Abstract
It is largely unknown how living organisms—especially vertebrates—survive and thrive in the coldness, darkness and high pressures of the hadal zone. Here, we describe the unique morphology and genome of Pseudoliparis swirei—a recently described snailfish species living below a depth of 6,000 m in the Mariana Trench. Unlike closely related shallow sea species, P. swirei has transparent, unpigmented skin and scales, thin and incompletely ossified bones, an inflated stomach and a non-closed skull. Phylogenetic analyses show that P. swirei diverged from a close relative living near the sea surface about 20 million years ago and has abundant genetic diversity. Genomic analyses reveal that: (1) the bone Gla protein (bglap) gene has a frameshift mutation that may cause early termination of cartilage calcification; (2) cell membrane fluidity and transport protein activity in P. swirei may have been enhanced by changes in protein sequences and gene expansion; and (3) the stability of its proteins may have been increased by critical mutations in the trimethylamine N-oxide-synthesizing enzyme and hsp90 chaperone protein. Our results provide insights into the morphological, physiological and molecular evolution of hadal vertebrates.
Similar content being viewed by others
Main
The deepest areas of the ocean (that is, those between 6,000 and 11,000 m) are commonly referred to as the hadal zone, and represent ~1–2% of the global benthic area1. They are among the most hostile environments on Earth, due to their high hydrostatic pressure, darkness, limited food resources, low temperatures and hypoxia2. The most conspicuous environmental constraint in the hadal zone is hydrostatic pressure, which increases by 10 atm per 100 m of depth, reaching ~1,000 atm in the deepest ocean trenches. Nevertheless, life thrives in these poorly explored realms. The first major trench-sampling campaigns were conducted during the early 1950s, and recent technological advances have prompted a renewed wave of hadal exploration, resulting in the discovery of hundreds of deep-dwelling species, including microbes, protists, worms, Porifera, Mollusca, Echinodermata, Crustacea, Cnidaria and fishes2,3.
The most common hadal vertebrate species are liparid snailfishes, which have the widest depth range of any marine fish family, with habitats ranging from intertidal to depths exceeding 8,100 m4,5. Liparid species have been found in seven trenches, indicating that snailfishes are a notably successful hadal fish family, extending deeper and reaching higher densities than other fish6,7. In addition, recent studies have shown that snailfish are top predators in the hadal food web and dominate the hadal fish fauna6,8. However, very little is known about the genetic basis and evolutionary history of snailfishes’ adaptation to deep-sea life.
During a recent expedition in the Mariana Trench—the world’s deepest known ocean trench—a previously unknown snailfish was observed in situ at a depth of 7,415 m, and was identified as a new species, Pseudoliparis swirei9. During a subsequent expedition, we successfully observed and collected P. swirei individuals using a baited video lander, and were able to sequence their genome. Here, we present comparative morphological, genomic and transcriptomic analyses of P. swirei that provide insights into genetic changes associated with adaptation to the deep sea.
Results and discussion
Morphological characterization of Mariana hadal snailfish (MHS)
MHS specimens were caught at a depth of about seven kilometres at multiple locations in the Mariana Trench (Fig. 1a) using the deep-sea landers Tianya and Haijiao, operated from the RV Tan Suo Yi Hao (Fig. 1b–d, Supplementary Table 1 and Supplementary Note 1). The fish were observed moving swiftly on the sea bed, foraging accurately and quickly (Supplementary Video 1). The MHS has a similar body size and shape to the related tide pool-dwelling species Tanaka’s snailfish (Liparis tanakae), but its skin is so transparent that its muscles and internal organs are clearly visible through the skin and abdominal wall (Fig. 1d–g and Supplementary Figs. 1–7). It also exhibits many other morphological adaptations to the hadal environment, including enlarged stomach, liver and eggs, thinner muscles and an incompletely ossified skeleton (Supplementary Note 1). Our specimens were identified as a new species9, P. swirei, based on morphological observations and DNA barcoding (Supplementary Note 1). The stomach of this MHS specimen was filled with 98 crustacean individuals (Supplementary Fig. 8), most of which were Hirondellea gigas. The dominance of H. gigas is consistent with an earlier report10.
a, ArcGIS Online map (left; Map data, 2019 Esri) and topographic base map (right; plotted using Generic Mapping Tools software (see Methods) of the collection locations within the Mariana Trench. b, RV Tan Suo Yi Hao. c, The deep-sea lander Tianya. d,e, In situ observations of the MHS (P. swirei). f, Fresh MHS specimen (holotype; Liparidae sp. 2 MT-2016; female). g, A Tanaka’s snailfish caught off the coast of Matsue city, Shimane Prefecture, Japan. Credit: photo of Tanaka’s snailfish taken by Takuya Morihisa in 2003.
De novo assembly of the MHS and sea surface snailfish reference genomes
We sequenced one MHS individual using a combination of single-molecule real-time sequencing and paired-end sequencing (Supplementary Figs. 9–11, Supplementary Tables 2 and 3 and Supplementary Note 2). The assembly consisted of 6,094 scaffolds, with a scaffold N50 of 418 kilobases (kb) (total length = 684 megabases (Mb)) and a contig N50 of 338 kb (total length = 682 Mb) (Supplementary Table 4 and Supplementary Fig. 12). A BUSCO assessment of single-copy orthologous genes indicated that the genome’s completeness was 91.7%, which is comparable to that achieved for other teleosts (Supplementary Table 5). To further assess the quality of the assembly, 40,154 transcripts were generated by sequencing messenger RNA from 28 samples of 15 tissues (Supplementary Table 6). Over 89% of the transcripts aligned to the genome along at least 90% of their length, confirming the assembly’s completeness (Supplementary Fig. 13). Additionally, 80% of the transcripts in which over 90% of the sequence aligned with the genome were located on single scaffold, demonstrating the contiguity of the assembly (Supplementary Fig. 13). We annotated 25,262 protein-coding genes (Supplementary Table 7), of which 23,043 (91.2%) were supported by transcriptome data. For comparative analyses, we also performed a de novo assembly of the Tanaka’s snailfish genome (Supplementary Fig. 12 and Supplementary Tables 3–5 and 7–9).
The genome of the MHS is about 21.9% (150 Mb) larger than that of Tanaka’s snailfish. This may be primarily due to expansions of repetitive sequences in the MHS (Supplementary Table 8). Other properties of the MHS genome, including its GC content, codon usage, gene length and exon number (Supplementary Fig. 14 and Supplementary Table 7) are similar to those of the ocean surface snailfish, suggesting that they probably do not contribute greatly to hadal adaptation.
Demographic history
We constructed a high-confidence species tree (Fig. 2a and Supplementary Fig. 15) for nine teleosts, including the MHS, Tanaka’s snailfish, stickleback, flatfish, pacific Bluefin tuna, fugu, platyfish, cod and zebrafish, using the coalescent method. The divergence time between the MHS and Tanaka’s snailfish was estimated to be about 20.22 million years ago (Ma) (Fig. 2a and Supplementary Fig. 16)—over 10 Myr before the formation of the Mariana Trench (estimated to have occurred 8–10 Ma11,12). A more extensive sampling effort including populations living at intermediate depths will be required to clarify how snailfish lived and adapted during the formation of the trench.
