Abstract
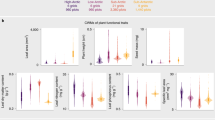
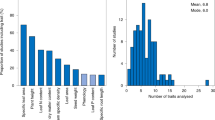
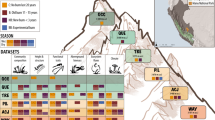
Plant functional traits directly affect ecosystem functions. At the species level, trait combinations depend on trade-offs representing different ecological strategies, but at the community level trait combinations are expected to be decoupled from these trade-offs because different strategies can facilitate co-existence within communities. A key question is to what extent community-level trait composition is globally filtered and how well it is related to global versus local environmental drivers. Here, we perform a global, plot-level analysis of trait–environment relationships, using a database with more than 1.1 million vegetation plots and 26,632 plant species with trait information. Although we found a strong filtering of 17 functional traits, similar climate and soil conditions support communities differing greatly in mean trait values. The two main community trait axes that capture half of the global trait variation (plant stature and resource acquisitiveness) reflect the trade-offs at the species level but are weakly associated with climate and soil conditions at the global scale. Similarly, within-plot trait variation does not vary systematically with macro-environment. Our results indicate that, at fine spatial grain, macro-environmental drivers are much less important for functional trait composition than has been assumed from floristic analyses restricted to co-occurrence in large grid cells. Instead, trait combinations seem to be predominantly filtered by local-scale factors such as disturbance, fine-scale soil conditions, niche partitioning and biotic interactions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data contained in sPlot (the vegetation-plot data complemented by trait and environmental information) are available on request, by contacting any of the sPlot consortium members, for submission of a paper proposal. The proposals should follow the Governance and Data Property Rules of the sPlot Working Group, which are available on the sPlot website (www.idiv.de/sPlot).
References
Warming, E. Lehrbuch der ökologischen Pflanzengeographie – Eine Einführung in die Kenntnis der Pflanzenvereine (Borntraeger, Berlin, 1896).
Garnier, E. et al. Plant functional markers capture ecosystem properties during secondary succession. Ecology 85, 2630–2637 (2004).
Ordoñez, J. C. A global study of relationships between leaf traits, climate and soil measures of nutrient fertility. Glob. Ecol. Biogeogr. 18, 137–149 (2009).
Garnier, E., Navas, M.-L. & Grigulis, K. Plant Functional Diversity – Organism Traits, Community Structure, and Ecosystem Properties (Oxford Univ. Press, Oxford, 2016).
Díaz, S. et al. The global spectrum of plant form and function. Nature 529, 167–171 (2016).
Moles, A. T. et al. Global patterns in plant height. J. Ecol. 97, 923–932 (2009).
Wright, I. J. et al. The worldwide leaf economics spectrum. Nature 428, 821–827 (2004).
Reich, P. B. The world-wide ‘fast–slow’ plant economics spectrum: a traits manifesto. J. Ecol. 102, 275–301 (2014).
Adler, P. B. et al. Functional traits explain variation in plant life history strategies. Proc. Natl Acad. Sci. USA 111, 740–745 (2014).
Marks, C. O. & Lechowicz, M. J. Alternative designs and the evolution of functional diversity. Am. Nat. 167, 55–67 (2006).
Grime, J. P. Trait convergence and trait divergence in herbaceous plant communities: mechanisms and consequences. J. Veg. Sci. 17, 255–260 (2006).
Muscarella, R. & Uriarte, M. Do community-weighted mean functional traits reflect optimal strategies?. Proc. R. Soc. B 283, 20152434 (2016).
Swenson, N. G. & Weiser, M. D. Plant geography upon the basis of functional traits: an example from eastern North American trees. Ecology 91, 2234–2241 (2010).
Fyllas, N. M. et al. Basin-wide variations in foliar properties of Amazonian forest: phylogeny, soils and climate. Biogeosciences 6, 2677–2708 (2009).
Swenson, N. G. et al. Phylogeny and the prediction of tree functional diversity across novel continental settings. Glob. Ecol. Biogeogr. 26, 553–562 (2017).
Swenson, N. G. et al. The biogeography and filtering of woody plant functional diversity in North and South America. Glob. Ecol. Biogeogr. 21, 798–808 (2012).
Wright, I. J. et al. Global climatic drivers of leaf size. Science 357, 917–921 (2017).
Mayfield, M. M. & Levine, J. M. Opposing effects of competitive exclusion on the phylogenetic structure of communities. Ecol. Lett. 13, 1085–1093 (2010).
