Abstract
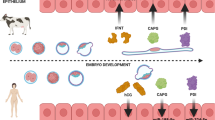
Among mammals, placental invasion is correlated with vulnerability to malignancy. Animals with more invasive placentation (for example, humans) are more vulnerable to malignancy. To explain this correlation, we propose the hypothesis of ‘Evolved Levels of Invasibility’ proposing that the evolution of invasibility of stromal tissue affects both placental and cancer invasion. We provide evidence for this using an in vitro model. We find that bovine endometrial and skin fibroblasts are more resistant to invasion than are their human counterparts. Gene expression profiling identified genes with high expression in human but not in bovine fibroblasts. Knocking down a subset of them in human fibroblasts leads to stronger resistance to cancer cell invasion. Identifying the evolutionary determinants of stromal invasibility can provide important insights to develop rational antimetastatic therapeutics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data for the human–cow transcriptome comparison with and without co-culture with trophoblast cells are available under GSE136299 in the Gene Expression Omnibus (GEO) database of NCBI, https://www.ncbi.nlm.nih.gov/geo/. The comparative fibroblast gene expression data are available under PRJNA564062 under SUB6229748 and SUB6264591 on the Sequence Read Archive (SRA) of NCBI, https://www.ncbi.nlm.nih.gov/sra.
Code availability
The computational methods are described and cited in Methods. No new code was used in this study.
Change history
16 January 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Chaturvedi, P. et al. Hypoxia-inducible factor-dependent breast cancer–mesenchymal stem cell bidirectional signaling promotes metastasis. J. Clin. Invest. 123, 189–205 (2013).
Colegio, O. R. et al. Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature 513, 559–563 (2014).
Zhang, W. & Huang, P. Cancer–stromal interactions: role in cell survival, metabolism and drug sensitivity. Cancer Bio. Ther. 11, 150–156 (2011).
Luga, V. & Wrana, J. L. Tumor–stroma interaction: revealing fibroblast-secreted exosomes as potent regulators of Wnt–planar cell polarity signaling in cancer metastasis. Cancer Res. 73, 6843–6847 (2013).
Murray, M. J. & Lessey, B. A. Embryo implantation and tumor metastasis: common pathways of invasion and angiogenesis. Semin. Reprod. Endocrinol. 17, 275–290 (1999).
Ferretti, C., Bruni, L., Dangles-Marie, V., Pecking, A. P. & Bellet, D. Molecular circuits shared by placental and cancer cells, and their implications in the proliferative, invasive and migratory capacities of trophoblasts. Hum. Reprod. Update 13, 121–141 (2007).
Ratcliffe, H. L. Incidence and nature of tumors in captive wild mammals and birds. Cancer Res. 17, 132–157 (1932).
Canfield, P. J., Hartley, W. J. & Reddacliff, G. L. Spontaneous proliferations in Australian marsupials—a survey and review. 2. Dasyurids and bandicoots. J. Comp. Pathol. 103, 147–158 (1990).
D’Souza, A. W. & Wagner, G. P. Malignant cancer and invasive placentation: a case for positive pleiotropy between endometrial and malignancy phenotypes. Evol. Mes. Public Health 2014, 136–145 (2014).
Perry, J. K., Lins, R. J., Lobie, P. E. & Mitchell, M. D. Regulation of invasive growth: similar epigenetic mechanisms underpin tumour progression and implantation in human pregnancy. Clin. Sci. 118, 451–457 (2009).
Bruning, A., Makovitzky, J., Gingelmaier, A., Friese, K. & Mylonas, I. The metastasis-associated genes MTA1 and MTA3 are abundantly expressed in human placenta and chorionic carcinoma cells. Histochem. Cell Biol. 132, 33–38 (2009).
Wildman, D. E. et al. Evolution of the mammalian placenta revealed by phylogenetic analysis. Proc. Natl Acad. Sci. USA 103, 3203–3208 (2006).
Mess, A. & Carter, A. M. Evolutionary transformation of fetal membrane characters in Eutheria with special reference to Afrotheria. J. Exp. Zool. B 306, 140–163 (2006).
Elliot, M. G. & Crespi, B. J. Phylogenetic evidence for early hemochorial placentation in eutheria. Placenta 30, 949–967 (2009).
Haeger, A., Krause, M., Wolf, K. & Friedl, P. Cell jamming: collective invasion of mesenchymal tumor cells imposed by tissue confinement. Biochim. Biophys. Acta 1840, 2386–2395 (2014).
