Abstract
Modern cartilaginous fishes are divided into elasmobranchs (sharks, rays and skates) and chimaeras, and the lack of established whole-genome sequences for the former has prevented our understanding of early vertebrate evolution and the unique phenotypes of elasmobranchs. Here we present de novo whole-genome assemblies of brownbanded bamboo shark and cloudy catshark and an improved assembly of the whale shark genome. These relatively large genomes (3.8–6.7 Gbp) contain sparse distributions of coding genes and regulatory elements and exhibit reduced molecular evolutionary rates. Our thorough genome annotation revealed Hox C genes previously hypothesized to have been lost, as well as distinct gene repertories of opsins and olfactory receptors that would be associated with adaptation to unique underwater niches. We also show the early establishment of the genetic machinery governing mammalian homoeostasis and reproduction at the jawed vertebrate ancestor. This study, supported by genomic, transcriptomic and epigenomic resources, provides a foundation for the comprehensive, molecular exploration of phenotypes unique to sharks and insights into the evolutionary origins of vertebrates.
Similar content being viewed by others
Main
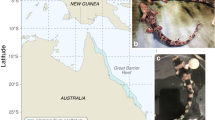
Cartilaginous fishes (Chondrichthyes) are divided into two subclasses, elasmobranchs (Elasmobranchii, including sharks, rays and skates) and chimaeras (Holocephali), and their common ancestor diverged from the rest of jawed vertebrates around 450 million years ago. More than a decade ago, the elephant fish, Callorhinchus milii, a member of the Holocephali that comprises approximately 50 species, was chosen for whole-genome sequencing because of its small genome size1. Since then, molecular comparative studies on vertebrates have largely relied on the C. milii genomic sequences as representative of cartilaginous fishes2, but the low fecundity and accessibility of live specimens have been a limitation. C. milii is often referred to as elephant ‘shark’ (or ghost ‘shark’), but true sharks belong to the subclass Elasmobranchii that comprises approximately 1,200 species. For elasmobranchs, however, no reliable genome-wide sequence resource allowing extensive molecular analyses has been established to date, in spite of some attempts3,4. Thus, there is an important need to obtain genomic information of elasmobranchs that will contribute to the elucidation of the molecular mechanisms underlying their unique traits of morphology, reproduction, sensing and longevity5, as well as thorough demographic analyses for conservation6,7. Here we report whole-genome analysis of three elasmobranch species (Fig. 1a–c), assisted by phylogenetics-oriented genome informatics. The utility of genome, transcriptome and epigenome data of prolific egg-laying (‘oviparous’) species provided by this study should expand the capacity for in-depth molecular investigation on elasmobranchs.
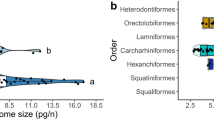
a–c, Species analysed in this study. d, Maximum-likelihood tree for vertebrate phylogeny using 475,700 residues in 935 orthologues. Bootstrap values were 100 for all nodes. e, Number of synonymous substitutions per site (KS) and evolutionary ages for selected branches (see Supplementary Note 12). The purple dots indicate the branches b1–b4 in d. f, Genome size and intron length. The correlation was confirmed with phylogenetically independent contrasts (for details, see Supplementary Note 10). Abbreviated species identifiers are included in d. g, Counts of orthologue groups containing duplicated genes only in either chondrichthyans (yellow), zebrafish (cyan) or human (magenta). TSGD, teleost fish-specific genome duplication. h, Breakdown of the whole-genome assemblies into different segments.
Results
Sequencing the large genomes of sharks
We focused on the brownbanded bamboo shark Chiloscyllium punctatum, for which we recently tabled embryonic stages8, and the cloudy catshark Scyliorhinus torazame. Their whole genomes, measured to be approximately 4.7 and 6.7 Gbp, respectively, were sequenced de novo to obtain assemblies including megabase-long scaffolds (Supplementary Note 1.1). We also assembled the genome of the whale shark Rhincodon typus using short sequence reads previously generated3 (Supplementary Note 1.2). Using these genome assemblies, we performed genome-wide gene prediction, assisted by transcript evidence and protein-level homology to other vertebrates. The obtained genome assemblies and gene models exhibit high coverage (Supplementary Fig. 1), and of these, the bamboo shark genome assembly achieved the highest continuity (N50 scaffold length, 1.9 Mbp) and completeness (97% of reference orthologues identified at least partially). Using the novel gene models, we constructed orthologue groups encompassing a diverse array of vertebrate species (see below). Our products outperform existing resources for elasmobranchs and provide the tools for genome-wide characterization of molecular evolution at the origin of jawed vertebrates and later in the chondrichthyan lineages.
Genome-wide trends in molecular evolution
We first examined genome-wide trends of molecular evolution, utilizing one-to-one orthologues in the constructed orthologue groups (Supplementary Note 7). Our comparisons of coding sequences detected a higher similarity in nucleotide and amino acid compositions of sharks to tetrapods and coelacanth than to actinopterygian fishes (Supplementary Fig. 2a,c). We performed a phylogenomic analysis using conserved protein-coding genes, which confirmed the phylogenetic positions of elasmobranchs and the reduced rate of molecular evolution in the entire chondrichthyan lineage (Fig. 1d). The reduced evolutionary rate was further scrutinized by comparing the numbers of synonymous substitutions per site (KS) between chondrichthyan and osteichthyan lineages (Fig. 1e). The result revealed that synonymous substitution rates for the chondrichthyan lineages were significantly smaller than those for almost all the osteichthyan lineages analysed (Supplementary Note 12), suggesting a reduced intrinsic mutation rate in the chondrichthyan lineages. Our cross-species comparison revealed a remarkable increase in the intron lengths of shark genomes and its correlation with genome size (Fig. 1f and Supplementary Note 10). Our analysis on the composition of orthologue groups did not detect massive gene duplications in the chondrichthyan lineage (Fig. 1g), which was supported by the inference of age distribution of paralogues (Supplementary Note 11). Thus, the increase of genome size in sharks is not attributable to additional whole-genome duplication.
Characterizing noncoding landscape
To characterize noncoding regions, we first scanned elasmobranch genomes for repetitive sequences including those unique to the species analysed (Supplementary Note 4). This identified long interspersed nuclear elements as the most abundant class of repetitive elements, exceeding the proportions of long terminal repeats and those unclassified into any existing repetitive element class, both of which were particularly expanded in the catshark (Fig. 1h). Overall, the genomic regions identified as repetitive elements, including simple repeats, amounted to half of the individual elasmobranch genome assemblies, and their abundance contributed to the observed variation in genome size (Fig. 1h).
Next, we surveyed the elasmobranch genomes for homologues of human conserved noncoding elements (CNEs), which yielded a much larger number of matches than in teleost genomes (Fig. 2a and Supplementary Note 13). Our analysis revealed some CNEs retained by elasmobranchs but missing in teleost fish and C. milii, which included a CNE in an intron of the Tbx4 gene (Fig. 2b) previously reported as the core lung mesenchyme-specific enhancer9. Its presence in a cartilaginous fish that lacks a lung homologue prompts a reexamination of its evolutionary significance. This finding also highlights the problem of using only a single holocephalan species as a representative of chondrichthyans, whether the CNEs missing in this species are lost during evolution or masked in gaps in the genome assembly.
a, Number of the retained CNEs identified by the human–chicken genome comparison. b, Elasmobranch homologue of the lung mesenchyme-specific enhancer (LME) in an intron of the mammalian Tbx4 gene. c, Expression profiles of the putative bamboo shark homologues of human lncRNAs expressed in at least one analysed tissue. See Supplementary Notes 13 and 14 for details.