Kraft, N. J. B. et al. Community assembly, coexistence and the environmental filtering metaphor. Funct. Ecol. 29, 592–599 (2015).
Barboni, D. et al. Relationships between plant traits and climate in the Mediterranean region: a pollen data analysis. J. Veg. Sci 15, 635–646 (2004).
Borgy, B. et al. Plant community structure and nitrogen inputs modulate the climate signal on leaf traits. Glob. Ecol. Biogeogr. 26, 1138–1152 (2017).
van Bodegom, P. M., Douma, J. C. & Verheijen, L. M. A fully traits-based approach to modeling global vegetation distribution. Proc. Natl Acad. Sci. USA 111, 13733–13738 (2014).
Moles, A. T. et al. Which is a better predictor of plant traits: temperature or precipitation? J. Veg. Sci. 25, 1167–1180 (2014).
Ordoñez, J. C. et al. Plant strategies in relation to resource supply in mesic to wet environments: does theory mirror nature?. Am. Nat. 175, 225–239 (2010).
Simpson, A. J., Richardson, S. J. & Laughlin, D. C. Soil–climate interactions explain variation in foliar, stem, root and reproductive traits across temperate forests. Glob. Ecol. Biogeogr. 25, 964–978 (2016).
Lienin, P. & Kleyer, M. Plant leaf economics and reproductive investment are responsive to gradients of land use intensity. Agric. Ecosyst. Environ. 145, 67–76 (2011).
Maire, V. et al. Habitat filtering and niche differentiation jointly explain species relative abundance within grassland communities along fertility and disturbance gradients. New Phytol. 196, 497–509 (2012).
Craine, J. M. et al. Global patterns of foliar nitrogen isotopes and their relationships with climate, mycorrhizal fungi, foliar nutrient concentrations, and nitrogen availability. New Phytol. 183, 980–992 (2009).
Güsewell, S. N:P ratios in terrestrial plants: variation and functional significance. New Phytol. 164, 243–266 (2004).
Reich, P. B. & Oleksyn, J. Global patterns of plant leaf N and P in relation to temperature and latitude. Proc. Natl Acad. Sci. USA 101, 11001–11006 (2004).
Scheiter, S., Langan, L. & Higgins, S. I. Next generation dynamic global vegetation models: learning from community ecology. New Phytol. 198, 957–969 (2013).
Boyle, B. et al. The Taxonomic Name Resolution Service: an online tool for automated standardization of plant names. BMC Bioinform. 14, 16 (2013).
Bremer, B. et al. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG III. Bot. J. Linn. Soc. 161, 105–121 (2009).
Schrodt, F. et al. BHPMF – a hierarchical Bayesian approach to gap-filling and trait prediction for macroecology and functional biogeography. Glob. Ecol. Biogeogr. 24, 1510–1521 (2015).
Kattge, J. et al. TRY ‒ a global database of plant traits. Glob. Change Biol. 17, 2905–2935 (2011).
Shan, H. et al. Gap filling in the plant kingdom – trait prediction using hierarchical probabilistic matrix factorization. In Proc. 29th Int. Conf. Machine Learning (ICML 2012) 1303−1310 (Omnipress, Madison, 2012).
Fazayeli, F. et al. Uncertainty quantified matrix completion using Bayesian Hierarchical Matrix factorization. In Proc. 13th Int. Conf. Machine Learning and Applications (ICMLA 2014) 312−317 (Institute of Electrical and Electronics Engineers, Danvers, 2014).
Borgy, B. et al. Sensitivity of community-level trait–environment relationships to data representativeness: a test for functional biogeography. Glob. Ecol. Biogeogr. 26, 729–739 (2017).
Herz, K. et al. Drivers of intraspecific trait variation of grass and forb species in German meadows and pastures. J. Veg. Sci. 28, 705–716 (2017).
Karger, D. N. et al. Climatologies at high resolution for the Earth’s land surface areas. Sci. Data 4, 170122 (2017).
Karger, D. N. et al. Climatologies at High Resolution for the Earth Land Surface Areas (Version 1.1) (World Data Center for Climate (WDCC) at DKRZ, 2016); http://chelsa-climate.org/downloads/
Dee, D. P. et al. The ERA-Interim reanalysis: configuration and performance of the data assimilation system. Q. J. R. Meteorol. Soc 137, 553–597 (2011).
