Abstract
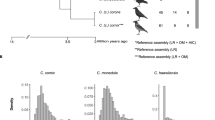
Songbirds have a species number close to that of mammals and are classic models for studying speciation and sexual selection. Sex chromosomes are hotspots of both processes, yet their evolutionary history in songbirds remains unclear. We characterized genomes of 11 songbird species, with 5 genomes of bird-of-paradise species. We conclude that songbird sex chromosomes have undergone four periods of recombination suppression before species radiation, producing a gradient of pairwise sequence divergence termed ‘evolutionary strata’. The latest stratum was probably due to a songbird-specific burst of retrotransposon CR1–E1 elements at its boundary, instead of the chromosome inversion generally assumed for suppressing sex-linked recombination. The formation of evolutionary strata has reshaped the genomic architecture of both sex chromosomes. We find stepwise variations of Z-linked inversions, repeat and guanine–cytosine (GC) contents, as well as W-linked gene loss rate associated with the age of strata. A few W-linked genes have been preserved for their essential functions, indicated by higher and broader expression of lizard orthologues compared with those of other sex-linked genes. We also find a different degree of accelerated evolution of Z-linked genes versus autosomal genes among species, potentially reflecting diversified intensity of sexual selection. Our results uncover the dynamic evolutionary history of songbird sex chromosomes and provide insights into the mechanisms of recombination suppression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Genome sequencing and RNA-seq data generated in this study have been deposited in the NCBI SRA under PRJNA491255. The raw genomic reads of Paradisaea raggiana are available in the CNGB Nucleotide Sequence Archive (https://db.cngb.org/cnsa; accession number CNP0000186). The genome assemblies are available under NCBI BioProject portal (PRJNA491255). The IDs of W-linked scaffolds are included in Supplementary Table 10.
Code availability
Custom scripts and pipelines used in this study have been deposited at Github (https://github.com/lurebgi/BOPsexChr).
References
World Bird List v.8.2 (IOC) (2018); https://www.worldbirdnames.org/ioc-lists/master-list-2/
Barker, F. K., Cibois, A., Schikler, P., Feinstein, J. & Cracraft, J. Phylogeny and diversification of the largest avian radiation. Proc. Natl Acad. Sci. USA 101, 11040–11045 (2004).
Ellegren, H. et al. The genomic landscape of species divergence in Ficedula flycatchers. Nature 491, 756–760 (2012).
Poelstra, J. W. et al. The genomic landscape underlying phenotypic integrity in the face of gene flow in crows. Science 344, 1410–1414 (2014).
Tuttle, E. M. et al. Divergence and functional degradation of a sex chromosome-like supergene. Curr. Biol. 26, 344–350 (2016).
Laine, V. N. et al. Evolutionary signals of selection on cognition from the great tit genome and methylome. Nat. Commun. 7, 10474 (2016).
Mank, J. E. Sex chromosomes and the evolution of sexual dimorphism: lessons from the genome. Am. Nat. 173, 141–150 (2009).
Coyne, J. A. Genetics and speciation. Nature 355, 511 (1992).
Charlesworth, B., Coyne, J. & Barton, N. The relative rates of evolution of sex chromosomes and autosomes. Am. Nat. 130, 113–146 (1987).
Smeds, L. et al. Evolutionary analysis of the female-specific avian W chromosome. Nat. Commun. 6, 7330 (2015).
Hooper, D. M. & Price, T. D. Chromosomal inversion differences correlate with range overlap in passerine birds. Nat. Ecol. Evol. 1, 1526–1534 (2017).
Hooper, D. M. & Price, T. D. Rates of karyotypic evolution in Estrildid finches differ between island and continental clades. Evolution 69, 890–903 (2015).
Backström, N. et al. A high-density scan of the Z chromosome in Ficedula flycatchers reveals candidate loci for diversifying selection. Evolution 64, 3461–3475 (2010).
Elgvin, T. O. et al. Hybrid speciation in sparrows II: a role for sex chromosomes?. Mol. Ecol. 20, 3823–3837 (2011).
Storchová, R., Reif, J. & Nachman, M. W. Female heterogamety and speciation: reduced introgression of the Z chromosome between two species of nightingales. Evolution 64, 456–471 (2010).
