Abstract
Microbial life has adapted to various individual extreme conditions; yet, organisms simultaneously adapted to very low pH, high salt and high temperature are unknown. We combined environmental 16S/18S ribosomal RNA gene metabarcoding, cultural approaches, fluorescence-activated cell sorting, scanning electron microscopy and chemical analyses to study samples along such unique polyextreme gradients in the Dallol–Danakil area in Ethiopia. We identified two physicochemical barriers to life in the presence of surface liquid water defined by (1) high chaotropicity–low water activity in Mg2+/Ca2+-dominated brines and (2) hyperacidity–salt combinations (pH ~0/NaCl-dominated salt saturation). When detected, life was dominated by highly diverse ultrasmall archaea that were widely distributed across phyla with and without previously known halophilic members. We hypothesize that a high cytoplasmic K+-level was an original archaeal adaptation to hyperthermophily, subsequently exapted during several transitions to extreme halophily. We detect active silica encrustment/fossilization of cells but also abiotic biomorphs of varied chemistry. Our work helps circumscribing habitability and calls for cautionary interpretations of morphological biosignatures on Earth and beyond.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sanger sequences have been deposited in GenBank (National Center for Biotechnology Information) with accession numbers MK894601–MK894820 and Illumina sequences in GenBank Short Read Archive with BioProject number PRJNA541281.
References
Harrison, J. P., Gheeraert, N., Tsigelnitskiy, D. & Cockell, C. S. The limits for life under multiple extremes. Trends Microbiol. 21, 204–212 (2013).
Merino, N. et al. Living at the extremes: extremophiles and the limits of life in a planetary context. Front. Microbiol. 10, 1785 (2019).
Johnson, S. S., Chevrette, M. G., Ehlmann, B. L. & Benison, K. C. Insights from the metagenome of an acid salt lake: the role of biology in an extreme depositional environment. PLoS ONE 10, e0122869 (2015).
Zaikova, E., Benison, K. C., Mormile, M. R. & Johnson, S. S. Microbial communities and their predicted metabolic functions in a desiccating acid salt lake. Extremophiles 22, 367–379 (2018).
Futterer, O. et al. Genome sequence of Picrophilus torridus and its implications for life around pH 0. Proc. Natl Acad. Sci. USA 101, 9091–9096 (2004).
Varet, J. in Geology of Afar (East Africa). Regional Geology Reviews (eds.Oberhänsli, R. et al.) Ch. 7 (Springer, 2018).
Franzson, H., Helgadóttir, H. M. & Óskarsson, F. Surface exploration and first conceptual model of the Dallol geothermal area, northern Afar, Ethiopia. In Proc. World Geothermal Congress (2015).
Darrah, T. H. et al. Gas chemistry of the Dallol region of the Danakil Depression in the Afar region of the northern-most East African Rift. Chem. Geol. 339, 16–29 (2013).
Holwerda, J. G. & Hutchinson, R. W. Potash-bearing evaporites in the Danakil area, Ethiopia. Econ. Geol. 63, 124–150 (1968).
Warren, J. K. Danakhil Potash, Ethiopia: Beds of Kainite/Carnallite, Part 2 of 4 (SaltWork Consultants, 2015).
Cavalazzi, B. et al. The Dallol geothermal area, northern Afar (Ethiopia): an exceptional planetary field analog on Earth. Astrobiology 19, 553–578 (2019).
Hallsworth, J. E. et al. Limits of life in MgCl2-containing environments: chaotropicity defines the window. Environ. Microbiol. 9, 801–813 (2007).
Stevenson, A. et al. Is there a common water-activity limit for the three domains of life? ISME J. 9, 1333–1351 (2015).
McKay, C. P. Requirements and limits for life in the context of exoplanets. Proc. Natl Acad. Sci. USA 111, 12628–12633 (2014).
Moissl-Eichinger, C., Cockell, C. & Rettberg, P. Venturing into new realms? Microorganisms in space. FEMS Microbiol. Rev. 40, 722–737 (2016).
