Abstract
Tuberaceae is one of the most diverse lineages of symbiotic truffle-forming fungi. To understand the molecular underpinning of the ectomycorrhizal truffle lifestyle, we compared the genomes of Piedmont white truffle (Tuber magnatum), Périgord black truffle (Tuber melanosporum), Burgundy truffle (Tuber aestivum), pig truffle (Choiromyces venosus) and desert truffle (Terfezia boudieri) to saprotrophic Pezizomycetes. Reconstructed gene duplication/loss histories along a time-calibrated phylogeny of Ascomycetes revealed that Tuberaceae-specific traits may be related to a higher gene diversification rate. Genomic features in Tuber species appear to be very similar, with high transposon content, few genes coding lignocellulose-degrading enzymes, a substantial set of lineage-specific fruiting-body-upregulated genes and high expression of genes involved in volatile organic compound metabolism. Developmental and metabolic pathways expressed in ectomycorrhizae and fruiting bodies of T. magnatum and T. melanosporum are unexpectedly very similar, owing to the fact that they diverged ~100 Ma. Volatile organic compounds from pungent truffle odours are not the products of Tuber-specific gene innovations, but rely on the differential expression of an existing gene repertoire. These genomic resources will help to address fundamental questions in the evolution of the truffle lifestyle and the ecology of fungi that have been praised as food delicacies for centuries.
Similar content being viewed by others
Main
Truffle fungi differentiate into subterranean fruiting bodies bearing spores sequestered in an inconspicuous globous-like mass of hyphae. Truffle-forming species have evolved in nearly every major group of fleshy fungi over 100 times independently within the Ascomycota and Basidiomycota and the majority of evolutionary transitions to a truffle morphology have occurred in lineages establishing ectomycorrhizal mutualistic symbiosis with plants1. This pattern suggests that symbiosis has been a major driver in the evolution of truffle diversity. The evolution of the hypogeous lifestyle across a diversity of truffle lineages also suggests that the transition from epigeous to hypogeous fruiting is driven by strong selection for traits, for example pungent odours, that promote animal dispersal2. The Tuberaceae family is one of the most diverse lineages of mainly truffle-forming fungi and is presumably one of the earliest diverging clades within the Pezizomycetes (Ascomycota)2. Truffles, such as the aromatic Périgord black truffle (T. melanosporum Vittad.), Burgundy truffle (T. aestivum Vittad.), Piedmont white truffle (T. magnatum Pico) and desert truffle (Te. boudieri Chatin), have been praised as food delicacies for centuries3. Despite their ecological importance and a long history as a gourmet food, there are still many unanswered questions concerning the biology, genetics and ecology of Tuberaceae4.
Previously, the genome of T. melanosporum was sequenced and revealed a set of unusual genomic features, including a very high proportion of transposable elements, a restricted set of plant cell wall degrading enzymes (PCWDEs) and rare gene duplications5. In addition, transcript profiling of fruiting bodies showed that the synthesis of volatile organic compounds (VOCs), forming the pervasive scent of truffles, is supported by a sustained expression of various VOC-related genes5. However, it is not known whether VOC metabolism is conserved within Tuberaceae6.
To gain insight into the evolution of ectomycorrhizal symbiosis, as well as truffle lifestyle within the Pezizomycetes, we sequenced additional symbiotic truffle-forming species (T. magnatum, T. aestivum, C. venosus Fr. and Te. boudieri) and compared their genomes to T. melanosporum5 and the saprotrophic black morel (Morchella importuna M. Kuo, O’Donnell & T.J. Volk), Ascobolus immersus Pers. and Pyronema confluens Tul & C. Tul7. Here we present the comparative analysis of these genomes, with particular emphasis on Tuberaceae-specific sequences, symbiosis- and fruiting-body-regulated genes, and genes coding for PCWDEs and VOC synthesis. Furthermore, gene expression in ectomycorrhizae and fruiting bodies from T. magnatum and T. melanosporum were compared to gain insights into how expression patterns changed along the evolution of these truffles.
Genome features and phylogeny of Tuberaceae
Nuclear genomes of the ectomycorrhizal truffles C. venosus, T. aestivum and T. magnatum in Tuberaceae and Te. boudieri in Terfeziaceae, as well as of saprotrophic Pezizomycetes M. importuna and A. immersus, were sequenced (Table 1 and Supplementary Information). Genomes ranged from 48 Mb (M. importuna) to 192 Mb (T. magnatum) (Table 1 and Fig. 1b). Tuberaceae genomes are significantly larger than Te. boudieri and saprotrophic Pezizomycetes genomes. Predicted protein-coding content ranged from 9,344 for T. aestivum to 17,986 for C. venosus (Fig. 1c and Table 1).
a, Phylogenetic relationship, genome size and number of gene models for the eight Pezizomycetes sequenced in this study. The maximum-likelihood phylogeny is constructed using 2,093 conserved, single-copy orthologous Pezizomycetes genes. Divergence time intervals and bootstrap values are displayed near tree nodes. b, Proportion of transposable elements and non-repeated sequences. c, Number of gene models. ECM, ectomycorrhizal; SAP, saprotrophic; TE, transposable element. Note that the ectomycorrhizal Te. boudieri may establish endomycorrhizal associations under certain growth conditions30.