Li, W. H., Yi, S. & Makova, K. Male-driven evolution. Curr. Opin. Genet. Dev. 12, 650–656 (2002).
Wang, Z. et al. Temporal genomic evolution of bird sex chromosomes. BMC Evol. Biol. 14, 250 (2014).
Ellegren, H. Characteristics, causes and evolutionary consequences of male-biased mutation. Proc. Biol. Sci. 274, 1–10 (2007).
Smeds, L., Qvarnström, A. & Ellegren, H. Direct estimate of the rate of germline mutation in a bird. Genome Res. 26, 1211–1218 (2016).
Vicoso, B. & Charlesworth, B. Effective population size and the faster-X effect: an extended model. Evolution 63, 2413–2426 (2009).
Mank, J. E., Nam, K. & Ellegren, H. Faster-Z evolution is predominantly due to genetic drift. Mol. Biol. Evol. 27, 661–670 (2010).
Wright, A. E. et al. Variation in promiscuity and sexual selection drives avian rate of Faster-Z evolution. Mol. Ecol. 24, 1218–1235 (2015).
Zhou, Q. et al. Complex evolutionary trajectories of sex chromosomes across bird taxa. Science 346, 1246338 (2014).
Bellott, D. W. et al. Avian W and mammalian Y chromosomes convergently retained dosage-sensitive regulators. Nat. Genet. 49, 387–394 (2017).
Charlesworth, D., Charlesworth, B. & Marais, G. Steps in the evolution of heteromorphic sex chromosomes. Heredity 95, 118–128 (2005).
Bachtrog, D. Y-chromosome evolution: emerging insights into processes of Y-chromosome degeneration. Nat. Rev. Genet. 14, 113–124 (2013).
Charlesworth, B. & Charlesworth, D. The degeneration of Y chromosomes. Philos. Trans. R. Soc. Lond. B 355, 1563–1572 (2000).
Bachtrog, D. The temporal dynamics of processes underlying Y chromosome degeneration. Genetics 179, 1513–1525 (2008).
Soh, Y. Q. et al. Sequencing the mouse Y chromosome reveals convergent gene acquisition and amplification on both sex chromosomes. Cell 159, 800–813 (2014).
Bellott, D. W. et al. Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508, 494–499 (2014).
Zhou, Q. & Bachtrog, D. Sex-specific adaptation drives early sex chromosome evolution in Drosophila. Science 337, 341–345 (2012).
Koerich, L. B., Wang, X., Clark, A. G. & Carvalho, A. B. Low conservation of gene content in the Drosophila Y chromosome. Nature 456, 949–951 (2008).
Wright, A. E., Harrison, P. W., Montgomery, S. H., Pointer, M. A. & Mank, J. E. Independent stratum formation on the avian sex chromosomes reveals inter-chromosomal gene conversion and predominance of purifying selection on the W chromosome. Evolution 68, 3281–3295 (2014).
Moghadam, H. K., Pointer, M. A., Wright, A. E., Berlin, S. & Mank, J. E. W chromosome expression responds to female-specific selection. Proc. Natl Acad. Sci. USA 109, 8207–8211 (2012).
Cortez, D. et al. Origins and functional evolution of Y chromosomes across mammals. Nature 508, 488–493 (2014).
Bergero, R., Forrest, A., Kamau, E. & Charlesworth, D. Evolutionary strata on the X chromosomes of the dioecious plant Silene latifolia: evidence from new sex-linked genes. Genetics 175, 1945–1954 (2007).
Gorelick, R. et al. Abrupt shortening of bird W chromosomes in ancestral Neognathae. Biol. J. Linn. Soc. Lond. 119, 488–496 (2016).
Rice, W. R. The accumulation of sexually antagonistic genes as a selective agent promoting the evolution of reduced recombination between primitive sex chromosomes. Evolution 41, 911–914 (1987).
Claramunt, S. & Cracraft, J. A new time tree reveals Earth historys imprint on the evolution of modern birds. Sci. Adv. 1, e1501005–e1501005 (2015).
Smeds, L. et al. Genomic identification and characterization of the pseudoautosomal region in highly differentiated avian sex chromosomes. Nat. Commun. 5, 5448 (2014).
Pala, I. et al. Evidence of a neo-sex chromosome in birds. Heredity 108, 264–272 (2012).