We also searched for elasmobranch homologues of human long noncoding RNAs (lncRNAs), which again revealed more candidate homologues than in teleost fishes (Supplementary Note 14). These were screened for transcript evidence in bamboo shark RNA-seq data and absence of homology to coding sequences. This screening resulted in the identification of 38 transcript contigs with variable degrees of spatial expression biases (Fig. 2c). These putative lncRNAs included a possible homologue of the Malat1 gene10[,11, whose presence in chondrichthyans was recently suggested only by a sequence similarity to a C. milii genomic region12. The inclusion of the putative bamboo shark Malat1 homologue in our result validates our screening procedure and more importantly, ascertains its noncoding transcription in a chondrichthyan species.
Overall, these findings indicate that despite the variable genome sizes and repetitive element compositions, elasmobranch genomes have undergone less modification in noncoding regions involved in gene regulation since the jawed vertebrate ancestor than is inferred by their evolutionary distance.
Evolution of Hox genes and clusters
Hox genes play crucial roles in embryogenesis and are organized into four clusters (Hox A–D) in osteichthyans (bony vertebrates) except for teleost fishes13. In the shark genomes, we found well-conserved Hox A, B and D clusters, which have identical gene repertories to their C. milii counterparts (Fig. 3a and Supplementary Fig. 5a) expressed in a temporally collinear manner (Fig. 3d). For a comparison of conformational regulation of Hox gene expression by CCCTC-binding factor (CTCF)14, we analysed the distribution of its binding sites with ChIP-seq (Methods and Supplementary Note 15). This comparison on the Hox A, B and D clusters revealed a high similarity between elasmobranchs and amniotes (Fig. 3b), which makes the whole gnathostomes including elasmobranchs distinct from the lamprey that has more CTCF binding sites within Hox clusters15 (Supplementary Fig. 4e). It is thus suggested that the jawed vertebrate ancestor already possessed the mechanism of CTCF-dependent conformational regulation of Hox genes documented for mammals16,17. We identified antisense transcripts in the genomic region of elasmobranchs containing Hoxa11 and -a13 as putative homologues of lncRNAs previously known only in tetrapods, Hoxa11-AS and Hottip (Fig. 3c and Supplementary Note 14). Although the acquisition of Hoxa11-AS is proposed to be linked with the fin-to-limb evolution18, our discovery of the elasmobranch counterparts indicates their early origins in the common ancestor of jawed vertebrates.
a, Structure of the bamboo shark Hox clusters. Coloured boxes denote coding exons of the Hox genes (see Supplementary Fig. 5m for scaffold IDs for Hox C). b, CTCF ChIP-seq peaks in the Hox A cluster for the bamboo shark and the chicken (for details, see Supplementary Note 16.1). c, Putative bamboo shark homologues of mammalian lncRNA, Hottip and HoxA11-AS revealed by RNA-seq read mapping for a stage 28 bamboo shark embryo. d, Temporal expression patterns of Hox A genes in whole catshark embryos. e, Molecular phylogeny of Hox11 inferred using 147 residues. f–h, Embryonic expression of catshark Hox11 genes at stage 27. Scale bars, 500 μm. i, Expression timings of the Hox11 genes in whole catshark embryos. j, Repetitiveness of the whole genomes and the Hox-containing genomic regions (see Methods and Supplementary Note 16.4 for details). The Hoxb10-b13 region was excluded. k, Putative Hox C gene repertories of diverse elasmobranchs. The ocellate spot skate Okamejei kenojei and zebra bullhead shark Heterodontus zebra genes were identified in transcriptome data23,24. Closed pink boxes, presence of intact genes; grey boxes, pseudogenization indicated by a stop codon inside the homeobox (see Supplementary Fig. 5c). The genes connected by a horizontal line form a tandem cluster on a genome scaffold.
Although the entire Hox C cluster was reportedly missing from elasmobranch genomes19,20, we identified putative Hox C genes in the genome and transcript sequences of the analysed shark species (Fig. 3a,e and Supplementary Fig. 5a–c). While our phylogenetic analyses supported their affiliations to Hox C, those genes showed extremely elevated evolutionary rates. Remarkably, none of the identified, putative shark Hox C genes comprised such a compact cluster as in other jawed vertebrates, spanning within a 100-kbp-long genomic region13—for example, catshark Hoxc11 was flanked by a 50-kbp-long stretch containing no other Hox gene (Supplementary Fig. 5a). In addition, although typical jawed vertebrate Hox clusters are almost free from repetitive elements21 (at most 2.9% in length for elasmobranch Hox A, B and D clusters; Fig. 3a,j), the elasmobranch genome scaffolds containing the putative Hox C genes have accumulated repetitive elements (at least 36.8%; Fig. 3j).
Our analysis on embryonic expression patterns indicated that the identified elasmobranch Hox C genes are still under spatiotemporal transcriptional regulation, which is typically exerting on clustered Hox genes13,22―Hoxc11 (Fig. 3f–i and Supplementary Fig. 5j–l) as well as Hoxc8 (Supplementary Fig. 5g–i) are expressed in the posterior regions concomitantly with their sister paralogues in the Hox A, B and D clusters. We further surveyed the transcriptome data of other elasmobranch species23,24, which uncovered more Hox C genes of the zebra bullhead shark (Fig. 3k). These findings demonstrate that Hox C genes were not lost in a cluster-wide deletion event in the elasmobranch ancestor as proposed previously19, but have eroded intermittently during elasmobranch evolution. Together, while the Hox A, B and D clusters exhibit the canonical, conservative nature even in elongated elasmobranch genomes, their Hox C cluster underwent remarkable lineage-specific, sequence-level modifications.
Encompassing genes secondarily lost in osteichthyan lineages
The constructed orthologue groups contained 304 genes that seemingly existed in the vertebrate ancestor are retained by elasmobranchs, but have disappeared in osteichthyan evolution (Supplementary Table 8). They included a member of the Fox gene family (designated as FoxG3) whose orthologues are retained only by non-tetrapod vertebrates (Fig. 4a). One of its close paralogues, FoxG1, functions as a key regulator of forebrain development in diverse animals25,26. The third paralogue, designated as FoxG2, was identified in only non-tetrapod vertebrates and some reptiles (Fig. 4a). Molecular phylogenetic analysis showed the triplication between FoxG1, -G2 and -G3 early in vertebrate evolution and among-lineage differential gene loss (Fig. 4a and Supplementary Note 17).
a, Molecular phylogeny of the FoxG genes using 192 residues. b, Comparison of characteristics in protein-coding and flanking sequences between the three FoxG genes (see Supplementary Note 17). Two left panels: the numbers of non-synonymous and synonymous substitutions per site (KA and KS, respectively) of bamboo shark and catshark orthologue pairs that were calculated with codeml in PAML v4.9c88. Three right panels: the values of the bamboo shark genes. c, Variable conservation levels of coding (grey) and flanking noncoding (orange) genomic regions of FoxG1, -G2 and -G3 between catshark (used as a reference) and bamboo shark. d–f, Embryonic expression of the three catshark FoxG genes at stage 24 (see Supplementary Fig. 6c–p for expression patterns at stage 27). VII+VIII, acoustico-facial ganglionic complex; X, vagal ganglion; tel, telencephalon; ret, retina; ot, otic vesicle; GC4, GC-content at four-fold degenerate sites. Scale bars, 500 µm.
While the determinant of loss or retention of gene duplicates is often imputed to their functions27, the effect of intrinsic genomic characteristics, independent of gene functions, has also been proposed as a cause28. As the less-derived shark genomes are expected to better reconstruct the differentiation process of ancient gene duplicates, we performed a multi-faceted comparison focusing on the shark FoxG paralogues. A highly conserved nature of FoxG1, retained in all the species analysed to date, was observed in not only its amino acid sequence but also in synonymous nucleotide and flanking noncoding genome sequences (Fig. 4b,c). This coding/noncoding association was detected for the divergent nature of FoxG2, while the level of sequence conservation of FoxG3 was intermediate (Fig. 4b,c). The among-paralogue variation was also observed in the GC-content of fourfold degenerate sites (GC4; Fig. 4b and Supplementary Table 15). More remarkably, the flanking sequences of the most divergent paralogue FoxG2 contain the most abundant repetitive elements and the highest GC-content (Fig. 4b). These local genomic characteristics may have facilitated the secondary loss of the coding genes embedded in the divergent genomic regions.