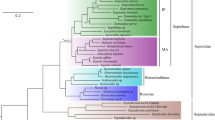
a, The phylogeny topology of the nine teleosts was reconstructed with coalescent methods based on both orthologues and syntenic block datasets. The branch lengths represent divergence times, while the grey rectangle at each node indicates the 95% confidence interval. b, Demographic history estimated by PSMC. The three blue lines represent the three collected MHS individuals, while the green line represents Tanaka’s snailfish. c, Comparison of mutation rates in the nine sequenced fish species based on 4D sites. d, Mutation rates of three species, estimated by syntenic alignment along the stickleback genome. The numbers around the outside represent the chromosome ID of the stickleback genome. The blue, green and orange dots indicate the mutation rates for each window in the MHS, Tanaka’s snailfish and sticklebacks, respectively. The green and orange dots almost overlap, while the blue dots are appreciably closer to the centre of the figure (corresponding to a lower mutation rate across the genome). μ indicates the mutation rate (×10–9 site–1 yr–1) of each window. e–g, Two-dimension kernel density distribution of Ka (e), Ks (f) and Ka/Ks (that is, ω; g). The MHS has much lower Ks values but similar Ka values, and so has a much greater Ka/Ks ratio.
Liparids are known to be the dominant fish in the hadal zone6 and they are the top predators8. Therefore, as a species of liparids, the MHS is likely to have a relatively large population size. Accordingly, its heterozygosity was ~0.36–0.51%, which is greater than that of Tanaka’s snailfish (0.26%) and comparable to other teleosts (Supplementary Fig. 17). Estimates of the dynamic effective population size (Ne) for both species indicated that the MHS had a larger population than the surface snailfish and underwent a significant population expansion around 50,000 years ago (Fig. 2b and Supplementary Fig. 18). This expansion was confirmed by multiple sequentially Markovian coalescent13 analyses (Supplementary Fig. 19), and might be related to some unknown geographic or environmental event. The divergence times among the three (sub)populations represented by the three individuals were estimated to be ~1.4 and ~2.9 Ma (Supplementary Fig. 16). These results suggest that the MHS population is quite large and has rich genetic diversity.
The MHS has a low rate of mutation across the genome, but a high rate of protein evolution
The branch length of the MHS was about one-third that for Tanaka’s snailfish in the maximum-likelihood tree (Supplementary Fig. 15). Among the nine species included in the tree, the MHS has the lowest mutation rate (Fig. 2c). This was not only true for the fourfold degenerate (4D) sites; the mutation rate of the MHS across the whole genome was also lower than for Tanaka’s snailfish and the stickleback (Fig. 2d). Previous studies have suggested that mutation rates are sensitive to many factors, including environmental energy14, metabolic rate15, life-history traits16 and, in particular, generation times17. Hadal species reportedly have comparatively low metabolic rates18, so the MHS may have a ‘slow life'. Coincidentally, we observed that the female MHS produced fewer but larger eggs than females of other snailfish species, suggesting that they may have a specialized reproduction strategy (for example, epimeletic behaviour and/or eggs that hatch as juveniles rather than larvae), which could further increase the generation time. It is thus plausible that the MHS has an extended generation time that contributes to its low mutation rate.
Despite the low nucleotide-level mutation rate of the MHS, its protein sequences appear to have evolved at a rate similar to other species. While the Ks value (the number of mutations per synonymous site) for the MHS was significantly lower than that for Tanaka’s snailfish, the two species had very similar Ka values (numbers of mutations per non-synonymous site), so the MHS had a significantly greater Ka/Ks ratio (that is, ω) (Fig. 2e–g). The high rate of protein evolution in the MHS was verified by comparing the ω distribution along the chromosomes of the stickleback genome (Supplementary Fig. 20). Overall, the MHS exhibited the largest ω value of the nine teleosts considered in this study (Supplementary Fig. 21). Its high proportion of mutations at non-synonymous sites could be due to factors such as positive selection or relaxation of selection19,20, since we have excluded the possibility of a small population size21. Additionally, the ratio of the heterozygosity of zerofold and fourfold degenerate sites in the MHS is lower than that in Tanaka’s snailfish, indicating a stronger positive selection effect in the MHS (Supplementary Fig. 22).
Molecular mechanisms underpinning the special phenotypes of the MHS
Vertebrates living on the surface of the Earth have closed skull spaces surrounded by hard bone, to protect the brain and maintain an appropriate intracranial pressure. However, closed skulls cannot maintain their structural integrity under the very high pressures of the hadal environment, necessitating an open system. Consequently, most multicellular hadal species are boneless creatures, such as Decapoda and Crustacea; only a few vertebrates, as well as species such as the MHS that exhibit adaptive structural features, can inhabit this zone2. Using micro-computed tomography, we found that the skull of the MHS is not completely closed (Fig. 3a,b and Supplementary Data 1 and 2), allowing internal and external pressure equalization. Moreover, most of the bones consist of cartilage rather than being ossified. Notably, we found that the osteocalcin gene—also known as the bone Gla protein (bglap) gene, which regulates tissue mineralization and skeletal development22,23,24—has a frameshift mutation that may cause premature termination of cartilage calcification in the MHS (Fig. 3c and Supplementary Fig. 23), which might cause its pseudogenization or severe modification. To evaluate the effects of disrupting bglap functionality in fish, the expression of bglap in the zebrafish (Danio rerio) was knocked down using two types of specific antisense morpholino (MO) oligonucleotides—one to prevent the proper splicing of exon 1 (bglap-e1i1-MO) and another to block the translation of bglap (bglap-ATG-MO) (Supplementary Fig. 24 and Supplementary Note 3). The amount of stained mineralized tissue in treated embryos at five days post-fertilization was markedly reduced compared with control-MO-injected fish (Fig. 3d–g, Supplementary Table 9 and Supplementary Fig. 24), suggesting that disrupting bglap expression indeed hinders skeletal development in fish, as has been observed in mammals22,23,24. Therefore, the premature termination of bglap in the MHS may be associated with this species’ unusual skull structure and reduced bone hardness.
a,b, Morphometric scans of the whole skeleton (a) and cranial bones (b) of the MHS holotype using micro-computed tomography. c, Gene structure (top), alignment of nucleotide and amino acid sequences (middle) and sequencing read depth (bottom; the numbers along the x-axis represent the position of the base at the scaffold) for the bglap gene. The premature termination (coloured red) of bglap is due to a single nucleotide insertion (coloured orange). The complete amino acid alignments of bglap are shown in Supplementary Fig. 23. d–f, Phenotypic differences in skeletal development between control-MO-injected zebrafish embryos (d) and embryos injected with bglap-e1i1-MO (e) and bglap-ATG-MO (f) at 5 days post-fertilization. In vivo visualization of the skeleton was achieved by adding a fluorescent dye (calcein) to the water housing the fish. Dyes that bind to calcified matrices can be used to label the entire skeleton. 5ba, fifth branchial arch; e, ethmoid plate; ec, ectopterygoid; m, Meckel’s cartilage; op, opercular bone; pq, palatoquadrate. Scale bars: 200 μm. g, Quantification of the relative fluorescence intensity (RFI) of the head skeleton bone mass. Columns heights and bars represent means ± s.e.m. (unpaired Student’s t-test, n = 10), ***P < 0.0001.