Ross, M. T. et al. The DNA sequence of the human X chromosome. Nature 434, 325–337 (2005).
O’Connor, R. E. et al. Chromosome-level assembly reveals extensive rearrangement in saker falcon and budgerigar, but not ostrich, genomes. Genome Biol. 19, 171 (2018).
Wright, A. E., Dean, R., Zimmer, F. & Mank, J. E. How to make a sex chromosome. Nat. Commun. 7, 12087 (2016).
Duret, L. & Galtier, N. Biased gene conversion and the evolution of mammalian genomic landscapes. Annu. Rev. Genomics Hum. Genet. 10, 285–311 (2009).
Kent, T. V., Uzunović, J. & Wright, S. I. Coevolution between transposable elements and recombination. Philos. Trans. R. Soc. B 372, 20160458 (2017).
Suh, A. et al. Mesozoic retroposons reveal parrots as the closest living relatives of passerine birds. Nat. Commun. 2, 443 (2011).
Friocourt, F. et al. Recurrent DCC gene losses during bird evolution. Sci. Rep. 7, 37569 (2017).
Patthey, C., Tong, Y. G., Tait, C. M. & Wilson, S. I. Evolution of the functionally conserved DCC gene in birds. Sci. Rep. 7, 42029 (2017).
Ligon, R. A. et al. Evolution of correlated complexity in the radically different courtship signals of birds-of-paradise. PLoS Biol. 16, e2006962 (2018).
Irestedt, M., Jonsson, K. A., Fjeldsa, J., Christidis, L. & Ericson, P. G. An unexpectedly long history of sexual selection in birds-of-paradise. BMC Evol. Biol. 9, 235 (2009).
Dunn, P. O., Whittingham, L. A. & Pitcher, T. E. Mating systems, sperm competition, and the evolution of sexual dimorphism in birds. Evolution 55, 161–175 (2001).
Diamond, J. Biology of birds of paradise and bowerbirds. Annu. Rev. Ecol. Syst. 17, 17–37 (1986).
Kirkpatrick, M. & Ryan, M. J. The evolution of mating preferences and the paradox of the lek. Nature 350, 33–38 (1991).
Suh, A. The specific requirements for CR1 retrotransposition explain the scarcity of retrogenes in birds. J. Mol. Evol. 81, 18–20 (2015).
Huang, N., Lee, I., Marcotte, E. M. & Hurles, M. E. Characterising and predicting haploinsufficiency in the human genome. PLoS Genet. 6, e1001154 (2010).
Stock, M. et al. Ever-young sex chromosomes in European tree frogs. PLoS Biol. 9, e1001062 (2011).
Vicoso, B., Emerson, J. J., Zektser, Y., Mahajan, S. & Bachtrog, D. Comparative sex chromosome genomics in snakes: differentiation, evolutionary strata, and lack of global dosage compensation. PLoS Biol. 11, e1001643 (2013).
Charlesworth, D. Evolution of recombination rates between sex chromosomes. Philos. Trans. R. Soc. Lond. B 372, 20160456 (2017).
Lemaitre, C. et al. Footprints of inversions at present and past pseudoautosomal boundaries in human sex chromosomes. Genome Biol. Evol. 1, 56–66 (2009).
Ellis, N. A. et al. The pseudoautosomal boundary in man is defined by an Alu repeat sequence inserted on the Y chromosome. Nature 337, 81–84 (1989).
Raudsepp, T. & Chowdhary, B. P. The eutherian pseudoautosomal region. Cytogenet. Genome Res. 147, 81–94 (2015).
Zamudio, N. et al. DNA methylation restrains transposons from adopting a chromatin signature permissive for meiotic recombination. Genes Dev. 29, 1256–1270 (2015).
Ben-Aroya, S., Mieczkowski, P. A., Petes, T. D. & Kupiec, M. The compact chromatin structure of a Ty repeated sequence suppresses recombination hotspot activity in Saccharomyces cerevisiae. Mol. Cell 15, 221–231 (2004).
Knief, U. et al. Fitness consequences of polymorphic inversions in the zebra finch genome. Genome Biol. 17, 199 (2016).
Campagna, L. et al. Repeated divergent selection on pigmentation genes in a rapid finch radiation. Sci. Adv. 3, e1602404 (2017).