Phylogenetic reconstruction using 2,093 concatenated conserved single-copy protein-coding genes from the eight Pezizomycetes corroborates previously reported topologies (Fig. 1a)2,8,9 with Tuber spp. clustering together and C. venosus being phylogenetically distinct from Tuber spp. within Tuberaceae2. We estimated the age of the most recent common ancestor (MRCA) of Pezizomycetes at 470 ± 67 million years (My), in accordance with previous estimations10. We also determined that Tuberaceae began diverging around 140 ± 10 Ma in the Early Cretaceous, which corresponds to the time when the Angiosperms evolved11.
A comparison of Tuberaceae genomes revealed no evidence of whole genome duplication events (that is, no segmental duplications). Despite the high proportion of scattered transposable elements and other repeated elements, a substantial gene co-linearity (microsynteny) among Tuberaceae genomes was apparent (Supplementary Fig. 1). As expected, there is a high correlation between the number of syntenic regions and the estimated age of divergence between the various clades (Supplementary Fig. 2).
Invasion of Tuberaceae genomes by transposable elements
Transposable element content ranged from 3.5% in A. immersus to 58% in T. magnatum and T. melanosporum (Table 1 and Supplementary Fig. 3). There was a strong association (Pearson’s, r = 0.83, P = 0.01) between genome size and the proportion of repeated elements across species (Fig. 1b and Supplementary Fig. 4), confirming that genome expansion in Tuberaceae is mainly driven by transposable element proliferation. Genome size, and consequently proportion of transposable elements, negatively correlates with the size of the gene repertoire (Pearson’s, r = −0.92, P = 0.001; Supplementary Fig. 3), highlighting the impact of transposable elements on the coding space. Gypsy long-terminal repeat (LTR) retrotransposons are the most frequent known transposable elements, representing up to 46% of the T. magnatum genome (Supplementary Fig. 3). Their age distribution shows that they have been proliferating over the past 5 to 6 Myr for T. melanosporum, T. aestivum and C. venosus, while LTR accumulation in T. magnatum is more ancient and occurred between 6 and 14 Ma (Supplementary Fig. 5). As a result of LTR sequence degeneration, very ancient invasions of LTR retrotransposons cannot be dated, but they were massive as they represent the largest proportion of current retrotransposon population (Supplementary Fig. 3). No evidence of repeat-induced point mutation was detected in sequenced Pezizomycetes (Supplementary Fig. 6). The presence of enzymes involved in DNA methylation (Supplementary Table 1) suggests that methylation induced premeiotically controls the activity of repeated sequences in these species12,13,14.
Core, dispensable and species-specific genes
To compare gene repertoires encoded by sequenced Pezizomycetes and identify species-specific gene families that might contribute to trait diversification, we clustered the predicted proteins to infer orthologous gene groups (orthogroups), including core genes (that is, occurring in the eight species), dispensable genes (that is, found in at least two species) and species-specific genes (that is, unique to a taxa). To define species-specific genes in a broader context, we then searched for orthologous sequences in >500 published fungal genomes. While the set of core genes is in the range of 2,443 to 2,511, the repertoire of species-specific genes is much lower in Tuber species compared to the other taxa (Supplementary Fig. 7). Species-specific genes, which are also referred to as taxonomically restricted genes, mostly encode proteins with no known function. They are thought to determine specific traits, such as adaptation to different environmental niches or preferential colonization of certain host plants15. The highest number of genes in both C. venosus and A. immersus compared to other Pezizomycetes is explained by their higher proportion of species-specific genes.
To determine whether there are significant differences between the shared and species-specific genes, we compared their protein length, exon and intron numbers, protein families database (Pfam) and transmembrane domain content, and signal peptide presence (Supplementary Table 2). According to our results, the averages of the above features of the two gene sets are significantly different. Species-specific genes exhibit the lowest values for most genetic features, suggesting that they evolve separately from the species-shared genes. The transcriptional RNA-Seq analysis revealed that 34% (T. melanosporum) to 87% (C. venosus) of species-specific genes have transcripts. Overall, the analysis of genetic features indicates that the two gene sets are noticeably distinct and that species-specific genes have a smaller gene size, fewer exons and a lower proportion of expressed genes than the conserved genes, suggesting a differential evolution, that is they might be evolutionarily young genes.