The environment 7,000 m under the sea is almost completely devoid of light. The MHS did not respond to the lights of our deep-sea lander, which is consistent with previous observations25. We therefore performed a comparative genomic analysis of changes in the crystallin and opsin genes of the hadal fish, revealing that it has lost several important photoreceptor genes (Supplementary Table 10 and Supplementary Figs. 25 and 26). Only five genes exhibited clear expression signals in the transcriptome data, three of which (rho, rgra and rgrb) were specifically expressed in the head (Supplementary Table 11). Rhodopsin, which is encoded by rho and regenerated by rgr26, is an extremely light-sensitive receptor protein found in rod cells that is responsible for low-light vision27. We hypothesize that the MHS may retain some photon-sensing ability or has gradually lost its visual ability—first losing colour perception, followed by the ability to perceive light in any form. Like other fish that lives in darkness, the MHS has lost its skin pigmentation and has become transparent28. We found that the most well-known pigmentation gene, mc1r, has been completely lost in this species (Supplementary Figs. 25 and 26).
Changes in cell membranes
The cell membrane is a lipid bilayer containing various proteins. High hydrostatic pressures reduce the fluidity of lipid bilayers and the reversibility of their phase transitions, ultimately leading to the denaturation and functional disorder of membrane-associated proteins29,30. Pressure also rigidifies membranes, impairing their transport functions31. Gene family analysis of the 9 teleosts included in our study revealed 310 significantly expanded gene families in the MHS (Supplementary Figs. 27 and 28 and Supplementary Table 12). The gene families exhibiting the most significant expansion were those associated with fatty acid metabolism (Fig. 4a and Supplementary Table 13). Phospholipids are major constituents of cellular membranes, and their fatty acid composition is regulated to maintain membrane order and fluidity. Biochemical studies have suggested that the membranes of deep-sea-adapted organisms contain a higher weight percentage of unsaturated fatty acids than the equivalent membranes of shallow-sea species32,33. It has been shown that docosahexaenoic acid (DHA) significantly alters many basic properties of membranes, including aryl chain order and ‘fluidity', elastic compressibility, permeability and protein activity at high pressure34. The last step of DHA biosynthesis is peroxisomal β-oxidation, and the protein acetyl-CoA acyltransferase encoded by acaa1 is the rate-limiting enzyme in this process. We found that the MHS genome has 15 copies of the acaa1 gene, while all other fully sequenced teleosts have only 5 copies (Fig. 4b and Supplementary Fig. 29). Another gene involved in DHA biosynthesis, fasn, also exhibited copy number increases in the MHS genome (Supplementary Fig. 30). These changes may increase the abundance of fluid membrane lipids, enabling survival in the world’s deepest ocean trench. Other significantly expanded categories include genes belonging to families with ion and solute transport-related functions, such as tfa and slc29a3 (Supplementary Fig. 30). This is consistent with a need to resist high-pressure-induced inhibition of fluid transport in hadal organisms35. The list of expanded gene families provides clues for future functional tests to reveal their correlation with the adaptation of the MHS to the extreme hydrostatic pressure.
a, Fatty acid synthesis and transport-related domains are expanded in the MHS compared with the other species tested (CO, cod; FF, flatfish; FU, Fugu; PB, Pacific bluefin tuna; PF, platyfish; SF, stickleback; TS, Tanaka’s snailfish; ZF, zebrafish). ‘Row z score' refers to the z score calculated for each value in the grid square within the row it belongs. This shows the significance of expansion. A complete list of expanded domains in the MHS is provided in Supplementary Table 13. b, Expanded genes (coloured blue) in the DHA biosynthesis pathway. In the chmical structure of DHA, the ω represents the methyl end, and the numbers indicate the positions of the carbon atoms. 22:6 indicates there are 22 carbons and 6 double bonds. c, Rapidly evolving Gene Ontology categories. These points represent mean Ka/Ks ratios for pairs of MHS and Tanaka’s snailfish genes by Gene Ontology category. Each point is coloured based on the significance (P value, one-sided binomial test) of the accumulation in non-synonymous mutations. A complete list of categories is provided in Supplementary Table 14.
The extensive deep-sea adaptations of the MHS are probably due to intense selective pressure acting on different gene families. Gene Ontology categories associated with significantly greater rates of protein evolution in the MHS compared with Tanaka’s snailfish include ‘ion transport’, ‘transmembrane transport’ and ‘calcium ion transport’ (Fig. 4c and Supplementary Table 14). The 86 MHS genes identified as positively selected genes (PSGs) (Supplementary Table 15) also exhibited functional enrichment with respect to ‘transmembrane transport’, ‘ATP binding’ and ‘ion transport’ (Supplementary Table 16). Among the PSGs, 79 have well-known functions, of which 18 are related to membrane transport systems, including 3 ATP-dependent transporters, 4 ion channel genes and 11 secondary transporter genes (Supplementary Table 15). Earlier studies showed that high pressure suppresses the activity of membrane transport genes, and that proteins such as Na+/K+-ATPases from deep-sea species are less pressure sensitive than those of sea-surface species30. The lineage-specific adaptive evolution of these genes in the MHS may thus indicate a role in maintaining transport activity and cell homeostasis36, helping the fish to thrive at high pressures. Analysis of the amino acid variations in these genes may yield insights into how transmembrane transport proteins adapt to high pressure.
Maintenance of protein activity
Hydrostatic pressure strongly inhibits protein function, affecting both folding and enzyme activity. Consequently, species living at great depths must maintain an intracellular milieu that preserves the intrinsic properties of proteins and confers pressure resistance2. Mechanisms based on physiological and structural adaptations have been proposed to explain the preservation of protein functionality in deep-sea organisms35,37.
The physiological adaptation mechanism involves accumulating small organic solutes such as trimethylamine N-oxide (TMAO) to preserve protein function at elevated hydrostatic pressures38. TMAO is a physiologically important protein stabilizer that can restore denatured proteins to their native structure39. Its abundance in teleosts increases with depth; deep-caught species have significantly higher TMAO levels in all tissues than shallow species40. Most teleost genomes contain five copies of the TMAO-generating enzyme flavin monooxygenase 3 (fmo3), four of which are tandem repeats (Fig. 5a and Supplementary Fig. 31). The first gene (fmo3a) of these four tandem-repeated copies was strongly expressed in the liver of the MHS (Supplementary Table 17). We found that the most strongly expressed copy of fmo3 of the MHS differs from species to species (Supplementary Table 17). It should be noted that this could be impacted by degraded transcriptome. Because these copies diverged long ago and the corresponding proteins’ structures differ appreciably, it is likely that different copies of fmo3 have different catalytic efficiencies. Interestingly, fmo3a was positively selected in the MHS. In addition, we predicted more putative promoters (five copies) upstream of this gene in the MHS than in Tanaka’s snailfish (one copy) or sticklebacks (two copies) (Supplementary Fig. 32). These changes in the gene’s protein-coding and regulatory sequences may help the MHS increase intracellular TMAO levels to enhance protein stability.
a, Model of the evolution of the gene encoding the TMAO-generating enzyme flavin monooxygenase 3. b, A maximum-likelihood tree and alignment of fish hsp90 sequences within five clades. Amino acids unique to the MHS are highlighted in red. c, Three-dimensional views of the hsp90 protein (hadal12091 is shown as an example; others are shown in Supplementary Fig. 36), highlighting MHS-specific amino acid substitutions. The homology model of the overall amino acid sequence is shown to the left; Ser202 atoms (coloured cyan) are shown as spheres. The expansion shows a model of the N-terminal domain, with the ligand-binding pocket coloured orange and shown in surface mode.