Pérez, E. & Chebude, Y. Chemical analysis of Gaet’Ale, a hypersaline pond in Danakil Depression (Ethiopia): new record for the most saline water body on Earth. Aquat. Geochem 23, 109–117 (2017).
Kotopoulou, E. et al. A polyextreme hydrothermal system controlled by iron: the case of Dallol at the Afar triangle. ACS Earth Space Chem. 3, 90–99 (2019).
Warren, J. K. Danakhil Potash, Ethiopia: Is the Present Geology the Key? Part 1 of 4 (SaltWork Consultants, 2015).
Tosca, N. J., Knoll, A. H. & McLennan, S. M. Water activity and the challenge for life on early Mars. Science 320, 1204–1207 (2008).
Stevenson, A. et al. Aspergillus penicillioides differentiation and cell division at 0.585 water activity. Environ. Microbiol. 19, 687–697 (2017).
Sheik, C. S. et al. Identification and removal of contaminant sequences from ribosomal gene databases: lessons from the census of deep life. Front. Microbiol. 9, 840 (2018).
Weyrich, L. S. et al. Laboratory contamination over time during low-biomass sample analysis. Mol. Ecol. Resour. 19, 982–996 (2019).
Narasingarao, P. et al. De novo metagenomic assembly reveals abundant novel major lineage of archaea in hypersaline microbial communities. ISME J. 6, 81–93 (2012).
Sorokin, D. Y. et al. Discovery of extremely halophilic, methyl-reducing euryarchaea provides insights into the evolutionary origin of methanogenesis. Nat. Microbiol. 2, 17081 (2017).
Sorokin, D. Y. et al. Methanonatronarchaeum thermophilum gen. nov., sp. nov. and ‘Candidatus Methanohalarchaeum thermophilum’, extremely halo(natrono)philic methyl-reducing methanogens from hypersaline lakes comprising a new euryarchaeal class Methanonatronarchaeia classis nov. Int J. Syst. Evol. Microbiol. 68, 2199–2208 (2018).
Borin, S. et al. Sulfur cycling and methanogenesis primarily drive microbial colonization of the highly sulfidic Urania deep hypersaline basin. Proc. Natl Acad. Sci. USA 106, 9151–9156 (2009).
Mwirichia, R. et al. Metabolic traits of an uncultured archaeal lineage–MSBL1–from brine pools of the red sea. Sci. Rep. 6, 19181 (2016).
Castelle, C. J. et al. Biosynthetic capacity, metabolic variety and unusual biology in the CPR and DPANN radiations. Nat. Rev. Microbiol. 16, 629–645 (2018).
Castelle, C. J. & Banfield, J. F. Major new microbial groups expand diversity and alter our understanding of the tree of life. Cell 172, 1181–1197 (2018).
Dombrowski, N., Lee, J. H., Williams, T. A., Offre, P. & Spang, A. Genomic diversity, lifestyles and evolutionary origins of DPANN archaea. FEMS Microbiol. Lett. 366, fnz008 (2019).
Petitjean, C., Deschamps, P., Lopez-Garcia, P. & Moreira, D. Rooting the domain archaea by phylogenomic analysis supports the foundation of the new kingdom proteoarchaeota. Genome Biol. Evol. 7, 191–204 (2014).
Golyshina, O. V. et al. ‘ARMAN’ archaea depend on association with euryarchaeal host in culture and in situ. Nat. Commun. 8, 60 (2017).
Minegishi, H. et al. Acidophilic haloarchaeal strains are isolated from various solar salts. Saline Syst. 4, 16 (2008).
Naor, A. & Gophna, U. Cell fusion and hybrids in archaea: prospects for genome shuffling and accelerated strain development for biotechnology. Bioengineered 4, 126–129 (2013).
Garcia-Ruiz, J. M. et al. Self-assembled silica-carbonate structures and detection of ancient microfossils. Science 302, 1194–1197 (2003).