Accelerated rate of genome diversification in Tuberaceae
The new genomes, and more specifically Tuberaceae and Terfeziaceae genomes, offer a unique opportunity to examine the evolution of truffle-related and ectomycorrhiza-related genes, and gene families across the Pezizomycetes clade. Reconstructing genome-wide duplication and contraction events in Pezizomycetes, as well as in a large sample of additional fungal species including representatives of the major subphyla, revealed a considerable heterogeneity in the temporal dynamics of genome diversification between the different clades (Fig. 2). Approximately 1,200 gene duplications are marking the origin of Pezizomycetes, followed by gradual genome contractions in all Pezizomycete lineages. Of note, as few as 84 gene duplications were inferred for the common ancestor of Te. boudieri and A. immersus (Terfeziaceae + Ascobolaceae), whereas considerable species-specific gene diversification was observed in these species, which may be due to the low sampling density in this clade. On the contrary, branches leading to Tuberaceae showed on average 400 to 1,000 gene duplications and 800 to 1,200 gene losses per branch. Notably, we found that the rate of gene duplication within the genus Tuber was 1.7- to 2.2-fold higher than in the other lineages (Supplementary Fig. 8). In addition to Tuber species, we inferred >7,000 gene duplications in the lineage leading to C. venosus, although most of the duplicated genes are of unknown function (Supplementary Fig. 7). This latter lineage shows the highest rate of genome diversification among Pezizomycetes (~10-fold faster than that estimated for the other clades), although further sampling of species will be necessary to understand how duplications happened along this lineage.
a,b, The expansion (green; a) and contraction (orange; b) of protein-coding gene repertoires in Tuberaceae and other Pezizomycetes (red box). The analysis was performed using the COMPARE pipeline27. The size of the bubbles corresponds to the ancestral number of genes inferred in that node. The scale bar units correspond to gene counts.
Altogether, Tuberaceae has a significantly accelerated evolution of its gene repertoire (analysis of variance (ANOVA), P = 0.015) compared to other ascomycetous species in this dataset, possibly reflecting an increased rate of genomic evolution underlying traits specific to either symbiosis and/or hypogeous fruiting-body formation. ANOVA within the entire hypogeous Ascomycetes dataset (that is, Tuberaceae + Terfeziaceae) also indicated a significantly higher rate of gene duplication compared to non-hypogeous ascomycetes (P = 0.014). A similar, albeit slightly weaker trend was observed for the gene loss rates in the Tuberaceae, which is significantly higher than in the other clades (P = 0.032), indicating once again an accelerated gene turnover in these particular species (but not in the hypogeous Ascomycetes in general, P = 0.061). Of note, we also inferred a significant number of terminal duplication events in Tuber species (1,000 to 1,200 per branch, yielding orphan genes) (Supplementary Fig. 8), suggesting a possible role of species-specific adaptive changes in the evolution of the genus.
To gain insights into the unique features of Tuberaceae and Tuber genomes, we compared the frequencies of Gene Ontology annotations of genes that showed duplications in these clades. Although most of the duplicated genes specific to Tuberaceae had no functional annotation, several functions were found to be overrepresented compared to other Pezizomycetes in particular genes involved in DNA binding, transposition, RNA metabolism and biosynthesis of aromatic and organic cyclic compounds (Supplementary Fig. 9). These compounds contribute to the pungent flavours released by truffle fruiting bodies.
Tuberaceae have a restricted set of PCWDEs
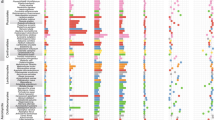
In sequenced Pezizomycetes, the total number of secreted carbohydrate-active enzymes involved in plant polysaccharide degradation ranged from 49 for the symbiotic Te. boudieri to 203 for the coprophilic A. immersus (Fig. 3). As expected, based on their saprotrophic lifestyle, genomes of A. immersus, M. importuna and P. confluens encode a large repertoire of PCWDEs, including lignin-, cellulose-, hemicellulose- and pectin-degrading enzymes, as well as multiple carbohydrate-binding modules, indicating the potential to degrade a large set of substrates found in decaying soil organic matter and plant debris (for example, herbivore dung) (Supplementary Table 3). In contrast, these PCWDE genes are lacking, or occur at a very low level, in Tuber spp. and Te. boudieri (Supplementary Table 3).
The double hierarchical heatmap shows gene copy numbers for PCWDEs in Pezizomycetes examined and a set of representative species of saprotrophic, pathogenic or symbiotic ascomycetes and basidiomycetes (see Supplementary Table 15 for the species list and their abbreviations). Black frames highlight Pezizomycetes taxa, whereas black arrows indicate Tuberaceae taxa and red dotted lines highlight PL1, PL4 and CE12 families. The ecology of each species is indicated at the right of the species abbreviations.
The set of PCWDEs in truffles is as low as the repertoire reported from previously sequenced ectomycorrhizal basidiomycetes16 (Fig. 3). This corroborates a convergent decrease in plant cell wall degrading capacity of ectomycorrhizal ascomycetous and basidiomycetous symbionts16,17, resulting in a limited ability to decompose soil (plant) organic matter and host plant cell wall materials. DNA decay is probably the inactivation mechanism specifically driving the loss of GH6 in T. melanosporum (Supplementary Material, Supplementary Figs. 10 and 11).