Structural adaptations of proteins to deep-sea conditions may include changes in amino acid substitution patterns and protein structure that counteract the effects of pressure on protein function41,42. To characterize these adaptations, we compared the MHS with other species with respect to the amino acid composition and substitution of all coding genes together (Supplementary Fig. 14 and Supplementary Tables 18 and 19) and each gene separately (Supplementary Fig. 33). No clear signal was identified in this analysis, suggesting that there is no global composition and substitution change that is present in all proteins. However, it has previously been reported that the evolutionary patterns of some proteins responded to hydrostatic pressure43,44. We further investigated whether any gene family of the MHS has convergent amino acid substitutions that are different from the ancestral genotypes at the homologous position (see Methods). The only gene family found to exhibit convergent amino acid changes in most of its family members with high confidence was hsp90; the same alanine-to-serine substitution occurred independently in four of five copies of the hsp90 protein of the MHS, at a site that is highly conserved in the corresponding proteins of humans, mice, chickens, chameleons and yeast (Fig. 5b and Supplementary Fig. 34). This convergent substitution was also found to be very rare under random conditions (Supplementary Fig. 35). Therefore, the recurrence and fixation of the substitution in such a conservative site suggest it is very likely to be beneficial for the adaptation of the MHS. Hsp90 is an evolutionarily conserved and highly abundant molecular chaperone that promotes the correct folding and activation of over 200 proteins, many of which are involved in essential cellular processes such as signal transduction, cell survival and responses to cellular stress45,46. We performed homology modelling using four MHS hsp90 isoforms, examining both the complete sequences and the amino (N)-terminal regions (representing the ATP-binding domains) separately46,47. The MHS hsp90 proteins feature an alanine-to-serine mutation in the relatively conserved motif FYSSX, which is predicted to exist as a short α-helix (Fig. 5c and Supplementary Fig. 36). In all cases, the mutated serine lies in close proximity to the ATP-binding pocket, and may contribute significantly to a local structural interaction affecting hsp90 activity (Fig. 5c). Further structural and chaperone function studies will shed light on this unique mutation’s structural and functional effects on the N-terminal regions of hsp90 proteins.
Conclusions
Advances in deep-diving and genome-sequencing technologies have allowed us to complete this study on the genetic basis of vertebrate adaptation to the extreme environment of deep-sea trenches. A Liparidae species discovered 6,000 m below the ocean surface was found to have adapted to life in the hadal zone over a period of only several million years. Although its mutation rate has declined, its rate of amino acid substitution was found to be high, allowing plasticity and adaptation. The species has undergone extensive internal and external adaptations to tolerate the immense pressures and other challenges of the deep-sea environment. Genomic analyses revealed molecular adaptations consistent with pressure-tolerant cartilage, loss of visual function and skin colour, enhanced cell membrane fluidity and transport protein activity, and increased protein stability. The numerous genetic changes identified in this study shed light on how vertebrate species can survive and thrive in the deep oceans.
Methods
Sample collection and identification
The MHS samples were collected from three sites in the Mariana Trench, at depths of 7,125, 7,034 and 6,879 m (Supplementary Note 1). On the basis of morphological observations, the specimens were identified as conspecific with P. swirei (Gerringer and Linley, 2017), also collected in the Mariana Trench, close to our collection sites9. One specimen—snailfish number 0 MT-2016 (named hadal01 in the main text)—was confirmed to be P. swirei by DNA barcoding analysis (Supplementary Note 1). The topographic base map in Fig. 1a was plotted using Generic Mapping Tools software48. The bathymetric data were integrated using high-resolution (~100 m) multibeam data collected by the cruises of the University of New Hampshire’s US Extended Continental Shelf Bathymetry Mapping Project in 201049 and Chinese TS09 in 2018. The ETOPO1 (ref. 50) bathymetric data were filled where high-resolution data do not exist in the ocean. Tanaka’s snailfish specimens were collected in the southern Yellow Sea in 2017 and identified as L. tanakae (Gilbert and Burke, 1912) on the basis of morphological observations (Supplementary Note 1).
Sequencing and assembly of MHS and Tanaka’s snailfish
For the MHS, a total of 55 gigabases of PacBio reads and 671 gigabases of Illumina reads were sequenced. The PacBio reads were used for initial assembly with FALCON pipeline, and the Illumina reads were used for extending, closing the gap and polishing the assembly (for details, see Supplementary Note 2.1). For Tanaka’s snailfish, a total of 1.15 terabases of Illumina reads were sequenced and the genome was assembled using Platanus version 1.24 (ref. 51). For details, see Supplementary Note 2.2.
Transcriptome sequencing and assembly
A total of 28 transcriptomes were generated from 15 tissues (abdominal skin, blood, bone, brain, brain fluid, cholecyst, gill, head, heart, liver, muscle, oesophagus, reproductive organ, spinal cord and stomach) from two MHS individuals collected from the second site. Total RNA was extracted from these individuals using TRIzol (Invitrogen) and subsequently purified using an RNeasy Mini Kit (Qiagen). Paired-end reads with insert sizes of 500 bp were generated using an Illumina HiSeq 2000 sequencing platform. The sequenced reads were filtered and trimmed by fastp52, then assembled using Binpacker53 with default parameters.
Genome annotation
Both homology-based and de novo predictions were used to identify repeat elements in the MHS and Tanaka’s snailfish genome sequences. For homology-based analysis, transposable elements were identified using RepeatMasker version 4.0.7 (ref. 54) and RepeatProteinMask version 1.36 with the Repbase transposable element library55. For de novo predictions, RepeatModeler version 1.0.11 (ref. 52) was used to construct a de novo transposable element library, which was then used to predict repeats with RepeatMasker. We also predicted tandem repeats using TRF version 4.0.4 (ref. 56).
We annotated the coding gene structure of the two genome sequences by integrating ab initio predictions, homology-based gene predictions and direct gene models produced by transcriptome assembly (only for the MHS). First, Augustus version 3.2.1 (ref. 57), GeneID version 1.4 (ref. 58), GlimmerHMM version 3.0.4 (ref. 59) and SNAP version 2013-11-29 (ref. 60) were used to generate ab initio predictions with internal gene models. Next, the protein sequences of seven species (cod, fugu, medaka, puffer, stickleback, zebrafish and human; ENSEMBL 89) were aligned to genome sequences with Exonerate. The MHS transcripts were assembled using both Binpacker version 1.0 (ref. 53) (de novo) and Hisat2 version 2.1.0 (ref. 61)/StringTie version 1.3.3b62 (reference-guided) with default parameters. We then integrated the two assemblies using Evidence Modeler (EVM) version 1.1.1 (ref. 63) with different weights for each. The integrated gene set was translated into amino acid sequences, which were used to search the InterPro database with InterProScan version 5.15 (ref. 64) to obtain Gene Ontology and PANTHER information for each gene, and the genes were further annotated using the KEGG databases65.
