Abstract
The copepod Tigriopus californicus shows extensive population divergence and is becoming a model for understanding allopatric differentiation and the early stages of speciation. Here, we report a high-quality reference genome for one population (~190 megabases across 12 scaffolds, and ~15,500 protein-coding genes). Comparison with other arthropods reveals 2,526 genes presumed to be specific to T. californicus, with an apparent proliferation of genes involved in ion transport and receptor activity. Beyond the reference population, we report re-sequenced genomes of seven additional populations, spanning the continuum of reproductive isolation. Populations show extreme mitochondrial DNA divergence, with higher levels of amino acid differentiation than observed in other taxa. Across the nuclear genome, we find elevated protein evolutionary rates and positive selection in genes predicted to interact with mitochondrial DNA and the proteins and RNA it encodes in multiple pathways. Together, these results support the hypothesis that rapid mitochondrial evolution drives compensatory nuclear evolution within isolated populations, thereby providing a potentially important mechanism for causing intrinsic reproductive isolation.
Similar content being viewed by others
Main
Intrinsic reproductive isolation occurs when gene interactions that are neutral or beneficial in their original genetic background become detrimental in a hybrid genetic background1. In recent years, there has been considerable focus on the origin and identity of these genetic incompatibilities. One category of hybrid conflict found across a diversity of eukaryotes is mitonuclear incompatibility2,3. In the shift from free-living microbe to organelle, mitochondria transferred most of their genome to the nucleus, leaving their function dependent on a large number (~1,500) of nuclear-encoded gene products4,5 and the handful of genes retained in the mitochondrial genome. Mitochondrial DNA (mtDNA) is particularly prone to accumulating deleterious mutations due to elevated mutation rates, limited recombination and the absence of sexual reproduction6. This favours a pattern of compensatory coevolution where nuclear genes must repeatedly evolve to rescue mitochondrial function. The recent proliferation of next-generation sequencing methods now allows genome-wide tests for signatures of mitonuclear coevolution across a diversity of taxa.
Here, we present a high-quality draft genome of the intertidal copepod Tigriopus californicus (Fig. 1a), with genomic comparisons across eight populations. Copepod genomes are important in their own right because copepods are among the most abundant organisms on the planet and play critical roles in both marine and freshwater environments7. Despite their diversity and abundance, only a few copepod draft genomes have been assembled and published8,9, two of which are in the genus Tigriopus (T. japonicus10 and T. kingsejongensis11). The T. californicus genome is especially important because the species is an emerging model for understanding hybrid breakdown12,13, with particular focus on nuclear-mitochondrial coevolution14,15,16,17,18,19. Mitonuclear conflicts may be a particularly important driver of postzygotic isolation in this system due to elevated mitochondrial substitution rates (synonymous site substitution rate ~55-fold higher for mtDNA relative to nuclear DNA20) and the absence of heteromorphic sex chromosomes21, meaning the species lacks the rapidly accumulating X-autosome interactions thought to be the first incompatibilities to arise in many other systems22,23,24.
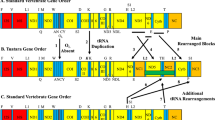
a, Image of an adult T. californicus female carrying an egg sac. Scale bar: 0.25 mm. b, Venn diagram of shared and unique gene clusters among T. californicus, Drosophila melanogaster, A. mellifera and Daphnia pulex. c, Neighbour-joining phylogeny of eight T. californicus populations inferred from an alignment of 150,000 nucleotide positions across 100 protein-coding genes. Arrows point to collection locations along the west coast of North America. AB, Abalone Cove; BB, Bodega Bay; BR, Bird Rock; CAT, Catalina Island; FHL, Friday Harbor; SC, Santa Cruz; SD, San Diego; SRQ, San Roque. Credit: photo in a, G. W. Rouse.
Results and discussion
The T. californicus genome is one of the most compact among copepods. A single cytophotometric study suggests a haploid genome of 244 megabases (Mb) (or 0.25 pg)25,26. However, we argue that this measurement may not be particularly accurate because the calibration standards used ranged from 2.5 to 6.3 pg; hence, extrapolation down to 0.5 pg (that is, the total DNA in a T. californicus nucleus) is probably not appropriate. Our high-coverage sequencing effort with long and short reads, as well as Hi-C technology, has consistently resulted in a total length estimate of 181–191.15 Mb (Supplementary Table 1). Moreover, based on the Benchmarking Universal Single-Copy Orthologs approach, the assembly captured 94.5% complete transcripts of the benchmarking arthropod gene set (1,007 out of 1,066) and 92.9% of the metazoan set (909 out of 978) (Supplementary Fig. 1), suggesting that the actual genome size is probably closer to 200 Mb. The missing sequence is probably enriched in repetitive DNA, which is known to hinder efforts to assemble eukaryotic genomes.
Although the assembly is fragmented into 459 scaffolds after removing those of bacterial origin (Supplementary Tables 1 and 2, and Supplementary Note), 99.3% of the sequence is contained in only 12 scaffolds that range in size from 13.38 to 18.07 Mb, with only 3.77 Mb (1.97%) of gaps within scaffolds. Markers from a linkage map27 readily identified these 12 scaffolds as the 12 autosomes, placing over 99% of the genome assembly in chromosomes. Repetitive elements occupy nearly 29% of the copepod genome, with long interspersed nuclear elements and long terminal repeats composing as much as 39% of all repeats (Supplementary Table 3). The MAKER2 pipeline28 predicted 15,646 gene models. Of these, 14,233 (91%) were supported by external transcriptomic data, such as BLASTN alignments of a high-coverage transcriptome assembly29 and mapping of RNA sequencing (RNA-Seq) reads (≥10 reads), and a high proportion (69%) showed significant homology to annotated proteins in Swiss-Prot (Supplementary Note).