Garcia-Ruiz, J. M., Melero-Garcia, E. & Hyde, S. T. Morphogenesis of self-assembled nanocrystalline materials of barium carbonate and silica. Science 323, 362–365 (2009).
Slonczewski, J. L., Fujisawa, M., Dopson, M. & Krulwich, T. A. Cytoplasmic pH measurement and homeostasis in bacteria and Archaea. Adv. Micro. Physiol. 55, 1–79 (2009).
Buetti-Dinh, A., Dethlefsen, O., Friedman, R. & Dopson, M. Transcriptomic analysis reveals how a lack of potassium ions increases Sulfolobus acidocaldarius sensitivity to pH changes. Microbiology 162, 1422–1434 (2016).
Gunde-Cimerman, N., Plemenitas, A. & Oren, A. Strategies of adaptation of microorganisms of the three domains of life to high salt concentrations. FEMS Microbiol. Rev. 42, 353–375 (2018).
Lee, C. J. D. et al. NaCl-saturated brines are thermodynamically moderate, rather than extreme, microbial habitats. FEMS Microbiol. Rev. 42, 672–693 (2018).
Colman, D. R., Lindsay, M. R. & Boyd, E. S. Mixing of meteoric and geothermal fluids supports hyperdiverse chemosynthetic hydrothermal communities. Nat. Commun. 10, 681 (2019).
López-García, P., Zivanovic, Y., Deschamps, P. & Moreira, D. Bacterial gene import and mesophilic adaptation in Archaea. Nat. Rev. Microbiol. 13, 447–456 (2015).
Hensel, R. & König, H. Thermoadaptation of methanogenic bacteria by intracellular ion concentration. FEMS Microbiol. Lett. 49, 75–79 (1988).
Shima, S., Thauer, R. K. & Ermler, U. Hyperthermophilic and salt-dependent formyltransferase from Methanopyrus kandleri. Biochem. Soc. Trans. 32, 269–272 (2004).
Cray, J. A., Russell, J. T., Timson, D. J., Singhal, R. S. & Hallsworth, J. E. A universal measure of chaotropicity and kosmotropicity. Environ. Microbiol. 15, 287–296 (2013).
Cray, J. A. et al. Chaotropicity: a key factor in product tolerance of biofuel-producing microorganisms. Curr. Opin. Biotechnol. 33, 228–259 (2015).
Fox-Powell, M. G., Hallsworth, J. E., Cousins, C. R. & Cockell, C. S. Ionic strength is a barrier to the habitability of Mars. Astrobiology 16, 427–442 (2016).
Stevenson, A. et al. Multiplication of microbes below 0.690 water activity: implications for terrestrial and extraterrestrial life. Environ. Microbiol. 17, 257–277 (2015).
R Development Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2017).
Lê, S., Josse, J. & Husson, F. FactoMineR: an R package for multivariate analysis. J. Stat. Softw. 25, 1–18 (2008).
Kassambara, A. & Mundt, F. factoextra: extract and visualize the results of multivariate data analyses. https://CRAN.R-project.org/package=factoextra (2017).
Magoc, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahe, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Guillou, L. et al. The protist ribosomal reference database (PR2): a catalog of unicellular eukaryote small sub-unit rRNA sequences with curated taxonomy. Nucleic Acids Res. 41, D597–D604 (2013).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Asnicar, F., Weingart, G., Tickle, T. L., Huttenhower, C. & Segata, N. Compact graphical representation of phylogenetic data and metadata with GraPhlAn. PeerJ 3, e1029 (2015).
Wang, H. C., Xia, X. & Hickey, D. Thermal adaptation of the small subunit ribosomal RNA gene: a comparative study. J. Mol. Evol. 63, 120–126 (2006).
Rodriguez-Valera, F., Ruiz-Berraquero, F. & Ramos-Cormenzana, A. Behaviour of mixed populations of halophilic bacteria in continuous cultures. Can. J. Microbiol. 26, 1259–1263 (1980).