Phylogeny reconstruction
Protein sequences from nine species (the MHS and Tanaka’s snailfish (assembled in this study), stickleback, fugu, platyfish, cod and zebrafish (V89; downloaded from ENSEMBL), flatfish (GCF_001970005.1; downloaded from the National Centre for Biotechnology Information) and Pacific bluefin tuna (Ver.1; downloaded from http://nrifs.fra.affrc.go.jp) were clustered with OrthoMCL version 2.0.9 (ref. 66) using default parameters, and 3,915 one-to-one orthologues were identified. Five species from ENSEMBL were chosen with the aim of covering more teleost groups (one species for one order). We chose flatfish and Pacific bluefin tuna because of their closer relationship to MHS. The protein sequences of each orthologue were aligned with MAFFT version 7.310 (ref. 67) using default parameters, and alignments of the coding sequences were generated with pal2nal version 14 (ref. 68) using default parameters. We then generated five datasets using the first, second and third base in each codon, 4D sites and whole coding sequence alignments. The five datasets were used to construct maximum-likelihood trees, separately, with RAxML version 8.2.10 (ref. 69) using the following parameters: -f a -m GTRGAMMAI -x 271828 -N 100 -p 31415, under the GTR + I model, which was suggested by jmodeltest2 (ref. 70). The maximum-likelihood tree for each gene was also constructed (as above) and plotted using Densitree71, to reveal phylogeny heterogeneity at the gene level. Then, a species tree was built with these gene trees using MP-EST version 2.0 (ref. 72). We also performed whole-genome synteny alignment for these nine teleosts using Last version 894 (ref. 73) and Multiz version 11.2 (ref. 74) with default parameters to generate another dataset. The 12-Mb one-to-one synteny alignment was used to construct a maximum-likelihood tree, and the 13,051 synteny blocks with a length larger than 200 bp were used to constructed a species tree. The divergence time was estimated using MCMCtree version 4.5 (ref. 75), with the topology of the species tree, 4D site alignments and three soft-bound calibration time points (zebrafish–stickleback: ~205–252 Ma; cod–stickleback: ~141–170 Ma; and snailfish–stickleback: ~32–73 Ma)76 based on previous studies.
Demographic history and genetic diversity
We inferred demographic histories of the MHS and Tanaka’s snailfish by applying the pairwise sequentially Markovian coalescence model (PSMC version 0.6.5-r67)77 to the complete diploid genome sequences. Consensus sequences were obtained using SAMtools version 1.3.1 (ref. 78) using the parameters ‘mpileup -q 20 -Q 20', and divided into non-overlapping 100-bp bins. Bases of low sequencing depth (less than one-third of the average depth) or high depth (twice the average depth) were masked. The analysis was performed using the following parameters: -N25 -t15 -r5 -p “4 + 25*2 + 4 + 6”. The mutation rate per site per year was set at 1.93 × 10−9 for the MHS and 6.77 × 10−9 for Tanaka’s snailfish; these values were estimated by r8s version 1.81 (ref. 79) with the penalized likelihood method. As no information about the snailfish generation time is available, we tested generation times of six months, one year, two years and three years for both species. We also performed an analysis with MSMC version 2.0.0 (ref. 13; an extension of pairwise sequential Markovian coalescent analysis) with default parameters, to infer a more recent demographic history for the MHS. All segregating sites were phased and imputed using fastPHASE version 1.1 (ref. 80) with default parameters, and the four above-mentioned combinations of generation times and mutation rates were evaluated.
The Illumina-sequenced reads from three MHS and one Tanaka’s snailfish were aligned to the genome sequences of the MHS with BWA version 0.7.15-r1140 (ref. 81) using the parameters: mem -t 16. Duplicated reads were filtered with SAMtools ‘rmdup’. Reads around indels were realigned by GATK version 3.6 (ref. 82) using default parameters, and the genotype of each site in every individual was called by SAMtools using the parameters: -t DP -A -q 20 -Q 20. We then used the mappability module in GEM version 20130406-045632 (ref. 83) using the parameters ‘-l 150' to extract 407 Mb of regions that could be uniquely mapped. Conservatively, we excluded polymorphic sites that were not bi-allelic or for which QUAL < 30. Finally, we masked any site that lacked 2- to 100-fold depth of aligned read coverage. The 4D sites were extracted and the divergence time of the four MHS individuals was estimated by MCMCtree with the same calibration as above.
Whole-genome alignment and mutation rate across the genome
We chose five species (the MHS, Tanaka’s snailfish, stickleback, flatfish and Pacific bluefin tuna) for whole-genome synteny alignment. We did not include more species because their divergence times were too long ago. Using the stickleback genome sequence as a reference, we performed synteny alignment for these five species with Last version 894 (ref. 73) using the parameters ‘-m100 -P 4 -E0.05', generating a total of 121 Mb (of which 66 Mb was informative for all species) of one-to-one alignment sequences with Multiz version 11.2 (ref. 74) using the default parameters.
We applied a sliding window (100 kb) along the synteny alignment to estimate the mutation rate across the genome. For each window, only neutral regions were retained (repetitive sequences and regions located within genes, or 3 kb upstream/downstream of them, were removed) to estimate branch lengths with RAxML and a given topology. The branch lengths were then used to estimate mutation rates for each branch with r8s and the previously estimated divergence time in the root node of these five species.
Strength of natural selection
A total of 18,620 genes were extracted from the synteny alignments, together with gene annotations based on the corresponding stickleback genes. Any gene not annotated in all five species at a given position in the synteny alignment was excluded from further analysis. We then filtered the alignments with Gblocks version 0.91b84 using default parameters, and excluded those with less than 150 bp of informative sites in all species, ultimately retaining 12,370 genes. The ratio of non-synonymous to synonymous substitutions (Ka/Ks) in each branch was estimated using the free ratio model of codeml in the PAML version 4.9e75 software package using default parameters. To enable comparisons with more species, we calculated the Ka/Ks ratios of the 3,915 one-to-one orthologues with codeml. For this part, the genes with Ks values above 2 in any branch due to the possibility of false alignment or pseudogenes were filtered.
To assess the ratio of diversity in neutral and functional sites, which should theoretically reflect the strength of natural selection, we first calculated the ratio of heterozygosity at zerofold relative to fourfold sites in the three MHS and one Tanaka’s snailfish. We identified a total of 24.2 Mb zerofold and 6.1 Mb fourfold sites with gene annotations in the MHS, and estimated the heterozygosity of each individual at these sites. We then calculated the Ka/Ks substitution ratio (based on heterozygous single nucleotide polymorphisms) within the four individuals. The non-synonymous and synonymous mutations were identified using SnpEff version 4.1 (ref. 85).
Putative gene loss
We identified genes putatively lost in the MHS using a four-step method. (1) The opsin- and pigment-related protein sequences (Supplementary Table 10) were downloaded from UniProt and searched against the MHS, Tanaka’s snailfish and stickleback protein sets with blastp86. (2) Genes absent in the MHS but present in the other two species were searched against the genome sequences and assembled transcripts with tblastn86. (3) The synteny alignment between the MHS and Tanaka’s snailfish was plotted to determine whether such genes had been partially or fully lost, or simply mis-annotated. Only fully lost genes were retained for further analysis. (4) The reads from the three MHS individuals and Tanaka’s snailfish were further mapped to the stickleback genome sequence (ENSEMBL V89) using BWA. For each putative gene, we plotted the read depth of the four individuals along the corresponding coding sequences in the stickleback genome. Genes were identified as lost in the MHS only if the reads from all three MHS individuals could not be mapped to the stickleback genome but the corresponding read from Tanaka’s snailfish could be mapped.
Bglap gene knockdown experiment and calcein staining
Antisense morpholino oligomers (Gene Tools) were microinjected into fertilized one-cell-stage embryos according to standard protocols87. The sequences of the bglap translation-blocking and splice-blocking morpholino oligomers were 5′-GGACTGTCAGGCTCTTCATATTCG-3′ (bglap-ATG-MO) and 5′-CACATACATGCACACTGACCTG-3′ (bglap-e1i1-MO), respectively. The sequence for the standard control morpholino was 5′-CCTCTTACCTCAGTTACAATTTATA-3′. The amounts of the morpholino used for injection were as follows: control-MO and bglap-e1i1-MO: 4 ng per embryo; bglap-ATG-MO: 2 ng per embryo. Calcein staining of morpholino-injected embryos was performed using 0.2% calcein solution 5 d post-fertilization of the zebrafish. For details of this protocol, see Supplementary Note 3.