Taking advantage of the access to embryonic samples, we analysed the spatial distribution of catshark FoxG gene expression during development (Fig. 4d–f and Supplementary Fig. 6c–p). FoxG2 was expressed in the acoustico-facial ganglionic complex (VII+VIII) and the vagal ganglion (X) (Fig. 4e). FoxG3 was expressed in an anterodorsal part of the retina, in addition to the FoxG2-positive domains (Fig. 4f), while FoxG1 expression was observed in the forebrain in addition to the FoxG3-positive domains (Fig. 4d). Together, the more prone a FoxG paralogue is to secondary loss, the more restricted is its expression domain in shark embryos. This among-paralogue comparison, enabled by the genomic resource of an egg-laying shark, confirms the association of the fates of gene duplicates with the variable natures of genomic regions containing those duplicates.
Early invention of homoeostatic machinery for gut–brain axis
To further characterize phenotypic traits refined in jawed vertebrates, we focused on gene repertories encoding endocrine hormones and their receptors that control growth, reproduction and homoeostasis. Our phylogenetic census in shark genomes and transcriptomes revealed potential orthologues of hormone and receptor genes previously unidentified in this taxon (Fig. 5a). These included prolactin (PRL1), orexin, kisspeptin, spexin, motilin and prolactin receptor implicated in fertility, appetite, digestion and sleep in mammals29,30,31, as well as osmolarity and gastrointestinal control in teleost fishes (Supplementary Note 18). For leptin whose putative orthologue was previously identified in a genomic sequence of C. milii32, we confirmed the orthology of the C. milii and elasmobranch genes to osteichthyan leptin genes, by means of molecular phylogeny and conserved synteny (Supplementary Note 18.10). Of these hormone and receptor genes, all but leptin were suggested to have existed in the vertebrate ancestor by the presence of possible cyclostome orthologues or gene duplication that probably occurred in genome expansion before the divergence of all extant vertebrates33 (Fig. 5a). This inference marks leptin, a key metabolic and neuroendocrine regulator in mammals34, and the signalling cascade through its receptor (LepR) as an invention in the jawed vertebrate lineage (Supplementary Note 18.10). In mammals, leptin is mainly expressed in adipose tissues35, and we could not identify any tissue with intensive expression of its orthologue in sharks that generally lack overt adipose tissues36 (Fig. 5b).
a, Cross-species comparison of peptide hormone gene repertories. The numbers of paralogues are indicated in the boxes. Green boxes, the genes whose orthology was inferred; beige boxes, those with ambiguous orthology; red rectangles, the members of the same gene families (see Supplementary Note 18). b, Expression profiles of hormone genes in an embryo and adult tissues of the catshark (see Supplementary Fig. 7a for a full heatmap). PAI (phasitocin) and ASP (aspargtocin) are two forms of oxytocin that are encoded by distinct loci38. PRL, prolactin; GALP, galanin-like peptide.
Overall, we identified the orthologues of almost all hormones and their receptors involved in the hypothalamo–pituitary and gastrointestinal systems documented mainly in mammals (Fig. 5a and Supplementary Note 18). Among these, the genes encoding oxytocin homologues have undergone a unique gene duplication in the elasmobranch lineage37,38, and our genomic and phylogenetic analysis indicated its intricate evolutionary history through intermittent gene conversions (Supplementary Note 18.3). The similarity in transcript localization of the identified shark hormone and receptor genes to mammalian counterparts, such as PRL1 in the pituitary and motilin in the intestine (Fig. 5b and Supplementary Fig. 7a), suggests the establishment of genetic components of the gut–brain axis39 before the last common ancestor of extant jawed vertebrates.
Sensory and neuronal gene repertories
Visual opsin gene repertories are often altered on adaptation to new habitats with dim light40,41. Previously, two short wavelength-sensitive opsin genes, SWS1 and SWS2, were found to be missing in the C. milii genome42. Our search in the elasmobranch genome assemblies ascertained the absence of not only these two but also the green/blue-sensitive opsin gene Rh2 (Fig. 6 and Supplementary Fig. 8). Moreover, long wavelength-sensitive opsin gene (LWS) is absent from the present cloudy catshark genome assembly, and thus rhodopsin (RHO) is the only visual opsin gene identified in it (Fig. 6 and Supplementary Note 19). Previously, the retention of only RHO was reported in some animals that adapted to fossorial, nocturnal or aquatic life43,44,45. In fact, the cloudy catshark inhabits not only inshore but also the deep sea46 (~300 m) and is a close relative of typical deep-sea dwellers47. Thus, the absence of LWS might be due to an evolutionary gene loss that was permitted in the catshark ancestor by its possible exclusive deep-sea habitat (Fig. 6). Adaptation to the deep sea, into which only blue light penetrates48, was previously corroborated for ray-finned fishes by the blueshifted absorption spectra of RHO pigments49. Our spectroscopic analysis of the RHO pigments revealed blueshifted spectra for not only the cloudy catshark (λmax, 484 nm) but also the whale shark (478 nm) that occasionally migrates down to the bathypelagic zone (~2,000 m) besides daytime surface feeding habits50 (Fig. 6). This study portrays the diversity of visual opsin gene repertories among elasmobranchs and illustrates the potential of in vitro molecular experiments supported by genomic sequence analysis, in understanding underwater ecology of inaccessible species.
Evolutionary history of visual opsin gene loss and duplication, including the duplicated C. milii LWS42, was suggested by phylogenetic analysis (Supplementary Fig. 8a). Dashed blank boxes indicate the absences of orthologues in the currently available genome assemblies. See Supplementary Note 19.1, for details of the habitat depths and the absorption spectra obtained by our spectroscopic analysis.
Our interest extended to olfactory receptor gene repertories that are often linked to adaptation to new lifestyles51. Although our present study does not include carnivorous epipelagic sharks that might have enhanced olfactory sensing, each of the shark species examined in the present study had only three olfactory receptor family genes (Supplementary Note 21), concordantly with the retention of few olfactory receptor family members by C. milii52. This finding indicates that at least the analysed shark species rely on a distinct molecular mechanism for olfaction from the conventional olfactory receptors.
Neuronal cell identities in mammalian brains are defined by the combinatorial expression of clustered protocadherin (Pcdh) genes53,54. Previously, the C. milii genome was shown to also contain a cluster of Pcdh genes55, but their expression profiles have remained unknown. Our study showed that elasmobranchs contain slightly higher numbers of Pcdh genes in markedly longer clusters than osteichthyans (Supplementary Fig. 9a,b and Supplementary Note 20). The bamboo shark transcriptome data demonstrated that most of the clustered Pcdh genes consist of both variable and constant exons and exhibit a biased expression pattern towards neural tissues, as previously shown for mammalian and teleost counterparts (Supplementary Fig. 9c). These findings, which are expected to be reinforced by single-cell analysis, suggest the early establishment of the mechanism for generating neuronal cell diversity through a Pcdh cluster in the last common ancestor of all extant jawed vertebrates.