Acknowledgements
We are grateful to O. Grunewald for co-organizing the Dallol expeditions, documenting field research and providing drone images, and also to Jean-Marie Hullot (in memoriam), Françoise Brenckmann and the Fondation Iris for funding the first field trip. We thank L. Cantamessa for the in situ logistics and discussions about local history. We acknowledge M. Tafari (Mekelle University), A. A. Aliyu and the Afar authorities for local assistance, as well as the Ethiopian army and the Afar police for providing security. We thank J. Barthélémy, E. Kotopoulou and J. Garcia-Ruiz for help and discussions during field trips. We thank H. Timpano and the UNICELL platform for cell sorting; A. Gutiérrez-Preciado for bioinformatic assistance; A. Kish and C. Faveau for allowing us to measure water activity of selected samples at the Muséum National d’Histoire Naturelle; E. Viollier for discussion on chemical analyses; C. Gille for help with cultures; G. Billo for script help to treat SEM pictures; and J. T. Díaz and P. T. Sanz for advice on statistical analyses. This research was funded by the French CNRS (National Center for Scientific Research) basic annual funding, the CNRS programme TELLUS INTERRVIE and the European Research Council (ERC) under the European Union’s Seventh Framework Programme (ERC grant no. 322669 to P.L.-G.). We thank the European COST Action TD1308 Origins for funding a short stay of A.I.L.-A. in Orsay. J.B. was financed by the French Ministry of National Education, Research and Technology.
Author information
Authors and Affiliations
Contributions
P.L.-G. and D.M. designed and supervised the research. P.L.-G. organized the scientific expeditions. J.B., P.L.-G., D.M., L.J. and J.M.L.-G. collected samples and took measurements in situ. J.B., P.L.-G. and P.B. carried out molecular biology analyses. J.B., A.I.L.-A. and D.M. performed culture, chemistry analyses and water–salt related measurements. A.I.L.-A. and J.B. performed statistical analyses. J.B., G.R. and D.M. analysed metabarcoding data. K.B. performed SEM and EDX analyses. J.M.L.-G. mapped geothermal activity and georeferenced all samples. L.J. and J.B. performed FACS-derived analyses. P.L.-G. and J.B. wrote the manuscript. All authors read and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
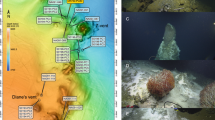
Extended Data Fig. 1 Aerial view of the main sampling sites in the Dallol area.
a, Dallol dome summit showing the acidic green-yellow-brown coloured hydrothermal ponds and active degassing areas during our 2017 sampling trip; the orange-shaded area shows the active hydrothermal zone in January 2016. b, Dallol West salt canyons and Black Mountain area. c, Black Lake. d, Yellow Lake and surroundings. Names of samples and sampling sites are indicated. The size of circles is proportional to the water volume collected or filtered for subsequent analyses. Aerial photographs were taken from a drone by O. Grunewald, except b, which is a Google Earth aerial image (09/03/2016) obtained by the Sentinel satellite (ESA Copernicus program) provided by Image © 2019 CNES/Airbus.
Extended Data Fig. 2 Views of different sampling sites in the Dallol dome and surroundings in the Danakil Depression.
a, DAL4 sampling site ponds; b, DAL5 pond and active degassing area; c, active hydrothermal springs in DAL9 ponds; d, in situ cell‐trap filtration at the 7DA7 sampling area; e, 7DA9 sampling site; f,7DA10 ponds showing increasingly darker and brownish colours along the oxidation gradient; g, water samples from the different 7DA10 ponds; h, DAL8 mineral precipitates; i,’proto-soil’-like salt crust (7YL-S1) near the Yellow Lake; j, Yellow Lake showing active degassing; k, YL3, salt-mud volcano in the Yellow Lake area; l, ‘Little Dallol’ hydrothermal very active area in 2016 on the way to the Black Mountain (in the distance; inlet, chimney emitting hydrocarbon‐rich fluids at 110 °C); m, Black Lake; n, PSBL2 (Black Lake area ponds); o, wet salt plain, influenced by hydrothermal activity, corresponding to PS3 sample area; p, the cave in the salt canyons where Gt, 7Gt and 8Gt samples were collected; q, salt canyons; r, Assale (Karum) lake. Sample names starting by 7 indicate collection in 2017. Pictures from all other samples/sampling sites were taken during the 2016 expedition.