Orthology analyses identified 4,803 orthologous gene groups that are shared among T. californicus and three arthropods with sequenced genomes (Fig. 1b and Supplementary Table 12). Between the two crustaceans (that is, T. californicus and Daphnia pulex), 5,767 gene groups are shared in total, and 433 groups are shared uniquely. Moreover, 661 gene groups, containing 2,526 genes, were identified as putatively specific to T. californicus. Among these, 77 Gene Ontology functional terms were found to be over-represented relative to all other T. californicus genes (Supplementary Table 4). We observed a striking proliferation of certain G-coupled protein receptors; the FMRFamide receptor gene (fmrfar) was found as a single copy in most arthropod genomes characterized, including Daphnia (Crustacea: Cladocera), but was detected as 14 copies in T. kingsejongensis and as at least 60 copies in T. californicus (Supplementary Note, Supplementary Fig. 2, and Supplementary Tables 12, 13, 15 and 17).
The T. californicus system provides an outstanding opportunity to investigate mechanisms of evolutionary divergence along the full continuum of speciation, as populations along its distribution exhibit varying levels of reproductive incompatibility when hybridized13,30. The genome sequences of seven additional T. californicus populations (Fig. 1c) were assembled by mapping Illumina reads from each population to the San Diego reference and reconstructing consensus sequences. Therefore, these assemblies do not capture structural differences among populations, but provide abundant sequence information for protein-coding regions (Supplementary Table 5).
We found extreme levels of mitochondrial divergence between natural populations of T. californicus, including one (San Roque) within a clade being described as the new species T. bajaensis3. Nucleotide divergence across the mitochondrial genome ranged from 9.5 to 26.5% relative to the reference San Diego mitochondrial genome, with an average of 19.6% across all populations (Supplementary Table 6). Among the 37 mitochondrial genes (Fig. 2a), the 13 protein-coding genes have the highest average population divergence. The 22 transfer RNA (tRNA) genes were also found to be highly variable across populations, ranging from 0 to 30% nucleotide divergence, with an average of 9%.
a, Pairwise nucleotide divergence (Dxy) for re-sequenced populations relative to the San Diego reference across all mtDNA-encoded genes. b, Estimates of dN/dS (top) and DoS (bottom) relative to the San Diego reference across the 13 protein-coding genes. Missing data are represented as grey squares. AB, Abalone Cove; BB, Bodega Bay; BR, Bird Rock; CAT, Catalina Island; FHL, Friday Harbor; SC, Santa Cruz; SRQ, San Roque. c, dN/dS (top) and DoS (bottom) for protein subunits of the OXPHOS complexes. For each complex, values were pooled across the seven interpopulation alignments. Superscript, lower-case letters above the bars denote the results of Mann-Whitney U-tests, followed by correction for multiple testing; complexes not sharing a letter are significantly different (FDR < 5%). Boxplots show the median, lower and upper quartiles, with whiskers extending to 1.5 times the interquartile range; single points are data beyond this range.
To investigate the evolutionary forces shaping differentiation, we examined deviations from neutrality using two statistical tests (the ratio of the rate of non-synonymous to the rate of synonymous nucleotide substitutions (dN/dS) and the direction of selection (DoS)). The mitochondrial genome-wide dN/dS ratio is well below 1 at 0.326, suggesting purifying selection as the principal force in mitochondrial evolution. Despite this, mitochondrial dN/dS in T. californicus is among the highest seen in animal mtDNA (1,855 species31). Analysis of nucleotide polymorphism confirms the broad pattern of purifying selection, revealing an average excess of amino acid polymorphism relative to divergence (DoS = −0.34).
A population comparison of individual protein-coding genes revealed that despite the overall pattern of purifying selection across the mitochondrial genome, some genes contain an excess of amino acid divergence (dN/dS > 1; DoS > 0; Fig. 2b). This suggests that despite the net evolutionary effects of linkage in a non-recombining genome and the cumulative action of purifying selection against amino acid polymorphisms, a handful of genes show evidence for diversifying selection (atp6, nad3, nad5 and nad6). Protein-coding genes for the different respiratory chain complexes experience different selection pressures within and between populations (dN/dS: Kruskal–Wallis rank sum test, P < 10−8; DoS: P = 0.001; Supplementary Tables 7 and 8). We find that dN/dS and DoS values are generally higher across complex I genes compared with other respiratory complexes (dN/dS: Mann-Whitney U-test, PMWU < 10–8; DoS: PMWU < 0.001; Fig. 2c), and we detected a strong signature of positive selection (ω = 4.39) on ~2% of the codons of one of the complex I genes (nd4l; Table 1). Despite showing low overall levels of dN/dS (Fig. 2c), an mtDNA-encoded component of complex IV (mt-co3) also exhibited statistical evidence for positive selection, albeit at a moderate level (ω = 1.36; Table 1).
We examined rates of coding sequence evolution via dN/dS levels in 13,538 predicted nuclear genes across the eight T. californicus population genomes. We used these estimates to test the hypothesis of compensatory evolution, which predicts that nuclear-encoded proteins that interact with mtDNA-encoded elements will show elevated rates of amino acid changes. Specifically, we performed four sets of comparisons: (1) mitochondrial versus cytosolic counterparts within ribosomal protein (RP) genes and (2) within aminoacyl tRNA synthetases (ARSs); (3) oxidative phosphorylation (OXPHOS) complexes I, III, IV and V versus complex II; and (4) within mitochondrially targeted proteins (MTPs), all genes in pathways predicted to interact with mtDNA (OXPHOS, mitochondrial RP (mt-RP), mitochondrial ARS (mt-ARS), mtDNA replication, transcription and elongation/initiation factors) versus all genes not in these groups (that is, those that probably form no interaction with any mtDNA-encoded product).