Discussion
Our study has provided an unprecedented set of genomic, transcriptomic and epigenomic data from three elasmobranch species, with the bamboo shark genome assembly achieving the highest continuity. We focused on unthreatened, oviparous species that allow captive breeding for continuous animal experimentation including embryonic operation. This is not feasible with other non-tetrapod vertebrates whose genomes are evolving relatively slowly, such as the coelacanths and the spotted gar. It would be intriguing to further explore the possible relationship of the large genome/gene sizes and low evolutionary rate of elasmobranchs with metabolic rate and/or longevity. Our results also highlighted some genomic elements retained by elasmobranchs but missing in holocephalans possibly because of the genome compaction in the latter lineage. Elasmobranchs have scarce repertories of opsin and olfactory receptor genes, possibly associated with their unique niche. Our study suggested that the jawed vertebrate ancestor was already equipped with the mechanism for generating neuronal cell diversity as well as the hormone gene repertories regulating homoeostasis and reproduction in mammals. Also, we showed that elasmobranchs have retained at least parts of the Hox C cluster, in which relict Hox C genes are under the typical Hox-like regulation in spite of relaxed genomic constraint on them. Our products will fuel diverse life science studies on sharks and evolutionary investigation about early vertebrates.
Methods
Animals
All samples of the brownbanded bamboo shark Chiloscyllium punctatum were supplied by captive breeding at the Osaka Aquarium Kaiyukan. Samples of the cloudy catshark Scyliorhinus torazame were supplied by captive breeding at the Aquarium Facility of RIKEN Center for Developmental Biology and Atmosphere and Ocean Research Institute of University of Tokyo. The developmental staging was performed according to existing literature for small-spotted catshark S. canicula56 and our original table for C. punctatum8. For the whale shark Rhincodon typus, whose genome sequence reads were publicly available2, only transcriptome sequencing and genome size estimation were performed in the present study, using blood sampled primarily for the purpose of regular health check-ups for captive animals from a male at the Okinawa Churaumi Aquarium (for transcriptome sequencing) and a female at the Osaka Aquarium Kaiyukan (for genome size estimate with flow cytometry), respectively. No wildlife was killed solely for this study. Animal handling and sample collections at the aquaria were conducted by veterinary staff without restraining the individuals57, in accordance with the Husbandry Guidelines approved by the Ethics and Welfare Committee of Japanese Association of Zoos and Aquariums. All other experiments were conducted in accordance with the Guideline of the Institutional Animal Care and Use Committee (IACUC) of RIKEN Kobe Branch (Approval ID: H16-11) or the Guideline for Care and Use of Animals at the University of Tokyo.
Genome sequencing and assembly
Genomic DNA was extracted from the liver of a 20-cm-long male juvenile brownbanded bamboo shark and a 4-cm-long whole cloudy catshark embryo of an unknown sex with phenol/chloroform as previously described58. The extracted genomic DNA was sheared with a S220 Focused-ultrasonicator (Covaris) to retrieve DNA fragments of variable length distributions (see Supplementary Table 1 for detailed amounts of starting DNA and conditions for shearing). The sheared DNA was used for paired-end library preparation with a KAPA LTP Library Preparation Kit (KAPA Biosystems). The optimal numbers of PCR cycles for individual libraries were determined with a Real-Time Library Amplification Kit (KAPA Biosystems) by preliminary qPCR-based quantification using an aliquot of adaptor-ligated DNAs. Small molecules in the prepared libraries were removed by size selection using Agencourt AMPure XP (Beckman Coulter). The numbers of PCR cycles and conditions of size selection for individual libraries are included in Supplementary Table 1. Mate-pair libraries were prepared using a Nextera Mate Pair Sample Prep Kit (Illumina), employing our customized iMate protocol59 (http://www.clst.riken.jp/phylo/imate.html). The detailed conditions of mate-pair library preparation are included in Supplementary Table 2. After size selection, the quantification of the prepared libraries was performed using a KAPA Library Quantification Kit (KAPA Biosystems). They were sequenced on a HiSeq 1500 (Illumina), operated by HiSeq Control Software v2.0.12.0 using a HiSeq SR Rapid Cluster Kit v2 (Illumina) and HiSeq Rapid SBS Kit v2 (Illumina), and MiSeq operated by MiSeq Control Software v2.3.0.3 using MiSeq Reagent Kit v3 (600 Cycles) (Illumina). Read lengths were 101, 127, 151 or 171 nt on HiSeq and 251 or 301 nt on MiSeq. Base calling was performed with RTA v1.17.21.3, and the fastq files were generated by bcl2fastq v1.8.4 (Illumina). Removal of low-quality bases from paired-end reads was processed by TrimGalore v0.3.3 with the options ‘--stringency 2 --quality 20 --length 25 --paired --retain_unpaired’. Mate-pair reads were processed by NextClip v1.160 with default parameters. De novo genome assembly and scaffolding employing the processed short reads were carried out by the program PLATANUS v1.2.161 with its default parameters. The assembly step employed paired-end reads and single reads whose pairs had been removed, and the scaffolding step employed paired-end and mate-pair reads. The gap closure step employed all of the single, paired-end and mate-pair reads. Resultant genomic scaffold sequences were screened for contaminating organismal sequences, PhiX sequences loaded as a control, mitochondrial DNA sequences, and those shorter than 500 bp, as performed previously28.
Measuring nuclear DNA contents
Nuclear DNA contents of the three species were measured as previously described28,62. We used cells prepared from the liver and blood of a 32-cm-long juvenile female bamboo shark, the blood of a 5-m-long live female whale shark reared in Osaka Aquarium Kaiyukan, and the liver and blood of a 16-cm-long juvenile male cloudy catshark (see above for the detail of sampling). Mouse embryonic fibroblast cells, used as a reference, were prepared from E14.5 embryos and cultured in DMEM media supplemented with 10% FBS, at 37 °C with 5% CO2. We also used human GM12878 cells as a reference, which were cultured in RPMI-1640 media (Thermo Fisher Scientific) supplemented with 15% FBS, 2 mM l-glutamine, and 1× antibiotic-antimycotic solution (Gibco) at 37 °C with 5% CO2. Liver tissues of bamboo shark and catshark were minced using scissors, rinsed once in shark saline solution (222.45 mM NaCl, 1.34 mM KCl, 2.38 mM NaHCO3 and 333 mM urea)63, and incubated in 0.125% trypsin-EDTA solution (1:1 mixture of 0.25% trypsin-EDTA (Thermo Fisher Scientific) and shark saline solution) for 15 min at 37 °C with gentle agitation to dissociate the cells. FBS was added to stop digestion with trypsin. The cell suspension was filtered through a 40 μm cell strainer (BD Bioscience) to remove cell clumps and debris. Blood of bamboo shark and catshark was sampled from the heart using a 1 ml syringe with a 21 G needle and immediately diluted 1:10 in shark saline solution containing 2 mM EDTA. After centrifugation at 500g for 5 min, blood cells were washed once in shark saline solution containing EDTA and counted. A total of 1 × 106 cells were collected by centrifugation at 500g for 5 min and permeabilized in shark saline solution containing 0.05% Triton X-100. DNA staining was performed by adding 1 ml of PI/RNase staining buffer (BD Bioscience). After a 15 min incubation at room temperature, cells were centrifuged at 1,500g for 5 min and resuspended in 400 μl of fresh propidium iodide (PI)/RNase staining buffer. Fluorescence intensities were measured with the excitation at 488 nm and the bandpass filter of 575/26 nm on a FACSCanto II cell sorter (BD Bioscience). Measurements were carried out with three technical replicates per sample, and the acquired values were averaged before DNA content calculations (Supplementary Table 4).
Completeness assessment of genome assemblies
Completeness of the genome assemblies was assessed with (1) CEGMA v2.564, (2) BUSCO v2.0.165 and (3) a manual curation-based census of Wnt genes (Supplementary Fig. 1). For both CEGMA and BUSCO, we employed not only the reference gene sets provided inherently with these program pipelines, but also the core vertebrate genes introduced particularly for vertebrates, especially species in isolated lineages such as elasmobranchs66. The assessments were executed on the gVolante web server67 (Supplementary Note 2) using a script released by us previously66. The genome-wide census of Wnt genes was performed with TBLASTN 2.2.31+68 searches in the elasmobranch genome assemblies using manually curated amino acid sequences of the C. milii Wnt homologues as queries, followed by a fine-scale exon search using the open reading frame sequences predicted on the individual elasmobranch genomes as queries.