Synes, N. W. & Osborne, P. E. Choice of predictor variables as a source of uncertainty in continental-scale species distribution modelling under climate change. Glob. Ecol. Biogeogr. 20, 904–914 (2011).
Enquist, B. et al. Scaling from traits to ecosystems: developing a general trait driver theory via integrating trait-based and metabolic scaling theories. Adv. Ecol. Res. 52, 249–318 (2015).
Buzzard, V. et al. Re-growing a tropical dry forest: functional plant trait composition and community assembly during succession. Funct. Ecol. 30, 1006–1013 (2016).
Rao, C. R. Diversity and dissimilarity coefficients: a unified approach. Theor. Popul. Biol. 21, 24–43 (1982).
Champely, S. & Chessel, D. Measuring biological diversity using Euclidean metrics. Environ. Ecol. Stat 9, 167–177 (2002).
Dowle, M. et al. data.table: Extension of data.frame. R Package Version 1.9.6 (2015); https://CRAN.R-project.org/package=data.table
Hawkins, B. A. et al. Structural bias in aggregated species-level variables driven by repeated species co-occurrences: a pervasive problem in community and assemblage data. J. Biogeogr. 44, 1199–1211 (2017).
Knijnenburg, T. A. et al. Fewer permutations, more accurate P-values. Bioinformatics 25, i161–i168 (2009).
Friendly, M. Corrgrams: exploratory displays for correlation matrices. Am. Stat. 56, 316–324 (2002).
Oksanen, J. et al. vegan: Community Ecology Package. R Package Version 2.3-3 (2016); https://CRAN.R-project.org/package=vegan
Lengyel, A., Chytrý, M. & Tichý, L. Heterogeneity-constrained random resampling of phytosociological databases. J. Veg. Sci 22, 175–183 (2011).
Baselga, A. The relationship between species replacement, dissimilarity derived from nestedness, and nestedness. Glob. Ecol. Biogeogr. 21, 1223–1232 (2012).
Garnier, E. et al. Towards a thesaurus of plant characteristics: an ecological contribution. J. Ecol. 105, 298–309 (2017).
Acknowledgements
The sPlot was initiated by sDiv, the Synthesis Centre of the German Centre for Integrative Biodiversity Research (iDiv) Halle-Jena-Leipzig, funded by the German Research Foundation (FZT 118) and is now a platform of iDiv. H.B., J.De., O.Pu, U.J., B.J.-A., J.K., D.C., F.M.S., M.W. and C.W. appreciate the direct funding through iDiv. For all further acknowledgements see the Supplementary Information.
Author information
Authors and Affiliations
Contributions
H.B. and U.J. wrote the first draft of the manuscript, with considerable input by B.J.-A. and R.F. Most of the statistical analyses and the production of the graphs were carried out by H.B.. H.B., O.Pu. and U.J. initiated sPlot as an sDiv working group and iDiv platform. J.De. compiled the plot databases globally. J.De., S.M.H., U.J., O.Pu. and F.J. harmonized vegetation databases. J.De. and B.J.-A. coordinated the sPlot consortium. J.K. provided the trait data from TRY. F.S. performed the trait data gap filling. O.Pu. produced the taxonomic backbone. B.J.-A., G.S. and E. Welk compiled environmental data and produced the global maps. S.M.H. wrote the Turboveg v3 software, which holds the sPlot database. J.L. and T.H. wrote the resampling algorithm. Many authors participated in one or more of the three sPlot workshops at iDiv where the sPlot initiative was conceived and planned, and evaluation of the data and first drafts were discussed. All other authors contributed data. All authors contributed to writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2, Supplementary Figures 1–11, and Supplementary Acknowledgements
Rights and permissions
About this article
Cite this article
Bruelheide, H., Dengler, J., Purschke, O. et al. Global trait–environment relationships of plant communities. Nat Ecol Evol 2, 1906–1917 (2018). https://doi.org/10.1038/s41559-018-0699-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0699-8
This article is cited by
-
Shifts in plant resource use strategies across climate and soil gradients in dryland steppe communities
Plant and Soil (2024)
-
Global multifaceted biodiversity patterns, centers, and conservation needs in angiosperms
Science China Life Sciences (2024)
-
A taxonomic snapshot of belowground organs in plants of Anatolian steppes
Folia Geobotanica (2024)
-
Plant Communities in Changing Environment
Biologia (2024)
-
A slow-fast trait continuum at the whole community level in relation to land-use intensification
Nature Communications (2024)