Our analyses revealed elevated rates of amino acid substitution in proteins that putatively interact with mtDNA-encoded products across multiple mitochondrial pathways. Consistent with a previous study of 2 T. californicus populations32, mt-RPs exhibited dN/dS levels that were 6.5 times higher than those of cytosolic RPs (cyt-RPs; Mann-Whitney U-test, PMWU < 10–15; Fig. 3a), with cyt-RPs having as many as 15 genes with no amino acid changes (dN/dS = 0). Differences in dN/dS may be caused by differences in functional constraint on proteins instead of compensatory evolution, with levels of gene expression being considered a major factor influencing the degree of constraint and having a strong inverse relationship with dN/dS33,34,35. The expression levels of cyt-RP genes were significantly higher than those of mt-RP genes (t = 55.8, d.f. = 124.4, P < 10–15; Supplementary Fig. 3 and Supplementary Note). After accounting for this difference in expression, the evolutionary rate among mt-RPs remained significantly elevated compared with cyt-RPs (analysis of covariance (ANCOVA), P1,130 = 0.0095). Among ARSs, those that interact with mtDNA-encoded tRNAs also had significantly higher dN/dS values than their cytosolic counterparts (PMWU = 5.87 × 10–6; Fig. 3b). In this case, after accounting for the markedly different levels of gene expression (t = 6.8, d.f. = 16, P = 1.73 × 10–7; Supplementary Fig. 3), dN/dS among the two groups did not differ significantly (ANCOVA, P1,130 = 0.136). Therefore, we cannot exclude the hypothesis that elevated dN/dS in mt-ARS is due to relaxed functional constraints. Similar results for ARS have been observed in flies and chickens36.
a–c, Estimates of dN/dS for cyt-RPs (n = 67) and mt-RPs (n = 67) (a), cyt-ARSs (n = 22) and mt-ARSs (n = 11) (b) and protein subunits of the OXPHOS enzyme complexes (n = 29, 4, 6, 11 and 12 for complexes I, II, III, IV and V, respectively) (c). a and b show the results of ANCOVAs that account for gene expression levels (**P < 0.01; NS, not significant). c shows the results of Mann-Whitney U-tests comparing complex II with each of the other complexes (*P < 0.05). Boxplots report the median, lower and upper quartiles, with whiskers extending to 1.5 times the interquartile range; single points are data beyond this range.
Within the OXPHOS pathway, a meaningful test of our hypothesis is the comparison of evolutionary rates between complex II and each of the other four complexes, since complex II is the only complex that does not contain an mtDNA-encoded subunit, and expression levels of complex II genes were not significantly different from those of the other complexes (t-tests, all P > 0.183; Supplementary Fig. 3). Our results are consistent with a pattern of compensatory evolution of nuclear-encoded OXPHOS components: complexes I, III and IV showed elevated dN/dS compared with complex II (PMWU = 0.018, 0.040 and 0.013, respectively), although the difference between complexes II and V was not significant (PMWU = 0.214) (Fig. 3c).
Finally, our annotation of nuclear-encoded MTPs allowed us to separate 2 broad groups of proteins within the organelle: nuclear-encoded proteins in pathways that involve mtDNA-encoded products (n = 147) and those in pathways that do not interact with such products (n = 458). This comparison includes proteins across multiple pathways and functions, considering only whether they putatively interact with mtDNA-encoded elements. We found that mtDNA-interacting proteins showed, on average, significantly higher molecular evolutionary rates than those that do not interact with mtDNA-encoded elements (PMWU = 8.37 × 10–5; Fig. 4), despite possible constraints as a result of them having higher transcription levels (t = 5.2, d.f. = 261, P = 4.1 × 10–6; Supplementary Fig. 3).
dN/dS estimates for proteins in pathways that interact with mtDNA-encoded elements (n = 147; Mito int.) and proteins in pathways that do not interact with mtDNA-encoded elements (n = 458; No int.). Mann-Whitney U-test, ***P = 8.4 × 10–4. Boxplots report the median, lower and upper quartiles, with whiskers extending to 1.5 times the interquartile range; single points are data beyond this range.
To allow dN/dS to vary among codons and scan the genomes for patterns of amino acid changes consistent with positive selection, we applied the CODEML ‘sites’ model37 across the 13,538 genes. This analysis revealed 775 genes (530 with BLAST annotation) with significant evidence for positive selection (false discovery rate (FDR) < 10%; Supplementary Table 9). These genes were distributed among many functional categories, with no over-representation of any Gene Ontology term. Nevertheless, we detected strong evidence for compensatory evolution to mtDNA divergence in adenosine triphosphate (ATP) synthesis and translation machineries, with four mt-RPs and five nuclear-encoded OXPHOS genes showing signatures of positive selection for at least one amino acid site (Table 1). Three of the OXPHOS genes belong to complex V, and one to each of complexes I and IV, consistent with elevated evolutionary rates of mtDNA-encoded proteins within these complexes. Coadaptation of these genes to interacting mtDNA OXPHOS proteins within populations is consistent with the highly reduced enzymatic activity of these complexes when divergent genomes are recombined38. The five OXPHOS genes highlighted here are therefore strong candidates for future functional studies39.
As mt-RPs associate closely with mtDNA-encoded ribosomal RNA (rRNA) in the assembly of mitochondrial ribosomes, adaptive evolution of mt-RP in response to rapid evolution in the rRNA may result in functional 'mismatches' between these interacting components in recombinant hybrids. We predict that low-fitness interpopulation hybrids will have mismatched mitonuclear genotypes at one or more of these mt-RPs, and at the cellular level, mtDNA gene translation may be strongly affected. Such a disruption of mitochondrial translation machinery was detected in Drosophila from an incompatibility between an mt-ARS and its mtDNA-encoded tRNA40. In T. californicus, we found no evidence of adaptive differentiation in mt-ARS (all P > 0.61), and we hypothesize that mt-RP coadaptation may play a larger role among hybrid incompatibilities in this species.