Repeat analysis
To obtain species-specific repeat libraries, RepeatModeler v1.0.869 was run on the genome assemblies of the individual species with default parameters. Detection of repeat elements in the genomes was performed by RepeatMasker v4.0.570, which employs National Center for Biotechnology Information (NCBI) RMBlast v2.2.27, using the custom repeat library obtained above. For gene prediction, the parts of genome sequences detected as repeats are soft-masked with the options ‘-nolow -xsmall’.
Construction of gene models
Construction of gene models on the cloudy catshark, whale shark and bamboo shark genomes was performed in this order, following the procedure previously reported15 (Supplementary Note 5). The gene prediction program Augustus v3.1 was employed with ‘trained’ species-specific parameters and hints based on RNA-seq reads and amino acid sequences of putative homologues from other vertebrates. To build homologue hints for the cloudy catshark, we used a set of 117,246 NCBI RefSeq protein sequences downloaded on 23 November 2015, including ‘known’ human proteins (39,582 sequences), chicken (6,189 sequences) and amniote vertebrates (44,675 sequences), as well as C. milii (NCBI Genome version 6.1.3, 26,800 sequences). For constructing gene models of the whale shark, we used a sequence set combining all predicted cloudy catshark peptide sequences along with the above-mentioned sequence set. Likewise, for the gene prediction of the bamboo shark, we incorporated the predicted whale shark peptide sequences into the sequence set used in gene prediction for the catshark. RNA-seq data used for exon hint construction is indicated in Supplementary Table 6.
RNA-seq and transcriptome data processing
Total RNAs were extracted with Trizol reagent (Thermo Fisher Scientific). Quality control of DNase I-treated RNA was performed with Bioanalyzer 2100 (Agilent Technologies). Libraries were prepared with TruSeq RNA Sample Prep Kit (Illumina) or TruSeq Stranded mRNA LT Sample Prep Kit (Illumina) as previously described66. The amount of starting total RNA and numbers of PCR cycles are included in Supplementary Table 6. The obtained sequence reads were trimmed for removal of adaptor sequences and low-quality bases with TrimGalore v0.3.3 as outlined above, and de novo transcriptome assembly was performed with the program Trinity v2.2.171 with the parameters ‘--SS_lib_type RF --min_kmer_cov 3’. Alignment of the trimmed RNA-seq reads to the genome assembly employed TopHat2 v2.0.11, followed by gene expression quantification with Cuffdiff v2.1.1, while read alignment to coding sequences employed bowtie2 v2.2.8 and eXpress v1.5.1.
Comparison of conserved noncoding elements (CNE)
A set of previously identified CNEs for the human genome hg19 was downloaded from UCNEbase72 (http://ccg.vital-it.ch/UCNEbase/data/download/fasta/hg19_UCNEs.fasta.gz). This set included 4,351 genomic segments in the human genome that exhibit >95% nucleotide sequence identity with counterparts in the chicken genome and are longer than 200 bp. Ten of the retrieved CNEs that include regions annotated as protein-coding in the hg38 genome assembly were removed with bedtools v2.25.073, and the remaining 4,341 sequences were queried with BLASTN 2.5.0+ in two different modes, namely ‘megablast’ and ‘dc-megablast’, against the genome assemblies of the bamboo shark and the cloudy catshark from this study and those of other vertebrate species (coelacanth, LatCha1; spotted gar, LepOcu1; western clawed frog, Xtropicalis_v7; zebrafish, GRCz10; medaka, MEDAKA1; Arctic lamprey, LetJap1; sea lamprey, Pmarinus_7.0). The number of best hits that were longer than 100 bp was counted for each of the two search modes. In the analysis of the enhancer in the Tbx4 locus, the program, LAST v75274, was used to detect conserved noncoding elements. Visualization of sequence similarity between species employed VISTA75 using a global pairwise alignment program Shuffle-LAGAN76.
Search for long noncoding RNA (lncRNA)
Human lncRNA sequences were downloaded from GENCODE database77 (release 25; https://www.gencodegenes.org/), which included 27,692 sequences. We removed sequences of antisense RNAs that overlapped open reading frames in the lncRNA database and repetitive sequences that were masked with RepeatMasker v4.0.5, as described above. By using the program BLASTN 2.5.0+ with the dc-megablast mode, we queried the refined set of lncRNAs against the genome assemblies of other vertebrates used above in CNE detection. Following the BLASTN searches, the best hits whose bit scores exceeded 60 were counted. We first made a transcriptome assembly from the bamboo shark RNA-seq data in Supplementary Table 6, using Trinity-v2.4.071 with the options ‘--SS_lib_type RF --trimmomatic’. Next, human lncRNAs were queried against the transcriptome assembly with BLASTN. Subsequently, we selected the best hits of the BLAST search whose bit scores exceeded 50 and removed those that were aligned with the opposite strands of human lncRNAs. We also queried the lncRNA candidates against the Augustus-predicted coding genes of the bamboo shark genome and removed sequences if the best hits were aligned with the forward strands of the predicted coding genes. To analyse the tissue distribution of the validated lncRNAs using RNA-seq data, we masked repeat elements in the transcriptome assembly with RepeatMasker and the repeat library built above for the bamboo shark genome, which was followed by read mapping performed as described above.
Antibody validation for chromatin immunoprecipitation (ChIP) assays
Western blotting for catshark CTCF protein was performed as previously described15, using protein extracts from tissues of a juvenile catshark (muscle and liver) and a human GM12878 cell line with antibodies for CTCF (Cell Signaling Technology, #3418 S in 1:2,000 dilution) and histone H3 (Wako, #304-34781 in 1:2,000 dilution). Immunoprecipitation was performed as previously described15 using the protein extract from the eye of a juvenile cloudy catshark. Protein identification was performed as described previously78, with nanoliquid chromatography tandem mass spectrometry using LTQ Orbitrap Velos Pro (Thermo Fisher Scientific), followed by data analysis with the MASCOT v2.6.1 software (Matrix Science).
ChIP-seq and data processing
Bamboo shark embryos at stage 27, cloudy catshark embryos at stage 27.5 and the stomach of a juvenile cloudy catshark were dissected and snap frozen in liquid nitrogen and kept at −80 °C until use. A whole embryo or a stomach of approximately 1 × 107 cells were used for ChIP with the above-mentioned anti-CTCF antibody. ChIP assays, as well as ChIP-seq and downstream data analysis, were performed as previously described15. Trimming of the obtained sequence reads, mapping against the genome assemblies, and peak calling were performed by TrimGalore v0.3.7, Bowtie v0.12.879 and MACS2 v2.0.1080, respectively. The peaks overlapping between replicates were identified by bedtools v2.19.173 and designated as ‘consensus peaks’. A subset of ‘consensus peaks’ with a fold enrichment value of no less than 10 was assigned as ‘significant peaks’. For the catshark stomach sample without a replicate, peak calling was performed using the embryonic sample as input. Significant peaks for the catshark stomach sample were determined with a fold enrichment value of no less than 10. CTCF core and upstream motifs enriched in the top 2,000 peak regions (peak summit ± 100 bp) were identified by MEME v4.10.081. FIMO v4.10.182 was then used to identify motif locations in the entire peak set (peak summit ± 100 bp). When multiple motifs were identified within a peak region, only the motif with the lowest p value was adopted for downstream analyses.
In situ hybridization
Catshark and bamboo shark embryos were fixed with 4% PFA/PBS, dehydrated with methanol series and stored in 100% methanol at −30 °C until use. In situ hybridization using whole-mount embryos and paraffin-embedded sections of 8 μm thickness was performed as previously reported83,84. Riboprobes were synthesized using complementary DNA amplified with gene specific primers in Supplementary Table 14 as templates. The regions for cDNA amplification were selected in untranslated or non-conserved coding parts of exons to avoid cross-hybridization between paralogues.