Shamir, E. R., Coutinho, K., Georgess, D., Auer, M. & Ewald, A. J. Twist1-positive epithelial cells retain adhesive and proliferative capacity throughout dissemination. Biol. Open 5, 1216–1228 (2016).
Wolf, K. et al. Multi-step pericellular proteolysis controls the transition from individual to collective cancer cell invasion. Nat. Cell Biol. 9, 893–904 (2007).
Lau, T. S. et al. A loop of cancer–stroma–cancer interaction promotes peritoneal metastasis of ovarian cancer via TNFalpha–TGFalpha–EGFR. Oncogene 36, 3576–3587 (2017).
Bremnes, R. M. et al. The role of tumor stroma in cancer progression and prognosis: emphasis on carcinoma-associated fibroblasts and non-small cell lung cancer. J. Thorac. Oncol. 6, 209–217 (2011).
Howard, J. D. et al. Notch signaling mediates melanoma–endothelial cell communication and melanoma cell migration. Pigment Cell Melanoma Res. 26, 697–707 (2013).
Kshitiz, Afzal, J., Kim, S. Y. & Kim, D. H. A nanotopography approach for studying the structure–function relationships of cells and tissues. Cell Adh. Migr. 9, 300–307 (2015).
Kim, D. H., Provenzano, P. P., Smith, C. L. & Levchenko, A. Matrix nanotopography as a regulator of cell function. J. Cell Biol. 197, 351–360 (2012).
Kondo, F. Assessment of stromal invasion for correct histological diagnosis of early hepatocellular carcinoma. Int. J. Hepatol. 2011, 241652 (2011).
Khalil, A. A. et al. Collective invasion in ductal and lobular breast cancer associates with distant metastasis. Clin. Exp. Metastasis 34, 421–429 (2017).
Friedl, P. & Alexander, S. Cancer invasion and the microenvironment: plasticity and reciprocity. Cell 147, 992–1009 (2011).
Cheung, K. J. et al. Polyclonal breast cancer metastases arise from collective dissemination of keratin 14-expressing tumor cell clusters. Proc. Natl Acad. Sci. USA 113, E854–863 (2016).
Ramsey, E. M. The Placenta: Human and Animal (Praeger, 1982).
Wooding, P. & Burton, G. Comparative Placentation: Structures, Functions and Evolution (Springer, 2008).
Orendi, K. et al. Placental and trophoblastic in vitro models to study preventive and therapeutic agents for preeclampsia. Placenta 32, S49–54 (2011).
Bertero, L. et al. Eighth Edition of the UICC Classification of Malignant Tumours: an overview of the changes in the pathological TNM classification criteria—what has changed and why? Virchows Arch. 472, 519–531 (2018).
Musser, J. M. & Wagner, G. P. Character trees from transcriptome data: origin and individuation of morphological characters and the so-called “species signal”. J. Exp. Zool. B 324, 588–604 (2015).
Wagner, G. P. Homology, Genes and Evolutionary Innovation (Princeton Univ. Press, 2014).
Liang, C., Musser, J. M., Cloutier, A., Prum, R. O. & Wagner, G. P. Pervasive correlated evolution in gene expression shapes cell and tissue type transcriptomes. Genome Biol. Evol. 10, 538–552 (2018).
Garma-Avina, A., Valli, V. E. & Lumsden, J. H. Cutaneous melanomas in domestic animals. J. Cutan. Pathol. 8, 3–24 (1981).
Meng, X. M., Nikolic-Paterson, D. J. & Lan, H. Y. TGF-beta: the master regulator of fibrosis. Nat. Rev. Nephrol. 12, 325–338 (2016).
Guido, C. et al. Metabolic reprogramming of cancer-associated fibroblasts by TGF-beta drives tumor growth: connecting TGF-beta signaling with “Warburg-like” cancer metabolism and L-lactate production. Cell Cycle 11, 3019–3035 (2012).
Vallee, A., Lecarpentier, Y., Guillevin, R. & Vallee, J. N. Interactions between TGF-beta1, canonical WNT/beta-catenin pathway and PPAR gamma in radiation-induced fibrosis. Oncotarget 8, 90579–90604 (2017).
Calon, A. et al. Stromal gene expression defines poor-prognosis subtypes in colorectal cancer. Nat. Genet. 47, 320–329 (2015).
Piersma, B., Bank, R. A. & Boersema, M. Signaling in fibrosis: TGF-beta, WNT, and YAP/TAZ Converge. Front. Med. 2, 59 (2015).