With the recent availability of genomic resources for two other Tigriopus species, we assessed patterns of coding sequence evolution on each lineage. While the ‘sites’ model examined above detects diversifying selection across all taxa in the phylogeny, ‘branch-sites’ models41 permit the detection of adaptive evolution that may have occurred episodically on specific branches. A total of 4,863 orthologues were analysed using ‘branch-sites’ in CODEML, with each of the six branches serving as a foreground branch (Supplementary Table 10). Branches leading to T. japonicus and T. kingsejongensis exhibited totals of 52 and 34 positively selected genes, respectively, with only one in each species interacting with mtDNA (Fig. 5). Notably, both of these mtDNA-interacting genes were associated with OXPHOS complex I, but they were different (ndufb9 in T. japonicus, and ndufa3 in T. kingsejongensis). No Gene Ontology terms were over-represented on branches leading to either of these two species. The two T. californicus populations examined exhibited only one and five positively selected genes on the branches separating these lineages. However, the branch leading to T. californicus showed evidence of positive selection on 591 genes (Fig. 5), and these were enriched with DNA metabolic processes such as replication (GO:0006261), recombination (GO:0006310), DNA repair (GO:0006281), and cellular response to DNA damage (GO:0006974) (Supplementary Table 11). Moreover, 28 nuclear-encoded MTPs exhibited positive selection, 7 of which are mtDNA-interacting (4 mt-RPs, 1 mt-ARS, 1 initiation factor, and the mitochondrial RNA polymerase; Supplementary Table 10). Although this elevated number may in part be due to increased statistical power as a result of having more than one taxon on the tested foreground clade, we argue that this is probably not the case, since the branch leading to the T. californicus plus T. japonicus clade exhibited only 34 selected genes, from which no mtDNA-interacting genes were detected (Fig. 5). Overall, these patterns suggest that the T. californicus genome underwent widespread adaptive evolution, involving mitochondrial transcription and translation pathways, after separation from T. japonicus. Functional sequence evolution among the T. californicus populations remained elevated in mt-RPs and OXPHOS genes, as suggested by the ‘sites’ analysis.
The tree was estimated in MEGA6 (ref. 73) from a concatenated alignment of 20 randomly selected genes, totalling 9,721 amino acid positions. Values above each branch show the number of genes detected to have significant positive selection following ‘branch-sites’ tests41 (total/mtDNA-interacting). SD, San Diego; SRQ, San Roque.
Nucleotide substitution rates in animal mtDNA are often >10× higher than rates within nuclear genomes of the same species. Since many nuclear DNA-encoded proteins are imported into the mitochondria and function by interacting with the mtDNA or its protein and RNA products, rapid evolution of mtDNA can become an important selective force acting on the evolution of nuclear genes. Here, we document extremely high levels of differentiation among mtDNAs across T. californicus populations and report the results of tests for the potential impact of such differentiation on the associated nuclear genomes. Previous examinations of this phenomenon were often restricted to the components of the OXPHOS pathway. Here, we test for similar effects among nuclear-encoded proteins that probably interact with other mtDNA-encoded elements, such as tRNAs (during translation), rRNAs (during ribosome assembly) and mtDNA itself (during replication and transcription).
We detected strong signatures of adaptive molecular evolution in mt-RPs and OXPHOS proteins. Although positive selection in these gene groups has been observed during the evolution of other arthropod species42, our study shows that these nuclear-encoded mitochondrial proteins are evolving rapidly among conspecific populations. We argue that adaptive evolution of these proteins represents compensatory changes within each population in response to its rapidly changing mtDNA, and that this divergence can contribute to nascent reproductive isolation among populations.
Methods
Tissue sampling and DNA isolation
T. californicus of mixed sex and life stages were collected from high intertidal rocky pools in San Diego (32° 44′ 44′′ N, 117° 15′ 18′′ W) and kept en masse in 1 l beakers with filtered seawater (0.2 µm pore filter) at 20 °C. To reduce the contribution of gut contents and bacteria to the DNA extract, the seawater was treated with an antibiotic mixture (ampicillin, 200 mg l–1; streptomycin, 50 mg l–1; spectinomycin, 50 mg l–1; kanamycin, 50 mg l–1; penicillin, 200 units ml–1; neomycin, 50 mg l–1; and chloramphenicol, 20 mg l–1) and the animals were moved to clean filtered seawater daily for 3 days, without food, before extraction. DNA was isolated from groups of hundreds of copepods using a Qiagen DNeasy Blood and Tissue Kit, and samples were pooled to ~100 µg of DNA.
Genome sequencing and assembly
Cofactor Genomics generated 3 short-insert libraries (200, 500 and 800 bp) and 2 long-insert libraries (2 and 5 kb), and then barcoded and sequenced the libraries in 1 lane using the Illumina HiSeq 2000 platform to obtain 2× 100 bp paired-end (short inserts) or mate-paired sequences (Supplementary Table 1). Illumina sequences from the five libraries were first filtered using an internal quality control script to remove the low-quality reads. For the 200 and 500 bp insert libraries, which have high coverage, the quality control parameters were more stringent. All raw reads with unknown base 'N' were removed. In addition, raw reads were removed if the sum of per-base error probability across all bases was greater than two. For other libraries, whose coverage is relative lower, reads with 'N' were kept. However, reads were deleted if the sum of per-base error probability was greater than three. We only kept read pairs for which both read ends passed the quality filter. Duplicated read pairs were identified by CD-HIT-DUP in the CD-HIT package with the parameters '-u 50 –e'. Here, the read pairs having near-identical sequences with up to 2 mismatches within the first 50 bases on both ends were considered duplicated. The duplicated read pairs with lower-quality scores were removed. The processed reads from these five libraries were assembled using AllPaths-LG (release 46212) following AllPaths’ standard workflow, including initial input file preparation and final assembly runs. The assembled scaffold sequences were further extended using the GapCloser programme in the SOAPdenovo package.
To improve contiguity of the T. californicus assembly, proximity ligation libraries based on the Chicago and Hi-C methods were prepared by Dovetail Genomics as described previously43,44. Briefly, the Chicago method started with high-molecular-weight DNA (~75 kb) while the Dovetail Hi-C method started with intact chromosomes in fresh copepod tissue, and chromatin was reconstituted in vitro and cross-linked with formaldehyde. While cross-linked, chromatin then underwent several steps of endonuclease digestion, biotinylation and ligation to preserve information on proximity and the association of fragments. DNA was then purified and fragmented for traditional Illumina library preparation and HiSeq sequencing. Finally, the Chicago sequence data were used to correct and improve the AllPaths assembly via the HiRise computational pipeline43, and the Hi-C data were then used to increase contiguity of the AllPaths plus Chicago assembly.