Estimation of gene expression in the MHS and other species
The sequenced transcriptome reads were aligned to the coding sequences using Bowtie 2 version 2.3.2 (ref. 88) with default parameters. After alignment, the count of mapped reads from each sample was derived and normalized to transcripts per million using custom scripts. Transcriptome data for zebrafish and sticklebacks were downloaded from the Sequence Read Archive database (Supplementary Table 17) and aligned to the corresponding non-redundant gene catalogue by keeping the longest open reading frame. However, it should be noted that decompression as the samples were brought to the surface may have reduced the accuracy of the gene expression measurements.
Gene family expansion/contraction
To evaluate gene family expansion and contraction in the MHS, we first used CAFE version 3.1 (ref. 89) with default parameters, which applies a maximum-likelihood framework, with results from the OrthoMCL pipeline90 with default parameters and estimated divergence times between species as input. A conditional P value was calculated for each gene family, and families with conditional P values lower than 0.05 were considered to have a significantly accelerated rate of expansion or contraction. Genes with >200 copies in 1 of the species were filtered out. We also annotated the protein sequences with Pfam91 using default parameters, and those with a z score above 1.96 and >5 members in the MHS were identified as expanded domains.
Identification of rapidly evolving Gene Ontology terms and PSGs
To identify rapidly evolving Gene Ontology terms in the MHS, which had a significantly higher Ka/Ks ratio than expected, we designed a new statistic that accounts for differences in Ka/Ks between two species (the MHS and Tanaka’s snailfish in this case) for a given Gene Ontology, as well as differences in Ka/Ks between that Gene Ontology and the genome background (for details, see Supplementary Note 4).
The 12,370 genes extracted from the synteny alignment described above were used to identify genes that have evolved under positive selection (PSGs) by applying the likelihood ratio test using the branch model implemented in the PAML package75. We first excluded genes with a Ks value above 2 in any branch due to the possibility of false alignment or pseudogenes. We then performed a likelihood ratio test comparing the two-ratio model (which calculates the Ka/Ks ratio for the lineage of interest and the background lineage) with the one-ratio model (which assumes a uniform Ka/Ks ratio across all branches) to determine whether the focal lineage is evolving significantly faster (false discovery rate-adjusted P < 0.05). We also required PSGs to have Ka/Ks > 1 in the focal lineage.
Amino acid preferences
Frequencies of amino acids in orthologues were calculated, and the significance of differences in these frequencies was tested by calculating z scores. We then tested the hypothesis that there may be significant differences in the frequencies of any two or three consecutive amino acids in MHS proteins relative to the mean frequency in orthologues from other species. In addition, the protein sequences in the ancestral node of the MHS and Tanaka’s snailfish were reconstructed with RAxML (version 8.2.10). We then counted the frequency of every type of amino acid replacement from the ancestor to the MHS or Tanaka’s snailfish, and subjected each replacement pattern in the two species to a two-sided binomial test using custom scripts.
Convergence within paralogues
We defined cases where the same amino acid replacement (that is, a replacement at the same position involving the same mutant amino acid) occurred independently in paralogous genes of a single species as instances of ‘convergence within paralogues', based on the approach adopted in common convergent analysis (which considers such events in orthologues from different species). Across the 9 studied species, 7,148 PANTHER gene families were identified with InterProScan, of which 3,058 are represented by at least 2 copies in the MHS. For each such gene family, the protein sequences were aligned using MAFFT with the default parameters. The phylogenetic tree and ancestral sequences were reconstructed with RAxML using the parameters ‘-m PROTGAMMAWAG'. Sites exhibiting the same amino acid substitution from the ancestral state in more than three gene copies within the MHS genome were identified as potential convergent sites. For each potential convergent site, we performed 100,000 Monte-Carlo simulations of protein sequence evolution with Seq-Gen (version 1.3.4)92, using the parameters ‘-n100000 -m WAG -wa -k1', based on the corresponding ancestral sequence and the protein’s phylogenetic tree to determine whether such convergence could plausibly have occurred by random chance.
Homology modelling of protein structures
To identify the presumable functional region formed by the amino acid sequence containing a serine mutation in four MHS hsp90 isoforms, we aligned their complete sequences with the high-resolution structures of yeast93 and human94 hsp90 isoforms using Clustal Omega version 1.2.4 (ref. 95). The results clearly indicate that each serine-substituted relevant fragment folds into an ATP-binding domain. We then probed the relative positions of the serine mutations in the three-dimensional structures by submitting the full-length and N-terminal region sequences of each isoform to the web-based server Phyre2 for homology modelling96. The ending residue in the N-terminal region was determined by sequence alignment to yeast heat-shock protein93. We chose the normal mode on the submitting page for all of the isoforms and downloaded the generated first-ranked model with the highest reported confidence and sequence coverage compared with the template thereafter. When we superimposed the generated pseudo-atomic models from the full-length and N-terminal sequences for each isoform using UCSF Chimera97, despite differences in several loop regions, the two models exhibited high similarity in the relatively rigid core structure, which consists of α-helices packed opposing an antiparallel β-sheet (comparisons for each isoform are shown in Supplementary Fig. 36); the serine is located in the fifth short α-helix. Further analysis of the N-terminal models using the protein cavity detection algorithm fpocket2 (ref. 98) indicated that the serine is in close proximity to a putative nucleotide-binding cavity in each isoform.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The sequence data have been deposited in the NCBI BioProject database with accession numbers PRJNA472845, PRJNA472846 (genome data) and PRJNA472245 (transcriptome data). The assemblies and annotation files have been deposited in GitHub (http://github.com/wk8910/hadal_snailfish).
Code availability
The custom scripts have deposited in GitHub (http://github.com/wk8910/hadal_snailfish).
References
Wolff, T. The hadal community, an introduction. Deep Sea Res. (1953) 6, 95–124 (1959).
Jamieson, A. J. The Hadal Zone: Life in the Deepest Oceans (Cambridge Univ. Press, 2015).
Wolff, T. The concept of the hadal or ultra-abyssal fauna. Deep Sea Res. Oceanogr. Abstr. 17, 983–1003 (1970).
Linley, T. D. et al. Fishes of the hadal zone including new species, in situ observations and depth records of Liparidae. Deep Sea Res. Pt I 114, 99–110 (2016).
Chernova, N. V. Family Liparidae Scopoli 1777, snailfishes. Calif. Acad. Sci. Annot. Checkl. Fishes 31, 1–72 (2004).
Fujii, T., Jamieson, A. J., Solan, M., Bagley, P. M. & Priede, I. G. A large aggregation of liparids at 7703 meters and a reappraisal of the abundance and diversity of hadal fish. Bioscience 60, 506–515 (2010).
Linley, T. D. et al. Bait attending fishes of the abyssal zone and hadal boundary: community structure, functional groups and species distribution in the Kermadec, New Hebrides and Mariana trenches. Deep Sea Res. Pt I 121, 38–53 (2017).
Blankenship, L. E. & Levin, L. A. Extreme food webs: foraging strategies and diets of scavenging amphipods from the ocean’s deepest 5 kilometers. Limnol. Oceanogr. 52, 1685–1697 (2007).
Gerringer, M. E., Linley, T. D., Jamieson, A. J., Goetze, E. & Drazen, J. C. Pseudoliparis swirei sp. nov.: a newly-discovered hadal snailfish (Scorpaeniformes: Liparidae) from the Mariana Trench. Zootaxa 4358, 161–177 (2017).