Of note, genes coding for pectate lyase (PL1), rhamnogalacturonan endolyase (PL4) and pectin acetylesterase (CE12) are present in ectomycorrhizal Pezizomycetes, whereas they have been lost in ectomycorrhizal basidiomycetes, indicating that the former symbionts may use these enzymes for the colonization of the middle lamella of host roots. Intriguingly, the PCWDE repertoire of C. venosus is closer to saprotrophic taxa (for example P. confluens) than to ectomycorrhizal Tuber spp. Its substantial PCWDE repertoire suggests that C. venosus mycelium can also acquire carbon directly from soil organic matter, although the natural abundance 15N/13C ratio of its fruiting body, a signature of carbon sources used for elaborating fruiting-body tissues, is typical of ectomycorrhizal symbionts18, that is host photoassimilates are the major source of carbon.
On the other hand, Tuberaceae has a richer set of copper-dependent lytic polysaccharide monooxygenases involved in chitin (AA11) degradation (Supplementary Table 3), which suggests their ability to partly acquire nitrogen and carbon from the soil fungal necromass19.
Shared symbiosis genes in truffle ectomycorrhizae
We found that 5,229 (75%) of the 6,952 orthologous T. magnatum/T. melanosporum genes were expressed in T. magnatum–Quercus robur and T. melanosporum–Corylus avellana ectomycorrhizae (Supplementary Fig. 12A), indicating that developmental and metabolic pathways associated with symbiosis in these truffles are very similar, despite their interaction with different host species. However, among orthologous genes, 218 and 253 displayed a species-specific expression in either T. melanosporum or T. magnatum, respectively (Supplementary Fig. 12A). Differential gene expression analysis of T. magnatum–Quercus robur ectomycorrhizae identified 514 upregulated and 928 downregulated genes, relative to free-living mycelium (fold-change > 5, false discovery rate (FDR), P < 0.05) (Supplementary Tables 4 and 5). Highly upregulated genes of known function include oxidoreductases (thioredoxin, cupredoxin, laccase, cytochrome P450), transcription factors, RNA metabolism-related proteins, and membrane transporters (carbohydrates, ammonium, amino acids, peptides) (Supplementary Table 4). As previously observed for T. melanosporum5, root colonization by T. magnatum triggers the induction of a limited set of PCWDEs. Genes coding for GH5 endoglucanases, GH28 polygalacturonase, CE12 carbohydrate esterase, GH43 arabinosidase and GH12 xyloglucan hydrolase are amongst the most upregulated genes (Supplementary Table 4), supporting their involvement in hemicellulose degradation during root apoplast colonization.
Most T. magnatum symbiosis-upregulated genes are conserved in other Pezizomycetes, including saprotrophic species (66%, clusters V and VI in Fig. 4a). These genes code for key metabolic and cellular functions (for example, membrane transporters, signalling proteins, oxidases, Zn-finger transcriptional factors) that are related to the symbiotic metabolism (for example, increased nutrient fluxes between symbionts) and symbiosis-related changes in hyphal development (Supplementary Table 4). Approximately one-third of symbiosis-induced genes are homologous to genes only present in Tuberaceae (clusters I, II and III). This large set of mycorrhiza-induced genes conserved in Tuberaceae indicates that similar ancient gene networks are involved in development and functioning of ectomycorrhizae in Tuberaceae species (for example, Supplementary Table 7). This is in agreement with the fact that the ancestral ecology for Tuberaceae is presumed to be ectomycorrhizal because all of the extant species possess this ecology, and the MRCA of the family was probably ectomycorrhizal with angiosperm hosts at the end of the Jurassic period (156 Ma)2.
a,b, The heatmap depicts a double hierarchical clustering of 514 ectomycorrhizae-upregulated (a) or 489 fruiting-body-upregulated (b) genes from T. magnatum (rows, fold-change > 5, FDR-corrected P < 0.05, n = 3) based on their BLASTP percentage sequence identity (colour scale at left) with their orthologues (if any) in selected fungal species (columns). Left, colour-coded gene clusters. Data were visualized and clustered using R (package HeatPlus). The hierarchical clustering was done using an Euclidian distance metric and Ward clustering method. For each cluster the percentages of putative functional categories are given as bargrams and the number and percentage of genes in each cluster are shown. Clusters significantly enriched for small secreted proteins (Fisher’s test, P < 0.01) are marked with an asterisk. KOG, eukaryotic orthologous groups.
A low proportion (15%) of T. magnatum symbiosis-upregulated genes are similarly induced in T. melanosporum–Corylus avellana ectomycorrhizae (Supplementary Fig. 12B, Supplementary Table 6)5,20. They are involved in core metabolic pathways required for an efficient symbiosis, such as carbohydrate, oligopeptide, amino acid and lipid transporters, or root colonization (for example, GH5 endoglucanase), but most of them (62–69%) have no known function (Supplementary Tables 6 and 7).