In an effort to fill assembly gaps, 1.68 million PacBio reads were obtained for the San Diego population and used to improve the final HiRise assembly using PBJelly45. PacBio sequencing was performed by the University of North Carolina High Throughput Genomic Sequencing Facility after first using BluePippin size selection to isolate larger fragments of DNA template. The reads were first filtered by mapping them to the putative bacteria scaffolds from our assembly (see Supplementary Note) and removing those that appeared to be bacterial. Over 1.5 million PacBio reads with a mean read length of 5,470 bp remained. Next, these filtered reads were mapped to the HiRise assembly using BLASR via PBJelly, and gaps were filled or ends extended for regions for which there was a depth of at least ten PacBio reads. Overfilled gaps (gaps that were larger than the estimated size in the HiRise assembly) were set to an arbitrary size of 51 bp when they could not be completely filled with this set of PacBio reads at this minimum coverage threshold of 10. The resulting assembly then went through two more rounds of gap filling and end extension using this procedure, yielding our current T. californicus assembly, SDv2.1. These three rounds of PBJelly dropped the percentage of Ns from 4.67 to 1.97%, decreased the number of contigs in scaffolds from 7,105 to 4,789, and increased the overall size of the assembly by 4.7 megabase pairs (Mb) while increasing the sequence in contigs by 9.7 Mb.
Genome annotation
We used the pipeline in MAKER2 (ref. 28) to annotate putative protein-coding regions of the T. californicus assembly. First, we generated a de novo repeat library for T. californicus using the programme RepeatModeler46 and its integrated tools (RECON47, TRF48 and RepeatScout49). We also trained two ab initio gene predictors, AUGUSTUS50 and SNAP51, using a high-quality subset of T. californicus transcripts29. Within MAKER, the genome was masked for repetitive and low-complexity regions, and protein and transcript sequences were aligned using BLAST scripts. For protein evidence, we supplied MAKER with complete proteomes of multiple metazoans, and for EST evidence, we used the high-coverage T. californicus from the same population29. Alignments were fine-tuned by Exonerate52, and putative exonic regions were identified by the trained gene-prediction algorithms. Three iterative runs of MAKER were performed, with gene predictions from each run serving as training sets for the following run. Finally, MAKER evaluated the consistency across these different forms of evidence, and then generated a final set of gene models. We assessed support of gene predictions by performing mapping of T. californicus RNA-Seq reads and transcripts, and using the Benchmarking Universal Single-Copy Orthologs approach53 to examine the completeness and contiguity of our assembly.
Functional annotation of gene models was performed by BLASTP searches of the UniProt/Swiss-Prot database, followed by assignment of Gene Ontology terms and identification of protein motifs and domains from InterProScan54. We performed further manual annotation of genes of interest, such as nuclear-encoded mitochondrial proteins and some G-protein-coupled receptors, by BLASTP alignments to well-annotated databases and computational predictions of signalling peptides. A detailed description of our genome annotation steps is included in the Supplementary Note.
Orthology among arthropod models
We employed clustering algorithms to determine orthology relationships among T. californicus and metazoan model genomes, and to detect possible gene family expansions in the copepod. In OrthoVenn55, orthology was assessed among the copepod, two insects (Drosophila melanogaster and Apis mellifera), and a branchiopod crustacean (D. pulex) (Supplementary Table 12). To assess orthology among many taxa (Supplementary Table 13), we used OrthoMCL56 to assign T. californicus proteins to pre-clustered orthologous groups curated in OrthoMCL-DB version 5 (ref. 57), which contains proteomes of 150 taxa. For both analyses, BLASTP e-value threshold and inflation values were left at default (e–5 and 1.5, respectively).
Sequencing and assembly of additional T. californicus populations
Copepods were collected from intertidal pools in San Roque (27° 11′ 12′′ N, 114° 23′ 52′′ W), Bird Rock (32° 48′ 54′′ N, 117° 16′ 23′′ W), Abalone Cove (33° 44′ 16′′ N, 118° 22′ 31′′ W), Catalina Island (33° 26.8′ N, 118° 28.6′ W), Santa Cruz (36° 56′ 58′′ N, 122° 02′ 49′′ W), Bodega Bay (38° 19′ 4′′ N, 123° 4′ 23′′ W) and Friday Harbor (48° 32′ 47′′ N, 123° 0′ 35′′ W). Animals were maintained as described previously. For each population, 250–300 adult individuals were pooled and DNA was isolated using the Qiagen DNeasy Blood and Tissue Kit or a phenol–chloroform procedure58.
Samples were prepared and sequenced by the University of North Carolina High Throughput Genomic Sequencing Facility as 100-bp paired-end libraries on the Illumina HiSeq 2000. Reads were trimmed for quality using PoPoolation59, discarding bases with a Phred score of <25 and <50 bp in length after trimming. Trimmed reads were mapped to the San Diego reference genome using BWA MEM60 with the following parameters changed from the default settings: -k 15 -B 3 -O 5 -E 0. Following mapping, reads with low mapping quality (MAPQ < 20) were removed. Both paired-end and orphans reads were mapped separately, and later merged with SAMtools61. References were extracted from the mapping file using SAMtools and BCFtools62. This procedure updates the San Diego reference with single-nucleotide polymorphisms (SNPs) from each of the other populations. To insure that all (or nearly all) SNPs were present in the reference for each of populations, this process was repeated, but this time reads were mapped to their respective reference using BWA MEM with default parameters. These references have the same coordinates as the San Diego reference, and SNP positions as well as any annotation coordinates can be directly compared between any of the populations.
Assembly and annotation of mitochondrial genomes
Reads of mitochondrial origin were identified by mapping the Illumina samples above to the geographically nearest published Tigriopus mitochondrial genome63 (Abalone Cove, San Diego or Santa Cruz; accessions DQ917373, DQ917374 and DQ913891, respectively) and the San Diego draft genome assembly using BBSplit from the BBMap suite64, with the parameters 'minratio = 0.56 minhits = 1 maxindel = 16000'. Mitochondrial reads were extracted from the alignments using Picard SamToFastq and used to generate incomplete assemblies using MITObim65. Gap filling was completed by adding population reads that were flagged as paired and unmapped in BBMap alignments to the San Diego draft genome, followed by up to 11 iterations of the MITObim pipeline. Putative open reading frames of protein-coding genes were identified using MITOS66 and visually confirmed by alignments with published T. californicus mitochondrial genomes63.