Extended Data Fig. 3 List and description of samples from the Dallol area analysed in this study and type of analyses performed.
DO, dissolved oxygen; ORP, oxido-reduction potential; SEM–/EDXS, scanning electron microscopy/energy-dispersive x-ray spectrometry; FACS, fluorescence-activated cell sorting analysis; n.a., not applicable; n.d. not determined. Refractometry-derived salinity refers to the percentage (w/v) of local salt composition (see Supplementary Tables 1 and 3 for elementary and ionic analyses) measured in situ. Salinity was also directly measured by weighting the total solids (dry weight experimentally measured in triplicates; SD, standard deviation).
Extended Data Fig. 4 Principal Component Analyses (PCA) of Dallol area sampling sites as a function of physicochemical parameters.
PCA of 29 samples according to their chemical composition; only relatively abundant elements (see Supplementary Table 1) are included in the analysis. A summary of this analysis is shown in Fig. 2f. b, PCA including the same variables as Fig. 2f but additionally including dissolved oxygen (DO). Measured parameters on site can be found in Extended Data Fig. 3. Coloured zones in PCA analyses correspond to the three major chemical zones identified in this study.
Extended Data Fig. 5 Chaotropicity, ionic strength and water activity for a selection of samples of the Dallol area.
Chaotropicity was measured experimentally (see Methods) and also calculated, together with ionic strength values were from dominant Na, K, Mg, Ca, Fe chemistry data; water activity values were measured using a probe (see Methods). Known limits for life for each parameter are listed at the top of the table. Samples beyond that threshold for one or more of those parameters are shaded in grey.
Extended Data Fig. 6 Sequence data and diversity measurements.
*Contaminant sequences included sequences identified in negative controls and/or high similarity to human-associated bacteria; s.e., standard error. Eventual mitochondrial and chloroplast 16S rRNA gene sequences were also removed at this step.
Extended Data Fig. 7 Phylogenetic tree of bacterial 16S rRNA gene sequences showing the phylogenetic placement of OTUs identified in the different Dallol area samples.
Sequences derived from metabarcoding studies are represented by blue lines (Illumina sequences); those derived from cloning and Sanger sequencing of environmental samples, cultures and FACS-sorted cells are labelled with a red dot. Reference sequences are in black. Concentric circles around the tree indicate the presence/absence of the corresponding OTUs in different groups of samples (groups shown in Fig. 3a). Only sequences not deemed contaminant (see Supplementary Table 5) were included in the tree. The full tree is provided as Supplementary Data 1.
Extended Data Fig. 8 Eukaryotic presence, diversity and relative abundance in Dallol area samples.
Histogram showing the phylogenetic affiliation and abundance of 18S rRNA gene amplicon reads of eukaryotes (upper panel) obtained with universal eukaryotic primers and the associated OTU diversity (lower panel). Only a few samples yielded amplicons; negative PCR controls were always negative. Sequences corresponding to macroscopic plants and fungi (probably derived from pollen or spores) were considered contaminant (light grey). The phylogenetic affiliation of dominant eukaryotic groups is colour-coded.