Shared fruiting-body-related genes in truffle
Although the vegetative mycelia of several Tuber species can be grown on agar medium and ectomycorrhizae produced by inoculated trees under controlled environmental conditions, truffle fruiting cannot be triggered in the laboratory. This is further complicated due to the heterothallic mating system of Tuber species (Supplementary materials, Supplementary Table 8). Therefore, fruiting bodies of T. magnatum were sampled in truffle grounds, their transcriptome analysed by RNA sequencing and compared to published T. melanosporum transcript profiling5. We found that 5,173 genes (74%) of orthologous T. magnatum/T. melanosporum genes were also expressed in T. magnatum fruiting bodies (Supplementary Fig. 12C). Most of these genes (5,010) are expressed in both truffles, indicating that developmental and metabolic pathways associated with the reproductive (sexual) stage of these two truffles are very similar. This is surprising, given that these species diverged ~100 Ma and belong to two separate clades2,8,9.
A total of 489 upregulated and 1,073 downregulated genes (fold-change > 5, FDR P < 0.05) were identified in T. magnatum fruiting bodies by comparison to free-living mycelium (Fig. 4b and Supplementary Tables 9, 10 and 12). Among upregulated genes, 121 (25%) are well conserved and share a sequence homology with sequenced Pezizomycetes (cluster VI); 163 (33%) have homologues in other Tuberaceae only (clusters I, III, IV and V). Among the most highly upregulated, shared genes, oxidoreductases (for example alcohol oxidase, cytochrome P450), membrane transporters and enzymes of sulfur metabolism (sulfate adenylyltransferase, adenylsulfate kinase) were found (Supplementary Tables 7 and 9).
As many as 178 genes (36%) are specific to T. magnatum (cluster VII). Up to 90% of the latter have no known function or conserved protein domain (Supplementary Table 12). Some of these species-specific genes may partly explain T. magnatum morphological and metabolic idiosyncrasies, although specificities in fruiting-body morphologies and truffle metabolism can also be driven by expression changes in a few conserved genes.
Expression of genes involved in the sexual reproduction machinery (that is, mating and meiosis) was assessed in several immature and mature T. melanosporum and T. magnatum fruiting bodies sampled in truffle orchards over several years. Despite a substantial transcript profile variability, probably reflecting the unavoidable heterogeneity inherent to environmental samples (Supplementary Fig. 13, Supplementary Table 13), the expression levels of these genes were overall remarkably similar between species and among samples.
Aroma biosynthesis in truffles
Truffle aroma comprises a complex mixture of VOCs6. By combining in silico analysis of biosynthetic pathways with transcriptome profiling, we found that genomes of the Tuberaceae, but also those from the other sequenced Pezizomycetes, encode genes of biosynthetic pathways (for example sulfate reduction, amino acid and fatty acid catabolism) leading to truffle VOCs (Supplementary Table 14). However, specific gene expression patterns were characterized in several taxa by a closer comparison of their transcript profiles. Expression of genes coding for enzymes and transporters involved in specific (often rate-limiting) steps of sulfur-amino acid metabolism, such as sulfate reduction, the homocysteine-methionine cycle and VOC synthesis from methionine, is particularly sustained in Tuber species (Fig. 5), in line with the fact that sulfur volatiles are key constituents of truffle VOCs and play a crucial role in their sexual reproduction, that is sulfur-related volatile organic compounds (S-VOCs) released by truffles are attractive to rodents and truffle flies6, which disperse their spores.
Sulfur assimilation pathway genes as well as putative (spontaneous) side-reactions leading to the synthesis of the most important S-VOCs are represented. For each enzyme, fruiting-body expression levels (as given in Supplementary Table 14) are indicated with a colour gradient ranging from black (‘no expression’) to violet (‘high expression’, ≥ 250 RPKM); whereas increased or decreased expression levels in fruiting bodies with respect to free-living mycelium are shown in red and green, respectively. Detailed information on each gene is reported in Supplementary Table 14. Cys (cysteine) and Met (methionine) are indicated in green boxes; S-VOCs are in light blue boxes. DMS, dimethylsulfide; DMTS, dimethyltrisulfide; DMDS, dimethyldisulfide; KMBA, α-keto-γ-(methylthio)butyric acid; HMBA, α-keto-γ-(methylthio)hydroxylbutyric acid; RPKM, reads per kilobase million; FB, fruiting body; FLM, free living mycelium; Ser, serine; SAM, S-adenosylmethionine; SAH, S-adenosyl-L-homocysteine; OAS, O-acetylserine; OAH, O-acetyl-L-homoserine; PAPS, 3'-phosphoadenylyl sulfate; APS, adenylyl sulfate. For gene abbreviations, see Supplementary Table 14. Dashed lines indicate spontaneous reaction.