Orthologue group construction
We employed the OMA platform85 for producing orthologue groups composed of diverse vertebrate species including the four cartilaginous fishes (Callorhinchus milii, Chiloscyllium punctatum, R. typus and S. torazame). We first retrieved all-against-all alignment results of the predicted peptides of the 19 osteichthyans included in Fig. 1d and sea lamprey from the OMA database (released in March 2017). OMA standalone v2.1.185 was run to perform additional all-against-all comparisons by incorporating the peptides of the four cartilaginous fishes and Arctic lamprey, which produced 31,498 hierarchical orthologous groups (HOGs).
Quantifying synonymous substitutions
For computation of numbers of synonymous substitutions per sites (KS; Supplementary Note 12), 1,656 one-to-one orthologues retained by the four cartilaginous fishes and ten osteichthyans were selected from the HOGs as follows (Supplementary Table 8). First, peptide sequences of the retrieved orthologues were aligned with MAFFT v7.299b86 with the option ‘-linsi’. The individual alignments were trimmed and back-translated into nucleotides with trimAl v1.4 rev1587 with the options ‘-automated1 -backtrans’ followed by removal of gapped sites using trimAl with the options ‘-nogaps’. Orthologue groups containing fewer than 50 aligned codons or a stop codon were discarded. For the selected orthologue groups, KS were computed with codeml in the PAML v4.9c88.
Phylogenetic tree inference
To reconstruct the species tree in Fig. 1d, we retrieved 935 one-to-one orthologue groups retained by all of the 25 vertebrates used in the HOGs, allowing for none or one missing orthologue for individual groups. The peptides of the individual orthologue groups were aligned with MAFFT v7.299b86 with the option ‘-linsi’. Unambiguously aligned sites were selected by trimAl v1.4 rev1587 with the option ‘-strictplus’ followed by a concatenation of these alignments into one. Phylogenetic tree inference was performed with the maximum-likelihood method using the program RAxML v8.2.889 with the options ‘-m PROTCATWAG -f a -# 100’, assuming the partition model for individual orthologue groups (the ‘-q’ option).
To infer individual gene family trees, amino acids sequences were retrieved from aLeaves90 incorporating Ensembl release 84. Multiple sequence alignment was performed with MAFFT with the option ‘-linsi’. The aligned sequence sets were processed using trimAl v1.4 rev1587 with the option ‘-automated1’. This was followed by another trimAl run with the option ‘-nogaps’ in the tree inference for Figs. 3e and 4a, and Supplementary Figs. 4a, 5b, 6a and 7s. Molecular phylogenetic trees were inferred by RAxML with the ‘-m PROTCATWAG -f a -# 1000’ options unless stated otherwise. Tree inference in the Bayesian framework was performed with the program PhyloBayes v4.1c91 with the options ‘-cat -dgam 4 -wag -nchain 2 1000 0.3 50’ unless stated otherwise. This was followed by an execution of bpcomp in the PhyloBayes v4.1c package with the option ‘-x 100’. The support values at the nodes of molecular phylogenetic trees included are, in order, bootstrap values and Bayesian posterior probabilities. The latter was shown only when the relationship at the node in the visualized tree was supported by the Bayesian inference.
Clustered Pcdh gene identification
The genomic scaffold sequences of elasmobranch sharks were first examined via TBLASTN v2.2.29+ using the amino acid sequences of individual clustered Pcdh genes of C. milii (retrieved from http://ensembl.fugu-sg.org, gene IDs: B0YN55-B0YN99, B0YNA0 and B0YNA1) and human clustered and non-clustered Pcdh genes (retrieved from the UCSC Genome Browser) to identify any prospective elasmobranch scaffolds containing clustered Pcdh genes. The regions exhibiting the homologies with the known C. milii and human clustered Pcdh proteins were utilized for gene prediction, which was accomplished by a coordination between GeneWise v2.2.3-rc792 and geneid v1.493. The predicted genes were further refined through manual inspection of exon–intron junctions and transcript evidence from RNA-seq data (Supplementary Table 6). For each species, HISAT2 v2.0.494 was run on each tissue sample with the options ‘-k 200 --known-splicesite-infile’ by inputting a list of splice sites extracted from the Pcdh gene annotation of the individual species. These alignments were passed to StringTie v1.3.095 to generate an additional set of tissue-specific gene models. These models were incorporated into the initial Pcdh gene annotation through StringTie, which was used to produce sets of read coverage tables using StringTie. The output file was utilized by Ballgown v2.6.096 to confirm transcription and splice site locations for each putative clustered Pcdh gene. The protein domain structures in the predicted clustered Pcdh genes were analysed using the HMMer v3.1b297 and SMART98.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Genome, transcriptome and ChIP sequence reads are deposited in the DNA Data Bank of Japan (DDBJ) under the accession number DRA006338. The genome assemblies of the brownbanded bamboo shark and cloudy catshark were deposited in DDBJ under the accession numbers BEZZ01000001–BEZZ01280241 and BFAA01000001–BFAA01458049, respectively. The whale shark genome assembly and the gene models of the three shark species are available at https://figshare.com/projects/sharkgenome1-phyloinfokobe/28863.
References
Venkatesh, B., Tay, A., Dandona, N., Patil, J. G. & Brenner, S. A compact cartilaginous fish model genome. Curr. Biol. 15, R82–R83 (2005).
Venkatesh, B. et al. Elephant shark genome provides unique insights into gnathostome evolution. Nature 505, 174–179 (2014).
Read, T. D. et al. Draft sequencing and assembly of the genome of the world’s largest fish, the whale shark: Rhincodon typus Smith 1828. BMC Genomics 18, 532 (2017).
Wyffels, J. et al. SkateBase, an elasmobranch genome project and collection of molecular resources for chondrichthyan fishes. F1000Research 3, 191 (2014).
Nielsen, J. et al. Eye lens radiocarbon reveals centuries of longevity in the Greenland shark (Somniosus microcephalus). Science 353, 702–704 (2016).
Dulvy, N. K. et al. Challenges and priorities in shark and ray conservation. Curr. Biol. 27, R565–R572 (2017).
Stein, R. W. et al. Global priorities for conserving the evolutionary history of sharks, rays and chimaeras. Nat. Ecol. Evol. 2, 288–298 (2018).
Onimaru, K., Motone, F., Kiyatake, I., Nishida, K. & Kuraku, S. A staging table for the embryonic development of the brownbanded bamboo shark (Chiloscyllium punctatum). Dev. Dyn. 247, 712–723 (2018).
Tatsumi, N. et al. Molecular developmental mechanism in polypterid fish provides insight into the origin of vertebrate lungs. Sci. Rep. 6, 30580 (2016).
Zhang, X., Hamblin, M. H. & Yin, K. J. The long noncoding RNA Malat1: its physiological and pathophysiological functions. RNA Biol. 14, 1705–1714 (2017).
Ji, P. et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 22, 8031–8041 (2003).
Zhang, B. et al. Identification and characterization of a class of MALAT1-like genomic loci. Cell Rep. 19, 1723–1738 (2017).
Duboule, D. The rise and fall of Hox gene clusters. Development 134, 2549–2560 (2007).
Ghirlando, R. & Felsenfeld, G. CTCF: making the right connections. Genes Dev. 30, 881–891 (2016).
Kadota, M. et al. CTCF binding landscape in jawless fish with reference to Hox cluster evolution. Sci. Rep. 7, 4957 (2017).
Narendra, V., Bulajic, M., Dekker, J., Mazzoni, E. O. & Reinberg, D. CTCF-mediated topological boundaries during development foster appropriate gene regulation. Genes Dev. 30, 2657–2662 (2016).