Extended Data Fig. 9 Multiparametric fluorescence analyses and fluorescence-activated cell sorting (FACS) analyses of representative Dallol area samples.
a, effect of DNA fluorescent dyes on background fluorescence emission; natural (sterile medium-only) and DNA dye-induced fluorescence in the sterile hypersaline SALT-YE medium used to dilute/sort Dallol samples. Fluorescence is plotted against the size of the analysed particles (forward scatter); events concentration is colour-coded, red being high concentration and blue, low concentration. DRAQ5 and SYTO13 introduced less background and were chosen for FACS of natural samples. The approximate background threshold (ca. 102) is indicated by a broken grey line. b, multiparametric fluorescence analyses of different Dallol samples before (left panels) and after (right panels) adding fluorescent DNA dyes. Events (particles) above background (red squares) were FACS-sorted and filtered on 0.1 µm pore-size filters prior to SEM observations. c, SEM photographs showing examples of sorted particles. Cells are observed in samples PS, Gt and 7Gt; halite crystals in 7DA7 and amorphous mineral particles in 7DA9 and 7YL. Arrows indicate ultrasmall cells. The scale bar is 1 µm.
Extended Data Fig. 10 Mineral phases observed by SEM-EDX in precipitates of typical abiotic morphology and ‘biomorphs’.
Biomorphs correspond to rounded-shaped crystalline morphs resembling cell structures (cocci, rods) and compatible with cellular sizes. Observed dominant phases are highlighted in bold.
Supplementary Information
Supplementary Information
Supplementary Figs. 1 and 2 and Tables 1–4.
41559_2019_1005_MOESM3_ESM.xlsx
Supplementary Table 5 Identification, phylogenetic affinity and relative abundance of prokaryotic OTUs. a, OTUs identified in samples that yielded amplicons in direct (PS3, PS, Gt, 7 Gt, 7 Gt-pp, Ass, Ass-PJ) PCR amplifications. b, OTUs identified in samples that yielded amplicons in seminested PCR amplification reactions. c, List of contaminant OTUs identified in ‘negative’ controls of nested PCR reactions. d, List of OTUs removed as potential contaminants owing to their similarity to typical dust/soil bacterial contaminants or human-associated biota. id, identifier; bh, best hit; db, database; cmr, cleaned merged reads; cult, cultivated organism; env, environmental organism; seq, sequence; pciden, percentage of identity; pcqcov; percentage of coverage with query sequence; mism, mismatch; aln, alignment; len, length. Supplementary Table 5 sections a to d are provided in independent sheets for readability.
41559_2019_1005_MOESM4_ESM.xlsx
Supplementary Table 6 Identification, phylogenetic affinity and relative abundance of eukaryotic OTU from Dallol area samples. a, Eukaryotic OTU corresponding to protists thriving in Danakil samples; only a small subset of samples yielded 18S rRNA gene amplicons. b, Potential contaminant sequences (likely dispersal of pollen/spores) (see sheet 6b). id, identifier; bh, best hit; db, database; cmr, cleaned merged reads; cult, cultivated organism; env, environmental organism; seq, sequence; pciden, percentage of identity; pcqcov; percentage of coverage with query sequence; mism, mismatch; aln, alignment; len, length.
Supplementary Dataset 1
Full tree of archaeal 16S rRNA gene fragments in Newick format.
Supplementary Dataset 2
Full tree of bacterial 16S rRNA gene fragments in Newick format.
Rights and permissions
About this article
Cite this article
Belilla, J., Moreira, D., Jardillier, L. et al. Hyperdiverse archaea near life limits at the polyextreme geothermal Dallol area. Nat Ecol Evol 3, 1552–1561 (2019). https://doi.org/10.1038/s41559-019-1005-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-1005-0
This article is cited by
-
Expanded phylogeny of extremely halophilic archaea shows multiple independent adaptations to hypersaline environments
Nature Microbiology (2024)
-
Extreme and Heterogeneous Conditions of the Desert Wetland Chott Ech Chergui (Algeria) Allow Isolating Halophilic, Alkalophilic and Thermophilic Bacteria
Wetlands (2023)
-
A metagenomic insight into the microbiomes of geothermal springs in the Subantarctic Kerguelen Islands
Scientific Reports (2022)