Irwin, D. E. Sex chromosomes and speciation in birds and other ZW systems. Mol. Ecol. 27, 3831–3851 (2018).
Prost, S. et al. Comparative analyses identify genomic features potentially involved in the evolution of birds-of-paradise. GigaScience https://doi.org/10.1093/gigascience/giz003 (2018).
Gnerre, S. et al. High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proc. Natl Acad. Sci. USA 108, 1513–1518 (2011).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience 1, 18 (2012).
Cantarel, B. L. et al. MAKER: an easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 18, 188–196 (2008).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19, ii215–ii225 (2003).
Marçais, G. et al. MUMmer4: A fast and versatile genome alignment system. PLoS Comput. Biol. 14, e1005944 (2018).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Genomics (q-bio.GN) https://arxiv.org/abs/1303.3997 (2013).
Singhal, S. et al. Stable recombination hotspots in birds. Science 350, 928–932 (2015).
Zhang, J., Li, C., Zhou, Q. & Zhang, G. Improving the ostrich genome assembly using optical mapping data. GigaScience 4, 24 (2015).
Xu, L. et al. Evolutionary dynamics of sex chromosomes of palaeognathous birds. Preprint at bioRxiv https://doi.org/10.1101/295089 (2018).
Harris, R. S. Improved pairwise alignment of genomic DNA. PhD thesis, The Pennsylvania State University (2017).
Kent, W. J. BLAT — the BLAST-like alignment tool. Genome Res. 12, 656–64 (2002).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Lechner, M. et al. Proteinortho: detection of (co-)orthologs in large-scale analysis. BMC Bioinformatics 12, 124 (2011).
Jetz, W., Thomas, G. H., Joy, J. B., Hartmann, K. & Mooers, A. O. The global diversity of birds in space and time. Nature 491, 444–448 (2012).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Zhang, G. et al. Comparative genomics reveals insights into avian genome evolution and adaptation. Science 346, 1311–1320 (2014).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Jiao, X. et al. DAVID-WS: a stateful web service to facilitate gene/protein list analysis. Bioinformatics 28, 1805–1806 (2012).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Yanai, I. et al. Genome-wide midrange transcription profiles reveal expression level relationships in human tissue specification. Bioinformatics 21, 650–659 (2005).
del Hoyo, J, . & Elliot, A. & Sargatal, J. & Christie, D. & de Juana, E. Handbook of the Birds of the World Alive (Lynx Editions, 2019).
Acknowledgements
We thank the Smithsonian Institute (G. Graves), Australian National Wildlife Collection, CSIRO Sustainable Ecosystems (L. Joseph), Museum Victoria, Australia (J. Sumner), Division of Vertebrate Zoology Yale University, Peabody Museum of Natural History (K. Zyskowski) for tissue samples; and E. Scholes for discussions on BOP. We also acknowledge the support from Science for Life Laboratory, the National Genomics Infrastructure (NGI), Uppmax. L.X. is supported by the uni:docs fellowship programme from University of Vienna. M.I. is supported by the Swedish Research Council (grant no. 621-2014-5113). Q.Z. is supported by National Natural Science Foundation of China (grant nos. 31722050 and 31671319), the Fundamental Research Funds for the Central Universities (grant no. 2018XZZX002-04) and start-up funds from Zhejiang University. The computational analyses were performed on CUBE cluster from Department of Computational System Biology of University of Vienna and Vienna Scientific Cluster.
Author information
Authors and Affiliations
Contributions
Q.Z. and M.I. conceived the project. L.X., Q.Z., G.A., V.P., Y.D., S.F., G.Z., M.B. and S. P. performed the analyses. Q.Z., L.X., A.S., L.C. and M.I. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Xu, L., Auer, G., Peona, V. et al. Dynamic evolutionary history and gene content of sex chromosomes across diverse songbirds. Nat Ecol Evol 3, 834–844 (2019). https://doi.org/10.1038/s41559-019-0850-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-0850-1
This article is cited by
-
Evolution and expression patterns of the neo-sex chromosomes of the crested ibis
Nature Communications (2024)
-
A high-quality genome assembly highlights the evolutionary history of the great bustard (Otis tarda, Otidiformes)
Communications Biology (2023)
-
Recurrent chromosome reshuffling and the evolution of neo-sex chromosomes in parrots
Nature Communications (2022)