Narendra, V. et al. CTCF establishes discrete functional chromatin domains at the Hox clusters during differentiation. Science 347, 1017–1021 (2015).
Kherdjemil, Y. et al. Evolution of Hoxa11 regulation in vertebrates is linked to the pentadactyl state. Nature 539, 89–92 (2016).
King, B. L., Gillis, J. A., Carlisle, H. R. & Dahn, R. D. A natural deletion of the HoxC cluster in elasmobranch fishes. Science 334, 1517 (2011).
Oulion, S. et al. Evolution of Hox gene clusters in gnathostomes: insights from a survey of a shark (Scyliorhinus canicula) transcriptome. Mol. Biol. Evol. 27, 2829–2838 (2010).
Fried, C., Prohaska, S. J. & Stadler, P. F. Exclusion of repetitive DNA elements from gnathostome Hox clusters. J. Exp. Zool. B Mol. Dev. Evol. 302, 165–173 (2004).
Montavon, T. & Duboule, D. Chromatin organization and global regulation of Hox gene clusters. Philos. Trans. R. Soc. Lon. B 368, 20120367 (2013).
Onimaru, K., Tatsumi, K., Shibagaki, K. & Kuraku, S. A de novo transcriptome assembly of the zebra bullhead shark, Heterodontus zebra. Sci. Data 5, 180197 (2018).
Tanegashima, C. et al. Embryonic transcriptome sequencing of the ocellate spot skate Okamejei kenojei. Sci. Data 5, 180200 (2018).
Danesin, C. & Houart, C. A Fox stops the Wnt: implications for forebrain development and diseases. Curr. Opin. Genet. Dev. 22, 323–330 (2012).
Hebert, J. M. & Fishell, G. The genetics of early telencephalon patterning: some assembly required. Nat. Rev. Neurosci. 9, 678–685 (2008).
Roux, J., Liu, J. & Robinson-Rechavi, M. Selective constraints on coding sequences of nervous system genes are a major determinant of duplicate gene retention in vertebrates. Mol. Biol. Evol. 34, 2773–2791 (2017).
Hara, Y. et al. Madagascar ground gecko genome analysis characterizes asymmetric fates of duplicated genes. BMC Biol. 16, 40 (2018).
Bernard, V., Young, J., Chanson, P. & Binart, N. New insights in prolactin: pathological implications. Nat. Rev. Endocrinol. 11, 265–275 (2015).
Takei, Y., Ando, H. & Tsutsui, K. Handbook of Hormones (Academic, Cambridge, 2016).
Sanger, G. J. & Furness, J. B. Ghrelin and motilin receptors as drug targets for gastrointestinal disorders. Nat. Rev. Gastroenterol. Hepatol. 13, 38–48 (2016).
Prokop, J. W. et al. Discovery of the elusive leptin in birds: identification of several ‘missing links’ in the evolution of leptin and its receptor. PLoS ONE 9, e92751 (2014).
Kuraku, S., Meyer, A. & Kuratani, S. Timing of genome duplications relative to the origin of the vertebrates: did cyclostomes diverge before or after? Mol. Biol. Evol. 26, 47–59 (2009).
Pan, W. W. & Myers, M. G. Jr. Leptin and the maintenance of elevated body weight. Nat. Rev. Neurosci. 19, 95–105 (2018).
Wolf, G. Leptin: the weight-reducing plasma protein encoded by the obese gene. Nutr. Rev. 54, 91–93 (1996).
Gesta, S., Tseng, Y. H. & Kahn, C. R. Developmental origin of fat: tracking obesity to its source. Cell 131, 242–256 (2007).
Acher, R., Chauvet, J., Chauvet, M. T. & Rouille, Y. Unique evolution of neurohypophysial hormones in cartilaginous fishes: possible implications for urea-based osmoregulation. J. Exp. Zool. 284, 475–484 (1999).
Hyodo, S., Tsukada, T. & Takei, Y. Neurohypophysial hormones of dogfish, Triakis scyllium: structures and salinity-dependent secretion. Gen. Comp. Endocrinol. 138, 97–104 (2004).
Sanger, G. J. & Lee, K. Hormones of the gut–brain axis as targets for the treatment of upper gastrointestinal disorders. Nat. Rev. Drug. Discov. 7, 241–254 (2008).
Bowmaker, J. K. & Hunt, D. M. Evolution of vertebrate visual pigments. Curr. Biol. 16, R484–R489 (2006).
Marshall, J., Carleton, K. L. & Cronin, T. Colour vision in marine organisms. Curr. Opin. Neurobiol. 34, 86–94 (2015).
Davies, W. L. et al. Into the blue: gene duplication and loss underlie color vision adaptations in a deep-sea chimaera, the elephant shark Callorhinchus milii. Genome Res. 19, 415–426 (2009).
Emerling, C. A. & Springer, M. S. Genomic evidence for rod monochromacy in sloths and armadillos suggests early subterranean history for Xenarthra. Proc. Biol. Sci. 282, 20142192 (2015).
Meredith, R. W., Gatesy, J., Emerling, C. A., York, V. M. & Springer, M. S. Rod monochromacy and the coevolution of cetacean retinal opsins. PLoS Genet. 9, e1003432 (2013).
Mohun, S. M. et al. Identification and characterization of visual pigments in caecilians (Amphibia: Gymnophiona), an order of limbless vertebrates with rudimentary eyes. J. Exp. Biol. 213, 3586–3592 (2010).
Ebert, D. A., Fowler, S. & Compagno, L. Sharks of the World: A Fully Illustrated Guide (Wild Nature Press, Plymouth, 2013).
Naylor, G. J. P. et al. in Biology of Sharks and Their Relatives (eds Carrier, J. C., Musick, J. A. & Heithaus, M. R.) Ch. 2, 31–56 (CRC Press, 2012).
Douglas, R. H. & Partridge, J. C. in Encyclopedia of Fish Physiology: From Genome to Environment Vol. 1 (ed A. Ferrell) 166–182 (Academic, Cambridge, 2011).
Yokoyama, S., Tada, T., Zhang, H. & Britt, L. Elucidation of phenotypic adaptations: molecular analyses of dim-light vision proteins in vertebrates. Proc. Natl Acad. Sci. USA 105, 13480–13485 (2008).
Tyminski, J. P., de la Parra-Venegas, R., Gonzalez Cano, J. & Hueter, R. E. Vertical movements and patterns in diving behavior of whale sharks as revealed by pop-up satellite tags in the Eastern Gulf of Mexico. PLoS ONE 10, e0142156 (2015).
Niimura, Y. Evolutionary dynamics of olfactory receptor genes in chordates: interaction between environments and genomic contents. Hum. Genomics 4, 107–118 (2009).
Niimura, Y. On the origin and evolution of vertebrate olfactory receptor genes: comparative genome analysis among 23 chordate species. Genome Biol. Evol. 1, 34–44 (2009).
Chen, W. V. & Maniatis, T. Clustered protocadherins. Development 140, 3297–3302 (2013).
Hirayama, T. & Yagi, T. Regulation of clustered protocadherin genes in individual neurons. Semin. Cell. Dev. Biol. 69, 122–130 (2017).
Yu, W. P. et al. Elephant shark sequence reveals unique insights into the evolutionary history of vertebrate genes: a comparative analysis of the protocadherin cluster. Proc. Natl Acad. Sci. USA 105, 3819–3824 (2008).
Ballard, W. W., Mellinger, J. & Lechenault, H. A series of normal stages for development of Scyliorhinus canicula, the lesser spotted dogfish (Chondrichthyes, Scyliorhinidae). J. Exp. Zool. 267, 318–336 (1993).
Ueda, K. et al. in The Elasmobranch Husbandary Manual II: Recent Advances in the Care of Sharks, Rays and their Relatives (eds Smith, M. et al.) 255–262 (Special Publication of the Ohio Biological Survey, 2017).