Intraspecific evolution of nuclear coding regions
For each of the eight T. californicus populations, we used the ‘extractfeat’ script within GenomeTools67 to extract the coding sequence from all genes using genomic coordinates from the gff files. We filtered out genes with too many missing data (‘N’s) by excluding those that had less than 150 bp of sequence, and retained only the longest isoform of each. We checked that the extracted sequences corresponded to the correct coding sequence, with frame + 1, by aligning them with BLASTX to the San Diego protein models.
We used the programme PRANK-codon68 to align sequences for each of the 14,000 orthologous coding sequences recovered (Supplementary Table 5). Poorly aligned regions were then removed with Gblocks version 0.91b (ref. 69) with parameters: -t = c -b1 = 5 -b2 = 7 -b3 = 6 -b4 = 9 -b5 = h. Alignments shorter than 150 bp were removed, leaving a total of 13,610 coding sequence alignments for analysis. For each alignment, we estimated overall dN/dS using the model M0 in the programme CODEML within PAML version 4.7 (ref. 70), and with a well-supported ‘species’ tree estimated in RAxML71. We used these gene-wide dN/dS estimates to test for general patterns consistent with the hypothesis of compensatory evolution. Using the RNA-Seq mapping performed above, we quantified transcription levels across the San Diego genes, and used ANCOVAs to account for expression in the hypothesis tests above.
We screened the genome for signals of positive selection on amino acid changes by employing codon-level evolutionary models of nucleotide substitution implemented in CODEML. We tested the ‘sites’ hypothesis by applying models M7 and M8 on each coding sequence alignment, and we compared the resulting log-likelihood ratios with a chi-squared distribution with two degrees of freedom37. After the analysis, we applied an FDR correction for multiple testing72. Further details and parameters are included in the Supplementary Note.
Molecular evolution and population genetic analysis of mtDNA
Multiple sequence alignment of each mtDNA gene was done in MEGA6 (ref. 73) by first aligning translated coding sequences according to the invertebrate mitochondrial code (transl_table = 5) and then replacing the amino acids with the original codons. Distance matrices of non-synonymous and synonymous substitutions were estimated in MEGA6 using the Nei–Gojobori model74. Substitution distance matrices for non-protein-coding genes were calculated from CLUSTAL multiple alignments using the Analyses of Phylogenetics and Evolution R package75. To estimate nucleotide diversity, mitochondrial reads from the re-sequenced populations were mapped back to their mitochondrial genome assembly using BBMap. Sequence pileups were created using SAMtools61, 4 bp regions surrounding indels were masked to avoid false positive SNPs and the pileups were re-sampled to equal depth of coverage (200×). High-quality SNPs (Phred > 20) were used to estimate mean Tajima's π, weighted for gene length, for each protein-coding gene using PoPoolation59. DoS was measured for each protein-coding gene according to Stoletzki and Eyre-Walker76. Finally, CODEML models M0, M7 and M8, as above, were also applied to the 13 protein-coding mtDNA genes.
Patterns of positive selection across the genus Tigriopus
We downloaded the annotated gene set of T. kingsejongensis11 and RNA-Seq raw data for T. japonicus (accession numbers SAMN05933003 and SAMN05933004) generated previously11. We assembled a de novo transcriptome for T. japonicus using Trinity77, and retrieved the correct reading frames using custom scripts. To identify orthologues, we performed reciprocal BLASTX searches among the three species. We identified 5,351 orthologues. For analyses of positive selection, we were interested in testing patterns of adaptive evolution across different lineages of Tigriopus species, including T. japonicus, T. kingsejongensis, our reference T. californicus from San Diego and the reproductively isolated T. californicus from San Roque. Therefore, the orthologues identified above were aligned across the four taxa, then quality-trimmed with PRANK68 and GBlocks69, as above. We removed any alignments that were shorter than 150 bp. Using the maximum likelihood method of Yang and Nielsen78, we computed pairwise dS for all genes and discarded those with dS > 5 in order to reduce the influence of site saturation or poor alignments. The final set of alignments contained 4,863 genes.
We concatenated 20 randomly picked alignments and estimated a maximum likelihood phylogeny in MEGA6 (ref. 73). In CODEML, we applied the branch-site model41, which allows dN/dS to vary among sites and among branches, and tested for evidence of positive selection on each of the 4,863 aligned coding sequences along each internal branch and tip. For each gene and each foreground branch, we calculated a log-likelihood ratio between a model allowing for positive selection and a null model in which dN/dS is fixed at 1, and these ratios were compared with a chi-squared distribution with one degree of freedom. After the analysis, we applied an FDR correction for multiple testing72.
Reporting Summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Raw Illumina and PacBio DNA sequencing reads have been deposited in the NCBI Sequence Read Archive database (San Diego reference: SRX469409–SRX469413; additional populations: SRX2746698–SRX2746704; PacBio reads for the San Diego reference: SRX3778522–SRX3778523). The T. californicus annotated genome assembly was deposited in the i5k Workspace of the National Agricultural Library (US Department of Agriculture), where it can be browsed, searched and downloaded: https://i5k.nal.usda.gov/Tigriopus_californicus.
References
Coyne, J. A. & Orr, H. A. Speciation (Sinauer Associates, Sunderland, 2004).
Burton, R. S., Pereira, R. J. & Barreto, F. S. Cytonuclear genomic interactions and hybrid breakdown. Annu. Rev. Ecol. Evol. Syst. 44, 281–302 (2013).
Chou, J.-Y. & Leu, J.-Y. The Red Queen in mitochondria: cyto-nuclear co-evolution, hybrid breakdown and human disease. Front. Genet. 6, 1–29 (2015).
Bar-Yaacov, D., Blumberg, A. & Mishmar, D. Mitochondrial-nuclear co-evolution and its effects on OXPHOS activity and regulation. Biochim. Biophys. Acta. 1819, 1107–1111 (2012).
Hill, G. E. Cellular respiration: the nexus of stress, condition, and ornamentation. Integr. Comp. Biol. 54, 645–657 (2014).
