower expression left, higher expression right). Positive correlation between expression and CpG methylation is restricted to invertebrate brain samples. Boxplot centre lines are medians, box limits are quartiles 1 (Q1) and 3 (Q3), whiskers are 1.5 × interquartile range (IQR).
Extended Data Fig. 5 Gene classification by CpA and CpG methylation levels.
(a) Distribution of gene body methylation levels on genes classified in deciles from lower to higher methylation levels. Few genes are CpG methylated in the honeybee (only 3 top deciles). The dynamic range of CpG gene body methylation of lampreys and birds differs from the rest of vertebrates, in which a vast majority of genes are highly methylated (>50%). Boxplot centre lines are medians, box limits are quartiles 1 (Q1) and 3 (Q3), whiskers are 1.5 × interquartile range (IQR). (b) Overlap between top and bottom decile genes classified by CpA and CpG gene body methylation levels. All deciles have the same size, thus overlap % captures the relative differences between categories in a comparable manner. (c) Level of conservation of gene sets classified by CpA and CpG gene body methylation levels. If a given orthologue is present in one subset of genes in only one species it is classified as a Singleton (1), whereas if it is found in the nine vertebrate species analyzed it is classified as 9. Each orthologue is counted once per species (for example if lamprey has 2 species-specific paralogues of one gene, it is only counted as 1).
Extended Data Fig. 6 Expression level of highly conserved CpA methylated genes.
Standardized expression level for genes conserved in at least 7 vertebrate species as belonging to the top decile of CpA methylated genes. Boxplot centre lines are medians, box limits are quartiles 1 (Q1) and 3 (Q3), whiskers are 1.5 × interquartile range (IQR).
Extended Data Fig. 7 Phylogeny and expression of DNMT3 enzymes.
(a) Maximum likelihood phylogenetic tree of DNMT3 orthologues across animals, representing the full version of that presented in Fig. 4a. Nodal supports represent 100 bootstrap nonparametric replications. Schematic protein domain configurations shown for each clade. PWWP, Pro-Trp-Trp-Pro motif domain (PF00855). AAD ATRX, DNMT3, DNMT3L domain. MT, cytosine Methyltransferase domain (PF00145). CH, Calponin Homology domain (PF00307). Asterisk highlights arctic lamprey sequences. Broken domains indicate that the domain has large deletions in the given clade. (b) Table with the steady-state transcriptional level of DNMT3A in vertebrate samples, and DNMT3 in invertebrate samples. Compared to previous analysis of the DNMT3 family, here we describe for the first time the presence of DNMT3L in non-mammalian genomes. These include non-avian reptiles (turtles, crocodiles and squamates) and two lamprey genomes. This indicates that DNMT3L was one of the ancestral onhologues product of the vertebrate ancestral WGD. Interestingly, both lampreys and tetrapod sequences show a truncated cytosine methyltransferase domain, which might indicate that the DNMT3L has been conserved despite its lack of catalytic activity.
Extended Data Fig. 8 Phylogeny and conservation of MBD4/MECP2.
(a) Maximum likelihood phylogenetic tree of the Methyl-CpG Binding Domain family in animals, representing the full-version of Fig. 4b. Nodal supports represent 100 bootstrap nonparametric replications. On the right, protein domain structure of each clade, as defined by Pfam domains. MBD, Methyl Binding Domain (PF01429). HhH-GPD, Thymine glycosylase (PF00730). MBDa, p55-binding region of MBD2/3 (PF16564). MBD_C, MBD2/3 C-terminal domain (PF14048). zf-CXXC, zinc finger (PF02008). CTD, MECP2 C-Terminal Domain. TRD, MECP2 Transcriptional Repression Domain. (b) Domain presence in MBD4/MECP2 orthologues in several invertebrate genomes. Lack of the Thymine glycosylase domain is likely due to incomplete gene annotation or genome assembly gaps.
Extended Data Fig. 9 Conservation of the MeCP2 protein domains.
(a) Amino acid multi-sequence alignment (MAFFT E-INS-i mode) of the Methyl-CpG Binding domain (MBD) from MeCP2, MBD4 and invertebrate MECP2/MBD4 sequences. The black square highlights the MBD domain as defined by Pfam. The red triangles indicate positions mutated in the human MECP2 gene that cause Rett Syndrome phenotypes52. (b) Amino acid multi-sequence alignment of the Transcriptional Repression Domain (TRD) from MeCP2, MBD4 and the homologous region (C-terminal of the MBD) of invertebrate MBD4/MECP2 proteins. NID stands for the N-CoR/SMRT interacting amino acids. Additional black squares highlight the AT-hook domains. Alignment visualised using Geneious software.
Extended Data Fig. 10 MBD4/MECP2 isoform expression in the european amphioxus.
Diagram representing the sequences used to uniquely map RNA-seq reads to each isoform across different tissues and developmental stages. Quantification of each isoform in each sample, normalised by gene length (TPM as per Kallisto quantification).
Supplementary information
Supplementary Information
Supplementary Figs. 1–3 and legends for Tables 1–3.
Supplementary Table 1
Gene Ontologies obtained using gProfileR. mCA_hi corresponds to the top deciles of CpA methylation, mCA_low corresponds to the bottom deciles of CpA methylation, mCG_hi corresponds to the top deciles of CpG methylation and mCG_low corresponds to the bottom deciles of CpG methylation.
Supplementary Table 2
List of genes conserved on the high CpA methylated decile across species.
Supplementary Table 3
‘Core set’ indicates the genomes that were searched in all cases. Additional genomes surveyed for specific searches are also highlighted in the Comment column. DNMT3B in amphibians and DNMT3L for non-mammals were searched against the NCBI non-redundant database.
Rights and permissions
About this article
Cite this article
de Mendoza, A., Poppe, D., Buckberry, S. et al. The emergence of the brain non-CpG methylation system in vertebrates. Nat Ecol Evol 5, 369–378 (2021). https://doi.org/10.1038/s41559-020-01371-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-020-01371-2
This article is cited by
-
Extensive DNA methylome rearrangement during early lamprey embryogenesis
Nature Communications (2024)
-
Ultrafast bisulfite sequencing detection of 5-methylcytosine in DNA and RNA
Nature Biotechnology (2024)
-
Methylation across the central dogma in health and diseases: new therapeutic strategies
Signal Transduction and Targeted Therapy (2023)
-
Comparative analysis of genome-scale, base-resolution DNA methylation profiles across 580 animal species
Nature Communications (2023)
-
Epigenetic machinery is functionally conserved in cephalopods
BMC Biology (2022)