Kuraku, S., Qiu, H. & Meyer, A. Horizontal transfers of Tc1 elements between teleost fishes and their vertebrate parasites, lampreys. Genome Biol. Evol. 4, 929–936 (2012).
Tatsumi, K., Nishimura, O., Itomi, K., Tanegashima, C. & Kuraku, S. Optimization and cost-saving in tagmentation-based mate-pair library preparation and sequencing. Biotechniques 58, 253–257 (2015).
Leggett, R. M., Clavijo, B. J., Clissold, L., Clark, M. D. & Caccamo, M. NextClip: an analysis and read preparation tool for Nextera Long Mate Pair libraries. Bioinformatics 30, 566–568 (2014).
Kajitani, R. et al. Efficient de novo assembly of highly heterozygous genomes from whole-genome shotgun short reads. Genome Res. 24, 1384–1395 (2014).
Hare, E. E. & Johnston, J. S. Genome size determination using flow cytometry of propidium iodide-stained nuclei. Methods Mol. Biol. 772, 3–12 (2011).
Huggel, H. Experimentelle untersuchungen ueber die automatie, temperaturabhaengigkeit und arbeit des embryonalen fischherzens, unter besonderer beruecksichtigung der salmoniden und scylliorhiniden. Z. Vgl Physiol. 42, 63–102 (1959).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Simao, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Hara, Y. et al. Optimizing and benchmarking de novo transcriptome sequencing: from library preparation to assembly evaluation. BMC Genomics 16, 977 (2015).
Nishimura, O., Hara, Y. & Kuraku, S. gVolante for standardizing completeness assessment of genome and transcriptome assemblies. Bioinformatics 33, 3635–3637 (2017).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Smit, A. F. A. & Hubley, R. RepeatModeler Open-1.0 (2008–2010); http://www.repeatmasker.org
Smit, A. F. A., Hubley, R. & Green, P. RepeatMasker Open-4.0 (2013–2015); http://www.repeatmasker.org
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Dimitrieva, S. & Bucher, P. UCNEbase-a database of ultraconserved non-coding elements and genomic regulatory blocks. Nucleic Acids Res. 41, D101–D109 (2013).
Quinlan, A. R. BEDTools: the Swiss-Army tool for genome feature analysis. Curr. Protoc. Bioinform. 47, 12 11–12 34 (2014). 11.
Frith, M. C. & Kawaguchi, R. Split-alignment of genomes finds orthologies more accurately. Genome. Biol. 16, 106 (2015).
Mayor, C. et al. VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 16, 1046–1047 (2000).
Brudno, M.et al. Glocal alignment: finding rearrangements during alignment. Bioinformatics 19 (Suppl. 1), i54–i62 (2003).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Sadaie, M., Shinmyozu, K. & Nakayama, J. A conserved SET domain methyltransferase, Set11, modifies ribosomal protein Rpl12 in fission yeast. J. Biol. Chem. 283, 7185–7195 (2008).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
O’Neill, P., McCole, R. B. & Baker, C. V. A molecular analysis of neurogenic placode and cranial sensory ganglion development in the shark, Scyliorhinus canicula. Dev. Biol. 304, 156–181 (2007).
Kuraku, S., Usuda, R. & Kuratani, S. Comprehensive survey of carapacial ridge-specific genes in turtle implies co-option of some regulatory genes in carapace evolution. Evol. Dev. 7, 3–17 (2005).
Train, C. M., Glover, N. M., Gonnet, G. H., Altenhoff, A. M. & Dessimoz, C. Orthologous Matrix (OMA) algorithm 2.0: more robust to asymmetric evolutionary rates and more scalable hierarchical orthologous group inference. Bioinformatics 33, i75–i82 (2017).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Kuraku, S., Zmasek, C. M., Nishimura, O. & Katoh, K. aLeaves facilitates on-demand exploration of metazoan gene family trees on MAFFT sequence alignment server with enhanced interactivity. Nucleic Acids Res. 41, W22–W28 (2013).
Lartillot, N., Rodrigue, N., Stubbs, D. & Richer, J. PhyloBayes MPI: phylogenetic reconstruction with infinite mixtures of profiles in a parallel environment. Syst. Biol. 62, 611–615 (2013).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–995 (2004).
Blanco, E., Parra, G. & Guigo, R. Using geneid to identify genes. Curr. Protoc. Bioinformatics 18, 4.3.1–4.3.28 (2007).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Letunic, I., Doerks, T. & Bork, P. SMART: recent updates, new developments and status in 2015. Nucleic Acids Res. 43, D257–D260 (2015).
Acknowledgements
We thank J. Donald for critical reading of the manuscript, K. Shirato, K. Yamamoto, S. Shibuya and members of the Evolutionary Morphology Laboratory, RIKEN for rearing catsharks, M. Andrabi for assistance in sequence data analysis, K. Tanimoto, H. Kiyonari, T. Yoshikuni, T. Ito, Y. Miyagawa and S. Sodeyama for providing materials. Our gratitude extends to K. Muguruma, Y. Murakami, F. Sugahara, J. Pascual-Anaya, R. Kusakabe, S. Higuchi, Y. Yamaguchi, W. Takagi, H. Kaiya, Y. Ishihama, S. Miyake, T. Kaku, T. Tanaka, D. Sipp, M. Tan, A. D. M. Dove, T. D. Read, D. Lagman, D. Ocampo Daza, D. Larhammar, Y. Uno and S. Mazan for insightful discussion. This study was supported by RIKEN and JSPS KAKENHI Grant Numbers 26650110, 26291065 and 17H03868 to S.Kuraku and S.H. and 17K07426 to S.Kuraku.
Author information
Authors and Affiliations
Contributions
S.Kuraku conceived the study. Y.H., K.Y., K.Tatsumi, K.Tanaka, F.M., Y.K., I.K., R.Nozu, R.Nakagawa, R.M., K.M., S.H., K.O., O.N., S.D.K., C.T., M.Kadota, M.Koyanagi and S.Kuraku performed experiments. Y.H., K.Y., K.O., S.D.K., M.Kadota, N.A., Y.K., A.T., M.Koyanagi, K.N., S.Kuratani and S.Kuraku interpreted data. Y.H., K.Y., S.D.K., M.Kadota, M.Koyanagi, S.H., K.S., K.O. and S.Kuraku drafted the manuscript. All authors contributed to the final manuscript editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes, Supplementary Figures and Supplementary Tables
Supplementary Data 1
Genomic scaffold sequences extracted from non-final genome assemblies
Supplementary Data 2
List of manually curated genes and scaffolds including curated genes extracted from non-final genome assemblies
Supplementary Data 3
Nucleotide sequences of putative bamboo shark lncRNAs
Supplementary Data 4
Nucleotide sequences of clustered protocadherins identified in the shark genomes
Supplementary Data 5
Amino acid sequences of clustered protocadherins identified in the shark genomes
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hara, Y., Yamaguchi, K., Onimaru, K. et al. Shark genomes provide insights into elasmobranch evolution and the origin of vertebrates. Nat Ecol Evol 2, 1761–1771 (2018). https://doi.org/10.1038/s41559-018-0673-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0673-5
This article is cited by
-
The evolution of tenascins
BMC Ecology and Evolution (2024)
-
Shark genome size evolution and its relationship with cellular, life-history, ecological, and diversity traits
Scientific Reports (2024)
-
Extensive MHC class IIβ diversity across multiple loci in the small-spotted catshark (Scyliorhinus canicula)
Scientific Reports (2023)
-
Evolution of lysine-specific demethylase 1 and REST corepressor gene families and their molecular interaction
Communications Biology (2023)
-
Genomic Characteristics of Okamejei kenojei and the Implications to Its Evolutionary Biology Study
Marine Biotechnology (2023)