Aanen, D. K., Spelbrink, J. N. & Beekman, M. What cost mitochondria? The maintenance of functional mitochondrial DNA within and across generations. Phil. Trans. R. Soc. B 369, 20130438 (2014).
Bron, J. E. et al. Observing copepods through a genomic lens. Front. Zool. 8, 22 (2011).
Madoui, M.-A. et al. New insights into global biogeography, population structure and natural selection from the genome of the epipelagic copepod Oithona. Mol. Ecol. 26, 4467–4482 (2017).
Eyun, S.-I. et al. Evolutionary history of chemosensory-related gene families across the Arthropoda. Mol. Biol. Evol. 34, 1838–1862 (2017).
Lee, J.-S. et al. The copepod Tigriopus japonicus genomic DNA information (574Mb) and molecular anatomy. Mar. Environ. Res. 69, S21–S23 (2010).
Kang, S. et al. The genome of the Antarctic-endemic copepod Tigriopus kingsejongensis. GigaScience 6, 1–9 (2017).
Burton, R. S. et al. Hybrid breakdown in developmental time in the copepod Tigriopus californicus. Evolution 44, 1814–1822 (1990).
Edmands, S. Heterosis and outbreeding depression in interpopulation crosses spanning a wide range of divergence. Evolution 53, 1757–1768 (1999).
Rawson, P. D. & Burton, R. S. Functional coadaptation between cytochrome c and cytochrome c oxidase within allopatric populations of a marine copepod. Proc. Natl Acad. Sci. USA 99, 12955–12958 (2002).
Harrison, J. S. & Burton, R. S. Tracing hybrid incompatibilities to single amino acid substitutions. Mol. Biol. Evol. 23, 559–564 (2006).
Ellison, C. K. & Burton, R. S. Interpopulation hybrid breakdown maps to the mitochondrial genome. Evolution 62, 631–638 (2008).
Ellison, C. K. & Burton, R. S. Genotype-dependent variation of mitochondrial transcriptional profiles in interpopulation hybrids. Proc. Natl Acad. Sci. USA 105, 15831–15836 (2008).
Ellison, C. K. & Burton, R. S. Cytonuclear conflict in interpopulation hybrids: the role of RNA polymerase in mtDNA transcription and replication. J. Evol. Biol. 23, 528–538 (2010).
Burton, R. S. & Barreto, F. S. A disproportionate role for mtDNA in Dobzhansky–Muller incompatibilities? Mol. Ecol. 21, 4942–4957 (2012).
Willett, C. S. Quantifying the elevation of mitochondrial DNA evolutionary substitution rates over nuclear rates in the intertidal copepod Tigriopus californicus. J. Mol. Evol. 74, 310–318 (2012).
Alexander, H. J., Richardson, J. M. L., Edmands, S. & Anholt, B. R. Sex without sex chromosomes: genetic architecture of multiple loci independently segregating to determine sex ratios in the copepod Tigriopus californicus. J. Evol. Biol. 28, 2196–2207 (2015).
Rieseberg, L. H. Chromosomal rearrangements and speciation. Trends Ecol. Evol. 16, 351–358 (2001).
Phillips, B. C. & Edmands, S. Does the speciation clock tick more slowly in the absence of heteromorphic sex chromosomes? BioEssays 34, 166–169 (2012).
Lima, T. G. Higher levels of sex chromosome heteromorphism are associated with markedly stronger reproductive isolation. Nat. Comm. 5, 4743 (2014).
Gregory, T. R. Animal Genome Size Database (2015); http://www.genomesize.com
Wyngaard, G. A. & Rasch, E. M. Patterns of genome size in the Copepoda. Hydrobiologia 417, 43–56 (2000).
Foley, B. R. et al. A gene-based SNP resource and linkage map for the copepod Tigriopus californicus. BMC Genom. 12, 568 (2011).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics 12, 491 (2011).
Barreto, F. S., Pereira, R. J. & Burton, R. S. Hybrid dysfunction and physiological compensation in gene expression. Mol. Biol. Evol. 32, 613–622 (2015).
Peterson, D. L. et al. Reproductive and phylogenetic divergence of tidepool copepod populations across a narrow geographical boundary in Baja California. J. Biogeogr. 40, 1664–1675 (2013).
Bazin, E., Glémin, S. & Galtier, N. Population size does not influence mitochondrial genetic diversity in animals. Science 312, 570–571 (2006).
Barreto, F. S. & Burton, R. S. Evidence for compensatory evolution of ribosomal proteins in response to rapid divergence of mitochondrial rRNA. Mol. Biol. Evol. 30, 310–314 (2013).
Pál, C., Papp, B. & Hurst, L. Highly expressed genes in yeast evolve slowly. Genetics 158, 927–931 (2001).
Subramanian, S. Gene expression intensity shapes evolutionary rates of the proteins encoded by the vertebrate genome. Genetics 168, 373–381 (2004).
Drummond, D. A., Bloom, J. D., Adami, C., Wilke, C. O. & Arnold, F. H. Why highly expressed proteins evolve slowly. Proc. Natl Acad. Sci. USA 102, 14338–14343 (2005).
Adrion, J. R., White, P. S. & Montooth, K. L. The roles of compensatory evolution and constraint in aminoacyl tRNA synthetase evolution. Mol. Biol. Evol. 33, 152–161 (2015).
Yang, Z., Nielsen, R., Goldman, N. & Pedersen, A. Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics 155, 431–449 (2000).
Ellison, C. K. & Burton, R. S. Disruption of mitochondrial function in interpopulation hybrids of Tigriopus californicus. Evolution 60, 1382–1391 (2006).
Barreto, F. S., Schoville, S. D. & Burton, R. S. Reverse genetics in the tide pool: knock-down of target gene expression via RNA interference in the copepod Tigriopus californicus. Mol. Ecol. Resour. 15, 868–879 (2014).
Meiklejohn, C. D. et al. An incompatibility between a mitochondrial tRNA and its nuclear-encoded tRNA synthetase compromises development and fitness in Drosophila. PLoS Genet. 9, e1003238 (2013).