Lan, Y. et al. Molecular adaptation in the world’s deepest-living animal: insights from transcriptome sequencing of the hadal amphipod Hirondellea gigas. Mol. Ecol. 26, 3732–3743 (2017).
Oakley, A. J., Taylor, B., Moore, G. F. & Goodliffe, A. Sedimentary, volcanic, and tectonic processes of the central Mariana Arc: Mariana Trough back-arc basin formation and the West Mariana Ridge. Geochem. Geophys. Geosyst. 10, Q08X07 (2009).
Robert, J. S., Matthew, J. F. & Simon, L. K. An overview of the Izu-Bonin-Mariana Subduction Factory. In Inside the Subduction Factory 175–222 (Geophysical Monograph Series Volume 138, American Geophysical Union, 2003).
Schiffels, S. & Durbin, R. Inferring human population size and separation history from multiple genome sequences. Nat. Genet. 46, 919–925 (2014).
Davies, T. J., Savolainen, V., Chase, M. W., Moat, J. & Barraclough, T. G. Environmental energy and evolutionary rates in flowering plants. Proc. Biol. Sci. 271, 2195–2200 (2004).
Martin, A. P. & Palumbi, S. R. Body size, metabolic rate, generation time, and the molecular clock. Proc. Natl Acad. Sci. USA 90, 4087–4091 (1993).
Bromham, L., Rambaut, A. & Harvey, P. H. Determinants of rate variation in mammalian DNA sequence evolution. J. Mol. Evol. 43, 610–621 (1996).
Thomas, J. A., Welch, J. J., Lanfear, R. & Bromham, L. A generation time effect on the rate of molecular evolution in invertebrates. Mol. Biol. Evol. 27, 1173–1180 (2010).
Brown, A. et al. Metabolic rates are significantly lower in abyssal Holothuroidea than in shallow-water Holothuroidea. R. Soc. Open Sci. 5, 172162 (2018).
Comeron, J. M. & Kreitman, M. The correlation between synonymous and nonsynonymous substitutions in Drosophila: mutation, selection or relaxed constraints? Genetics 150, 767–775 (1998).
Subramanian, S. Significance of population size on the fixation of nonsynonymous mutations in genes under varying levels of selection pressure. Genetics 193, 995–1002 (2013).
Xue, Y. et al. Mountain gorilla genomes reveal the impact of long-term population decline and inbreeding. Science 348, 242–245 (2015).
Gavaia, P. J. et al. Osteocalcin and matrix Gla protein in zebrafish (Danio rerio) and Senegal sole (Solea senegalensis): comparative gene and protein expression during larval development through adulthood. Gene Expr. Patterns 6, 637–652 (2006).
Kavukcuoglu, N. B., Patterson-Buckendahl, P. & Mann, A. B. Effect of osteocalcin deficiency on the nanomechanics and chemistry of mouse bones. J. Mech. Behav. Biomed. Mater. 2, 348–354 (2009).
Li, J., Zhang, H., Yang, C., Li, Y. & Dai, Z. An overview of osteocalcin progress. J. Bone Miner. Metab. 34, 367–379 (2016).
Jamieson, A. J. et al. Liparid and macrourid fishes of the hadal zone: in situ observations of activity and feeding behaviour. Proc. Biol. Sci. 276, 1037–1045 (2009).
Chen, P. et al. A photic visual cycle of rhodopsin regeneration is dependent on Rgr. Nat. Genet. 28, 256–260 (2001).
Nathans, J. Rhodopsin: structure, function, and genetics. Biochemistry 31, 4923–4931 (1992).
McGaugh, S. E. et al. The cavefish genome reveals candidate genes for eye loss. Nat. Commun. 5, 5307 (2014).
Chong, P. L., Cossins, A. R. & Weber, G. A differential polarized phase fluorometric study of the effects of high hydrostatic pressure upon the fluidity of cellular membranes. Biochemistry 22, 409–415 (1983).
Kato, M., Hayashi, R., Tsuda, T. & Taniguchi, K. High pressure-induced changes of biological membrane. Study on the membrane-bound Na(+)/K(+)-ATPase as a model system. Eur. J. Biochem. 269, 110–118 (2002).
Casadei, M. A., Manas, P., Niven, G., Needs, E. & Mackey, B. M. Role of membrane fluidity in pressure resistance of Escherichia coli NCTC 8164. Appl. Environ. Microbiol. 68, 5965–5972 (2002).
Cossins, A. R. & MacDonald, A. G. Homeoviscous theory under pressure: II. The molecular order of membranes from deep-sea fish. Biochim. Biophys. Acta 776, 144–150 (1984).
Fang, J., Barcelona, M. J., Nogi, Y. & Kato, C. Biochemical implications and geochemical significance of novel phospholipids of the extremely barophilic bacteria from the Marianas Trench at 11,000 m. Deep Sea Res. Pt I 47, 1173–1182 (2000).
Yano, Y., Nakayama, A., Ishihara, K. & Saito, H. Adaptive changes in membrane lipids of barophilic bacteria in response to changes in growth pressure. Appl. Environ. Microbiol. 64, 479–485 (1998).
Simonato, F. et al. Piezophilic adaptation: a genomic point of view. J. Biotechnol. 126, 11–25 (2006).
Campanaro, S. et al. Laterally transferred elements and high pressure adaptation in Photobacterium profundum strains. BMC Genomics 6, 122 (2005).
Somero, G. N. Protein adaptations to temperature and pressure: complementary roles of adaptive changes in amino acid sequence and internal milieu. Comp. Biochem. Physiol. B 136, 577–591 (2003).
Yancey, P. H., Blake, W. R. & Conley, J. Unusual organic osmolytes in deep-sea animals: adaptations to hydrostatic pressure and other perturbants. Comp. Biochem. Physiol. A 133, 667–676 (2002).
Ma, J., Pazos, I. M. & Gai, F. Microscopic insights into the protein-stabilizing effect of trimethylamine N-oxide (TMAO). Proc. Natl Acad. Sci. USA 111, 8476–8481 (2014).
Yancey, P. H., Gerringer, M. E., Drazen, J. C., Rowden, A. A. & Jamieson, A. Marine fish may be biochemically constrained from inhabiting the deepest ocean depths. Proc. Natl Acad. Sci. USA 111, 4461–4465 (2014).
Somero, G. N. Adaptations to high hydrostatic pressure. Annu. Rev. Physiol. 54, 557–577 (1992).
Yafremava, L. S., Di Giulio, M. & Caetano-Anolles, G. Comparative analysis of barophily-related amino acid content in protein domains of Pyrococcus abyssi and Pyrococcus furiosus. Archaea 2013, 680436 (2013).
Yancey, P. H. Adaptations to hydrostatic pressure in protein structure and organic osmolytes in deep-sea animals. High Pressure Biosci. Biotechnol. 1, 90–95 (2007).
Siebenaller, J. & Somero, G. N. Pressure-adaptive differences in lactate dehydrogenases of congeneric fishes living at different depths. Science 201, 255–257 (1978).
Ritchie, H., Jamieson, A. J. & Piertney, S. B. Heat-shock protein adaptation in abyssal and hadal amphipods. Deep Sea Res. Pt II 155, 61–69 (2018).
Schopf, F. H., Biebl, M. M. & Buchner, J. The HSP90 chaperone machinery. Nat. Rev. Mol. Cell Biol. 18, 345–360 (2017).