Akhmetshina, A. et al. Activation of canonical Wnt signalling is required for TGF-beta-mediated fibrosis. Nat. Commun. 3, 735 (2012).
Menke, J. et al. Circulating CSF-1 promotes monocyte and macrophage phenotypes that enhance lupus nephritis. J. Am. Soc. Nephrol. 20, 2581–2592 (2009).
Takada, I., Kouzmenko, A. P. & Kato, S. Wnt and PPARgamma signaling in osteoblastogenesis and adipogenesis. Nat. Rev. Rheumatol. 5, 442–447 (2009).
Ahn, E. H. et al. Spatial control of adult stem cell fate using nanotopographic cues. Biomaterials 35, 2401–2410 (2014).
Hubbi, M. E. et al. A nontranscriptional role for HIF-1alpha as a direct inhibitor of DNA replication. Sci. Signal. 6, ra10 (2013).
Suhail, Y. et al. Modeling intercellular transfer of biomolecules through tunneling nanotubes. Bull. Math. Biol. 75, 1400–1416 (2013).
Zhang, P., Takeuchi, K., Csaki, L. S. & Reue, K. Lipin-1 phosphatidic phosphatase activity modulates phosphatidate levels to promote peroxisome proliferator-activated receptor gamma (PPARgamma) gene expression during adipogenesis. J. Biol. Chem. 287, 3485–3494 (2012).
Park, H. W. et al. Alternative Wnt signaling activates YAP/TAZ. Cell 162, 780–794 (2015).
Meredith, R. W. et al. Impacts of the cretaceous terrestrial revolution and KPg extinction on mammal diversification. Science 334, 521–524 (2011).
de Magalhaes, J. P. & Costa, J. A database of vertebrate longevity records and their relation to other life-history traits. J. Evol. Biol. 22, 1770–1774 (2009).
Seluanov, A. et al. Hypersensitivity to contact inhibition provides a clue to cancer resistance of naked mole-rat. Proc. Natl Acad. Sci. USA 106, 19352–19357 (2009).
Tian, X. et al. High-molecular-mass hyaluronan mediates the cancer resistance of the naked mole rat. Nature 499, 346–349 (2013).
Nunney, L. Lineage selection and the evolution of multistage carcinogenesis. Proc. Biol. Sci. 266, 493–498 (1999).
Caulin, A. F., Graham, T. A., Wang, L. S. & Maley, C. C. Solutions to Peto’s paradox revealed by mathematical modelling and cross-species cancer gene analysis. Phil. Trans. R. Soc. Lond. B 370, 20140222 (2015).
Abegglen, L. M. et al. Potential mechanisms for cancer resistance in elephants and comparative cellular response to DNA damage in humans. J. Am. Med. Assoc. 314, 1850–1860 (2015).
Sulak, M. et al. TP53 copy number expansion is associated with the evolution of increased body size and an enhanced DNA damage response in elephants. eLife 5, e11994 (2016).
Costanzo, V., Bardelli, A., Siena, S. & Abrignani, S. Exploring the links between cancer and placenta development. Open Biol. 8, 180081 (2018).
Curik, I. et al. Inbreeding and melanoma in Lipizzan horses. Agric. Conspec. Sci. 65, 181–186 (2000).
Pandey, M. K. et al. Benign melanocytoma in a non-descript cow: a case report. Indian J. Anim. Res. 50, 632–633 (2016).
Priester, W. A. & Mantel, N. Occurrence of tumors in domestic animals. Data from 12 United States and Canadian colleges of veterinary medicine. J. Nat. Cancer Inst. 47, 1333–1344 (1971).
Bego, T. et al. Association of PPARG and LPIN1 gene polymorphisms with metabolic syndrome and type 2 diabetes. Med. Glas. 8, 76–83 (2011).
Wong, N. A. & Pignatelli, M. Beta-catenin—a linchpin in colorectal carcinogenesis? Am. J. Pathol. 160, 389–401 (2002).
Zhao, B., Li, L., Lei, Q. & Guan, K. L. The Hippo–YAP pathway in organ size control and tumorigenesis: an updated version. Genes Dev. 24, 862–874 (2010).
Padua, D. & Massague, J. Roles of TGFbeta in metastasis. Cell Res. 19, 89–102 (2009).
Wendt, M. K., Tian, M. & Schiemann, W. P. Deconstructing the mechanisms and consequences of TGF-beta-induced EMT during cancer progression. Cell Tissue Res. 347, 85–101 (2012).