C. venosus shows a low expression of almost all of the S-VOC-related genes. This finding suggests that the unique flavour of the pig truffle may originate from a different set of pathways. In addition, the expression patterns of genes involved in the Ehrlich pathway, leading to alcohol and aldehyde VOCs, appear to be quite distinctive in most Tuber species (Supplementary Table 14). For instance, the higher expression levels of genes coding for branched-chain and aromatic amino acid transferases probably explain the preferential occurrence of specific VOCs (for example veratrole, anisole, 3-methyl-1-butanal) in T. melanosporum compared to the other truffles. Unexpectedly, specific VOC compositions of individual truffle species may thus be largely explained by the differential expression of selected subsets of metabolic genes, while variation in gene content and/or gene copy number appear to play a relatively minor role in Tuber aroma formation.
The analysis of the expression profiles of genes involved in fatty acid synthesis revealed significant similarities between Tuber species and M. importuna, yet failed to explain the quite unique volatile ester profile of T. melanosporum (Supplementary Table 14). Similarly, the production of 2,4-dithiapentane by T. magnatum could not be traced down to a specific gene repertoire, corroborating recent results showing that VOCs may actually be produced by the community of bacteria, yeasts and moulds specifically associated with different truffle species21. Future challenges include the experimental assessment of the catalytic properties of specific enzymes involved in VOC biosynthesis and the elucidation of the role played by fruiting-body-associated microorganisms in aroma formation.
Conclusions
In Pezizomycetes, the transition from saprotrophic nutrition mode and epigeous fruiting body to the ectomycorrhizal lifestyle and truffle habit occurred at least twice in the MRCAs of Tuberaceae and Terfeziazeae. From a genomic point of view, Tuber species appear quite similar overall, with a strikingly high abundance of transposons, low protein-coding gene repertoires, restricted sets of PCWDEs, highly expressed genes involved in VOC synthesis, and conserved genes with no known function, preferentially expressed in fruiting bodies, and presumably associated with their formation. Some of the observed differences at the genome and transcriptome levels have probably contributed to the evolution of their specific phenotypic features. Our findings also suggest that the complex cocktail of Tuber VOCs are not the result of specific gene innovations, but mostly rely on the differential expression of existing gene repertoire. The present Tuberaceae genomes have provided significant insight into our understanding of the evolution of complex developmental and morphological traits. It also led to the identification of a number of genes specifically related to ectomycorrhizae and truffle development. A detailed understanding of the ontogenetic and metabolic roles played by these genes will require an in-depth functional characterization of their protein products.
Methods
Strains and fungal material used for genome sequencing
Genomic DNA used for sequencing T. aestivum, T. magnatum and C. venosus genomes were extracted from fruiting bodies (ascomata) sampled in truffle grounds at Montdoré (Haute-Saône, France), Montemagno (Piedmont, Italy) and Charmes State Forest (Vosges, France), respectively. Cultures of free-living mycelium were used for M. importuna, A. immersus and Te. boudieri. Strain RN42 (CBS 140666) of A. immersus was grown as described in Rhounim et al22. Strain S1 (ATCC MYA-4762) of Te. boudieri was grown on cellophane sheet placed on a solid Fontana medium in the dark at 25°C for 1 month23.
Genome sequencing, assembly and annotation
T. melanosporum and P. confluens genomes have previously been published5,7. Genomes of A. immersus, M. importuna, Te. boudieri and C. venosus were sequenced by the US Department of Energy Joint Genome Institute (JGI) (Table 1) using a combination of Illumina fragment (270 base pair (bp) insert size) and 4 kb long mate-pair (LMP) libraries, and assembled using ALLPATHS-LG24. T. aestivum genome was sequenced by the Genoscope (Institut de Génomique, Evry, France) using 454 and Illumina sequencing technologies, and assembled using Newbler (Roche). Final consensus was polished as described in Aury et al.25. T. magnatum genome was sequenced by using 454 and Illumina sequencing platforms (Beckman Genomics) and a long jump insert (20 kb) mate-pair library (Eurofins), and assembled using ALLPATHS-LG and GapCloser. Gene prediction for T. melanosporum and T. aestivum was performed using the Genoscope annotation pipeline (see Martin et al.5 and Dubarry et al.26 for details), whereas gene predictions for genomes sequenced at JGI, as well as T. magnatum, were annotated using the JGI annotation pipeline (see Kohler et al.16 for details) and made available via the JGI MycoCosm database (jgi.doe.gov/fungi27). Transcriptomes of all species produced at JGI were sequenced using Illumina, assembled using Rnnotator and used for genome annotation (see Kohler et al.16 for further details). According to BUSCO (Benchmarking Universal Single-Copy Orthologs)28,29 over 94% of the fungal single-copy orthologues were found in gene repertoires (Table 1). We also checked the mappability of expressed sequence tag sequences generated by RNA-Seq analysis of full-length complementary DNA libraries, and found that >97% of them could be mapped to the assemblies, indicating a nearly complete coverage of the protein-coding gene regions.