Zhang, J. Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Mol. Biol. Evol. 22, 2472–2479 (2005).
Roux, J. et al. Patterns of positive selection in seven ant genomes. Mol. Biol. Evol. 31, 1661–1685 (2014).
Putnam, N. H. et al. Chromosome-scale shotgun assembly using an in vitro method for long-range linkage. Genome Res. 26, 342–350 (2016).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
English, A. C. et al. Mind the gap: upgrading genomes with Pacific Biosciences RS Long-Read Sequencing Technology. PLoS ONE 7, e47768 (2012).
Smit, A. & Hubley, R. RepeatModeler Open-1.0 (2014); http://www.repeatmasker.org/RepeatModeler/
Bao, Z. & Eddy, S. R. Automated de novo identification of repeat sequence families in sequenced genomes. Genome Res. 12, 1269–1276 (2002).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21, i351–i358 (2005).
Stanke, M. & Morgenstern, B. AUGUSTUS: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Res. 33, W465–W467 (2005).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Slater, G. S. C. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6, 31 (2005).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Quevillon, E. et al. InterProScan: protein domains identifier. Nucleic Acids Res. 33, W116–W120 (2005).
Wang, Y., Coleman-Derr, D., Chen, G. & Gu, Y. Q. OrthoVenn: a web server for genome wide comparison and annotation of orthologous clusters across multiple species. Nucleic Acids Res. 43, W78–W84 (2015).
Li, L., Stoeckert, C. J. & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Fischer, S. et al. Using OrthoMCL to assign proteins to OrthoMCL-DB groups or to cluster proteomes into new ortholog groups. Curr. Protoc. Bioinformatics 6, 6.12.1–6.12.19 (2011).
Sambrook, J. & Russell, D. W. Purification of nucleic acids by extraction with phenol:chloroform. Cold Spring Harb. Protoc. 2006, pdb.prot4455 (2010).
Kofler, R. et al. PoPoolation: a toolbox for population genetic analysis of next generation sequencing data from pooled individuals. PLoS ONE 6, e15925 (2011).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Burton, R. S., Byrne, R. J. & Rawson, P. D. Three divergent mitochondrial genomes from California populations of the copepod Tigriopus californicus. Gene 403, 53–59 (2007).
Bushnell, B. BBMap (2014); http://www.repeatmasker.org/RepeatModeler/
Hahn, C., Bachmann, L. & Chevreux, B. Reconstructing mitochondrial genomes directly from genomic next-generation sequencing reads—a baiting and iterative mapping approach. Nucleic Acids Res. 41, e129 (2013).
Bernt, M. et al. MITOS: improved de novo metazoan mitochondrial genome annotation. Mol. Phylogenet. Evol. 69, 313–319 (2013).
Gremme, G., Steinbiss, S. & Kurtz, S. GenomeTools: a comprehensive software library for efficient processing of structured genome annotations. IEEE/ACM Trans. Comput. Biol. Bioinform. 10, 645–656 (2013).
Löytynoja, A. & Goldman, N. Phylogeny-aware gap placement prevents errors in sequence alignment and evolutionary analysis. Science 320, 1632–1635 (2008).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. 57, 289–300 (1995).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Nei, M. & Gojobori, T. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol. Biol. Evol. 3, 418–426 (1986).
Paradis, E. Analysis of Phylogenetics and Evolution Using R (Springer, New York, 2012).
Stoletzki, N. & Eyre-Walker, A. Estimation of the neutrality index. Mol. Biol. Evol. 28, 63–70 (2011).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Yang, Z. & Nielsen, R. Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol. Biol. Evol. 17, 32–43 (2000).
Acknowledgements
This work was supported by US National Science Foundation grants (IOS1154321 to S.E.; IOS1155030 to R.S.B.; and IOS1155325 to C.S.W.) and Oregon State University faculty startup funds to F.S.B. The authors thank S. Morgan and R. J. Pereira for help with sample collection.
Author information
Authors and Affiliations
Contributions
F.S.B., E.T.W., T.G.L., C.S.W., S.E. and R.S.B. contributed to the design of the project, collection of biological samples, and sequence data acquisition. W.L. contributed to initial genome sequence assembly. F.S.B., E.T.W. and C.S.W. contributed to genome annotation. F.S.B., E.T.W., T.G.L. and C.S.W. contributed to computational and statistical analyses. F.S.B., E.T.W., T.G.L., C.S.W., S.E. and R.S.B. contributed to data interpretation and writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Note, Supplementary Figures and Supplementary Tables
Supplementary Data
Supplementary Tables 9, 10, 12–17. Supplementary Table 9. Results of sites test among eight T. californicus populations. Supplementary Table 10. Results of branch-sites tests among Tigriopus species. Supplementary Table 12. Orthology of T. californicus proteins and three arthropods. Supplementary Table 13. Orthology of T. californicus proteins and nine metazoans. Supplementary Table 14. Classification of genome scaffolds of microbial origin. Supplementary Table 15. Clustering of putative FMRFamide receptors in T. californicus. Supplementary Table 16. BLASTP searches of arthropods proteomes for FMRFaR. Supplementary Table 17. Self-BLAST of FMRFaR against all T. californicus proteins
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Barreto, F.S., Watson, E.T., Lima, T.G. et al. Genomic signatures of mitonuclear coevolution across populations of Tigriopus californicus. Nat Ecol Evol 2, 1250–1257 (2018). https://doi.org/10.1038/s41559-018-0588-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0588-1
This article is cited by
-
New genes helped acorn barnacles adapt to a sessile lifestyle
Nature Genetics (2024)
-
High heterogeneity in genomic differentiation between phenotypically divergent songbirds: a test of mitonuclear co-introgression
Heredity (2023)
-
Mitonuclear incompatibility as a hidden driver behind the genome ancestry of African admixed cattle
BMC Biology (2022)
-
Genome of a giant isopod, Bathynomus jamesi, provides insights into body size evolution and adaptation to deep-sea environment
BMC Biology (2022)
-
Mito-nuclear selection induces a trade-off between species ecological dominance and evolutionary lifespan
Nature Ecology & Evolution (2022)