Weeraratna, A. T. et al. Wnt5a signaling directly affects cell motility and invasion of metastatic melanoma. Cancer Cell 1, 279–288 (2002).
Kurayoshi, M. et al. Expression of Wnt-5a is correlated with aggressiveness of gastric cancer by stimulating cell migration and invasion. Cancer Res. 66, 10439–10448 (2006).
Kim, J. Y., Kim, G., Lim, S. C. & Choi, H. S. LPIN1 promotes epithelial cell transformation and mammary tumourigenesis via enhancing insulin receptor substrate 1 stability. Carcinogenesis 37, 1199–1209 (2016).
Mendelsohn, R. et al. Complex N-glycan and metabolic control in tumor cells. Cancer Res. 67, 9771–9780 (2007).
Carter, A. M. Evolution of placental function in mammals: the molecular basis of gas and nutrient transfer, hormone secretion, and immune responses. Physiol. Rev. 92, 1543–1576 (2012).
Kirkun, G. et al. A novel immortalized human endometrial stroma cell line with normal progestational response. Endocrinology 145, 2291–2296 (2004).
Haeger, J. D., Hambruch, N., Dilly, M., Froehlich, R. & Pfarrer, C. Formation of bovine placental trophoblast spheroids. Cells Tissues Organs 193, 274–284 (2011).
Hambruch, N., Haeger, J. D., Dilly, M. & Pfarrer, C. EGF stimulates proliferation in the bovine placental trophoblast cell line F3 via Ras and MAPK. Placenta 31, 67–74 (2010).
Wagner, G. P., Kin, K. & Lynch, V. J. Measurement of mRNA abundance using RNA-seq data: RPKM measure is inconsistent among samples. Theory Biosci. 131, 281–285 (2012).
The Gene Ontology Consortium The gene ontology resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330–D338 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Paradis, E., Claude, J. & Strimmer, K. APE: Analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004).
Paradis, E. Analysis of Phylogenetics and Evolution with R 2nd edn (Springer, 2012).
Acknowledgements
This work was funded by National Cancer Institute Center grant no. U54-CA209992.
Author information
Authors and Affiliations
Contributions
G.P.W. and A.L. conceptualized this project. K., J.D.M. and C.L. were responsible for data curation. K., C.L. and G.P.W. conducted the formal analysis. A.L. and G.P.W. acquired the funding. K., E.E., J.D.M. and A.H. undertook the investigation. K. and H.N.K. were responsible for the methodology. A.L. and G.P.W. were responsible for project admininstration. J.-D.H., C.P., T.H., T.O., T.S., M.P. and D.F.A. obtained resources. C.L. worked on software. G.P.W. and A.L. supervised the project. K., E.M.E. and G.P.W. validated the results. K. and C.L. worked on visualization. K., G.P.W. and A.L. wrote the first draft and all other authors were involved in reviewing and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Nanotextured stromal invasion assay quantitatively and sensitively measures collective intrusion of cells into stroma.
(A) Time stamped images at 0 and 18 hours showing extent of invasion by DiI-labelled 1205Lu cells (red) into BJ5ta stromal fibroblast monolayer (unlabelled) on flat and nanotextured substrata; Quantification shown in Fig. 1d. (B-C) Sensitive measurement of differences in invasion by nanotextured stromal invasion platform; Time stamped images at 0, 6, 12 and 18 hours showing extent of invasion by DiI-labelled non-malignant WM35 (B), and malignant 1205Lu (C) cells into BJ5ta monolayer shown in both flat, and nanotextured cell patterned assay; Quantification shown in Fig. 1e.
Extended Data Fig. 2 Human and bovine trophoblasts exhibit different invasive potential into endometrial stroma.
Time stamped images showing extent of invasion of human choriocarcinoma-derived trophoblasts, J3 (red) (A), and bovine trophoblasts, F3 (red) (B) into their respective endometrial stromal fibroblast monolayers at 0, and 48 hours.
Extended Data Fig. 3 Human and bovine ESFs respond differently to co-culture with trophoblasts.