Comparative genomic analyses
Gene family clustering, COMPARE analysis, phylogenetic analysis, annotation of functional categories, in silico analysis of VOC biosynthetic pathways, synteny analyses, RNA extraction, RNA sequencing and data analysis are described in the Supplementary Information.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Genome assemblies and gene annotations are available at JGI MycoCosm website by following these links: A. immersus v1.0, https://genome.jgi.doe.gov/Ascim1; C. venosus v1.0, https://genome.jgi.doe.gov/Chove1; M. importuna v1.0, https://genome.jgi.doe.gov/Morco1; P. confluens v1.0, https://genome.jgi.doe.gov/Pyrco1; Te. boudieri v1.1, https://genome.jgi.doe.gov/Terbo2; T. aestivum v1.0, https://genome.jgi.doe.gov/Tubae1; T. magnatum v1.0, https://genome.jgi.doe.gov/Tubma1; T. melanosporum v1.2, https://genome.jgi.doe.gov/Tubme1v2. Assemblies and annotations of T. aestivum and T. melanosporum are available at the European Nucleotide Archive under accession numbers PRJEB11238 and PRJEA38847, respectively. Other genome assemblies are archived at NCBI under the following accession numbers: A. immersus v1.0 (PRJNA346638), C. venosus v1.0 (PRJNA20409), M. importuna v1.0 (PRJNA217205), P. confluens v1.0 (PRJNA65573),Te. boudieri v1.1 (PRJNA200599) and T.magnatum v1.0 (PRJNA439374).
References
Tedersoo, L., May, T. W. & Smith, M. E. Ectomycorrhizal lifestyle in fungi: global diversity, distribution, and evolution of phylogenetic lineages. Mycorrhiza 20, 217–263 (2010).
Bonito, G. et al. Historical biogeography and diversification of truffles in the Tuberaceae and their newly identified Southern Hemisphere sister lineage. PLoS ONE 8, e52765 (2013).
Mello, A., Murat, C. & Bonfante, P. Truffles: much more than a prized and local fungal delicacy. FEMS Microbiol. Lett. 260, 1–8 (2006).
Le Tacon, F. et al. Certainties and uncertainties about the life cycle of the Périgord black truffle Tuber melanosporum. Ann. Forest Sci. 73, 105–117 (2016).
Martin, F. et al. Périgord black truffle genome uncovers evolutionary origins and mechanisms of symbiosis. Nature 464, 1033–1038 (2010).
Splivallo, R., Ottonello, S., Mello, A. & Karlovsky, P. Truffle volatiles: from chemical ecology to aroma biosynthesis. New Phytol. 189, 688–699 (2011).
Traeger, S. et al. The genome and development-dependent transcriptomes of Pyronema confluens: a window into fungal evolution. PLoS Genet. 9, e1003820 (2013).
O’Donnell, K., Cigelnik, E., Weber, N. S. & Trappe, J. M. Phylogenetic relationships among ascomycetous truffles and the true and false morels inferred from 18S and 28S ribosomal DNA sequence analysis. Mycologia 89, 48–65 (1997).
Percudani, R., Trevisi, A., Zambonelli, A. & Ottonello, S. Molecular phylogeny of truffles (Pezizales: Terfeziaceae, Tuberaceae) derived from nuclear rDNA sequence analysis. Mol. Phylogenet. Evol. 13, 169–180 (1999).
Prieto, M. & Wedin, M. Dating the diversification of the major lineages of Ascomycota (Fungi). PLoS ONE 8, e65576 (2013).
Fitzpatrick, C. R. et al. Assembly and ecological function of the root microbiome across angiosperm plant species. Proc. Natl Acad. Sci. USA 115, E1157–E1165 (2018).
Clutterbuck, J. A. Genomic evidence of repeat-induced point mutation (RIP) in filamentous ascomycetes. Fungal. Genet. Biol. 48, 306–326 (2011).
Montanini, B. et al. Non-exhaustive DNA methylation-mediated transposon silencing in the black truffle genome, a complex fungal genome with massive repeat element content. Genome Biol. 15, 10–1186 (2014).
Amselem, J., Lebrun, M.-H. & Quesneville, H. Whole genome comparative analysis of transposable elements provides new insight into mechanisms of their inactivation in fungal genomes. BMC Genomics 16, 141 (2015).
Tautz, D. & Domazet-Lošo, T. The evolutionary origin of orphan genes. Nat. Rev. Genet. 12, 692–702 (2011).
Kohler, A. et al. Convergent losses of decay mechanisms and rapid turnover of symbiosis genes in mycorrhizal mutualists. Nat. Genet. 47, 410–415 (2015).
Peter, M. et al. Ectomycorrhizal ecology is imprinted in the genome of the dominant symbiotic fungus Cenococcum geophilum. Nat. Commun. 7, 12662 (2016).
Hobbie, E. A., Weber, N. S. & Trappe, J. M. Mycorrhizal vs saprotrophic status of fungi: the isotopic evidence. New Phytol. 150, 601–610 (2001).
Drigo, B., Anderson, I. C., Kannangara, G. S. K., Cairney, J. W. G. & Johnson, D. Rapid incorporation of carbon from ectomycorrhizal mycelial necromass into soil fungal communities. Soil Biol. Biochem. 49, 4–10 (2012).