Prodromou, C. Mechanisms of Hsp90 regulation. Biochem. J. 473, 2439–2452 (2016).
Wessel, P. & Smith, W. H. F. New, improved version of generic mapping tools released. Eos 79, 579 (1998).
Gardner, J. V. The West Mariana Ridge, western Pacific Ocean: geomorphology and processes from new multibeam data. GSA Bulletin 122, 1378–1388 (2010).
Amante, C. & Eakins, B. W. ETOPO1 1 Arc-Minute Global Relief Model: Procedures, Data Sources and Analysis (Department of Commerce, NOAA, National Oceanic and Atmospheric Administration & National Environmental Satellite, Data, and Information Service, 2009).
Kajitani, R. et al. Efficient de novo assembly of highly heterozygous genomes from whole-genome shotgun short reads. Genome Res. 24, 1384–1395 (2014).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Liu, J. et al. BinPacker: packing-based de novo transcriptome assembly from RNA-Seq data. PLoS Comput. Biol. 12, e1004772 (2016).
Smit, A., Hubley, R. & Green, P. RepeatMasker Open-4.0 (Institute for Systems Biology, 2013); http://www.repeatmasker.org
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, 435–439 (2006).
Alioto, T., Picardi, E., Guigo, R. & Pesole, G. ASPic-GeneID: a lightweight pipeline for gene prediction and alternative isoforms detection. Biomed. Res. Int. 2013, 502827 (2013).
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics 20, 2878–2879 (2004).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-Seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 9, R7 (2008).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, 353–361 (2017).
Fischer, S. et al. Using OrthoMCL to assign proteins to OrthoMCL-DB groups or to cluster proteomes into new ortholog groups. Curr. Protoc. Bioinformatics 35, 6.12.1–6.12.19 (2011).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Suyama, M., Torrents, D. & Bork, P. PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res. 34, 609–612 (2006).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: more models, new heuristics and high-performance computing. Nat. Methods 9, 772 (2012).
Bouckaert, R. R. DensiTree: making sense of sets of phylogenetic trees. Bioinformatics 26, 1372–1373 (2010).
Liu, L., Yu, L. & Edwards, S. V. A maximum pseudo-likelihood approach for estimating species trees under the coalescent model. BMC Evol. Biol. 10, 302 (2010).
Kielbasa, S. M., Wan, R., Sato, K., Horton, P. & Frith, M. C. Adaptive seeds tame genomic sequence comparison. Genome Res. 21, 487–493 (2011).
Blanchette, M. et al. Aligning multiple genomic sequences with the threaded blockset aligner. Genome Res. 14, 708–715 (2004).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Benton, M. J. & Donoghue, P. C. Paleontological evidence to date the tree of life. Mol. Biol. Evol. 24, 26–53 (2007).
Li, H. & Durbin, R. Inference of human population history from individual whole-genome sequences. Nature 475, 493–496 (2011).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Sanderson, M. J. r8s: inferring absolute rates of molecular evolution and divergence times in the absence of a molecular clock. Bioinformatics 19, 301–302 (2003).
Scheet, P. & Stephens, M. A fast and flexible statistical model for large-scale population genotype data: applications to inferring missing genotypes and haplotypic phase. Am. J. Hum. Genet. 78, 629–644 (2006).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Marco-Sola, S., Sammeth, M., Guigo, R. & Ribeca, P. The GEM mapper: fast, accurate and versatile alignment by filtration. Nat. Methods 9, 1185–1188 (2012).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Nasevicius, A. & Ekker, S. C. Effective targeted gene ‘knockdown’ in zebrafish. Nat. Genet. 26, 216–220 (2000).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
De Bie, T., Cristianini, N., Demuth, J. P. & Hahn, M. W. CAFE: a computational tool for the study of gene family evolution. Bioinformatics 22, 1269–1271 (2006).
Li, L., Stoeckert, C. J. Jr & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, 279–285 (2016).
Rambaut, A. & Grassly, N. C. Seq-Gen: an application for the Monte Carlo simulation of DNA sequence evolution along phylogenetic trees. Comput. Appl. Biosci. 13, 235–238 (1997).
Meyer, P. et al. Structural and functional analysis of the middle segment of hsp90: implications for ATP hydrolysis and client protein and cochaperone interactions. Mol. Cell 11, 647–658 (2003).
Verba, K. A. et al. Atomic structure of Hsp90-Cdc37-Cdk4 reveals that Hsp90 traps and stabilizes an unfolded kinase. Science 352, 1542–1547 (2016).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Le Guilloux, V., Schmidtke, P. & Tuffery, P. Fpocket: an open source platform for ligand pocket detection. BMC Bioinformatics 10, 168 (2009).
Acknowledgements
The authors are grateful to J. Chen, J. Li, D. Cai, Y. Xin and H. Zhou for collecting specimens. We thank all crew members for help with specimen preparation and photography. We also thank T. Morihisa for authorizing the photography of Tanaka’s snailfish. This study was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB060101 and XDB13020100), grants from the Talents Team Construction Fund of Northwestern Polytechnical University (to Q.Q. and W.W.), grants from the National Natural Science Foundation of China (numbers 41876179 and 31601858), the National Program for Support of Top-notch Young Professionals (to Q.Q.), the National Key R&D Program of China (2018YFC0309800), the Hong Kong Research Grants Council Area of Excellence Scheme (AoE/M-403/16), the 1000 Talent Project of Shaanxi Province (to W.W., Q.Q. and K.W) and the Wuhan Branch of the Supercomputing Center at the Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
S.H., Q.Q. and W.W. conceived the study. Z.G., G.Y. and L.H. collected the materials. S.H., Y.S., X.G., S.L. and H.Z. performed the morphology laboratory work. K.W. and Y.Yuan performed the genome assembly and genome annotation. Q.Q. and K.W. designed the evolutionary analyses. K.W., Y.Yang, X.P., W.Xu and C.Z. performed the evolutionary analyses. L.Z. and C.L. simulated the protein structures. G.L., K.H., Y.L., L.Y., X.S., C.F., Y.D. and W.Xiao. performed the bglap knockdown experiment. S.H., Q.Q., W.W. and K.W. wrote the manuscript with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing Interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes 1–4, Supplementary Figs. 1–36 and Supplementary Tables 1–19
Supplementary Video
In situ video of Mariana hadal snailfish
Supplementary Data 1
Three-dimensional images of micro-CT scans from high-precision skeletons
Supplementary Data 2
Three-dimensional images of micro-CT scans from low-precision skeletons
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, K., Shen, Y., Yang, Y. et al. Morphology and genome of a snailfish from the Mariana Trench provide insights into deep-sea adaptation. Nat Ecol Evol 3, 823–833 (2019). https://doi.org/10.1038/s41559-019-0864-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-0864-8
This article is cited by
-
Insights into aging mechanisms from comparative genomics in orange and silver roughies
Scientific Reports (2024)
-
Advances in environmental DNA monitoring: standardization, automation, and emerging technologies in aquatic ecosystems
Science China Life Sciences (2024)
-
The chromosome-level genome and key genes associated with mud-dwelling behavior and adaptations of hypoxia and noxious environments in loach (Misgurnus anguillicaudatus)
BMC Biology (2023)
-
Bioinspired soft robots for deep-sea exploration
Nature Communications (2023)
-
A chromosome-level genome assembly of a deep-sea starfish (Zoroaster cf. ophiactis)
Scientific Data (2023)