(A) Experimental plan to isolate endometrial cells after co-culture with respective trophoblast; (B) Volcano plot showing fold change in genes between bovine versus human endometrial stromal fibroblasts, along with their significance depicted in P value. (C) P-value distribution of t-tests comparing human ESFs with and without co-culture with HTR8 trophoblast cells. (D) P-value distribution of t-tests comparing bovine ESFs with and without co-culture with F3 trophoblast cells. In both C and D, note that thin right hand tail of the distribution, which indicates that a large number of genes are differentially expressed in response to the presence of the corresponding trophoblast cells. (E-F) Scatter plots showing relative TPM values of genes between (E) hESF and hESF co-cultured with HTR8, and (F) bESF and bESF co-cultured with F3; Red dots refer to individual gene transcripts abundance significantly different in between the compared conditions.
Extended Data Fig. 4 Human and bovine endometrial stromal fibroblasts do not appear to differentially respond to their respective trophoblasts by regulating cell-cell adhesion.
Gene ontology analysis of individual genes belonging to GO_Adherens (A), GO_Endothelial Barrier (B), GO_Fibroblast Migration (C), and Kegg ontology for Wnt Signalling (D) for hESF and bESF shown with their relative TPM values; Also are shown the coefficient of determinant, R2 for the linear regression between TPM values of hESF and bESF in a given gene-sets, along with the standard deviation of the residuals, Sy.x. (E) Ingenuity Pathway Analysis of selected signalling pathways differentially activated in hESF and bESF; Shown are –log(p-values) of genes in the respective pathways differentially activated, while the colour depicts the z-score.
Extended Data Fig. 5 siRNA-induced gene knockdown reduces transcript levels in hESFs and BJ5ta cells.
qRT–PCR results showing the percentage knockdown of transcript levels in hESF (A) and BJ5ta (B), calculated using ΔΔCt method, compared using GAPDH as housekeeping gene with scrambled siRNA as control. For both A, and B, n = 4 biological replicates.
Supplementary information
Supplementary Video 1
bESF versus F3-mCherry: imaging of mCherry labelled F3 bovine trophoblast cells invading a monolayer of unlabelled bovine endometrial stromal cells. The total imaging duration is 44 h.
Supplementary Video 2
bESF versus F3-Phase: phase-contrast-based imaging of mCherry labelled F3 bovine trophoblast cells invading a monolayer of unlabled bovine endometrial stromal cells. The total imaging duration is 44 h.
Supplementary Video 3
hESF versus J3-mCherry: imaging of mCherry labelled J3 human choriocarcinoma cells invading a monolayer of human endometrial stromal cells. The total imaging duration is 44 h.
Supplementary Video 4
hESF versus J3-Phase: phase-contrast-based imaging of mCherry labelled J3 human choriocarcinoma cells invading a monolayer of human endometrial stromal cells. The total imaging duration is 44 h.
Supplementary Video 5
hESF versus BeWo: imaging of cytotracker green labelled HTR8 invading into a monolayer of unlabelled hESFs. The total imaging duration is 24 h.
Supplementary Video 6
hESF versus HTR8: imaging of cytotracker green labelled BeWo invading into a monolayer of unlabelled hESFs. The total imaging duration is 24 h.
Supplementary Video 7
hESF versus F3: imaging of cytotracker green labelled F3 invading into a monolayer of unlabelled hESFs. The total imaging duration is 24 h.
Supplementary Video 8
bESF versus BeWo: imaging of cytotracker green labelled HTR8 invading into a monolayer of unlabelled bESFs. The total imaging duration is 24 h.
Supplementary Video 9
bESF versus HTR8: imaging of cytotracker green labelled BeWo invading into a monolayer of unlabelled bESFs. The total imaging duration is 24 h.
Supplementary Video 10
bESF versus F3: imaging of cytotracker green labelled F3 invading into a monolayer of unlabelled bESFs. The total imaging duration is 24 h.
Rights and permissions
About this article
Cite this article
Kshitiz, Afzal, J., Maziarz, J.D. et al. Evolution of placental invasion and cancer metastasis are causally linked. Nat Ecol Evol 3, 1743–1753 (2019). https://doi.org/10.1038/s41559-019-1046-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-1046-4
This article is cited by
-
A common allele increases endometrial Wnt4 expression, with antagonistic implications for pregnancy, reproductive cancers, and endometriosis
Nature Communications (2024)
-
Lactate in breast cancer cells is associated with evasion of hypoxia-induced cell cycle arrest and adverse patient outcome
Human Cell (2024)
-
Deciphering the Epigenetic Landscape: Placental Development and Its Role in Pregnancy Outcomes
Stem Cell Reviews and Reports (2024)
-
A spatially resolved timeline of the human maternal–fetal interface
Nature (2023)
-
Prioritization of potential causative genes for schizophrenia in placenta
Nature Communications (2023)