Hacquard, S. et al. Laser microdissection and microarray analysis of Tuber melanosporum ectomycorrhizas reveal functional heterogeneity between mantle and Hartig net compartments. Environ. Microbiol. 15, 1853–1869 (2013).
Vahdatzadeh, M., Deveau, A. & Splivallo, R. The role of the microbiome of truffles in aroma formation: a meta-analysis approach. Appl. Environ. Microbiol. 81, 6946–6952 (2015).
Rhounim, L., Grégoire, A., Salama, S. & Faugeron, G. Clustering of multiple transgene integrations in highly-unstable Ascobolus immersus transformants. Curr. Genet. 26, 344–351 (1994).
Roth-Bejerano, N., Livne, D. & Kagan-Zur, V. Helianthemum—Terfezia relations in different growth media. New Phytol. 114, 235–238 (2006).
Gnerre, S. et al. High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proc. Natl Acad. Sci. USA 108, 1513–1518 (2011).
Aury, J.-M. et al. High quality draft sequences for prokaryotic genomes using a mix of new sequencing technologies. BMC Genomics 9, 603 (2008).
Dubarry, M. et al. Gmove a Tool for Eukaryotic Gene Predictions Using Various Evidences (F1000Research, 2016).
Grigoriev, I. V. et al. MycoCosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res. 42, D699–D704 (2014).
Nagy, L. G. et al. Latent homology and convergent regulatory evolution underlies the repeated emergence of yeasts. Nat. Commun. 5, 4471 (2014).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with singlecopy orthologs. Bioinformatics 31, 3210–3212 (2015).
Turgeman, T. et al. The role of pre-symbiotic auxin signaling in ectendomycorrhiza formation between the desert truffle Terfezia boudieri and Helianthemum sessiliflorum. Mycorrhiza 26, 287–297 (2016).
Acknowledgements
The research was funded by the French National Research Agency through the projects SYSTRUF (grant nos. ANR-14-CE06-0020 and ANR-09-STRA-10) and SYMWOOD (Laboratory of Excellence Advanced Research on the Biology of Tree and Forest Ecosystems, grant no. ANR-11-LABX 0002 01). The work conducted by the US Department of Energy Joint Genome Institute, a DOE Office of Science User Facility, within the framework of the 1000 Fungal Genome project, was supported by the Office of Science of the US Department of Energy under contract no. DE-AC02-05CH11231. The Genoscope is partly supported by France Génomique (grant no. ANR-10-INBS-09). Computer facilities at INRA-Nancy are funded by the Region Lorraine Research Council and the European Commission (European Regional Development Fund). Financial support by the University of Parma (local funding, FIL programme) to B.M. and S.O. and by the Interuniversity Consortium for Biotechnologies to S.O., and by Piedmont Region to P.B. is also gratefully acknowledged. L.Ž. and P.Ba. were supported by the Czech Science Foundation (grant no.16–08916S). M.N. was supported by the German Research Foundation (grant no. NO407/7-1) and would like to thank U. Kück for support at the Department of General and Molecular Botany. We would like to thank the Agritruffe and Robin Nurseries companies for providing trees inoculated by T. melanosporum and T. magnatum, respectively. We are grateful to G. Chevalier, V. Gavazza and J.-P. Maurice for harvesting T. aestivum, T. magnatum and C. venosus fruiting bodies. We also thank F. Sillo and E. Zampieri for obtaining and analysing the initial RNA-seq dataset from T. magnatum fruiting body.
Author information
Authors and Affiliations
Contributions
F.M.M. and C.M. conceived, managed and coordinated the project. F.M.M., C.M., P.B., L.N., B.H., P.B., S.O., M.N., F.P. and J.S. planned and managed the research. I.V.G. supervised the sequencing and annotation at the JGI. J.M.A. and P.W. supervised the sequencing and annotation at the Genoscope. T.P., B.N., A. Kuo, J.C., A. Kohler, K.K., E.M., R.B., C.D.S., B.M., M.H., E.L., K.W.B., B.B., N. C., A.C., R.B.D., L.F., J.G., M.I., E.A.L., A.L., J.P., F.M., S.M., A.M., V.M., F.P., A.R., C.R., Y.S., R.S., S.T., M.W., L.Z., D.W. and A.Z. performed experiments or sequencing and annotation, and collected, analysed or interpreted the data; F.M.M. and C.M. drafted and wrote the manuscript. B.M., E.L., A.M., A.R., R.S., F.P., S.O., P.B., J.S., B.H., L.N., P.W., I.G. and P.B. contributed to sections of the manuscript. All authors read and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Results and Figures 1–13
Supplementary Tables
Supplementary Tables 1–15
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Murat, C., Payen, T., Noel, B. et al. Pezizomycetes genomes reveal the molecular basis of ectomycorrhizal truffle lifestyle. Nat Ecol Evol 2, 1956–1965 (2018). https://doi.org/10.1038/s41559-018-0710-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0710-4